Abstract
Synchronization phenomenon characterize the collective behavior of complex systems. In recent years, data-driven approaches have become relevant because they can identify and quantify synchronized activity. Here, we study networks of mutually coupled Kuramoto phase oscillators adopting the normalized persistent entropy, NPE, based on persistent homology, as a synchronization quantifier, focusing our analysis on zero-dimensional homology groups of closed and open triads. We show that, as the coupling intensity increases, the NPE identifies transitions from incoherent to synchronized states in networks with different structural connectivity. We also use the NPE to analyze experimental data of mutually coupled electronic oscillators with Rössler-type dynamics, and identify transitions toward synchronization as the coupling intensity increases. Our results show that the NPE is a practical tool to quantitatively identify changes in the dynamics of networks of coupled oscillators.
Similar content being viewed by others
Introduction
Synchronization is a phenomenon that arises when two or more systems evolve in a coordinated way, and occurs in a wide range of natural and artificial (or “man-made”) systems1,2. In natural systems, controlled experiments are rarely feasible because usually it is not possible to precisely control the mechanisms that are ultimately responsible for coordinated activity. Therefore, the usual approach to studying synchronization is resorting to simulated or experimental systems3,4,5, where the coupling parameter that determines the intensity of interaction or the exchange of information between subsystems can be varied in a controlled way.
One of the most remarkable works in this direction was done in 1975 by Yoshiki Kuramoto6, who considered a population of mutually interacting phase oscillators, with a coupling parameter controlling the strength of the interactions, whereby transition from independent (incoherent) to synchronized state of the oscillators phases at a given coupling value occurs, whose practicality and adaptability has made possible to study the synchronization phenomenon in different fields, including physics3,7, biology8,9,10,11, neuroscience12,13,14,15, engineering16,17,18, economics19,20,21, among many others.
A way to study a system composed by many interacting elements (a complex system), as considered by Kuramoto, is representing it as a complex network22,23,24,25,26,27, a graph theory based strategy, where the vertices represent the subsystems (nodes) and the links (edges) symbolize functional or structural relationships between them, providing a mathematical framework to analyze the collective behavior of non-linear dynamics complex systems. In a network, the degree distribution configures the information flow between the nodes. For example, random networks are known to have a characteristic connectivity scale, which means that most nodes have a similar number of links. A representative model of these kind of networks was introduced in 1959 by Paul Erdös and Alfréd Rényi by randomly linking the nodes of a network with a certain connection probability28. When the network is large enough, it adopts a Poisson-type degree distribution, well represented by an average connectivity. This model allowed to advance the understanding of some complex systems29,30; however, many complex systems are often represented by networks whose degree distributions do not have a characteristic scale, in which a large number of nodes have only one or very few links, while very few nodes, called hubs, have many links31,32. These networks, where the degree distribution follows a power law, are known as Scale-Free24,33,34.
Another popular approach to the study of complex systems from the perspective of non-linear dynamics is based on the use of measures of entropy and complexity35,36,37,38. This approach has proven to be particularly useful for univariate time series analysis39,40,41,42,43,44,45. Extensions based on mutual information and transfer entropy46 allow bivariate time series analysis and the exploration of synchronization phenomena in terms of pairwise interactions47,48,49,50,51.
A data-driven approach that overcomes this limitation is persistent homology52,53,54,55,56, a topological data analysis tool57,58,59 that represents complex, high-dimensional data (such as one, two or several time series) as a cloud of points in an appropriate space to construct a simplicial complex. Persistent homology infers the topological properties of a dataset by filtering it at multiple scales and such an advantage has been applied in many fields60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76. The simplicial complex is the collection of simplexes (topological invariants) formed in the filtration of the cloud of points, generating homology groups (topological characteristics). Each homology group consists of many elements, which are referred to as “classes” (the number of classes in a homology group is its Betti number), that are generated (and extinguished) during the filtration. The persistence diagram of such a homology group are the birth-and-death coordinates of the classes. Thus, the persistence diagram contains the topological properties of the dataset under study and consequently of the underlying system. When the state of the system that generates the data changes, the point cloud changes, and the persistence diagrams will also change. Thus, by tracking changes in persistence diagrams, it is possible to identify different states and their transitions in the system under study77,78.
Persistent homology’s ability to handle high-dimensional data allows us to use it to simultaneously analyze the activity of multiple network nodes. The characterization of the transition from incoherent to synchronized states is typically done in terms of global quantifiers, such as Kuramoto order parameter, that involve network-wide information3,4,6. However, local structures, such as closed triads (a set of three nodes \(\{v_1,v_2,v_3\}\) connected by three edges, \(\{v_1,v_2\}, \{v_1,v_3\}, \{v_2,v_3\}\)) can be sensitive to variations in the network state and can provide useful information to detect such changes.
In this work, we leverage the power of persistent homology to analyze high-dimensional data, bridging the gap between studies that consider multiple interactions between pairs of nodes and studies that consider global quantifiers of the entire network. Our goal is to identify synchronization transitions by studying local connectivity structures as a benchmark to characterize the collective behavior of systems whose dynamics can be modeled by oscillator networks.
We focus the study on Kuramoto oscillators6 mutually coupled in networks with Erdös-Rényi (ER), Scale-Free (SF) and Random (RN) topologies, also applying our approach to an experimental dataset (EN) of 28 mutually coupled electronic circuits with Rössler-type dynamics arranged in a network configuration79. We analyze the transition from incoherent to synchronized states in the network (system) through the temporal evolution of closed triads (“triangles”) and sets of three disconnected nodes (“triplets”), as a function of the coupling intensity. While a triangle is a set of three nodes connected by three edges, a “triplet” is a set of three nodes that are not connected by any edge.
Specifically, we analyze, for the networks of Kuramoto oscillators, the temporal evolution (activity) expressed as the cosine \(\cos (\theta _i(t))\) of the phases \(\theta _i(t)\) of oscillators triads [\(\cos (\theta _i(t))\), \(\cos (\theta _j(t))\), \(\cos (\theta _k(t))\)]. In the experimental network case, we analyzed the voltages measured in three circuits, \(x_i(t)\), \(x_j(t)\), and \(x_k(t)\). We represent the evolution of a triangle or a triplet as three-dimensional (3D) point cloud defined by \(\cos (\theta _i(t))\), \(\cos (\theta _j(t))\), and \(\cos (\theta _k(t))\), for the Kuramotto oscillators, and by \(x_i(t)\), \(x_j(t)\), and \(x_k(t)\), for the electronic circuits. We detect topological changes in the 3D cloud point as the coupling strength increases using the normalized persistent entropy (NPE)78,80. NPE quantifies the diversity of the homology classes persistencies in a homology group (see Methods). NPE is based on the persistent entropy introduced by Rucco et al. in 2014, an extension of Shannon entropy in the persistent homology context81.
