Abstract
Glucocorticoid-Induced TNFR-related protein (GITR) is a costimulatory molecule involved in the proliferation and effector functions of CD8 + and CD4 + T-cells. Recently, it has gained attention as a novel target for HIV immunotherapy. However, reports on its expression in people living with HIV (PLWH), as well as functional studies of GITR ligands effects in the context of HIV remain scarce. Here, we performed a thorough immune characterization of GITR expression in PLWH following HIV peptide stimulation. We found a prevalence of an effector memory phenotype on HIV-specific GITR-expressing CD8 + T-cells that correlated with viral control. Costimulation with a hexameric GITR ligand (GITRL) showed a modest improvement on antiviral function. Characterization of CD4 T-cells revealed an association between GITR expression among regulatory T-cells and viral control, as well as the prevalence of a central and effector memory T-cell phenotype. Remarkably, GITRL costimulation enhanced viral transcription without increasing the reservoir size, positioning GITR as an interesting target for the development of novel latency reversal agents.
Similar content being viewed by others
Introduction
Human Immunodeficiency Virus (HIV) is a retrovirus responsible for causing a persistent infection that results in a progressive decline in the immune system1. So far, an estimated 85 million people worldwide acquired HIV-1, and nearly 40 million died from related causes. Despite the dramatic success in reducing mortality and morbidity, antiretroviral treatment (ART) is unable to eradicate the virus. This is due to the persistence of inducible, intact and replicative competent provirus integrated in the genome of several cell types-mainly resting memory CD4 T-cells (CD4TCs)-which constitute the viral reservoir2. However, evidence from a few exceptional cases of ART-free viral remission suggests a cure is possible3,4,5,6. These inspiring cases, along with the acknowledgment of the pharmacological, economic and logistical limitations of ART, have led to a global consensus that finding a cure is a high priority7. Different approaches to achieve an eradicative cure have been proposed. The “shock and kill” strategy involves reactivation of viral production from latently infected cells and their subsequent elimination by HIV-specific CD8 T-cells (CD8TCs)8. However, it is well reported that CD8TCs exhibit signs of exhaustion-such as limited proliferative capacity and cytokine expression9,10-which remain under ART, reducing their capacity to clear infected cells. Another limitation is the poor antigen induction by existing latency-reversing agents (LRAs) and its off-target effects11,12.
Thus, development of new components capable of potentiating latency reversal and enhancing anti-HIV response functionality is needed. The glucocorticoid-induced TNFR-related protein (GITR) is a type-I transmembrane receptor, member of the TNFR superfamily (TNFRSF)13 that has gained attention due to its role in the improvement of T-cell effector function. GITR is constitutively expressed at high levels on regulatory T-cells (Tregs), at lower levels on naïve and memory T-cells, and is upregulated upon T-cell activation14,15. Its ligand is expressed by antigen presenting cells, and its signaling culminates in NF-κB activation16. Several in vitro studies have shown that upon TCR costimulation, GITR triggering enhances proliferation, cytokine production and effector function of CD8 and CD4TCs17,18.
Despite promising results from in vitro studies, most clinical trials evaluating GITR agonist monoclonal antibodies for the treatment of solid tumors have shown limited therapeutic activity. This could be the result of antibody structural restrictions that make them unsuitable for stimulating the well-defined trimeric receptors of the TNFRSF. Fc-Flag-TNC-GITR ligand (GITRL) is a hexavalent agonist developed by Wyzgol et al.19 in an attempt to overcome these constraints20. GITRL can promote IL-8 production on cell lines; however, it has not been evaluated on primary cells nor in the context of HIV infection.
Here, we performed an exhaustive immunophenotypic characterization of GITR-expressing CD8TCs, conventional CD4TCs and regulatory CD4TCs from people living with HIV (PLWH) who were either receiving ART or not at the moment of enrollment. Furthermore, we evaluated the impact of GITRL on the effector function of HIV-specific CD8TCs and on viral reactivation from latently infected CD4TCs.
Materials and methods
Study participants
53 PLWH were included in this study: 42 receiving ART (On-ART) and 11 not receiving ART (Off-ART). Immunophenotypic characterization was performed on 22 participants (11 Off-ART and 11 On-ART) (Table S1). Functional and latency reversal assays were performed on 31 PLWH On-ART (Table S2).
Ethical considerations
This study was approved by the institutional review board Comité de Ética Humana, Facultad de Medicina, Universidad de Buenos Aires, Argentina. All participants provided written informed consent. All methods were performed in accordance with the Declaration of Helsinki.
Samples
Cryopreserved peripheral blood mononuclear cell (PBMC) samples from Off-ART PLWH were formally requested and obtained from the Argentinean Biobank of Infectious Diseases (BBEI). On-ART participants were enrolled at Fundación Huésped and 40 mL of whole blood was collected in EDTA-containing tubes at enrollment. Upon centrifugation, plasma was recovered, and stored at − 80 °C. PBMCs were isolated by Ficoll-Hypaque density gradient centrifugation (Ficoll-PaqueTM, USA) and cryopreserved in liquid nitrogen for phenotypic and functional assays. Plasma viral load (VL) was determined by Abbott Real Time HIV-1 assay (Abbott Park, USA). CD4 + and CD8 + T-cell counts were determined by flow cytometry (FACSCalibur, BD, USA).
Experimental assays
Post-stimulation characterization of GITR expression on CD4 and CD8TCs, and GITR expression kinetics were performed by flow cytometry (Supplementary Table S3). A hexameric GITR ligand fusion protein was kindly provided by Dr. Wajant19 and used for functional assays: GITRL binding, GITRL-mediated modulation of T-cell functionality, proliferation, reservoir size and latency reversion were evaluated and analyzed as described in Supplementary data.
Data analysis
Data were assessed using parametric or non-parametric methods, based on whether the normality assumption was met. Data were analyzed using GraphPad Prism v8.4 and SPICE 6.0 software (https://niaid.github.io/spice/). Tests were considered significant when p < 0.05. For correlation analysis, p-values were adjusted for multiple comparisons using a hypothesis-driven false discovery rate (FDR) procedure, according to the Benjamini and Hochberg method Adjusted p-values were considered significant when < 0.1.
Results
Cohort description
22 PLWH (11 Off-ART and 11 On-ART) were included in the first part of this study. Clinical characteristics are depicted in Table S1. At enrollment, all Off-ART participants had detectable viral loads (VL, median = 5.2log10 HIV RNA copies/mL) and all On-ART participants had undetectable VL. The On-ART group showed significantly higher CD4 counts and CD4/CD8 ratios. Additionally, samples from 31 On-ART participants were obtained for functional assays (Table S2).
GITR is expressed in T-cells from PLWH off and On-ART and correlates with parameters related to viral control
PBMCs were cultured for 3 (Off-ART) or 14 (On-ART) days with Nef or p24 peptide pools (or a cytomegalovirus, Epstein-Barr and influenza virus peptide pool (CEF) as control) and GITR expression was evaluated in bulk CD8TCs, HIV-specific CD8TCs, conventional CD4TCs (Tconv) and Treg (Fig. S1). The longer culture time for the On-ART group was necessary for the enrichment of HIV-specific cells for analysis, based on the notion that the frequency of this population decays significantly following ART initiation.
