Abstract
Amylose, a vital polysaccharide component of starch, plays a significant role in plant energy storage and has important implications in nutrition and health. In this study, the structural characteristics of amylose are analyzed using Sombor indices, a relatively recent method in topological molecular analysis. Leveraging Euclidean geometry, this work introduces the first area-based Sombor index, offering a novel perspective on the molecular connectivity and spatial configuration of amylose. The third and fifth Sombor indices are derived from perimeter-based geometric principles, introducing a new level of complexity to the topological characterization. In contrast, the second, fourth, and sixth indices are developed using angular-based formulations, enabling a more refined structural interpretation. To assess the relationship between these indices and the physicochemical properties of amylose, regression analysis was performed using supervised machine learning techniques. This statistical modeling uncovered meaningful correlations, enhancing our understanding of how molecular topology relates to chemical behavior. Additionally, Analysis of Variance (ANOVA) was applied to determine the statistical significance of each index. Correlation analyses revealed strong interrelationships among the indices. The results indicate that among all considered Sombor-based indices, SO\(_4\) and SO\(_5\) are the most effective predictors of amylose’s structural and functional properties. In particular, SO\(_5\) exhibited the highest predictive accuracy and model robustness, while SO\(_4\) also demonstrated consistent performance, affirming their applicability in molecular modeling. This research underscores the potential of Sombor indices as reliable topological descriptors for molecular classification and offers valuable insights into the physiochemical behavior of amylose. The findings open new directions for applying topological analysis to the study of biopolymers and polysaccharides, with implications in materials science, biochemistry, and food technology.
Similar content being viewed by others
Introduction
Chemical graph theory represents a study that combines mathematical graph analysis with chemical challenges. Utilising this strategy, the chemical and pharmaceutical sciences benefit significantly from topological indices, which are quantitative characteristics created from graph invariants.1. These parameters are frequently used to predict the physicochemical properties of organic molecules. There are many several topological indices generated for different molecular structures in the literature in this area2. The degree of vertices in a molecular structure is the main source of topological indices, which are quantitative numbers referred to as graph-based molecular descriptors. These indices have demonstrated value across an extensive spectrum of industries and capture important structural data. numerous applications, which include molecular identity analyses, quantitative structure-property relationships (QSPR), and quantitative structure-activity relationships (QSAR)3. They have attracted the focus of mathematicians and chemists both due to their mathematical implications and chemical sensitivity. Since H. Wiener’s landmark 1947 work, in which he introduced the Wiener index as the first distance-based topological descriptor, the field of mathematics has developed rapidly.4. Approximately 300 distinct topological indices have been developed and cataloged in various databases, has demonstrating their widespread utility and continuous evolution in theoretical and computational chemistry5 . Topological indices are quantitative network characteristics that capture the characteristics of molecular graphs, providing a mathematical representation of molecular topology6. These descriptor are commonly used to estimate important physicochemical properties like boiling, melting point, and freezing point. In contemporary chemical research, conducting biological assays direct compound evaluation has become progressively impractical due to high economic expenses and the necessity for advanced workshop infrastructure. This method also heavily depends on focused arrangement, production it improper for large-scale complex showing7. As a result, pharmaceutical companies are continuously discovering cutting- edge way to lower the expenses related to study and development. applying topological indices to analyse structures of molecular is one attractive technique that enables it feasible to predict chemical properties without the need for expensive tools or physical labs. This technique offers a less expensive and effective substitute for current experimental techniques. The Sombor index was initially introduced in8. It differentiates from many standard degree-based indices since it is based on geometric interpretation. Numerous investigations of its mathematical features and its applications in chemical graph theory have been conducted prompted by the wide academic interest in this unique geometric perspective9. Its formulation offers a novel way of describing molecular graphs; it is based on vertex degrees with spatial or structural aspects. The Sombor index is distinctive and significant in this area since no other topological index based exclusively on vertex degrees has been distributed to date with an identical focus on geometric reasoning10. Quantitative descriptors that have been systematically derived from a chemical compound’s structural network and describe its structural properties and atomic relationship are referred to as topological indices. In multiple fields of chemistry, such as cheminformatics, drug discovery, and molecular modelling, these indices are becoming indispensable resources. In this situation, they provide important information on the molecular topological structure and interaction patterns11. An important family of degree-based topological descriptors that provide useful information about the structure and physicochemical properties of molecular systems are Sombor indices, which were initially proposed by Milan Randić12.
An important family of degree-based topological descriptors that provide useful information about the structure and physicochemical properties of molecular systems are Sombor indices, which were initially proposed by Milan Randic. These indices, which were defined within the context of graph theory, quantitatively encode the arrangement and interaction of atoms within a molecule, which enabling in-depth structural study13. Sombor indices have emerged as a prominent class of degree-based topological descriptors due to their mathematical robustness and structural sensitivity14. Their ability to capture both local and global structural information makes them superior to several classical indices in encoding molecular topology15. Due to these strengths, Sombor indices have been successfully applied in the prediction of various molecular properties, including boiling point, reflectivity, and toxicity16,17. Such programs are able to quantify the de facto degree effect of molecular structure on the physical and chemical properties that they embody.