In our previous work78 we used NPE to analyze the behavior of the voltages of 28 electronic circuits arranged as a network of coupled oscillators with a Rössler-type chaotic dynamics79, and also, of 28 Kuramoto phase oscillators mutually coupled in a network configuration with identical connectivity to the network of experimental electronic oscillators. We represented the evolution of pairs of nodes (for the electronic oscillators, \(x_i(t)\) and \(x_j(t)\), and for the Kuramoto oscillators, \(\cos (\theta _i(t))\) and \(\cos (\theta _j(t))\)), as two-dimensional (2D) point clouds. We found that, in both cases, NPE was sensitive to changes in the point clouds due to changes in the network’s coupling strength. We also found that NPE was sensitive to the nodes’ connectivity degrees and the distance between pairs of nodes.
As a natural continuation of our previous work, here we focus the analysis on 3D point clouds that represent the temporal evolution of triangles and triples. We use the zero homology group –that corresponds to the connected components (\(H_0\))- because in78 we found that both, \(H_0\) and \(H_1\) provide usable information for these data, varying similarly with coupling strength, but \(H_1\) shows a greater degree of variability. In78 we performed the filtration for a maximum edge length, \(\epsilon\), fixed, generating only one simplicial complex filtrated up to such a maximum edge length. However, keeping the maximum edge length fixed can mask interesting geometric properties of the data at small scales because the resulting simplicial complex encloses the properties at all the scales. Therefore, here we analyze how NPE varies as a function of the maximum edge length, which is adaptively chosen to guarantee that no point is a connected component itself (i.e., when the filtration parameter, \(\epsilon\), reaches its maximum value, we have just one connected component in the simplicial complex).
We find, for the Kuramoto oscillators and also for the electronic circuits, that the variation of NPE with \(\epsilon\), calculated from 3D point clouds, that represent the temporal evolution of triangles and triplets, reveals changes in the dynamics of the network as the coupling increases. Therefore, our results suggest that NPE can be a useful analysis tool in fields where it is important to identify and quantify synchronization transitions.
Results
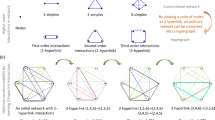
Figure 1 displays a representation of the four networks studied [panels (a)-(d)], and illustrates the procedure [panel (e)] used to calculate the normalized persistent entropy, NPE, of a set of three nodes. For the analysis of the Kuramoto oscillators, we first generate networks using algorithms that return networks with Erdös-Rényi [ER,Fig. 1(a)], scale-free [SF, Fig. 1(b)], and random topology [RN, Fig. 1(c)], respectively. The model, algorithms and parameters are described in Methods. As well as the experimental network [EN, Fig. 1(d)] reproduced by79, consisting of electronic circuits with Rössler-type dynamics in chaotic state. In the four networks, the links are undirected and the coupling intensity (\(\lambda\) for the Kuramoto oscillators and \(\kappa\) for the electronic oscillators) is uniform. The evolution of the ith Kuramoto oscillator is described by an observable whose time series is \(\cos (\theta _i(t))\), while the evolution of the ith electronic circuit is described by the measured voltage, \(x_i(t)\).
Graphical representation of the networks used in this work: (a) Erdös-Rényi (ER, 30 nodes and 48 links), (b) scale-free (SF, 30 nodes and 56 links), (c) random network (RN, 30 nodes and 40 links), and (d) experimental network (EN, 28 nodes and 42 links, colored nodes and edges illustrate a triangle in the network). (e) Illustration of the procedure used to calculate the normalized persistent entropy of zero homology groups, \(NPE(H_0)\), for a point cloud defined by three time series associated to three nodes (oscillators), \(x_1\), \(x_2\), and \(x_3\). The filtration procedure is applied by increasing the maximum edge length, \(\epsilon _i\), to obtain the barcodes (the difference death-born times of each homology class) of the resulting simplicial complex. For each \(\epsilon _i\), the \(NPE_{\epsilon _i}(H_0)\) is calculated with Eq. (2) (see Methods).
We characterized the transition from an incoherent to a synchronized state in the four networks shown in Fig. 1(a)-(d) as follows: for Kuramoto’s dynamics (networks shown in Fig. 1(a)-(c)), we constructed 3D point clouds (top of Fig. 1(e)) using simulated time series of three nodes that are either adjacent (triangles) or non-adjacent (triplets). Specifically, the time series used to construct 3D point clouds consist of the simulated temporal evolution of the cosine of the phases of the nodes. Each time series was normalized to have a zero mean and a unitary standard deviation. Then, we performed the filtration (as described in Methods) for several maximum edge lengths \(\epsilon _i\), as illustrated in bottom of the panel 1(e), computing the NPE of the \(H_0\) homology group, at each \(\epsilon _i\). This procedure that allows us to obtain information of the point cloud at different scales, by analyzing how NPE varies as a function of the maximum edge length. For each network, to analyze different synchronization states (from incoherent to synchronized), we repeated this procedure for different coupling values, performing, for each coupling value, 10 simulations with different initial conditions, but keeping the network structure constant. In the experiments79, the network structure was also constant, and only the coupling strength was varied. We also preprocessed the experimental data by normalizing each time series (the temporal evolution of the voltage in each electronic circuit) to zero mean and unit standard deviation.
First, we study the transition to synchronization in the three networks of Kuramoto oscillators, in terms of the variation of NPE with the coupling strength, setting the filtration parameter \(\epsilon =0.15\) as the maximum edge length because at this value we ensure that the simplicial complex is a single connected component. We calculate NPE for all closed triads (triangles) present in the networks, and for the same number of open triads (triplets). We also calculate the order parameter R, which is the standard metric to quantify synchronization in Kuramoto networks, described in Methods.
The calculation of R is performed for triangles and triplets, with the data averaged over all triangles in the network and a matching number of triplets. We also calculate R for the entire network. The results are presented in Fig. 2(a)-(c), where we see how R changes as the coupling strength \(\lambda\) increases for the ER, SF and RN networks. In the three networks, for both, triangles and triplets, R begins at an intermediate value and subsequently increases to a value approximating 1 for sufficiently large coupling. If the coupling strength is not too large, as shown in the insets, the value of R calculated from triangles is usually higher than the value of R calculated from triplets, and differences are more pronounced in the RN network.
In contrast to a gradual increase of R, the value of NPE calculated from triangles and triplets in ER and SF networks shows a steep increase as \(\lambda\) increases and reaches a comparable saturation value for large \(\lambda\). As seen in Fig. 2(f), in the RN network NPE grows gradually. While in ER and SF networks almost no difference is observed between the NPE of triangles and triples, in the RN network, NPE grows slower for triplets than for triangles. Therefore, NPE analysis of local structures (triangles and triples) uncovers new information: the existence of a steep transition in the ER and SF networks that is not detected by the usual quantifier, the Kuramoto order parameter R. Furthermore, the differences in average NPE values observed in Figs. 2(d)-(f) may allow, for certain coupling strengths, to differentiate triangles from triples in a RN network (since triples tend to have lower average values than triangles) and may also allow to differentiate triangles and triples in an RN network from those in SF and ER networks (as for coupling strengths just above the steep transition, the average NPE values of triangles and triples in the SF and ER networks are higher than in the RN network). In the case of the RN, the NPE exhibits a smooth transition, likely due to limited connectivity. This coincides with previous reports of a slow transition toward synchronization in Watts-Strogatz networks82, which interpolate between regular and random networks. It is interesting to note that, for the SF network, an oscillatory behavior of NPE is observed when the coupling is not too strong [inset in Fig. 2(e)]. We speculate that this feature may be due to multistability or an unstable synchronized state; however, further studies are needed to clarify the origin of the oscillatory behavior.