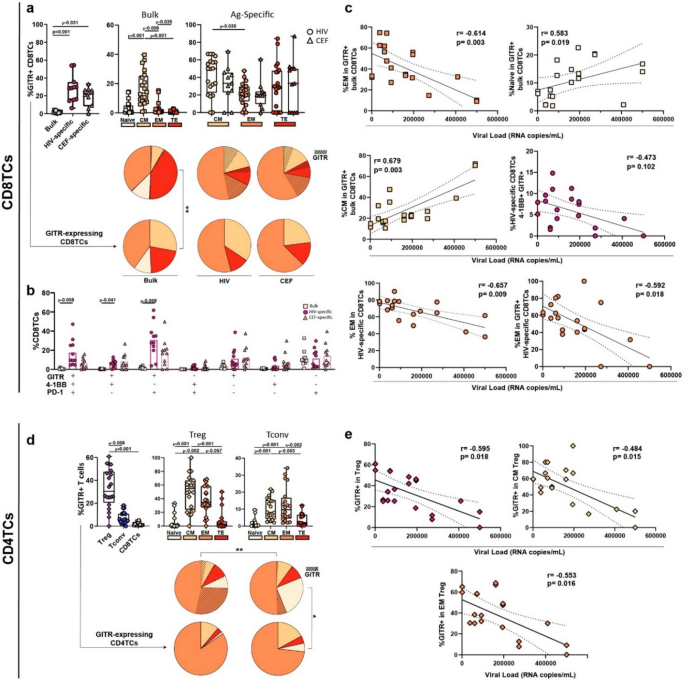
In the Off-ART group, post-stimulation GITR expression within HIV- and CEF-specific CD8TC subpopulations was significantly higher compared to CD8TC bulk compartment (Fig. 1a, upper left). Within bulk CD8TCs, the central memory (CM) subset was the most enriched in GITR (median = 13.50%, IQ25-75 = 7.56–25.48) (Fig. 1a, upper center). The median frequencies of virus-specific CD8TCs were HIV = 0.375 and CEF = 0.13. The mean frequencies of each memory subset within HIV-specific CD8TCs were CM = 20.91, EM = 67.59, TE = 11.48 while within CEF-specific these were CM = 17.15, EM = 72.25, TE = 10.59. Within antigen-specific cells, the percentage of GITR-expressing HIV-specific CM cells was higher than EM cells. No significant differences in GITR expression were found across memory subsets for CEF-specific CD8TCs (Fig. 1a, upper right), nor in global memory distribution between HIV-specific and CEF-specific CD8TCs (Fig. 1a upper pies).
We then analyzed the distribution of memory/effector subpopulations within GITR-expressing CD8TCs (Fig. 1a, lower pies). This distribution was significantly different to the memory distribution of global CD8TCs (upper pie), with a higher contribution of the CM subset. The effector memory (EM) subset dominated in bulk GITR-expressing CD8TCs, reaching a mean = 40.07%, and was even higher in the HIV-specific and CEF-specific compartments (54.01% and 57.79%).
Next, we evaluated the single-expression and coexpression of GITR, PD-1 and 4-1BB (costimulatory receptor, member of TNFRSF). A significantly higher post-stimulation coexpression of GITR+/4-1BB+/PD-1+ was found in HIV-specific CD8TCs compared to bulk. GITR+/4-1BB+ and GITR+/PD-1+ coexpression was also higher in HIV-specific cells than in bulk CD8TCs (Fig. 1b).
Correlation analyses (Fig. 1c and supplementary Table S4) depicted that %EM in bulk GITR + CD8TCs inversely correlated with VL. Conversely, %Naive and %CM in bulk GITR + CD8TCs positively correlated with VL. Within HIV-specific CD8TCs, %GITR+/4-1BB+ cells negatively correlated with VL and %EM in GITR-expressing cells also correlated negatively with VL, although this association could be driven by the overall negative correlation between %EM and VL (Supplementary Table S5, Box 8, 10 and 11). Finally, %GITR in terminal effector (TE) and %TE in GITR + positively correlated with CD4/CD8 and CD4TC counts, respectively (Supplementary Table S5, Box 8 and 12).
In the CD4TC compartment, GITR expression was significantly higher in Treg compared with Tconv and bulk CD8TCs (Fig. 1d left panel). Within Tregs, the CM compartment was the most enriched in GITR (54.55%, IQ25-75 = 23.88–66.70), followed by EM (34.85%, IQ25-75 = 27.23–58.30) (Fig. 1d, center). In Tconv, GITR enrichment was higher in CM (8.09%, IQ25-75 = 4.12–15.43) and EM (7.42%, IQ25-75 = 4.15–14.15) compared with Naïve and TE (Fig. 1d right). The global distribution of subsets was significantly different between Treg and Tconv, with a higher contribution of GITR + EM within Treg (Fig. 1d upper pies). Within GITR-expressing CD4TCs, most Treg and Tconv had an EM phenotype (81.23% and 73.24%) (Fig. 1d, lower pies) and global distribution of bulk Tconv (upper pie) was significantly different compared to GITR-expressing Tconv (lower pie), with a contraction of the Naive subset in the latter.
Correlation analyses depicted negative correlations between VL and %GITR-expressing cells in bulk, CM and EM Tregs (Fig. 1e and supplementary Table S6). No other relevant significant associations were found (Supplementary Figs. S3a, S4a–c).
GITR expression on bulk and HIV-specific T-cells from PLWH Off-ART. (a) Box plots represent the proportion of GITR + events within bulk and antigen-specific (Ag-specific) CD8TCs (left) and within each memory subset (bulk: center and Ag-specific: right). The color code shown at the bottom mirrors the one shown in the pie charts. Upper pie graphs depict the CD8TC memory distribution and contribution of GITR + events on bulk and Ag-specific cells. Lower pie graphs depict the memory distribution of the bulk and Ag-specific GITR + CD8TCs. (b) Bar plot represents the GITR, 4-1BB and PD-1 single and coexpression pattern in bulk, HIV- and CEF-specific CD8TCs. (c) Correlation analysis between GITR expression across memory subsets in bulk/HIV-specific CD8TCs and viral load. (d) Box plots represent the proportion of GITR + events within regulatory (Treg), conventional (Tconv) CD4TCs and bulk CD8TCs (left); and within each memory subset of Treg and Tconv CD4TCs (center and right, respectively). Upper pie graphs depict the CD4TC memory distribution and contribution of GITR + events on Treg and Tconv cells. Lower pie graphs depict the memory distribution of the GITR + Treg and GITR + Tconv. (e) Correlation analyses between GITR expression across different memory subsets in Treg and viral load. CM: central memory, EM: effector memory, TE: terminal effector. N = 11 individuals, 20 HIV responses (10 Nef + 10 p24) and 11 CEF responses. In box plots, boxes extend from Q1 to Q3, horizontal bars within boxes represent median values and whiskers extend from min to max. In bar plots, bar height represents mean. When analyzing differences between CEF and HIV, the Nef and p24 responses were averaged and grouped under the HIV response to obtain the same number of responses as CEF and apply the appropriate statistical test. When analyzing differences within a single response, the full dataset was used. P-values were calculated using the Friedman test for paired samples, followed by Dunn’s post-hoc correction for multiple comparisons. When analyzing global distribution (pies) permutation tests were applied using SPICE software. In all plots, squares represent bulk CD8TCs, circles represent HIV-specific CD8TCs, triangles represent CEF-specific CD8TCs, diamonds represent Treg and hexagons represent Tconv. r and p values were computed by Spearman´s correlation and p-values were adjusted for multiple comparisons using a hypothesis-driven false discovery rate (FDR) procedure, according to the Benjamini and Hochberg method.