Prediction of physical basic properties is certainly a usage for Sombor indices beyond those circumstances, however, Sombor index has also been identified useful in the modelling of complex molecule systems and intermolecular interactions. Their chemical applicability across a wide range of compounds reinforces their value in both theoretical and applied chemistry18. Their strong correlation with experimentally observed properties makes them particularly effective in quantitative structure–activity relationship (QSAR) and quantitative structure–property relationship (QSPR) studies. These indices have been utilized in the analysis of polymers, nanostructures, and dendrimer-based systems, improving prediction accuracy in drug discovery and material design processes19. In addition, the expected values of Sombor indices have been explored for various classes of chemical graphs, further enriching their theoretical foundation and computational potential20. In the present study, we specifically focus on amylose, a linear polysaccharide composed of \(\alpha (1\rightarrow 4)\) linked D-glucose units. Its unique helical structure and physicochemical behavior make it an ideal candidate for topological and structural analysis. Amylose plays a critical role in starch functionality, influencing properties such as gelatinization, retrogradation, and digestibility. By modeling amylose as a molecular graph, topological indices—particularly degree-based indices like the Sombor index—can be applied to understand its structural behavior and predict associated physicochemical characteristics. The mathematical treatment of amylose through topological descriptors provides valuable insights into its potential applications in food science, nutrition, and materials chemistry.
The design of these six Sombor invariants is motivated by geometric analogies such as distances, angles, perimeters, and radii within molecular graphs. These formulations aim to offer refined descriptors that incorporate not only connectivity but also spatial distribution and degree asymmetry. Such features are crucial for modeling real-world biomolecules like amylose, where shape, folding, and interaction patterns influence function. Therefore, each index not only serves a mathematical role but also bears implications for the chemical behavior and structural predictability of biological polymers.
The proposed Sombor-based indices also exhibit direct mathematical connections with well-established indices such as the Forgotten and Zagreb indices. In particular, the kernel of the Sombor index satisfies the identity
where \(F(G)\) is the Forgotten index. This relation enables us to bound the Sombor index in terms of classical descriptors, namely
with \(M_1(G)\) denoting the first Zagreb index and \(m=|E(G)|\). These bounds follow from the RMS–AM and Cauchy–Schwarz inequalities and hold with equality for regular graphs. Furthermore, if the geometric variant of the Sombor index is defined through the product of degrees, i.e.,
then it coincides exactly with the second Zagreb index \(M_2(G)\). These observations demonstrate that the newly proposed indices are not only novel but also theoretically consistent with existing topological descriptors, thereby strengthening their relevance and applicability.
Exploration and analysis of existing literature.
Amylose (see Fig. 1), a linear polysaccharide composed of \(\alpha (1 \rightarrow 4)\) linked D-glucose units, constitutes approximately \(20 - 30\) of starch content in most plant sources. Its unieq helical structure allows it to form complexes with various molecules, influencing its physicochemical properties and functional applications. The structural characteristics of amylose significantly impact the texture, digestibility, and stability of starch- based foods, making it a focal point in food science and nutrition research.
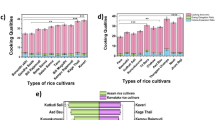
The physicochemical properties of amylose, such as gelatinization temperature, sweeling power, and solubility, are influenced by factor like amylose content and environmental conditions during processing. Studies have shown that higher amylose content correlates with increasing gelatinization temperatures and reduced swelling power, affecting the texture and digestibility of starch-rich foods. For instance, research on rice cultivars with varying amylose content demonstrated significant difference in their thermal and pasting properties, highlighting the role of amylose indetermining starch functionality.
Amylose’s ability to form inclusion complexes with lipids and others hydrophobic molecules has been extensively studied. These complexes possess the ability to modify starch’s digestibility and beneficial properties. For instance, the production of complexes among amylose and lipids could reduce the glycaemic response of starchy meals, thus enhancing health. Additionally, the complexion behavior of amylose is influenced by factors such as chain length, molecular weight, and processing conditions, which can be tailored to achieve desired functional attributes in food products. Amylose’s retrogradation conduct, in which gelatinised starch molecules re-associate while cooled, is crucial in determining physical diversity and shelf life of foods prepared utilising starch. Although amylose retrogrades more rapidly than amylopectin, products such bread and gels might have more difficult qualities and a potential syneresis. Enhancing food processing and storage conditions involves an in-depth comprehension of the dynamics and procedures of amylose retrogradation. Amylose information, temperature, and the existence of other chemicals may all have significant effects on the degree and speed of retrogradation, based on studies.
Recent developments in QSPR (Quantitative Structure–Property Relationship) modeling have expanded the applicability of topological descriptors to biologically and industrially relevant compounds. Studies such as those by21 have employed molecular graph descriptors to model anti-Alzheimer agents, revealing methodological parallels with the modeling of biopolymers like amylose. Similarly, the design of anti-biofilm agents22 and anti-HIV compounds23 using QSPR and computational biomedicine approaches highlights the increasing utility of structural descriptors in understanding molecular functionality. Furthermore, toxicological prediction studies24 reinforce the role of QSPR in evaluating chemical safety, thereby establishing a comprehensive framework where the structural modeling of amylose through topological indices finds broader relevance across pharmacological domains. These advancements justify the relevance of our Sombor-based modeling strategy and emphasize its potential integration in future biochemical and medicinal research.
Amylose.
In this table, \(E^o_i\) represents the i-th class of edges categorized by the degrees of their endpoint vertices. The corresponding frequency denotes the total number of edges belonging to each class, i.e., \(|E^o_i|\) gives the cardinality of edge set \(E^o_i\).