Order parameter R (top row) and NPE (bottom row) vs. the coupling strength, \(\lambda\), for triangles (solid symbols) and triplets (empty symbols) of networks of Kuramoto oscillators: (a, d) ER, (b, e) SF and (c, f) RN. The error bars represent the standard deviation computed over 10 triangles and triplets, and over 10 simulations of each network. In (a), (b), and (c), the black solid circles represent the value of R computed over all the nodes of the network. The insets show in detail the variation of R and NPE for low coupling strengths.
Next, we compare the variation of NPE in the three Kuramoto networks with the variation of NPE in the experimental network (EN). The results are presented in Fig. 3, which displays NPE calculated from the triangles, as a function of the coupling strength. We consider different values of \(\epsilon\), shown in color code, with the darkest green representing the value used in Fig. 2. For the four networks, as \(\epsilon\) decreases (from green to red) NPE increases, and this occurs for all coupling strengths considered. If the coupling strength is sufficiently large, in the experimental network NPE remains nearly constant, for all \(\epsilon\)s. In the Kuramoto networks, for large \(\epsilon\) (green lines) NPE remains nearly constant (ER and SF networks) or grows gradually (RN network) but, for small values of \(\epsilon\) (red lines), NPE displays some fluctuations, even when the coupling is large. These fluctuations capture the variability of geometric information of the point cloud at small scales, which is significantly less pronounced in the point clouds defined by the experimental data. We speculate that this difference is due to the fact that the experimental detection system filters out small voltage fluctuations.
NPE of triangles vs. coupling strength (\(\lambda\) or \(\kappa\) for generated and experimental data, respectively) for several values of the filtration parameter, \(\epsilon\) (in color code), for the ER (a), SF (b), RN (c), and EN (d) networks. The symbols represent the average NPE, while the error bars represent the standard deviation. For the simulated these values were calculated over 10 triangles and performing 10 simulations of each network. For the experimental data the computations are performed over 5 triangles, because the network only contains 5 triangles.
In Fig. 3(a) and (b) we also note that NPE shows a sharp variation at \(\lambda \approx 0.4\) that is consistent with the variation seen in Fig. 2(d) and (e). The variation of NPE when \(\lambda\) increases above \(\approx 0.4\) depends on \(\epsilon\): for the smallest values of \(\epsilon\) (red lines), NPE abruptly decreases, while, for the largest values of \(\epsilon\) (green lines), it abruptly increases. The change in NPE is robust because it occurs for all \(\epsilon\)s, but is more pronounced for the smallest and largest values. In contrast, for the RN and EN networks, as seen in Fig. 3(c) and (d), the variation of NPE is gradual.
The results that were presented in Fig. 3 are displayed in Fig. 4 in an alternative way, by plotting the value of NPE as a function of \(\epsilon\) and representing the coupling strength in the color code. Here red colors indicate low coupling and blue colors high coupling. Displaying NPE as a function of \(\epsilon\) reveals that, for all networks, the minimum value of \(\epsilon\) that we can use (such that no point is a connected component itself) decreases with the coupling (note that blue lines start at \(\epsilon =10^{-3}\)). Also, it can be seen in Fig. 4 that the way NPE varies regarding \(\epsilon\) changes with the coupling strength. For very low coupling (dark red curves), NPE decreases with \(\epsilon\). This is seen in the four networks and is interpreted as due to the fact that the nodes’ dynamics (and therefore, the topology of the point clouds), is independent of the network topology. For larger coupling (orange and light blue curves), differences in the variation of NPE with \(\epsilon\) can be observed for the different networks. Differences are seen even for the strongest coupling values (dark blue curves) than render the oscillators’ dynamics highly synchronized.
NPE of triangles vs. filtration parameter (the maximum edge length used to construct the simplicial complex), for several values of the coupling strength (in color code, \(\lambda\) or \(\kappa\) for simulated and experimental data, respectively) for the networks: (a) ER, (b) SF, (c) RN, (d) EN. The symbols represent the mean value and the vertical bars represent the standard deviation calculated over 10 triangles for ER, SF, and RN, and over 5 triangles in EN network.
To illustrate how the ”shape” of the point cloud changes as a function of the coupling strength, Fig. 5 depicts the 3D scatter plot for ER (top row) as representative case of Kuramoto dynamics, as well as for the EN (bottom row) for Rössler dynamics, at different coupling strength values. As the coupling parameter increases (moving the oscillators from incoherent to synchronized state), such condition leads to changes in the shape of the point cloud. Consequently, the diversity of the lifetimes of the homology class (\(H_0\)) changes, and those changes are quantified by the NPE.
In both cases (Kuramoto and Rossler dynamics), Figure 5 shows the 3D point cloud constructed with the time series of three nodes forming a triangle at coupling values \(\lambda =0.00\) [Fig. 5(a), incoherent state], \(\lambda =0.08\) [Fig. 5 (b), weakly synchronized state], and \(\lambda =0.30\) [Fig. 5 (c), synchronized state] in ER case as representative cases of Kuramoto networks. Figure 5 also shows the point cloud of a triangle for the experimental network at coupling strengths \(\kappa =0.00\) [Fig. 5 (d), incoherent state], \(\kappa =0.02\) [Fig. 5 (e), weakly synchronized state], and \(\kappa =0.10\) [Fig. 5 (f), synchronized state].
As Fig. 5 shows, the configuration of the point cloud emanating from the triangle undergoes qualitatively distinguishable shape as the system (network) goes from independent to a coordinated state. When the system is in an incoherent state (either \(\lambda =0.00\) or \(\kappa =0.00\)), the points are located sparsely in space. For \(\lambda =0.08\) or \(\kappa =0.02\) (when the system starts to change from incoherent to a synchronized state), the point cloud is concentrated in each region. Finally, when the system reaches a synchronized state (\(\lambda =0.30\) or \(\kappa =0.10\)) the point cloud of the oscillators adopts a defined shape. Such changes are encoded in the persistence diagram (the collection of persistencies or lifetimes of homology classes), which is captures by the NPE (see Figs. 2 and 3).
Representative point clouds illustrating three oscillators forming a triangle. (Top row) Point cloud from a triangle in the Erdos-Renyi network at coupling strengths \(\lambda =0.00\) (a), \(\lambda =0.08\) (b), and \(\lambda =0.30\) (c), as representative case for Kuramoto dynamics. (Bottom row) Point cloud from a triangle in the Experimental network of electronic oscillators at intensity couplings \(\kappa =0.00\) (d), \(\kappa =0.08\) (e), and \(\kappa =0.10\) (f).