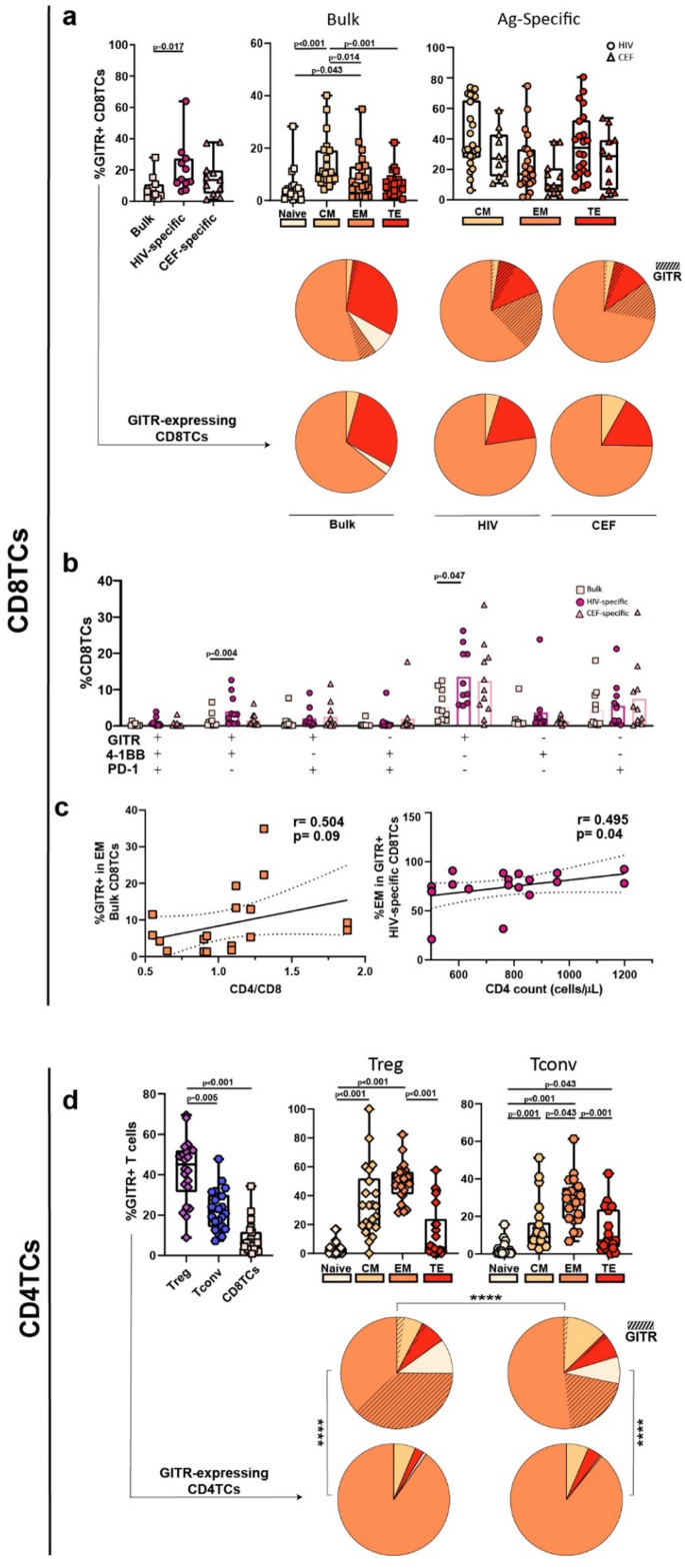
After exploring GITR expression on T-cells from PLWH Off-ART, we extended the analysis to PLWH On-ART. Similar to the Off-ART group, the percentage of GITR + cells was higher among HIV-specific CD8TCs compared with bulk CD8TCs (Fig. 2a upper left). When analyzing GITR expression across memory/effector subsets within bulk CD8TCs, the CM subset showed the greatest GITR enrichment (10.25% IQ25-75 = 7.83–19.18) (Fig. 2a upper center). The median of frequencies of virus-specific CD8TCs were HIV = 19.7, CEF = 9.85. The mean frequencies of each memory subset within HIV-specific CD8TCs were CM = 2.54, EM = 81.35, TE = 16.09, while within CEF-specific these were CM = 3.35, EM = 84.79, TE = 11.86. Within post-stimulated HIV-specific CD8TCs, the TE and CM phenotypes were the most enriched in GITR (34.10% IQ25-75 = 17.38–52.25 and 33.25% IQ25-75 = 27.55–65.28, respectively), followed by EM (16.85% IQ25-75 = 10.63–33.20) (Fig. 2a upper right). The same hierarchy (TE = CM > EM) was observed in CEF-specific CD8TCs. There was no difference between global memory distribution between HIV-specific and CEF-specific CD8TCs (Fig. 2a, upper pie). Among GITR-expressing cells (Fig. 2a, lower pie), the EM phenotype dominated in bulk CD8TCs, with a mean of 71.16%, and was also higher in the HIV- and CEF-specific compartments (74.05% and 75.36%, respectively), with no differences in the global memory distribution between GITR-expressing HIV-specific and CEF-specific cells.
The analysis of the post-stimulation single-expression and coexpression of GITR, 4-1BB and PD-1 revealed a significantly higher percentage of GITR+ and GITR+/4-1BB+ CD8TCs in HIV-specific compartment compared with bulk CD8TCs (Fig. 2b). Correlation analyses showed that the percentage of GITR + cells within EM CD8TCs positively correlated with CD4/CD8 ratios and, within GITR + HIV-specific CD8TCs, the %EM cells directly correlated with CD4TC counts (Fig. 2c; supplementary Tables S7 and S8). No other relevant significant associations were found (Supplementary Table S9, supplementary Figs. S3b and S4d).
On-ART CD4TC analyses also revealed significantly higher GITR expression in Tregs versus Tconv and bulk CD8TCs (Fig. 2d left panel). In both Treg and Tconv, the EM phenotype was the most enriched in GITR (51.50% IQ25-75 = 40.93–56.55 and 28.75% IQ25-75 = 18.75–35.40, respectively), followed by CM (33% IQ25-75 = 18.93–51.98 and 9.03% IQ25-75 = 5.14–16.70, respectively) (Fig. 2d, center and right box charts). Memory global distribution was significantly different between Treg and Tconv, with a higher contribution of GITR + EM within Treg (Fig. 2d upper pies). In both GITR-expressing Tregs and Tconv, most cells (90.14% and 89.18%, respectively) exhibited an EM phenotype (Fig. 2d, lower pies), with significant differences in global memory distribution compared to bulk subsets (upper pies).