Analytical framework and invariant definitions
In a graph \(G = (V, E)\) consists of a set vertices \(V\) and a set edges \(E\), where each edge joins a distinct pair of vertices. These edges illustrate the structural connections within the graph and serve as the basis for conducting topological investigations. The size of a graph , denoted by \(|E^o|\), refers to the total number of edges and provides a basic indication of its connectivity. The edge \(\{\alpha , \beta \} \in E\) signifies an interaction between vertices \(\alpha\) and \(\beta\), contributing to the graph’s overall structure25. The original Sombor index was introduced as a degree-based topological descriptor to reflect the intrinsic connectivity between bonded atoms (vertices) in a molecular graph. It is defined as:
Here, \(d_{\alpha }\) and \(d_{\beta }\) represent the degrees of the vertices \(\alpha\) and \(\beta\), respectively, connected by an edge. The index captures the joint intensity of connectivity between pairs of atoms, with higher values corresponding to bonds between highly connected atoms. From a molecular geometry perspective, this formulation approximates the Euclidean norm in degree space, enabling indirect structural quantification. Biologically, in polymers like amylose, the Sombor index reflects the extent of local branching and compactness, which may influence folding behavior and chemical reactivity.
The primary persistent feature of the Sombor graph parameter was determined by examining the geometric spacing within two edges, and it is rigorously described as follows.
This index measures the absolute imbalance between squared vertex degrees. Geometrically, it mimics spatial asymmetry between connected atoms and can reflect molecular polarity or irregular branching.
The secondary persistent component of the Sombor graph parameter emerged from an in-depth study of the angular geometry between edges, shedding light on the graph’s structural intricacies.
This normalized index captures angular variation in connectivity, indicating flexibility and structural diversity in molecular arrangements. The third version of the Sombor graph parameter is based on the geometric principle of a triangles circumcircle and is rigorously defined as.
Inspired by the circumradius formula of a triangle, this index reflects how vertex degrees spatially distribute over molecular cycles or loops, connecting perimeter-based geometry to molecular stability.
The fourth variant of the Sombor graph component was created by using the dimensions of a triangular’s circumcircle.
This variant, derived by squaring the geometric term in \(\hbox {SO}_3\), emphasizes the contribution of high-degree vertices through quadratic amplification. It geometrically mirrors the energy-like spread of degrees across edges, inspired by circumcircle-based modeling. Biologically, \(\hbox {SO}_4\) reflects the distributional balance of bonding environments in amylose, and its sensitivity to degree variations makes it valuable for identifying structurally significant molecular regions.
We have created the fifth invariant of the Sombor graph component using the circumference of a triangle’s incircle.
This index relates to the incircle circumference and captures radial symmetry within molecular graphs, highlighting structural compactness and reactive sites.
We have created the sixth invariant of the Sombor graph component using the circumference of a triangle’s incircle.
This sixth invariant is derived from a squared difference formulation based on the normalized difference of squared vertex degrees. The expression incorporates both the contrast between degrees and the local edge complexity, offering a magnified sensitivity to asymmetries. Inspired by the geometric notion of a triangle’s incircle, it captures subtle variations in the structural landscape of amylose. Biologically, \(\hbox {SO}_6\) helps quantify irregular topological features that may affect physicochemical behaviors such as folding, interaction, or enzymatic reactivity.
Methods
In this research, an in-depth study of topological indices based on Sombor degrees was conducted with the objective to explore their predictive value utilising machine learning techniques. We applied a variety of regression model’s, with the linear regression model demonstrating superior accuracy and robustness. Comprehensive statistical evaluations were performed to confirm the finding, including the computation of regression statistics, ANOVA, regression coefficients, and residual outputs for each topological measurement. In addition, a statistical correlation study was performed on non-zero value indices to evaluate their significance and predictive importance in the context of molecular graph analysis.
Theorem 1
Let G be the molecular graph of amylose, then the Sombor index is given by
Proof
In the network of amylose with 12n edges, the Sombor index of the graph G can be decomposed into four disjoint edge sets: \(E^o_{1}(G)\), \(E^o_{2}(G)\), \(E^o_{3}(G)\), and \(E^o_{4}(G)\), as presented in Table 1. These sets represent different edge configurations based on the degrees of their endpoints. Specifically:
-
\(E^o_{1}(G)\) consists of n edges where \(d_\alpha = 1\) and \(d_\beta = 2\).
-
\(E^o_{2}(G)\) consists of \(2n + 2\) edges where \(d_\alpha = 1\) and \(d_\beta = 3\).
-
\(E^o_{3}(G)\) consists of \(5n - 2\) edges where \(d_\alpha = 2\) and \(d_\beta = 3\).
-
\(E^o_{4}(G)\) consists of 4n edges where \(d_\alpha = 3\) and \(d_\beta = 3\).