The differences between the three Kuramoto oscillator networks are more clearly seen when comparing the behavior of NPE in terms of \(\epsilon\) at fixed coupling strength, as displayed in Fig. 6. We show the average NPE of triangles for the ER, SF and RN networks, for different values of the coupling strength. While for that for \(\lambda =0\), Fig. 6(a), as expected, it is not possible to distinguish the connectivity structure, when the coupling strength is such that the oscillators are partially synchronized, \(\lambda =0.5\), Fig. 6(b), the variation of NPE with \(\epsilon\) reveals differences, particularly of RN with respect to ER and SF. On the other hand, when the coupling strength is large and the oscillators are highly synchronized, \(\lambda =1.5\), Fig. 6(c), the variation of NPE with \(\epsilon\) is the same in the three networks. This is due to the fact that the cloud points are very similar, because when the oscillators are synchronized, and consequently, the coupling term vanishes in the dynamics equation, regardless of their network connectivity configuration (see Eq. 3). For the experimental Rössler network, Fig. 4(d), the variation of NPE with \(\epsilon\) is qualitatively similar to the variation observed in the three Kuramoto networks, in particular, to the variation observed in the ER network (Fig. 4(a)).
NPE vs. the filtration parameter (\(\epsilon\)) of triangles for ER (green symbols) SF (blue symbols) and RN (maroon symbols) networks of Kuramoto oscillators at coupling strength \(\lambda =0.00\) (a), \(\lambda =0.50\) (b), and \(\lambda =1.50\) (c). The symbols represent the average NPE, and the vertical bars represent the standard deviation, both values computed over 10 triangles and 10 simulations of each network.
Lastly, we compare the NPE calculated from triangles and triplets, as a function of the coupling parameter. As shown in Fig. 7, where maximum edge length was set at \(\epsilon =0.15\), when the nodes are uncoupled (i.e., either \(\lambda =0\), or \(\kappa =0\)), there is no distinction between triangles and triplets. As the coupling strength increases, differences between triangles and triplets can be seen in the RN network [panel 7(a)] and in the EN network [panel 7(b)] (no significant differences were found in the ER and SF networks). Therefore, our results suggest that NPE may allow, depending on the network topology and the coupling strength, to discriminate between triangles and triplets. However, we remark that we have only studied four networks, and to demonstrate the generality of these findings, there is the need to simulate a much larger number of different networks, which is computationally very demanding and out of the scope of our work, because we focus on obtaining information about the network state, from NPE analysis of local structures.
NPE vs. the coupling parameter (either \(\lambda\) or \(\kappa\)) of triangles and triplets of RN (a) and EN (b) networks computed at filtration parameter \(\epsilon =0.15\). For the RN network, the green symbols corresponds to the average NPE calculated over 10 triangles, and the black symbols, to the average NPE computed over 10 triplets. The errorbars represent one standard deviation. For the EN network, the average value and the standard deviation are computed over 5 triangles (because the experimental network only contains 5 triangles).
Discussion
We have presented a methodology, based on persistent entropy, to study synchronization phenomena in networks of coupled oscillators. A comprehensive analysis was conducted on experimental data obtained from a set of mutually coupled chaotic electronic circuits, and on synthetic data obtained from simulations of Kuramoto oscillators mutually coupled in an Erdös-Rényi, a scale-free, and a random network.
Specifically, with have used the normalized persistent entropy, NPE, to quantify geometrical information encoded in three-dimensional point clouds that represent the temporal evolution of local network structures, triangles and triples. This methodology has allowed us to identify gradual or steep transitions that are not detected by the order parameter, R, calculated for all the nodes of the network, or calculated for only triangles and triplets.
Furthermore, we have shown that this methodology can detect differences depending on the network connectivity, since the SF and ER networks exhibit a steeper transition compared to the random network.
By studying the behavior of NPE during the synchronization transition, we have found that the changes in the NPE values as the coupling increases are generally consistent with those of the order parameter R, which is the conventional synchronization indicator. However, NPE is more sensitive to changes in coupling strength, and we speculate that this is because it is calculated not from the phases of individual oscillators (as R), but from three-dimensional point clouds that represent the cosine of the phases of the oscillators forming such a point cloud.
However, an important drawback is that the calculation of NPE is computationally demanding, a widespread problem in the context of persistent homology. On the other hand, an advantage of the NPE approach is that the filtration parameter, \(\epsilon\), allows to tune the maximum edge distance in the simplicial complex construction, which in turn can allow to reduce the computational cost. An interesting finding of this study is that, for certain coupling strengths, triangles and triplets in the random network have, on average, different NPE values, and these differences may allow them to be distinguished. It may also be possible, for certain coupling strengths, to distinguish based on NPE values, triangles and triplets in the random network from those in the Erdös-Rényi network or in the scale-free network. Furthermore, based on the results obtained, it is hypothesized that the transition from incoherent to synchronized states maybe identified in a manner that varies according to the specific topology under consideration. However, both aspects–the possibility of distinguishing between triangles and triplets in network connectivity and synchronization behavior based on network connectivity–require further investigation to take advantage of NPE analysis under different coupling conditions, network sizes, and topologies. As future research directions, it will be also interesting to apply this methodology to real-world data, such as biomedical, financial, and climatic data, among others.
Methods
Topological data analysis is a set of methods for studying high-dimensional datasets in terms of its shape, with persistent homology63,66,68as its most versatile tool to extract and characterize geometrical properties by studying homology groups formed in simplicial complexes when a filtration procedure is performed. Here, we will be presenting some general concepts; more detailed discussion of the theoretical underpinnings can be found in Refs52,53,83,84,85,86,87,88., by mentioning a few.