GITR expression on bulk and HIV-specific T-cells from PLWH On-ART. (a) Box plots represent the percentage of GITR + events within bulk and antigen-specific (Ag-specific) CD8TCs (left) and within each memory subset (bulk, center and Ag-specific, right). The color code shown at the bottom mirrors the one shown in the pie charts. Upper pie graphs depict the CD8TC memory distribution and contribution of GITR + events on bulk and Ag-specific cells. Lower pie graphs depict the memory distribution of the bulk and Ag-specific GITR + CD8TCs. (b) Bar plots represent the GITR, 4-1BB and PD-1 single and coexpression pattern in bulk, HIV- and CEF-specific CD8TCs. (c) Correlation analysis between percentage of GITR + in EM bulk CD8TCs and %EM in GITR + HIV-specific CD8TCs with CD4/CD8 ratio and CD4TC counts. (d) Box plots represent the proportion of GITR + events within regulatory (Treg), conventional (Tconv) CD4TCs and bulk CD8TCs (left), and within each memory subset of Treg and Tconv CD4TCs (center and right, respectively). Upper pie graphs depict the CD4TC memory distribution and contribution of GITR + events on Treg and Tconv cells. Lower pie graphs depict the memory distribution of the GITR + Treg and GITR + Tconv. CM: central memory, EM: effector memory, TE: terminal effector. N = 11 individuals. For bar charts: 22 HIV responses (11 Nef + 11 p24) and 11 CEF responses. For correlation analyses: 20 HIV responses (10 Nef + 10 p24). In box plots, boxes extend from Q1 to Q3, horizontal bars within boxes represent median values and whiskers extend from min to max. In bar plots, bar height represents mean. When analyzing differences between CEF and HIV, the Nef and p24 responses were averaged and grouped under the HIV response to obtain the same number of responses as CEF and apply the appropriate statistical test. When analyzing differences within a single response, the full dataset was used. P-values were calculated using the Friedman test for paired samples, followed by Dunn’s post-hoc correction for multiple comparisons. When analyzing global distribution (pies) permutation tests were applied using SPICE software. In all plots, squares represent bulk CD8TCs, circles represent HIV-specific CD8TCs, triangles represent CEF-specific CD8TCs, diamonds represent Treg, and hexagons represent Tconv. r and p values were computed by Spearman´s correlation and p-values were adjusted for multiple comparisons using a hypothesis-driven false discovery rate (FDR) procedure, according to the Benjamini and Hochberg method.
Overall, we showed that GITR is expressed in T-cells from PLWH Off- and On-ART. Given the potential role of GITR in HIV control and the fact that most GITR-expressing CD8TCs are EM cells, we explored enhancing antiviral function through a hexameric GITR ligand. Additionally, due to the observed GITR enrichment on CM and EM CD4TCs, we evaluated the HIV latency-reversal properties of GITRL.
GITR expression peaks at 96 h post-stimulation on CD8 and CD4TCs from PLWH On-ART and it is specifically recognized by a hexameric GITR ligand
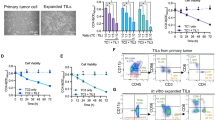
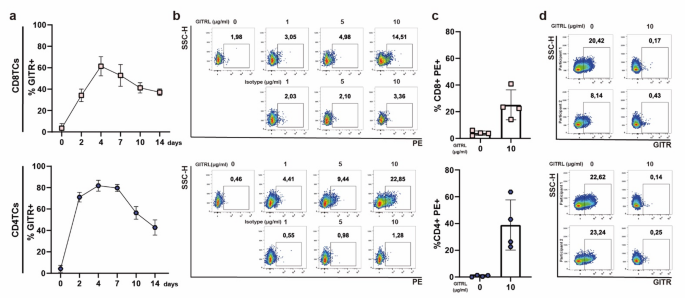
A post-activation kinetic analysis of PBMCs revealed similar GITR expression patterns between CD8 and CD4TCs, with rapid upregulation up to day 4 (Fig. 3a). After establishing GITR peak expression, we assessed the binding capacity of GITRL on CD8 and CD4TCs at that time point. A dose-response relationship between GITRL concentration and fluorescence signal was observed on activated CD8 and CD4TCs (Fig. 3b–c) and an interference assay confirmed GITRL specificity for its receptor (Fig. 3d). A similar binding assay on resting PBMCs in the absence of stimuli for 4 days showed comparable results (Fig. S2a). Finally, using the Flag tag in the GITRL construct, we repeated the binding assay in activated CD4TCs with an anti-Flag mAb M2, observing the same pattern (Fig. S2b).
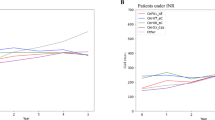
GITR expression kinetics and GITRL binding on activated CD8 (upper panel) and CD4TCs (lower panel) from PLWH On-ART. (a) GITR kinetic expression. Squares and circles represent the mean percentage of GITR + CD8 and CD4TCs. Whiskers represent the standard deviation. N = 4 participants. (b) GITRL binding on activated CD8 and CD4TCs. Representative flow cytometry dot plot from one participant, showing the proportion of PE + events after GITRL or isotype incubation, developed with αhuman IgG1-biotin + Streptavidin-PE. (c) Pooled data (N = 4 participants) from GITRL binding assay. Percentage of PE + CD4 and CD8TCs determined after T-cell activation and treatment with 0 and 10 ug/mL of GITRL. Bar height represents the mean and whiskers represent the standard deviation. (d) Results from GITR-GITRL interference assay. Flow cytometry dot plots from 2 participants show the percentage of GITR + events within activated CD8 and CD4TCs after 5 h of incubation in the absence (0 µg/mL) or presence (10 µg/mL) of GITRL.
GITRL costimulation enhances IFN-γ production and degranulation in HIV-specific CD8TCs from PLWH On-ART in the short-term, but it has no effect on their long-term cytolytic function
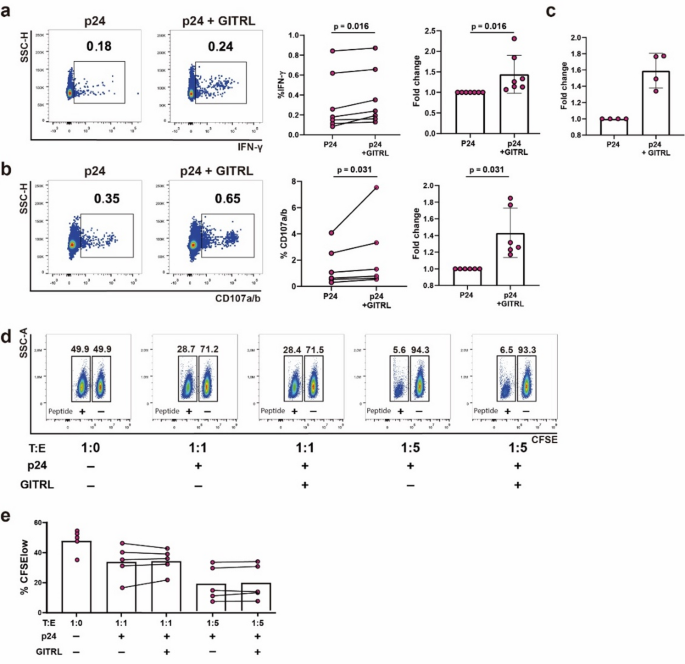
To assess GITRL effect on enhancing HIV-specific responses, PBMCs from 7 On-ART PLWH were treated with a p24 peptide pool plus GITRL for 4 days. After a brief restimulation, the frequency of IFN-γ + and CD107a/b + CD8TCs significantly increased in the GITRL-stimulated condition (Fig. 4a–b). A similar trend was observed for IFN-γ levels measured by ELISA in culture supernatants (Fig. 4c).
To evaluate if this improvement in cytokine production and degranulation had an effect on CD8TC cytolytic capacity, a VITAL killing assay was performed, following our group’s established protocol21. Figure 4d shows flow-cytometry dot-plots obtained in one representative assay, which indicates a specific progressive loss of antigen-pulsed CFSElow target cells as effectors are added at increasing ratios. Treatment of effector cells with GITRL during the expansion with p24 peptides had no effect on the cytotoxic function of CD8TCs in either effector: target condition (Fig. 4e). Moreover, a CFSE-based proliferation assay on PBMCs stimulated with p24 peptides showed no significant difference in the percentage of proliferating CD8TCs between conditions with or without GITRL after 7 days of stimulation (Supplementary Fig. 5).