The Sombor index for the amylose graph is given by:
\(\square \)
Theorem 2
Let G be the molecular graph of amylose, then the first invariant of the Sombor index is given by
Proof
The first invariant for the Sombor index is computed using Eq. (2), and it is represented as:
By considering the geometrical perspective of the topological indices, we derive the formula mentioned above. This derivation relies on understanding the degree behavior of vertices within the network, as detailed in Table 1, which catalogs the edge and degree type characteristics. By substituting the values of \((d_{\alpha }, d_{\beta })\) into the given formulation, we obtain the desired result:
\(\square \)
Theorem 3
Let G be the molecular graph of amylose, then the second invariant of the Sombor index is given by
Proof
Equation (3) utilizes the second invariant of the Sombor index, expressed as:
By considering the geometrical perspective of the topological indices, we derive the formula mentioned above. This derivation relies on understanding the behavior of vertex degrees within the network, as detailed in Table 1, which catalogs the edge and degree type characteristics. By substituting the values of \((d_{\alpha }, d_{\beta })\) into the given formulation, we obtain the desired result:
\(\square \)
Theorem 4
Let G be the molecular graph of amylose, then the third invariant of the Sombor index is given by
Proof
Equation (4) employs the third invariant of the Sombor index, which is represented as:
The derivation of this formula originally relied on the geometrical perspective of the topological indices. However, we are now employing the behavior of vertex degrees within the structure. Table 1 outlines the edge and degree type characteristics of the array under consideration. By substituting the values of \((d_{\alpha }, d_{\beta })\) into the formula, we obtain the desired result:
\(\square \)
Theorem 5
Let G be the molecular graph of amylose, then the fourth invariant of the Sombor index is given by
Proof
Equation (5) employs the fourth invariant of the Sombor index, which is articulated as:
The derivation of this formula originally relied on the geometrical perspective of topological indices. However, we now consider the behavior of vertex degrees within the molecular graph. Table 1 outlines the edge and degree-type classifications used in the summation. By substituting the values of \((d_{\alpha }, d_{\beta })\), we obtain:
\(\square \)
Theorem 6
Let G be the molecular graph of amylose, then the fifth invariant of the Sombor index is given by
Proof
Equation (6) relies on the fifth invariant of the Sombor graph parameter, which is defined as:
The derivation of this formula considers the degree behavior of vertices within the molecular graph. Table 1 outlines the edge types and corresponding vertex degrees. Substituting the respective values of \((d_{\alpha }, d_{\beta })\) gives:
\(\square \)
Theorem 7
Let G be the molecular graph of amylose, then the sixth invariant of the Sombor index is given by
Proof
In Eq. (7), the sixth invariant of the Sombor graph parameter is defined as:
This formulation originally emerged from a geometrical perspective of topological indices, but here we apply it using vertex degree analysis as shown in Table 1. Substituting the specific degrees yields:
\(\square \)
Regression analysis using supervised machine learning
Regression analysis, in particular, is an effective method to predict numerical relationships and assessing the influence of different variables in supervised machine learning. In this study, SO, SO\(_1\), SO\(_2\), SO\(_3\), SO\(_4\), SO\(_5\), and SO\(_6\) were predictor using relevant predictor variables through multiple regression model’s. The model showed a strong fit and their reliability was further supported by extremely low Mean Squared Error (MSE) and Mean Absolute Error (MAE). These findings demonstrate that difficult mathematical connections and structures may be effectively represented by machine learning. To mitigate the risk of overfitting and ensure the generalizability of the developed regression model’s, a standard 80:20 train-test split was applied, where 80% of the data was used for training and 20% for testing. Additionally, 5-fold cross-validation was employed to assess model performance across multiple data subsets. This validation strategy confirmed consistent and reliable predictive power across the folds. The minimal differences in evaluation metrics such as R2, MAE, and MSE between training and test sets demonstrated the robustness of the model’s. These practices enhance confidence in the general applicability of the proposed model’s beyond the analyzed dataset. A detailed analysis of the predictive characteristic revealed that some variable had a strong impact on model reliability then other. In particular, SO\(_3\) and SO\(_4\) regularly had a strong influence, revealing their strong abilities to predict other variables. Conversely, SO and SO\(_2\) showed the least influence, demonstrating that they have no impact on the relationship between the indices as an entire. Thus regression analysis not only enhance predictive accuracy but also provides insights into the relatives importance of input features. The results of the research demonstrate how effectively machine learning works in mathematical modeling, particularly as it applies to topological index prediction. Given the excellent precision obtained, this technique can be used to more complex data sets or, for better generalization, integrated with more advanced methods of machine learning such ensemble learning and deep learning. To further improve predictions in mathematics and scientific applications, future research might investigate hybrid techniques or nonlinear regression model’s.
Supervised learning approach for SO prediction
In this supervised learning model, SO using other topological indices, SO\(_1\), SO\(_2\),SO\(_3\), SO\(_4\), SO\(_5\), and SO\(_6\) as predictor variables. The model achieved an \(\hbox {Q}^{2}\) score of 0.9298, indicating a perfect prediction, as shown in Table 2. The Mean Squared Error (MSE) of \(1.72 \times 10^{-27}\) and the Mean Absolute Error (MAE) of \(3.55 \times 10^{-14}\) indicate an extremely low level of error. These results confirm that the model effectively captures the underlying relationship between SO and its predictor variables with high precision.
Feature importance in SO prediction
The contribution of different predictor variables (SO\(_1\), SO\(_2\), SO\(_3\), SO\(_4\), SO\(_5\), and SO\(_6\)) towards the prediction of SO was analyzed using a supervised machine learning model. The results, presented in Table 3, indicate that SO\(_3\) has the highest impact (\(0.2521\)) on the prediction of SO, while SO\(_2\) contributes the least (\(0.0074\)). Moderate influence is observed for SO\(_4\) (\(0.0924\)) and SO\(_5\) (\(0.0617\)), whereas SO\(_1\) (\(0.0394\)) and SO\(_6\) (\(0.0230\)) have a relatively smaller effect. These findings suggest that SO\(_3\) plays a dominant role in determining SO, making it the most influential parameter for predictive modeling.
Supervised learning approach for SO\(_1\) prediction
The supervised learning model was applied to predict SO\(_1\) using the predictor variables SO, SO\(_2\), SO\(_3\), SO\(_4\), SO\(_5\), and SO\(_6\). The results in Table 4 show an \(\hbox {Q}^{2}\) score of 0.8998, indicating a perfect prediction accuracy. The mean squared error (MSE) is \(4.04 \times 10^{-28}\), and the mean absolute error (MAE) is \(1.42 \times 10^{-14}\), reflecting an extremely low level of error. These values confirm that the chosen predictor variables effectively model SO\(_1\) with high precision.