Given a point set \(V=\{v_1, \cdots , v_N\}\) of N elements, an abstract simplicial complex \(K=\{\sigma _k\}\) is the nested collection of simplexes \(\sigma _k=\{v_1, \cdots , v_k\}\) (\(\sigma _k \subseteq V\)), which are convex hulls of \(k+1\) affinely independent points. If \(\sigma '=k'+1\) is a simplex formed by \(k'+1\) affinely independent points, and \(\sigma ' \subseteq \sigma\), consequently, \(\sigma ' \in K\), and \(\sigma '\) is said a face of \(\sigma\). The dimension of a \(k-simplex\) \(\sigma\) is \(dim(\sigma )=k\), and the dimension of a K simplicial complex is the maximum dimension of any \(k-simplex\) in K. A chain \(c=\sum _{i=1}^{m_k}\alpha _i \sigma _i\) is the formal sum of a collection of \(m_k\) simplexes, where \(\alpha _i\) are the coefficients associated to the \(i-th\) simplex \(\sigma _i\) of dimension k. Two chains \(c=\sum _{i=1}^{m_k}\alpha _i \sigma _i\) and \(c'=\sum _{i=1}^{m_k}\alpha _i' \sigma _i\) can be added to form a chain group \(\text {C}=c+c'=\sum _{i=1}^{m_k}(\alpha _i + \alpha _i') \sigma _i\). Two chain groups of different dimensions are related by a boundary operator \(\partial _k:\text {C}_k \rightarrow \text {C}_{k-1}\) that sends a chain group \(\text {C}_k\) to its lower dimensional \(\text {C}_{k-1}\). A \(k-homology\) group \(\text {{\textbf {H}}}_K=\text {{\textbf {Z}}}_k / \text {{\textbf {B}}}_k\) is the quotient group between \(k-th\) cycle group \(\text {{\textbf {Z}}}_k=\ker \partial _k\), and the boundary group \(\text {{\textbf {B}}}_k=\text {im } \partial _{k-1}\). To continue this context, we will restrict to geometrical simplicial complex and \(Vietoris-Rips\) complex (K(VR)). It means that to construct a simplicial complex we will have a Euclidean distance metric \(\epsilon> 0\) and the \(K_{\epsilon }(VR)=\{\sigma _0, \cdots , \sigma _k\}\) is the collection all simplexes \(\sigma _k = \{v_1, \cdots , v_k\}\) such that \(\Vert v_i-v_j\Vert _2 \le \epsilon\) (\(\forall v_i, v_j \in \sigma _k \subseteq V\)). Thus, we can interpret a \(0-simplex\) as a point, a \(1-simplex\) as a line, and a \(2-simplex\) as a triangle, extending to its higher dimensional analogs, and in the simplicial complex, we can interpret the \(0-homology\) group \(H_0\) as the connected components, \(1-homology\) group \(H_1\) the one-dimensional holes, extending to the higher dimensional analogs. In practical cases, finding a distance \(\epsilon\) to construct a simplicial complex whose homology groups are informative about the geometrical properties of a dataset is difficult. A solution is varying the distance measure over its possible values (\(0 \le \epsilon \le \epsilon _{max}\)). This procedure is known as a filtration \(\mathcal {F}: \emptyset =K_{\epsilon _1} \subseteq K_{\epsilon _2} \subseteq \cdots \subseteq K_{\epsilon _n} = K\). In a filtration, born (b) and die (d) a finite number n of homology groups \(H_k\) of dimension k, whose collection \(H_k=\{(b_j^{H_k},d_j^{H_k})\}\) (\(1 \le j \le n\)) is the persistence diagram \(PD(H_k)\) of \(H_k\), also known as the Betti numbers. The difference \(\ell _j^{H_k}=d_j^{H_k}-b_j^{H_k}\) is the persistence or lifetime of the \(j-th\) class of a homology group \(H_k\) in the \(PD(H_k)\), the collection \(B(H_k)=\{\ell _j^{H_k}\}\) is the associated barcodes, which contains information about geometrical properties of the dataset under the filtration.
Normalized persistent entropy
Given the barcodes \(B(H_k)=\{\ell _j (H_k)\}\) endowed with n persistences \(\{\ell _j (H_k)\}\) (\(1 \le j \le n\)) associated to the persistence diagram \(PD(H_k)\) of a homology group \(H_k\) of dimension k, a way to quantify the unequal abundance of the persistences in such \(B(H_k)\) is through persistent entropy \(PE(H_k)\)computing according to the following expression81:
where \(p(\ell _m (H_k))=\ell _m (H_k)/L (H_k)\), and \(L (H_k)=\sum _{m=1}^n \ell _m (H_k)\), such that the Eq. 1 measures the average uncertainty of the persistences.
In our previous work78, a variant of normalized persistent entropy80 (NPE) was considered, by weighting the PE generated by the \(B(H_k)\) of a homology group \(H_k\) of dimension k by its total persistence, L,
The present study aims to detect changes in synchronization states by studying how the NPE of the zero homology group, \(H_0\), changes under two conditions: (i) as the maximum value of the filtering parameter changes, and (ii) by observing its dependence on local structures of the specific networks. We refer to local structures to sets such as triangles (groups of three nodes connected to each other) and triplets (groups of three nodes without direct links between them). The general procedure is shown in Figure 1.
With regard to the first point, NPE is calculated for a value of the filtering parameter that extends up to a parameter cutoff or maximum edge length. This is conventionally taken as the maximum distance, \(\epsilon\), between any two points in such a point cloud (\(\epsilon\) can take values in the interval \(0 \le \epsilon < \infty\)). The result of this filtering is a simplicial complex that encompasses micro, meso, and macro geometric property information of the data. Since this key information may change for different scales in the filtering parameter, we perform the filtering at different values of the distance parameter. These range from \(\epsilon\) close to zero to a maximum value, \(\epsilon _{max}\). The simplicial complex formed at \(\epsilon _{max}\) includes all points in the point cloud. It is important to note that at \(\epsilon _{max}\) no point is a connected component in itself. This approach ensures that all information from zero-dimensional homology groups is incorporated while reducing computational demands. In addressing the second point, we analyze the effect of considering groups of three interconnected oscillators that we refers as triangles, and groups of three nodes with no links between them, that also are no part of a triangle, which we refers as triplets. This is: given a graph G endowed with N vertices \(V=\left\{ v_1, \cdots , v_N\right\}\) (oscillators or nodes), and \(E=\left\{ (v_i, v_j), (v_k, v_l), \cdots \right\}\) edges (\(1 \le i,j,k,l \le N; i \ne j\ne l\ne k\)), a triangle is formed by the set of vertices \(V_{triangle} = \left\{ v_i, v_j, v_k \right\} \in V\) such that the set of edges \(\left\{ (v_i, v_j), (v_i, v_k), (v_j, v_k)\right\} \in E\). On the other hand, a triplet is a non-adjacent set of vertices \(V_{triplet} = \left\{ v_l, v_m, v_n \right\} \in V\), such that the set of edges \(\left\{ (v_l, v_m), (v_l, v_n), (v_m, v_n)\right\} \notin E\) and \(V_{triplet} \cap V_{triangle} = \emptyset\). The time series of these structures (triangles and triplets) were used to represent a three-dimensional point cloud. Subsequently, the maximum edge length value to construct the simplexes forming the corresponding simplicial complex, designated as the filtering parameter, denoted as \(\epsilon\), is varied to compute the NPE of the zero homology groups (\(H_0\)) for each maximum \(\epsilon\). A comparison of the NPE behavior of triangles versus triplets reveals how information is shared locally between them when direct links exist or are lacking at different scales of the data.
Model
We simulated the temporal evolution of a network of N Kuramoto phase oscillators whose governing equations are7,
where \(\theta _{{i}}\) and \(\omega _{i}\) are the phase and frequency, respectively, of the ith oscillator, \(\lambda\) is the global coupling strength and A is the symmetric adjacency matrix of the network.
The standard synchronization quantifier is the order parameter6,7:
where M is the number of oscillators (in Fig. 2 we calculate R with all the oscillators, \(M=N\), or only with \(M=3\)), and \(\langle |\cdot | \rangle _T\) indicates the average over the time interval T (\(1\le t \le T\))7.