Effect of GITRL costimulation on HIV-specific CD8TCs from PLWH On-ART. Effect of GITRL costimulation on (a) IFN-γ production and (b) degranulation. Within panels A and B, dot plots depict the results from a representative donor. Scatter plots depict pooled data from 7 donors. Left scatter plot with lines shows percentages of IFN-γ + or CD107a/b + events while right scatter plot with bars shows the fold change between conditions. p values were calculated using Wilcoxon test. (c) Scatter plot with bar graph of pooled data from 4 participants showing the fold change of IFN-γ concentration from supernatants of PBMCs after 4 days of p24 peptide stimulation, in the absence or presence of 10 µg/mL GITRL. In all scatter plots bar height represents mean value and whiskers represent standard deviation. (d) Dot plots obtained in one representative VITAL assay. At the 1:0 target-to-effector (T: E) ratio (far left panel), an even distribution of peptide-loaded (CFSElow) targets and non-loaded targets (CFSEhigh) is observed. From left to right, increasing ratios of effectors CD8TCs (1:1 and 1:5) expanded in the absence or presence of GITRL are shown. (e) Scatter plot with bar graph of pooled data from 5 donors showing percentages of peptide-loaded target (CFSElow) at each condition, where bar height represents the mean value.
GITRL costimulation promotes CD4TC proliferation but it has no effect on the HIV reservoir size
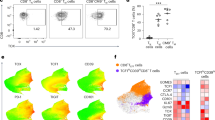
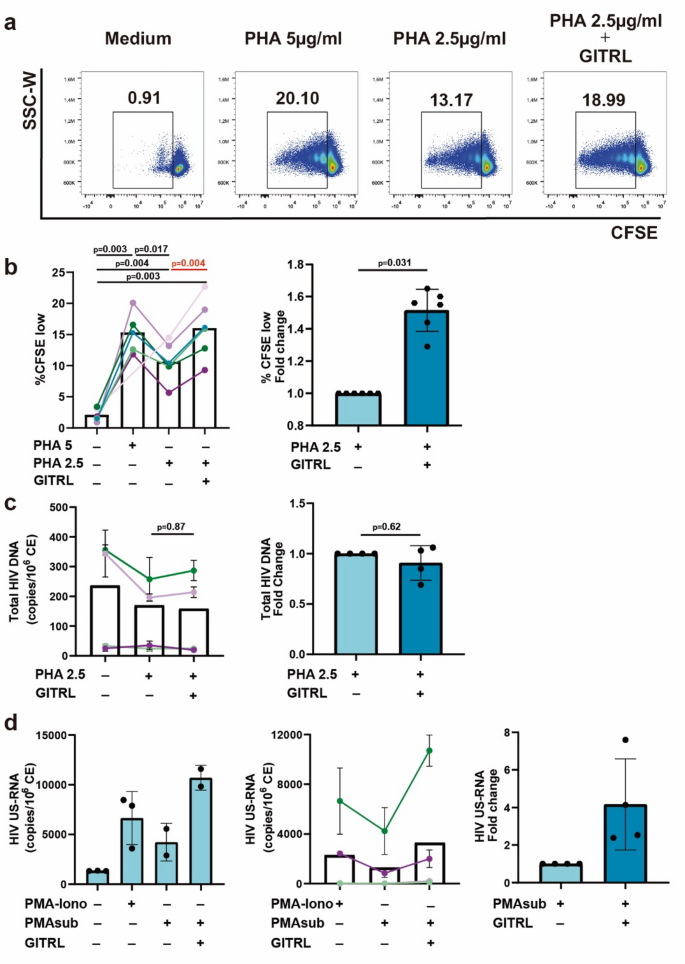
We examined GITRL effect on the proliferation of purified CD4TCs from 6 On-ART donors. After 7 days of co-culture with suboptimal concentrations of PHA plus GITRL (10 µg/mL), a significant increase in proliferating CD4TCs (1.5x) was observed compared to PHA alone (Fig. 5a–b). To evaluate if GITRL costimulation had an impact on the viral reservoir size, we measured cell-associated total HIV DNA across unstimulated, suboptimal PHA and suboptimal PHA + GITRL conditions from 4 donors (Fig. 5c). Both PHA alone and PHA + GITRL conditions showed a non-significant decline in HIV DNA, probably due to replication-mediated death of cells harboring viral DNA or to a higher duplication rate of non-infected cells. Interestingly, we observed no significant differences between conditions with or without GITRL, suggesting that the effect on proliferation is comparable between HIV-infected and non-infected cells and that GITRL costimulation does not impact the size of the HIV reservoir.
GITRL costimulation reverts HIV latency
HIV transcription is dependent on NF-κB activation22. Given GITR signaling relevance in this process23,24 we hypothesized that GITR costimulation could have an effect on HIV latency reversion. To test this, we performed an ex vivo latency reversal assay. Purified CD4TCs from 4 On-ART participants were treated for 16 h with a suboptimal PMA concentration (2ng/mL) plus GITRL (10 µg/mL) or PMA (50ng/mL) and Ionomicyn (1 µg/mL) as a positive control. While PMA alone did not induce the same level of viral transcription observed in the PMA-Ionomycin condition (Fig. 5d), co-stimulation with GITRL led to a 4.16-fold increase in HIV US-RNA, suggesting a role of GITR costimulation in viral reactivation and positioning this pathway as an interesting target for the development of new latency reversal agents.
GITRL-mediated effect on CD4TC proliferation, HIV reservoir size and HIV latency reversion. (a) Representative flow cytometry dot plot showing the frequency of proliferating CD4TCs after 7 days of costimulation with optimal (5 µg/mL) and suboptimal (2.5 µg/mL) concentrations of PHA plus 10 µg/mL of GITRL. Proliferating cells were identified as CFSElow. (b) Pooled data from 6 donors. Percentages of proliferating CD4TCs for each condition (left) and fold change of suboptimal PHA condition plus GITRL (right). Bar height stands for mean values and whiskers represent standard deviation. p-values were calculated using a mixed-effects model for repeated measures with missing values, applying Tukey’s correction for multiple comparisons (left) and one sample t test (right). (c) Total HIV DNA evaluation in proliferating CD4TCs after 7 days of costimulation with suboptimal concentrations of PHA in the absence or presence of GITRL. Left: Total HIV DNA levels were reported as the number of HIV copies per 106 cell equivalents (CE). Each point represents the mean of three experimental replicates. Right: fold change of suboptimal PHA condition plus GITRL. Bar height stands for mean values and whiskers represent standard deviation. p values were calculated with Friedman test followed by Dunn´s post-hoc test (left) or one sample Wilcoxon test (right). (d) Left: scatter plot with bar graph from a representative donor showing the levels of HIV US-RNA after 16 h of costimulation with optimal (PMA = 50 ng/mL + Ionomicyn 1 µg/mL; PMA-iono) and suboptimal (PMA = 2 ng/mL; PMAsub) policlonal stimuli, in the absence or presence of GITRL. Each condition was evaluated in triplicates. Replicates positive for the no reverse transcriptase control condition were excluded. HIV US-RNA levels were reported as the number of RNA copies per 106 cell equivalents (CE). Center: pooled data from 4 donors. Each point represents the mean value of experimental replicates. Right: Fold change of suboptimal PMA condition plus GITRL. Bar height stands for mean values and whiskers represent standard deviation.