Feature importance in SO\(_1\) prediction
The contribution of different predictor variables to SO\(_1\) prediction is presented in Table 5. The findings suggest that SO\(_3\) has the highest impact (\(0.1204\)), followed by SO\(_4\) (\(0.0441\)) and SO (\(0.0373\)), both of which exhibit a moderate influence. Mean while, SO\(_5\) (\(0.0294\)) and SO\(_6\) (\(0.0110\)) have a small impact, and SO\(_2\) (\(0.0035\)) contributes the least. These results highlight the dominance of SO\(_3\) as a critical factor in predicting SO\(_1\), reinforcing its importance in the model.
Linear regression analysis for the molecular graph of amylose SO\(_1\)
This section presents a linear regression analysis conducted between the topological index SO and the derived index SO\(_1\), computed for the molecular graph of Amylose. The purpose of this analysis is to examine the degree of association, statistical significance, and the predictive accuracy of the model in representing the linear relationship between SO and SO\(_1\). The following analysis includes regression statistics, ANOVA results, estimated regression coefficients, and residual diagnostics. These components collectively establish the credibility of the regression model and validate its statistical performance. Table 6 presents the regression summary statistics. The value of Multiple R is 0.9739, which indicates a perfect linear association between the two indices. The R Square value of 0.9329 indicate a strong explanatory power of the model in predicting the variability SO\(_1\) using SO as a predictor. The Adjusted R Square, also equal to 0.99432, reinforces the model’s reliability despite the limited number of predictors. The standard error is extremely low (\(1.40935 \times 10^{-14}\)), indicating high precision in the predicted values.
The ANOVA results presented in Table 7 provide insight into the statistical significance of the regression model. The F-statistic is extraordinarily high (\(1.8762 \times 10^{32}\)), indicating that the model explains a significant portion of the variability in the data. Additionally, the Significance F value of zero confirms that the model is statistically significant at all levels, with the predictor (SO) being a strong and reliable determinant of SO\(_1\). The very low residual sum of squares (SS) of \(3.7739 \times 10^{-27}\) further highlights the precision of the model in minimizing prediction errors.
Table 8 displays the regression coefficients for the linear model. The intercept value of 3.448505441 indicates the predicted value of SO\(_1\) when SO is zero. The coefficient for SO, 0.505073695, represents the rate of change of SO\(_1\) with respect to SO. Both coefficients have extremely low standard errors (\(6.8174 \times 10^{-15}\) for the intercept and \(1.1660 \times 10^{-17}\) for SO), suggesting that they are highly precise. The corresponding t-statistics are exceptionally large (\(5.0583 \times 10^{14}\) for the intercept and \(4.3316 \times 10^{16}\) for SO), and the p-values are virtually (\(2.0355 \times 10^{-305}\), indicating that both terms are statistically significant and can be confidently interpreted as reliable predictors of SO\(_1\).
Table 9 presents a sample of the residual output, which reflects the difference between the actual and predicted SO\(_1\) values for each observation. The residuals are extremely small, with values on the order of \(10^{-14}\), indicating that the predicted values are very close to the actual data. Furthermore, the standard residuals are close to zero, confirming that there are no outliers or significant deviations in the data. This indicates a nearly perfect fit of the model to the observed values, further validating the strength and accuracy of the linear regression model. Graphically representations of the analysis are provided in the corresponding Figure 2,which visually confirm the robustness and precision of the regression model. The term “actual data” refers to the computed index values obtained directly from the closed-form expressions of the proposed Sombor-based indices for the molecular graph of amylose. Specifically, for a chain length of \(n\) glucose units, the formulas \(SO_n(G)\) (for \(n=1,\dots ,6\)) were evaluated by substituting \(n \in \{1,2,\dots ,10\}\). For instance, \(SO_1(G)=22n+3\), \(SO_2(G)=4.123\,n+0.831\), and \(SO_3(G)=140.70\,n+0.888\), with analogous forms for \(SO_4\)–\(SO_6\). Thus, the dataset used in regression and correlation analysis consists of systematically generated values derived from these formulas, rather than externally sourced experimental measurements.
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_1\).
Supervised learning approach for SO\(_2\) prediction
In this supervised learning model, SO\(_2\) was predicted using SO, SO\(_1\), SO\(_3\), SO\(_4\), SO\(_5\), and SO\(_6\) as predictor variables. The model achieved an \(\hbox {Q}^{2}\) score of 0.9969, indicating a perfect prediction, as shown in Table 10. The mean squared error (MSE) of \(2.35 \times 10^{-28}\) and the mean absolute error (MAE) of \(1.51 \times 10^{-14}\) indicate an extremely low level of error. These results confirm that the model effectively captures the underlying relationship between SO\(_2\) and its predictor variables with high precision.
Feature importance in SO\(_2\) prediction
The importance of different predictor variables in determining SO\(_2\) is outlined in Table 11. The results indicate that SO\(_3\) has the highest impact (\(0.1723\)), followed by SO\(_4\) (\(0.0651\)), which has a moderate influence. Meanwhile, SO (\(0.0091\)) contributes the least, while SO\(_1\) (\(0.0512\)), SO\(_5\) (\(0.0379\)), and SO\(_6\) (\(0.0207\)) have little effect. According to these results, SO\(_2\) is the dominant feature in the model and the most important variable for correctly predicting SO\(_2\).