We generated three types of networks, Erdös-Rényi (ER), scale free (SF) and random networks (RN). The ER and SF were generated using the graph generator module of NetworkX, implemented in Python. The ER graph was generated with a probability connection of \(p=0.13\). The SF graph has a hub (oscillator 1) endowed with 12 links, and vertices (oscillators) 4, 5, and 6 have 7, 9, and 8 links, respectively; the remaining nodes have at most five links. The network that we refer to as RN consists of vertices randomly linked without any probabilistic linking preference. All the simulated networks have \(N=30\) nodes, and we ensured that they were connected networks, having 10 triangles on each network. The Kuramoto model was simulated for \(0 \le \lambda \le 4\), and \(1 \le t \le 2^{14}\) time steps, with time steps increments of \(10^{-2}\), sub-sampling it to intervals of \(2^{3}\) time steps intervals, having final time series of length \(T=2^{11}\). The frequencies were drawn from a Gaussian distribution with zero mean and standard deviation \(S=0.2\) and for each network, we performed 10 simulations with different frequencies.
Experimental data
We also analyzed experimental data generated by Sevilla-Escoboza and Buldú79. In this dataset the network has \(N=28\) nodes with 5 triangles inside its connectivity, and each node corresponds to an electronic circuit that displays Rössler-like dynamics. The circuits are diffusively coupled through the y variable, and coupling strength is controlled globally by a coupling parameter, \(\kappa\). The data are the voltages recorded \(3\times 10^4\) for each node at a given coupling value \(\kappa\) (\(0.00\le \kappa \le 1.00\), with increments of \(\Delta \kappa = 10^{-2}\)). We analyzed \(2^{11}\) data points taken from the interval [12952, 17048] of the records to avoid border transient effects but ensuring enough activity to characterize the synchronization state of the oscillators. In both cases, simulated and experimental data, the time series were normalized to have zero mean and unitary standard deviation.
Data availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Pikovsky, A., Rosenblum, M. & Kurths, J. Synchronization: A Universal Concept in Nonlinear Sciences (Cambridge University Press, 2003).
Strogatz, S. Sync: The Emerging Science of Spontaneous Order (Hyperion Press, 2003).
Boccaletti, S., Kurths, J., Osipov, G., Valladares, D. & Zhou, C. The synchronization of chaotic systems. Physics Reports 366, 1–101. https://doi.org/10.1016/S0370-1573(02)00137-0 (2002).
Arenas, A., Díaz-Guilera, A., Kurths, J., Moreno, Y. & Zhou, C. Synchronization in complex networks. Physics Reports 469, 93–153. https://doi.org/10.1016/j.physrep.2008.09.002 (2008).
Bayani, A. et al. The transition to synchronization of networked systems. Nature Communications 15, 4955. https://doi.org/10.1038/s41467-024-48203-6 (2024).
Kuramoto, Y. Self-entrainment of a population of coupled non-linear oscillators. In International Symposium on Mathematical Problems in Theoretical Physics, (ed. Araki, H.) 420–422. https://doi.org/10.1007/BFb0013365(Springer, 1975).
Acebrón, J. A., Bonilla, L. L., Pérez Vicente, C. J., Ritort, F. & Spigler, R. The kuramoto model: A simple paradigm for synchronization phenomena. Rev. Mod. Phys. 77, 137–185. https://doi.org/10.1103/RevModPhys.77.137 (2005).
Strogatz, S. H. & Stewart, I. Coupled oscillators and biological synchronization. Scientific American 269, 102–109 (1993).
Glass, L. Synchronization and rhythmic processes in physiology. Nature 410, 277–284. https://doi.org/10.1038/35065745 (2001).
Repp, B. H. & Su, Y.-H. Sensorimotor synchronization: A review of recent research (2006 2012). Psychonomic Bulletin & Review 20, 403–452. https://doi.org/10.3758/s13423-012-0371-2 (2013).
Wang, Z. Cell Cycle Progression and Synchronization: An Overview, 3–23 (Springer, 2022).
Singer, W. Neuronal synchrony: A versatile code for the definition of relations?. Neuron 24, 49–65. https://doi.org/10.1016/S0896-6273(00)80821-1 (1999).
Fell, J. & Axmacher, N. The role of phase synchronization in memory processes. Nature reviews neuroscience 12, 105–118. https://doi.org/10.1038/35065745 (2011).
Dumas, G., Lachat, F., Martinerie, J., Nadel, J. & George, N. From social behaviour to brain synchronization: Review and perspectives in hyperscanning. IRBM 32, 48–53. https://doi.org/10.1016/j.irbm.2011.01.002 (2011).
Jiruska, P. et al. Synchronization and desynchronization in epilepsy: controversies and hypotheses. The Journal of Physiology 591, 787–797. https://doi.org/10.1113/jphysiol.2012.239590 (2013).
Motter, A. E., Myers, S., Anghel, M. & Nishikawa, T. Synchronization and control of power grids. Nature Physics 9, 191–197. https://doi.org/10.1038/nphys2535 (2013).
Jaalam, N., Rahim, N., Bakar, A., Tan, C. & Haidar, A. M. A comprehensive review of synchronization methods for grid-connected converters of renewable energy source. Renewable and Sustainable Energy Reviews 59, 1471–1481. https://doi.org/10.1016/j.rser.2016.01.066 (2016).
Kiss, I. Z. Synchronization engineering. Current Opinion in Chemical Engineering 21, 1–9. https://doi.org/10.1016/j.coche.2018.02.006 (2018).
Harding, D. & Pagan, A. Synchronization of cycles. Journal of Econometrics 132, 59–79. https://doi.org/10.1016/j.jeconom.2005.01.023 (2006).
Ikeda, Y., Aoyama, H., Iyetomi, H. & Yoshikawa, H. Direct evidence for synchronization in japanese business cycles. Evolutionary and Institutional Economics Review 10, 315–327. https://doi.org/10.14441/eier.A2013016 (2013).
Camargo, V. E., Amaral, A. S., Crepaldi, A. F. & Ferreira, F. F. Synchronization and bifurcation in an economic model. Chaos: An Interdisciplinary Journal of Nonlinear Science 32, 103112. https://doi.org/10.1063/5.0104017 (2022).
Strogatz, S. H. Exploring complex networks. nature 410, 268–276. https://doi.org/10.1038/35065725 (2001).
Evans, T. Complex networks. Contemporary Physics 45, 455–474. https://doi.org/10.1080/00107510412331283531 (2004).
Boccaletti, S., Latora, V., Moreno, Y., Chavez, M. & Hwang, D.-U. Complex networks: Structure and dynamics. Physics Reports 424, 175–308. https://doi.org/10.1016/j.physrep.2005.10.009 (2006).
Newman, M. E. J. Resource Letter CS 1: Complex Systems. American Journal of Physics 79, 800–810. https://doi.org/10.1119/1.3590372 (2011).