Discussion
Enhancement of HIV-specific T-cell response through immunomodulatory agents is a promising approach towards HIV remission7. Although costimulation through the GITR pathway has lately gained attention for this purpose, limited data exists on GITR expression patterns in HIV-specific CD8TCs and CD4TCs from PLWH.
Here, we conducted a comprehensive immunophenotypic analysis of GITR expression among T-cells from PLWH Off- and On-ART. As expected14,15, GITR was significantly upregulated on HIV-specific CD8TCs. It is worth noting that, compared to CEF-specific, HIV-specific CD8TCs are typically dysfunctional. Thus, the lack of a clear difference in GITR expression between these populations suggests that GITR expression levels may not be directly linked to their functional status. In the Off-ART group, GITR upregulation was accompanied by 4-1BB and PD-1 coexpression, while in the On-ART group, upregulation was accompanied only by 4-1BB coexpression, indicating PD-1 downmodulation. While the downmodulation of PD-1 in long-term expansion protocols has already been described21,25,26,27, in our model GITR and 4-1BB expression persisted after 14 days of expansion. This observation suggests that their expression kinetics may differ from those of PD-1, although direct kinetic studies would be required to confirm this.
Preserving the distribution of HIV-specific memory/effector phenotype is crucial for viral control28,29. EM cells accumulate during viremic infection30,31 as a consequence of chronic antigenic exposure32 or, in vitro, after long-term peptide stimulation27,33. Here, we showed that, although the CM phenotype was the most enriched subset in GITR expression on bulk CD8TCs, most HIV-specific GITR-expressing CD8TCs showed a predominance of the EM phenotype, a subset associated with an effective suppression of replication, both in vitro28 and in SIV vaccination models34.
Studies in mouse models have highlighted the role of GITR in enhancing antiviral responses against chronic viral infections35,36,37. Here, we found several associations between GITR expression on EM bulk and HIV-specific CD8TCs with lower viral loads and higher CD4 counts. Positive associations were also observed in the TE subset, a phenotype characterized by potent cytotoxic activity and the ability to inhibit HIV replication in vitro31. To our knowledge this is the first study to report a link between post-stimulation GITR expression on effector memory CD8 + T cells and correlates of HIV control, underscoring the potential of this receptor as a target for HIV immunotherapy. Moreover, the association between higher %EM and lower viral loads in the Off-ART group, together with the observation that most GITR-expressing HIV-specific CD8 T cells exhibited this phenotype after expansion, supports the rationale for GITR-based immunomodulation of ex vivo–expanded CD8 T cells. For instance, strategies involving adoptive cell transfer therapy7 could be a suitable scenario to evaluate the potential of GITR stimulation as a therapeutic approach.
Regarding regulatory CD4TCs, most GITR + Treg corresponded to the EM subset, known for exhibiting greater in vitro activity compared to CM or naïve Tregs38. The role of Tregs in HIV infection remains, however, debated39,40 While we found no direct association between Treg percentages and clinical parameters, our data suggest a possible role of GITR expression in this subset, supported by the negative correlation between VL and GITR expression in bulk, CM, and EM Tregs. However, given the correlative nature of our analysis, it is also plausible that viremia influences GITR expression on Tregs; therefore, directionality cannot be determined from the present data.
This phenotypical analysis yielded that GITR could have a role in HIV infection. However, the differential GITR expression across cellular subsets, their unbalanced contribution to HIV control and the fact that GITR is preferentially expressed on susceptible cells underscore the complexity of these interactions. To explore this further, we conducted assays to evaluate the potential of a hexameric GITR ligand to enhance CD8TC antiviral function and to reverse viral latency. This hexameric fusion protein holds the promise to overcome limitations of other monomeric ligands that have already failed in clinical trials20.
Remarkably, GITRL enhanced IFN-γ production and degranulation of HIV-specific CD8TCs after 4 days of costimulation. In contrast, Lahey et al.41 found an increased cytokine production only among CD4TCs but not in CD8TCs from PLWH Off-ART. This discrepant result may be explained by different stimulation conditions since these authors performed an overnight GITR pre-stimulation without TCR triggering. In conditions resembling our study but using a different GITR agonist, Gubser et al.15 observed a significant increase in the percentage of IFN-γ + CD8TCs.
Although GITRL improved HIV-specific CD8TC functionality in the short term, a 14-day expansion with GITRL showed no increase in CD8TC cytotoxicity in the VITAL assay. This aligns partially with findings by Gubser et al.15, who hypothesize that CD8TC functionality may decline due to a decrease in cytokine-producing EM cells and the expansion of a dysfunctional PD-1 + TIM + GITR + population. Moreover, the absence of a discernible effect of GITRL on HIV-specific CD8TC proliferation may account for the overall lack of functional enhancement, although it remains possible that the assay employed here did not possess sufficient sensitivity to detect such an effect. Further studies on CD8TC antiviral responses following GITR costimulation should include multiple rounds of costimulation or shorter time points, along with additional checkpoint modulation, particularly PD-126,42.
Within the CD4TC compartment, GITR expression was enriched in CM and EM subsets in both Off- and On-ART PLWH, with most GITR-expressing CD4TCs harboring an EM phenotype. This is meaningful, since CM cells contribute to the viral reservoir due to a higher infection rate, enhanced self-renewal and greater proliferation capacity43, while EM cells exhibit the highest frequencies of intact and inducible proviruses44,45,46. Since GITR signaling triggers NF-κB translocation47 and activation16,48—a key factor in regulating HIV transcription—we hypothesized that GITRL may contribute to latency reversion.
We demonstrated for the first time that GITRL costimulation induced viral reactivation, at least at the transcriptional level. It has been widely reported that the vast majority of the proviral composition corresponds to defective proviruses, containing internal deletions, deleterious stop codons, hypermutations, among others49. Thus, further studies should investigate the nature of these transcripts through transcriptional profiling, full-length sequencing or transcriptomic analysis. Moreover, at the proteomic level, measuring and characterizing viral proteins by ultrasensitive p24 assays or quantitative viral outgrowth assay should be performed.
As expected, we observed a clear effect of GITRL costimulation on CD4TC proliferation. Gubser et al. have correctly pointed out that this outcome could potentially enhance persistence of the HIV reservoir through clonal expansion on CD4TCs. However, similar to results obtained in other models of PHA-mediated expansion50, our quantification of total HIV DNA showed no differences in the relative size of the viral reservoir after GITRL costimulation, likely due to the balanced proliferation of CD4TCs, regardless of whether these cells harbor viral DNA.
Limitations of this work included: (i) The immune phenotyping of Off- and On-ART groups were not comparable due to different stimulation times. (ii) HIV-specific cells were defined using CD25+/Ki67+ coexpression instead of the more reliable HIV-tetramer protocol. (iii) Cytokine/degranulation production after GITRL costimulation was assessed at a different temporality than the VITAL assay, which may explain the divergence in experimental outputs. (iv) The need to use a prolonged stimulation protocol for T-cell expansion in On-ART samples. Although this approach is a useful tool to detect HIV-specific responses post-ART, it may also alter cell phenotype (skewing it toward an effector status) and function. (v) The relatively small number of biological replicates in the latency reversion experiment. Although the fact that all individuals analyzed exhibited the same trend provides additional confidence in the result, conclusions should still be interpreted with caution given the limited sample size. Moreover, latency reversal experiment only included the quantification of HIV transcripts, excluding viral protein and particle production.