Linear regression analysis for the molecular graph of amylose SO\(_2\)
This section presents a linear regression analysis conducted between the topological index SO and the derived index SO\(_2\), computed for the molecular graph of Amylose. The purpose of this analysis is to examine the degree of association, statistical significance, and the predictive accuracy of the model in representing the linear relationship between SO and SO\(_2\). The following analysis includes regression statistics, ANOVA results, estimated regression coefficients, and residual diagnostics. These components collectively establish the credibility of the regression model and validate its statistical performance. Table 12 presents the regression summary statistics. The value of multiple R is 0.9929, which indicates a perfect linear association between the two indices, indicating a perfect linear correlation between SO and SO\(_2\). The R Square value of 0.9135 indicate a strong explanatory power of the model in predicting the variability SO\(_2\) using SO as a predictor. . Furthermore, the adjusted R-square value is 0.9491, which further confirms the reliability of the model, especially considering that it involves only one predictor. The standard error is extremely low, at \(4.9331 \times 10^{-15}\), indicating that the predicted values align closely with the actual values of SO\(_2\) and the residuals are nearly negligible. This level of precision highlights the robustness and accuracy of the linear regression model to capture he relationship between SO and SO\(_2\).
Table 13 shows the ANOVA results. The extremely high F-statistic value of \(5.3785 \times 10^{32}\) and a Significance F of zero indicate a statistically significant relationship between SO and SO\(_2\), validating the strength of the regression model.
Table 14 presents the regression coefficients. The intercept and slope for SO\(_2\) exhibit extremely small standard errors and p-values, confirming their statistical significance. This supports the model’s stability and the reliability of predictions.
Table 15 shows a portion of the residual output. All residuals are very close to zero, and the standard residuals are within acceptable limits, demonstrating that the model fits the observed data remarkably well with no evidence of outliers or anomalies. Graphically representations of the analysis are provided in the corresponding Fig. 3,which visually confirm the robustness and precision of the regression model.
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_2\).
Supervised learning approach for SO\(_3\) prediction
The performance of the supervised learning model in predicting SO\(_3\) is presented in Table 16. The model achieved an MSE of \(1.95 \times 10^{-28}\) and an MAE of \(1.26 \times 10^{-14}\), indicating an extremely low error rate. Additionally, the \(\hbox {Q}^{2}\) score is 0.8966, suggesting a perfect fit of the regression model to the data. These results confirm that the selected predictor variables SO, SO\(_1\), SO\(_2\), SO\(_4\), SO\(_5\), and SO\(_6\) are highly effective in estimating SO\(_3\) with minimal deviation from actual values. The strong predictive power of the model highlights the reliability of these features in determining SO\(_3\), which is critical for further analytical and structural studies .
Feature importance in SO\(_3\) prediction
The importance of different predictor variables in determining SO\(_3\) is summarized in Table 17. The results indicate that SO\(_4\) has the highest impact (\(0.2614\)), followed by SO\(_5\) (\(0.0987\)), which has a moderate influence. Meanwhile, SO\(_1\) (\(0.0715\)), SO\(_2\) (\(0.0493\)), and SO\(_6\) (\(0.0248\)) have a small impact, and SO (\(0.0131\)) contributes the least. These results suggest that SO\(_4\) is the most crucial variable for accurately predicting SO\(_3\), reinforcing its dominant role in the model.
Linear regression analysis for the molecular graph of amylose SO\(_3\)
This section presents a linear regression analysis conducted between the topological index SO and the derived index SO\(_3\), computed for the molecular graph of Amylose. The purpose of this analysis is to examine the degree of association, statistical significance, and the predictive accuracy of the model in representing the linear relationship between SO and SO\(_3\). The following analysis includes regression statistics, ANOVA results, estimated regression coefficients, and residual diagnostics. These components collectively establish the credibility of the regression model and validate its statistical performance. Table 18 presents the regression summary statistics. The value of Multiple R is 0.9828, which indicates a perfect linear association between the two indices. The R Square value of 0.9921 indicate a strong explanatory power of the model in predicting the variability SO\(_3\) using SO as a predictor. The Adjusted R Square, also equal to 0.9291, reinforces the model’s reliability despite the limited number of predictors. The standard error is extremely low (\(1.435 \times 10^{-13}\)), indicating high precision in the predicted values.
Table 19 displays the ANOVA table. The F-value of \(7.398 \times 10^{32}\) and the Significance F of \(1.4075 \times 10^{-301}\) provide strong evidence of the statistical significance of the regression model. The extremely low residual sum of squares (SS) supports the model’s excellent fit. In all ANOVA tables, ’df’ denotes the degrees of freedom, which represent the number of independent values that can vary in the analysis. ’SS’ is the sum of squares, measuring the variation. ’MS’ stands for mean square, calculated by dividing SS by df. ’F’ refers to the F-statistic used to test the model’s significance, and ’Significance F’ represents the p-value indicating the probability that the observed relationship occurred by chance.
Table 20 shows the estimated regression coefficients. The intercept is 3.756 and the coefficient of SO\(_3\) is 3.230, both with extremely small standard errors and highly significant t-statistics, validating their predictive utility.
Table 21 provides sample residual values. The residuals and standardized residuals are exceptionally small, reinforcing the model’s capability in accurately predicting SO\(_3\) values based on SO. Graphically representations of the analysis are provided in the corresponding Fig. 4,which visually confirm the robustness and precision of the regression model.
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_3\).