Liu, Y.-Y. & Barabási, A.-L. Control principles of complex systems. Rev. Mod. Phys. 88, 035006. https://doi.org/10.1103/RevModPhys.88.035006 (2016).
Ladyman, J. & Wiesner, K. What is a complex system? (Yale University Press, 2020).
Erdos, P. L. & Rényi, A. On random graphs. Publicationes Mathematicae 6, 290–297. https://doi.org/10.5486%2FPMD.1959.6.3-4.12 (1959).
Watts, D. J. & Strogatz, S. H. Collective dynamics of small-world networks. nature 393, 440–442. https://doi.org/10.1038/30918 (1998).
Newman, M., Barabási, A.-L. & Watts, D. J. The Structure and Dynamics of Networks (Princeton University Press, Princeton, 2006).
Albert, R., Jeong, H. & Barabási, A.-L. Diameter of the world-wide web. nature 401, 130–131. https://doi.org/10.1038/43601 (1999).
Barabási, A.-L., Ravasz, E. & Vicsek, T. Deterministic scale-free networks. Physica A Statistical Mechanics and its Applications 299, 559–564. https://doi.org/10.1016/S0378-4371(01)00369-7 (2001).
Albert, R. & Barabási, A.-L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–97. https://doi.org/10.1103/RevModPhys.74.47 (2002).
Newman, M. E. J. The structure and function of complex networks. SIAM Review 45, 167–256. https://doi.org/10.1137/S003614450342480 (2003).
Eckmann, J. P. & Ruelle, D. Ergodic theory of chaos and strange attractors. Rev. Mod. Phys. 57, 617–656. https://doi.org/10.1103/RevModPhys.57.617 (1985).
Pincus, S. M. Approximate entropy as a measure of system complexity. Proceedings of the National Academy of Sciences 88, 2297–2301. https://doi.org/10.1073/pnas.88.6.2297 (1991).
Richman, J. S. & Moorman, J. R. Physiological time-series analysis using approximate entropy and sample entropy. American Journal of Physiology-Heart and Circulatory Physiology 278, H2039–H2049. https://doi.org/10.1152/ajpheart.2000.278.6.H2039 (2000).
Costa, M., Goldberger, A. L. & Peng, C.-K. Multiscale entropy analysis of complex physiologic time series. Phys. Rev. Lett. 89, 068102. https://doi.org/10.1103/PhysRevLett.89.068102 (2002).
Pincus, S. Approximate entropy (ApEn) as a complexity measure. Chaos An Interdisciplinary Journal of Nonlinear Science 5, 110–117. https://doi.org/10.1063/1.166092 (1995).
Lake, D. E., Richman, J. S., Griffin, M. P. & Moorman, J. R. Sample entropy analysis of neonatal heart rate variability. American Journal of Physiology-Regulatory, Integrative and Comparative Physiology 283, R789–R797. https://doi.org/10.1152/ajpregu.00069.2002 (2002).
Bandt, C. & Pompe, B. Permutation entropy: a natural complexity measure for time series. Phys. Rev. Lett. 88, 174102 (2002).
Zhang, T., Yang, Z. & Coote, J. H. Cross-sample entropy statistic as a measure of complexity and regularity of renal sympathetic nerve activity in the rat. Experimental Physiology 92, 659–669. https://doi.org/10.1113/expphysiol.2007.037150 (2007).
Rosso, O. A., Larrondo, H. A., Martin, M. T., Plastino, A. & Fuentes, M. A. Distinguishing noise from chaos. Physical Review Letters 99, 154102 (2007).
Delgado-Bonal, A. & Marshak, A. Approximate entropy and sample entropy: A comprehensive tutorial. Entropy 21. https://doi.org/10.3390/e21060541 (2019).
Zabaleta-Ortega, A., Carrizalez-Velazquez, C., Obregón-Quintana, B. & Guzmán-Vargas, L. Multiscale simplicial complex entropy analysis of heartbeat dynamics. Entropy 27, https://doi.org/10.3390/e27050467 (2025).
Schreiber, T. Measuring information transfer. Phys. Rev. Lett. 85, 461–464 (2000).
Quian Quiroga, R., Kraskov, A., Kreuz, T. & Grassberger, P. Performance of different synchronization measures in real data: A case study on electroencephalographic signals. Phys. Rev. E 65, 041903. https://doi.org/10.1103/PhysRevE.65.041903 (2002).
Barreiro, M., A. C., M. & Masoller, C. Inferring long memory processes in the climate network via ordinal pattern analysis. Chaos 21, 013101 (2011).
Terrien, J., Germain, G., Marque, C. & Karlsson, B. Bivariate piecewise stationary segmentation; improved pre-treatment for synchronization measures used on non-stationary biological signals. Medical Engineering & Physics 35, 1188–1196. https://doi.org/10.1016/j.medengphy.2012.12.010 (2013).
Aguilar-Velázquez, D. & Guzmán-Vargas, L. Critical synchronization and 1/f noise in inhibitory/excitatory rich-club neural networks. Scientific Reports 9, 1–13. https://doi.org/10.1038/s41598-018-37920-w (2019).
Zabaleta-Ortega, A., Mercado-Fernández, T., Reyes-Ramírez, I., Angulo-Brown, F. & Guzmán-Vargas, L. Statistical interdependence between daily precipitation and extreme daily temperature in regions of mexico and colombia. Entropy 26, https://doi.org/10.3390/e26070558 (2024).
Edelsbrunner, H., Letscher, D. & Zomorodian, A. Topological persistence and simplification. Discrete Comput Geom 28(511), 533. https://doi.org/10.1007/s00454-002-2885-2 (2002).
Zomorodian, A. & Carlsson, G. Computing persistent homology. In Proceedings of the Twentieth Annual Symposium on Computational Geometry, 347 356, https://doi.org/10.1145/997817.997870 (Association for Computing Machinery, New York, NY, USA, 2004).
Carlsson, G., Zomorodian, A., Collins, A. & Guibas, L. J. Persistence barcodes for shapes. International Journal of Shape Modeling 11, 149–187. https://doi.org/10.1142/S0218654305000761 (2005).
Edelsbrunner, H. & Harer, J. Persistent homology-a survey. Contemporary mathematics 453, 257–282. https://doi.org/10.1090/conm/453/08802 (2008).
Edelsbrunner, H. & Morozov, D. Persistent homology: theory and practice. California Digital Library (2013).
Carlsson, G. Topology and data. Bulletin of the American Mathematical Society 46(255), 308. https://doi.org/10.1090/S0273-0979-09-01249-X (2009).
Zomorodian, A. Topological data analysis. Advances in Applied and Computational Topology 70, 39. https://doi.org/10.1090/psapm/070 (2012).
Wasserman, L. Topological data analysis. Annual Review of Statistics and Its Application 5, 501–532. https://doi.org/10.1146/annurev-statistics-031017-100045 (2018).