In summary, we performed an extensive characterization of GITR-expressing T-cells of PLWH Off- and On-ART, showing several correlations between GITR expression and HIV control. Moreover, we proved the correct functionality of a novel GITRL that exerted a limited enhancement of CD8TC antiviral response. Interestingly, we observed viral reactivation at a transcriptional level from latently infected CD4TCs after GITRL costimulation, without increasing the total reservoir size, positioning GITR as an interesting target for the development of novel LRAs, although in the context of the shock-and-kill strategy, this should be accompanied by more potent enhancers of the antiviral response. Since GITR stimulation requires concomitant TCR signaling, and global T cell activators such as PHA would not be safe in a clinical context, these results support the rationale for exploring GITRL in combination with clinically tolerated LRAs or within therapeutic vaccination strategies for HIV latency reversal.
Data availability
The datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
References
Bekker, L-G. et al. HIV infection. Nat. Rev. Dis. Primers. 9, 42. https://doi.org/10.1038/s41572-023-00452-3 (2023).
McMyn, N. F. et al. The latent reservoir of inducible, infectious HIV-1 does not decrease despite decades of antiretroviral therapy. J. Clin. Invest. 133 https://doi.org/10.1172/JCI171554 (2023).
Hütter, G. et al. Long-term control of HIV by CCR5 Delta32/Delta32 stem-cell transplantation. N Engl. J. Med. 360, 692–698. https://doi.org/10.1056/NEJMoa0802905 (2009).
Jensen, B-E-O. et al. In-depth virological and immunological characterization of HIV-1 cure after CCR5∆32/∆32 allogeneic hematopoietic stem cell transplantation. Nat. Med. 29, 583–587. https://doi.org/10.1038/s41591-023-02213-x (2023).
Turk, G. et al. A possible sterilizing cure of HIV-1 infection without stem cell transplantation. Ann. Intern. Med. 175, 95–100. https://doi.org/10.7326/L21-0297 (2022).
Sáez-Cirión, A. et al. Sustained HIV remission after allogeneic hematopoietic stem cell transplantation with wild-type CCR5 donor cells. Nat. Med. 1–11. https://doi.org/10.1038/s41591-024-03277-z (2024).
Deeks, S. G. et al. Research priorities for an HIV cure: International AIDS society global scientific strategy 2021. Nat. Med. 27, 2085–2098. https://doi.org/10.1038/s41591-021-01590-5 (2021).
Kim, Y., Anderson, J. L. & Lewin, S. R. Getting the kill into shock and kill: Strategies to eliminate latent HIV. Cell. Host Microbe 23, 14–26. https://doi.org/10.1016/j.chom.2017.12.004 (2018).
Attanasio, J. & Wherry, E. J. Costimulatory and coinhibitory receptor pathways in infectious disease. Immunity 44, 1052–1068. https://doi.org/10.1016/j.immuni.2016.04.022 (2016).
Rutishauser, R. L. & Trautmann, L. CD8 + T-cell responses in HIV controllers: Potential implications for novel HIV remission strategies. Curr. Opin. HIV AIDS 17, 315–324. https://doi.org/10.1097/COH.0000000000000748 (2022).
Fidler, S. et al. Antiretroviral therapy alone versus antiretroviral therapy with a kick and kill approach, on measures of the HIV reservoir in participants with recent HIV infection (the RIVER trial): A phase 2, randomised trial. Lancet 395, 888–898. https://doi.org/10.1016/S0140-6736(19)32990-3 (2020).
Curty, G., Iniguez, L. P., Soares, M. A., Nixon, D. F. & de Mulder Rougvie, M. Off-target effect of activation of NF-κB by HIV latency reversal agents on transposable elements expression. Viruses 14, 1571. https://doi.org/10.3390/v14071571 (2022).
Wang, F. et al. Structures of mouse and human GITR-GITRL complexes reveal unique TNF superfamily interactions. Nat. Commun. 12, 1378. https://doi.org/10.1038/s41467-021-21563-z (2021).
Nocentini, G., Ronchetti, S., Cuzzocrea, S. & Riccardi, C. GITR/GITRL: More than an effector T cell co-stimulatory system. Eur. J. Immunol. 37, 1165–1169. https://doi.org/10.1002/eji.200636933 (2007).
Gubser, C. et al. GITR activation ex vivo impairs CD8 T cell function in people with HIV on antiretroviral therapy. iScience 26, 108165. https://doi.org/10.1016/j.isci.2023.108165 (2023).
Nocentini, G., Cari, L. & Riccardi, C. GITR: A modulator of regulatory and effector T-cell activity, crucial in tumour rejection and autoimmune diseases. 1–11 (Wiley, 2017). https://doi.org/10.1002/9780470015902.a0026242.
van Olffen, R. W. et al. GITR triggering induces expansion of both effector and regulatory CD4 + T cells in vivo. J. Immunol. 182, 7490–7500. https://doi.org/10.4049/jimmunol.0802751 (2009).
Liao, G. et al. GITR engagement preferentially enhances proliferation of functionally competent CD4 + CD25 + FoxP3 + regulatory T cells. Int. Immunol. 22, 259–270. https://doi.org/10.1093/intimm/dxq001 (2010).
Wyzgol, A. et al. Trimer stabilization, oligomerization, and antibody-mediated cell surface immobilization improve the activity of soluble trimers of CD27L, CD40L, 41BBL, and glucocorticoid-induced TNF receptor ligand. J. Immunol. 183, 1851–1861. https://doi.org/10.4049/jimmunol.0802597 (2009).
Davar, D. & Zappasodi, R. Targeting GITR in cancer immunotherapy—There is no perfect knowledge. Oncotarget 14, 614–621. https://doi.org/10.18632/oncotarget.28461 (2023).
Salido, J. et al. Phenotype, polyfunctionality, and antiviral activity of in vitro stimulated CD8 + T-cells from HIV + subjects who initiated cART at different time-points after acute infection. Front. Immunol. 9, 2443. https://doi.org/10.3389/fimmu.2018.02443 (2018).
Nabel, G. & Baltimore, D. An inducible transcription factor activates expression of human immunodeficiency virus in T cells. Nature 326, 711–713. https://doi.org/10.1038/326711a0 (1987).
Nocentini, G. et al. A new member of the tumor necrosis factor/nerve growth factor receptor family inhibits T cell receptor-induced apoptosis. Proc. Natl. Acad. Sci. U S A. 94, 6216–6221. https://doi.org/10.1073/pnas.94.12.6216 (1997).
Daniels, M. A., Luera, D. & Teixeiro, E. NFκB signaling in T cell memory. Front. Immunol. 14, 1129191. https://doi.org/10.3389/fimmu.2023.1129191 (2023).
Day, C. L. et al. PD-1 expression on HIV-specific T cells is associated with T-cell exhaustion and disease progression. Nature 443, 350–354. https://doi.org/10.1038/nature05115 (2006).
Trautmann, L. et al. Upregulation of PD-1 expression on HIV-specific CD8 + T cells leads to reversible immune dysfunction. Nat. Med. 12, 1198–1202. https://doi.org/10.1038/nm1482 (2006).
SalidoJP et al. Early skewed differentiation and PD-1 expression in CD4 + cells relate to immune dysfunction and viral persistence in individuals living with HIV 1 year post-cART initiation. J. Virus Eradication https://doi.org/10.1016/S2055-6640(20)31026-8 (2019).