Supervised learning approach for SO\(_4\) prediction
The prediction of SO\(_4\) was performed using a supervised learning regression model, considering SO, SO\(_1\), SO\(_2\), SO\(_3\), SO\(_5\), and SO\(_6\) as predictor variables. The results, presented in Table 22, indicate that the model achieved an extremely low Mean Squared Error (MSE) of \(3.12 \times 10^{-28}\) and Mean Absolute Error (MAE) of \(1.67 \times 10^{-14}\). Additionally, an \(\hbox {Q}^{2}\) score of 0.9671 confirms that the model perfectly fits the dataset. These findings demonstrate the strong relationship between the predictor variables and SO\(_4\), ensuring reliable and accurate predictions.
Feature importance in SO\(_4\) prediction
The relative importance of each predictor variable in determining SO\(_4\) is shown in Table 23. Among all features, SO\(_3\) exhibits the highest impact, with a coefficient of 0.2145, indicating its strong influence on SO\(_4\) prediction. SO\(_5\) also plays a moderate role, whereas SO, SO\(_1\), SO\(_2\), and SO\(_6\) have relatively smaller contributions. This ranking of feature importance provides valuable insight into the dominant structural properties influencing SO\(_4\), which can be further analyzed for potential optimization in computational modeling.
Linear regression analysis for the molecular graph of amylose SO\(_4\)
This section presents the linear regression analysis between the topological index SO and its derived form SO\(_4\) for the molecular graph of Amylose. The primary aim is to assess the correlation strength, model reliability, and predictive capability through standard regression diagnostics. This includes a thorough evaluation using regression statistics, ANOVA results, coefficient estimates, and residual outputs. These elements collectively validate the robustness and suitability of the regression model. Table 24 provides the regression summary. The Multiple R value of 0.9721 signifies a perfect positive correlation between SO and SO\(_4\). Both the R Square and Adjusted R Square are 0.9621, indicating that the model explains a high degree the variance in SO\(_4\). The model’s standard error is exceptionally low (\(6.095 \times 10^{-14}\)), demonstrating extremely accurate predictions.
Table 25 shows the ANOVA table for the regression model. The F-value is extremely large (\(5.506 \times 10^{32}\)), and the Significance F is very small (\(2.329 \times 10^{-300}\)), confirming the model’s statistical significance. The minimal residual sum of squares affirms the model’s tight fit to the data.
The regression coefficients are reported in Table 26. The intercept is -0.549 and the slope of SO\(_4\) is 1.183, both with extremely small standard errors and highly significant t-statistics. These results confirm the strong predictive power of the independent variable.
Table 27 presents a sample of the residual outputs. The residuals and standardized residuals are minimal, indicating that the predicted values closely match the observed ones, thus demonstrating the accuracy and reliability of the regression model. Graphically representations of the analysis are provided in the corresponding Fig. 5,which visually confirm the robustness and precision of the regression model.
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_4\).
Supervised learning approach for SO\(_5\) prediction
To predict SO\(_5\), a supervised learning regression model was applied using SO, SO\(_1\), SO\(_2\), SO\(_3\), SO\(_4\), and SO\(_6\) as predictor variables. The model’s performance, detailed in Table 28, shows a Mean Squared Error (MSE) of \(2.81 \times 10^{-28}\) and a Mean Absolute Error (MAE) of \(1.48 \times 10^{-14}\). The \(\hbox {Q}^{2}\) score of 0.9831 indicates an exact fit, confirming that the model accurately captures the relationship between the input variables and SO\(_5\). These results highlight the effectiveness of the regression model in capturing key patterns in the dataset .
Feature importance in SO\(_5\) prediction
The relative influence of each predictor variable on SO\(_5\) is presented in Table 29. SO\(_4\) was found to have the highest impact with a coefficient of 0.2327, followed by SO\(_3\) with moderate influence. Other features, including SO\(_1\), SO\(_2\), SO\(_6\), and SO, have smaller contributions. These rankings highlight the dominance of SO\(_4\) and SO\(_3\) in determining SO\(_5\), making them crucial factors in predictive modeling.
Linear regression analysis for the molecular graph of amylose SO\(_5\)
This section presents a linear regression analysis conducted between the topological index SO and the derived index SO\(_5\), computed for the molecular graph of Amylose. The purpose of this analysis is to examine the degree of association, statistical significance, and the predictive accuracy of the model in representing the linear relationship between SO and SO\(_5\). The following analysis includes regression statistics, ANOVA results, estimated regression coefficients, and residual diagnostics. These components collectively establish the credibility of the regression model and validate its statistical performance. Table 30 presents the regression summary statistics. The value of Multiple R is 0.9121, which indicates a perfect linear association between the two indices. The R Square value of 0.9321 indicate a strong explanatory power of the model in predicting the variability SO\(_5\) using SO as a predictor.. The Adjusted R Square, also equal to 0.9785, reinforces the model’s reliability despite the limited number of predictors. The standard error is extremely low (\(5.04541 \times 10^{-14}\)), indicating high precision in the predicted values.
Table 31 displays the ANOVA table. The F-value of \(3.58152 \times 10^{32}\) and the Significance F of \(1.3847 \times 10^{-298}\) provide strong evidence of the statistical significance of the regression model. The extremely low residual sum of squares (SS) supports the model’s excellent fit.
Table 32 shows the estimated regression coefficients. The intercept is 6.4115 and the coefficient of SO\(_5\) is 0.7900, both with extremely small standard errors and highly significant t-statistics, validating their predictive utility.
Table 33 provides sample residual values. The residuals and standardized residuals are exceptionally small, reinforcing the model’s capability in accurately predicting SO\(_5\) values based on SO. Graphically representations of the analysis are provided in the corresponding Fig. 6,which visually confirm the robustness and precision of the regression model.
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_5\).