Nielson, J. L. et al. Topological data analysis for discovery in preclinical spinal cord injury and traumatic brain injury. Nature Communications 6, 8581. https://doi.org/10.1038/ncomms9581 (2015).
Stolz, B. J., Harrington, H. A. & Porter, M. A. Persistent homology of time-dependent functional networks constructed from coupled time series. Chaos 27, 047410. https://doi.org/10.1063/1.4978997 (2017).
Mittal, K. & Gupta, S. Topological characterization and early detection of bifurcations and chaos in complex systems using persistent homology. Chaos: An Interdisciplinary Journal of Nonlinear Science 27, 051102. https://doi.org/10.1063/1.4983840 (2017).
Motta, F. Topological Data Analysis: Developments and Applications, 369–391 (Springer International Publishing, 2018).
Sizemore, A. E., Phillips-Cremins, J., Ghrist, R. & Bassett, D. S. The importance of the whole: topological data analysis for the network neuroscientist, https://doi.org/10.48550/ARXIV.1806.05167 (2018).
Garside, K., Henderson, R., Makarenko, I. & Masoller, C. Topological data analysis of high resolution diabetic retinopathy images. PLoS ONE 14, e0217413. https://doi.org/10.1371/journal.pone.0217413 (2019).
Ravishanker, N. & Chen, R. Topological data analysis (tda) for time series (2019). arXiv:1909.10604.
Xu, X., Cisewski-Kehe, J., Green, S. & Nagai, D. Finding cosmic voids and filament loops using topological data analysis. Astronomy and Computing 27, 34–52. https://doi.org/10.1016/j.ascom.2019.02.003 (2019).
Carlsson, G. Topological methods for data modelling. Nature Reviews Physics 2(697), 708. https://doi.org/10.1038/s42254-020-00249-3 (2020).
Carlsson, G. & Vejdemo-Johansson, M. Topological Data Analysis with Applications 2nd edn, (Cambridge University Press, 2021).
Christian, P. et al. Topological data analysis of black hole images. Phys. Rev. D 106, 023017. https://doi.org/10.1103/PhysRevD.106.023017 (2022).
Skaf, Y. & Laubenbacher, R. Topological data analysis in biomedicine: A review. Journal of Biomedical Informatics 130, 104082. https://doi.org/10.1016/j.jbi.2022.104082 (2022).
Muszynski, G., Kashinath, K., Kurlin, V., Wehner, M. & Prabhat. Topological data analysis and machine learning for recognizing atmospheric river patterns in large climate datasets. Geoscientific model development 12, 613–628 https://doi.org/10.5194/gmd-12-613-2019 (2019).
Strommen, K., Chantry, M., Dorrington, J. & Otter, N. A topological perspective on weather regimes. Climate Dynamics 60, 1415–1445. https://doi.org/10.1007/s00382-022-06395-x (2023).
Gidea, M. & Katz, Y. Topological data analysis of financial time series: Landscapes of crashes. Physica A 491, 820–834. https://doi.org/10.1016/j.physa.2017.09.028 (2018).
Akingbade, S. W., Gidea, M., Manzi, M. & Nateghi, V. Why topological data analysis detects financial bubbles?. Commun. Nonlinear Sci. Numer. Simulat. 128, 107665. https://doi.org/10.1016/j.cnsns.2023.107665 (2024).
Guzman-Vargas, L., Zabaleta-Ortega, A. & Guzman-Saenz, A. Simplicial complex entropy for time series analysis. Scientific Reports 13. https://doi.org/10.1038/s41598-023-49958-6 (2023).
Shah, W. H., Jaimes-Reátegui, R., Huerta-Cuellar, G., García-López, J. & Pisarchik, A. Persistent homology approach for uncovering transitions to chaos. Chaos, Solitons & Fractals 192, 116054. https://doi.org/10.1016/j.chaos.2025.116054 (2025).
Zabaleta-Ortega, A., Masoller, C. & Guzmán-Vargas, L. Topological data analysis of the synchronization of a network of R ssler chaotic electronic oscillators. Chaos: Interdiscip. J. Nonlinear Sci. 33, 113110. https://doi.org/10.1063/5.0167523 (2023).
Sevilla-Escoboza, R. & Buld, J. M. Synchronization of networks of chaotic oscillators: Structural and dynamical datasets. Data in Brief 7, 1185–1189. https://doi.org/10.1016/j.dib.2016.03.097 (2016).
Myers, A., Munch, E. & Khasawneh, F. A. Persistent homology omf complex networks for dynamic state detection. Physical Review E 100, 022314 (2019).
Rucco, M., Castiglione, F., Merelli, E. & Pettini, M. Characterisation of the idiotypic immune network through persistent entropy. In Proceedings of ECCS 2014 (eds Battiston, S. et al.) 117–128, https://doi.org/10.1007/978-3-319-29228-1_11 (Springer International Publishing, Cham, 2016).
Rossi, K. L., Budzinski, R. C., Boaretto, B. R. R., Muller, L. E. & Feudel, U. Shifts in global network dynamics due to small changes at single nodes. Phys. Rev. Res. 5, 013220. https://doi.org/10.1103/PhysRevResearch.5.013220 (2023).
Munkres, J. Elements of algebraic topology. Library of Congress Cataloguing in Publication Data 2nd edn (Addison-Wesley Publishing Company, 1984).
Hatcher, A. Algebraic topology 1st edn, (Cornell University Press, 2001).
Zomorodian, A. & Carlsson, G. The theory of multidimensional persistence. Proceedings of the twenty-third annual symposium on Computational geometry June, 184 193, https://doi.org/10.1145/1247069.1247105 (2007).
Edelsbrunner, E. & Harer, J. Computational topology: an introduction. QA3-611-E353 (American Mathematical Society, 2010).
Munkres, J. Topology. British Library Cataloguing-in-Publication Data 2nd edn, (Pearson Education Limited, 2014).
Dey, T. K. & Wang, Y. Computational Topology for Data Analysis (Cambridge University Press, 2022).
Acknowledgements
Á.Z-O. acknowledges CONAHCYT-México for the financial support during the PhD program. C.M. acknowledges the Spanish Ministerio de Ciencia, Innovación y Universidades (PID2021-123994NB-C21), the Institució Catalana de Recerca i Estudis Avançats (Academia) and the Agencia de Gestió d’Ajuts Universitaris i de Recerca (AGAUR, SGR2021). L.G.-V acknowledges COFAA-IPN, EDI-IPN and CONAHCYT-México.
Author information
Authors and Affiliations
Contributions
Á.Z-O., L.G-V., and C.M. conceived the study and designed the methodology, Á.Z-O. conducted the numerical experiments, Á.Z-O., L.G-V., and C.M. analysed the results. Á.Z-O. formal analysis. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zabaleta-Ortega, Á., Masoller, C. & Guzmán-Vargas, L. Unveiling synchronization transitions in networks of coupled oscillators through persistent homology of local structures. Sci Rep 15, 42772 (2025). https://doi.org/10.1038/s41598-025-27083-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27083-w