Buckheit, R. W. 3, Salgado, M., Silciano, R. F., Blankson, J. N. & rd,, Inhibitory potential of subpopulations of CD8 + T cells in HIV-1-infected elite suppressors. J. Virol. 86, 13679–13688. https://doi.org/10.1128/JVI.02439-12 (2012).
Takata, H. et al. Delayed differentiation of potent effector CD8 + T cells reducing viremia and reservoir seeding in acute HIV infection. Sci. Transl. Med. 9 https://doi.org/10.1126/scitranslmed.aag1809 (2017).
Champagne, P. et al. Skewed maturation of memory HIV-specific CD8 T lymphocytes. Nature 410, 106–111. https://doi.org/10.1038/35065118 (2001).
Ghiglione, Y. et al. Early skewed distribution of total and HIV-specific CD8 + T-cell memory phenotypes during primary HIV infection is related to reduced antiviral activity and faster disease progression. PLoS One 9, e104235. https://doi.org/10.1371/journal.pone.0104235 (2014).
Masopust, D. & Picker, L. J. Hidden memories: Frontline memory T cells and early pathogen interception. J. Immunol. 188, 5811–5817. https://doi.org/10.4049/jimmunol.1102695 (2012).
Salido, J. et al. Pre-cART immune parameters in people living with HIV might help predict CD8 + T-cell characteristics, inflammation levels, and reservoir composition after effective cART. Pathog. Immun. 6, 60–89. https://doi.org/10.20411/pai.v6i2.447 (2021).
Hansen, S. G. et al. Profound early control of highly pathogenic SIV by an effector memory T-cell vaccine. Nature 473, 523–527. https://doi.org/10.1038/nature10003 (2011).
Snell, L. M., Lin, G. H. Y., McPherson, A. J., Moraes, T. J. & Watts, T. H. T-cell intrinsic effects of GITR and 4-1BB during viral infection and cancer immunotherapy: Effects of GITR and 4-1BB on T cells. Immunol. Rev. 244, 197–217. https://doi.org/10.1111/j.1600-065X.2011.01063.x (2011).
Clouthier, D. L., Zhou, A. C. & Watts, T. H. Anti-GITR agonist therapy intrinsically enhances CD8 T cell responses to chronic lymphocytic choriomeningitis virus (LCMV), thereby circumventing LCMV-induced downregulation of costimulatory GITR ligand on APC. J. Immunol. 193, 5033–5043. https://doi.org/10.4049/jimmunol.1401002 (2014).
Pascutti, M. F. et al. Enhanced CD8 T cell responses through GITR-mediated costimulation resolve chronic viral infection. PLoS Pathog. 11, e1004675. https://doi.org/10.1371/journal.ppat.1004675 (2015).
Mailloux, A. W. et al. Expansion of effector memory regulatory T cells represents a novel prognostic factor in lower risk myelodysplastic syndrome. J. Immunol. 189, 3198–3208. https://doi.org/10.4049/jimmunol.1200602 (2012).
Chevalier, M. F. & Weiss, L. The split personality of regulatory T cells in HIV infection. Blood 121, 29–37. https://doi.org/10.1182/blood-2012-07-409755 (2013).
Jiao, Y. et al. The decrease of regulatory T cells correlates with excessive activation and apoptosis of CD8 + T cells in HIV-1-infected typical progressors, but not in long-term non-progressors. Immunology 128, e366–e375. https://doi.org/10.1111/j.1365-2567.2008.02978.x (2009).
Lahey, T. P., Loisel, S. D. & Wieland-Alter, W. Glucocorticoid-induced tumor necrosis factor receptor family-related protein triggering enhances HIV-specific CD4 + T cell cytokine secretion and protects HIV-specific CD4 + T cells from apoptosis. J. Infect. Dis. 196, 43–49. https://doi.org/10.1086/518613 (2007).
Mylvaganam, G. H. et al. Combination anti-PD-1 and antiretroviral therapy provides therapeutic benefit against SIV. JCI Insight 3 https://doi.org/10.1172/jci.insight.122940 (2018).
Chomont, N. et al. HIV reservoir size and persistence are driven by T cell survival and homeostatic proliferation. Nat. Med. 15, 893–900. https://doi.org/10.1038/nm.1972 (2009).
Fromentin, R. & Chomont, N. HIV persistence in subsets of CD4 + T cells: 50 shades of reservoirs. Semin Immunol. 51, 101438. https://doi.org/10.1016/j.smim.2020.101438 (2021).
Duette, G. et al. The HIV-1 proviral landscape reveals that Nef contributes to HIV-1 persistence in effector memory CD4 + T cells. J. Clin. Invest. 132 https://doi.org/10.1172/JCI154422 (2022).
Hiener, B. et al. Identification of genetically intact HIV-1 proviruses in specific CD4 + T cells from effectively treated participants. Cell. Rep. 21, 813–822. https://doi.org/10.1016/j.celrep.2017.09.081 (2017).
Ronchetti, S. et al. Glucocorticoid-induced TNFR-related protein lowers the threshold of CD28 costimulation in CD8 + T cells. J. Immunol. 179, 5916–5926. https://doi.org/10.4049/jimmunol.179.9.5916 (2007).
Bae, E. M. et al. Reverse signaling initiated from GITRL induces NF-kappaB activation through ERK in the inflammatory activation of macrophages. Mol. Immunol. 45, 523–533. https://doi.org/10.1016/j.molimm.2007.05.013 (2008).
Bruner, K. M., Hosmane, N. N. & Siliciano, R. F. Towards an HIV-1 cure: Measuring the latent reservoir. Trends Microbiol. 23, 192–203. https://doi.org/10.1016/j.tim.2015.01.013 (2015).
Takata, H. et al. Modeling HIV-1 latency using primary CD4 + T cells from virally suppressed HIV-1-infected individuals on antiretroviral therapy. J. Virol. 93. https://doi.org/10.1128/JVI.02248-18 (2019).
Acknowledgements
We acknowledge Dra. Virginia González Polo for flow cytometry technical assistance, Dr. H. Wajant for the provision of GITRL plasmid, the INBIRS staff who contributed in sample collection and especially all participants who made this study possible.
Funding
This work was supported by grants from the Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT, Grant # PICT2019-00504) and from the University of Buenos Aires (Grant UBACYT2020/20020190100194BA). The funders had no role in the study design, data collection and interpretation, or the decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
Alejandro Czernikier: Investigation, Formal analysis, Writing (original draft, review & editing), Visualization, Conceptualization, Methodology. Lucia Baquero: Investigation and Writing (review & editing). Paula Benencio, Laura Alejandra Osorio Ocampo: Investigation. Jimena Salido: Investigation, Methodology and Writing (review & editing). Ana Gun: Resources. María Ines Figueroa: Resources. Natalia Laufer: Resources. Yanina Ghiglione: Investigation, Writing (review & editing). María Fernanda Pascutti: Supervision, Writing (review & editing), Methodology, Conceptualization. Gabriela Turk: Supervision, Project administration, Funding acquisition, Resources, Writing (review & editing), Methodology, Conceptualization.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Generative AI and AI-assisted technologies in the writing process
During the preparation of this work the authors used ChatGPT in order to improve the readability and language of the manuscript as none of the authors are native English speakers. After using this tool, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Czernikier, A., Baquero, L., Benencio, P. et al. GITR triggering has limited impact on HIV-specific CD8 T-cell function but enhances HIV transcription reactivation from latently infected CD4 T-cells. Sci Rep 15, 43002 (2025). https://doi.org/10.1038/s41598-025-26952-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26952-8