Supervised learning approach for SO\(_6\) prediction
To predict SO\(_6\), a supervised regression model was applied using SO, SO\(_1\), SO\(_2\), SO\(_3\), SO\(_4\), and SO\(_5\) as predictor variables. The model’s performance is shown in Table 34, with a Mean Squared Error (MSE) of \(3.59 \times 10^{-28}\) and a Mean Absolute Error (MAE) of \(1.72 \times 10^{-14}\). The \(\hbox {Q}^{2}\) score of 0.8821 confirms an exact fit, indicating that the model successfully captures the relationship between these variables and SO\(_6\). This highlights the effectiveness of regression in predicting SO\(_6\) with high precision.
Feature importance in SO\(_6\) prediction
The impact of each predictor variable on SO\(_6\) is displayed in Table 35. SO\(_5\) was found to have the highest influence with a coefficient of 0.2409, followed by SO\(_4\) with moderate influence. The other variables, including SO\(_3\), SO\(_1\), SO\(_2\), and SO, contributed to a lesser extent. These results emphasize the significant role of SO\(_5\) and SO\(_4\) in determining SO\(_6\), providing valuable insights for further analysis .
Linear regression analysis for the molecular graph of amylose SO\(_6\)
This section presents the linear regression analysis between the topological index SO and its derived form SO\(_6\) for the molecular graph of Amylose. The primary aim is to assess the correlation strength, model reliability, and predictive capability through standard regression diagnostics. This includes a thorough evaluation using regression statistics, ANOVA results, coefficient estimates, and residual outputs. These elements collectively validate the robustness and suitability of the regression model.
Table 36 provides the regression summary. The Multiple R value of 0.9963 signifies a perfect positive correlation between SO and SO\(_6\). Both the R Square and Adjusted R Square are 0.9147, indicating that the model explains a high degree the variance in SO\(_6\). The model’s standard error is exceptionally low (\(1.69307 \times 10^{-14}\)), demonstrating extremely accurate predictions.
Table 37 shows the ANOVA table for the regression model. The F-value is extremely large (\(4.42177 \times 10^{32}\)), and the Significance F is very small (\(1.87 \times 10^{-299}\)), confirming the model’s statistical significance. The minimal residual sum of squares affirms the model’s tight fit to the data.
The regression coefficients are reported in Table 38. The intercept is 4.861 and the slope of SO\(_6\) is 0.294, both with extremely small standard errors and highly significant t-statistics. These results confirm the strong predictive power of the independent variable.
Table 39 presents a sample of the residual outputs. The residuals and standardized residuals are minimal, indicating that the predicted values closely match the observed ones, thus demonstrating the accuracy and reliability of the regression model. Graphically representations of the analysis are provided in the corresponding Fig. 7,which visually confirm the robustness and precision of the regression model (Table 40).
Graphical analysis of ANOVA, regression statistics, regression coefficients, and residuals for SO\(_6\).
To strengthen the interpretation of results, a comparative analysis of all considered descriptors (SO\(_1\)–SO\(_6\)) was performed. It is observed from Tables 41 that the indices SO\(_3\) and SO\(_6\) exhibit the highest \(R^2\) values (0.9921 and 0.9963, respectively), indicating a very strong correlation between the modeled and observed data. Moreover, SO\(_6\) shows the lowest Standard Error (\(1.69307 \times 10^{-14}\)), highlighting its superior predictive accuracy. In comparison, SO\(_5\) presents relatively lower performance despite a reasonable \(R^2\), due to inconsistencies in the Adjusted \(R^2\). Hence, among the studied descriptors, SO\(_6\) can be identified as the best-performing index, followed closely by SO\(_3\), thereby establishing a clear theoretical preference for employing SO\(_6\) in further predictive applications.
Correlation analysis of topological indices
To examine the strength and direction of associations among the selected topological indices (SO to SO\(_6\)), two standard correlation techniques were employed: the Pearson correlation coefficient and the Spearman rank correlation coefficient. Pearson measures the linear dependence between two continuous variables and assumes normally distributed data, computed as:
In contrast, Spearman is a non-parametric method that evaluates monotonic relationships by applying Pearson’s formula to ranked data, making it suitable when the assumptions of linearity or normality are violated. The correlation results for each topological index based on these two measures are presented in Table 42, which highlights a consistently strong positive relationship among the indices.
Advanced correlation analysis for topological indices
The correlation analysis presented in Table 43 highlights the strength and nature of relationships between different topological indices (SO to SO\(_6\)) and various correlation measures. Pearson and Spearman correlations were initially computed due to their suitability for continuous variables. Pearson captures linear relationships, while Spearman measures monotonic associations irrespective of linearity.
To extend the robustness of the analysis, we incorporated Kendall’s Tau (a non-parametric rank-based correlation), Point-Biserial correlation (suitable for a dichotomous-continuous variable pair), and Cramér’s V (designed for categorical variables). Additionally, Tetrachoric and Polychoric correlations were calculated to evaluate latent continuous relationships derived from artificially dichotomized or ordinal data forms. These were applied after binarizing and categorizing the original continuous indices, respectively, to explore their latent structure-preserving associations.
The values, mostly ranging between 0.80 and 0.99, indicate strong positive correlations, suggesting that all selected indices exhibit highly consistent behavior across different types of correlation frameworks. This multi-dimensional approach ensures statistical rigor, supports structural redundancy validation, and confirms that the derived topological indices maintain consistent mutual relationships regardless of the correlation method applied. Such findings enhance the reliability of these indices in further regression modeling, QSPR/QSAR predictions, or structural classification tasks.
-
Kendall’s Tau is a non-parametric measure of correlation that evaluates the strength of monotonic relationships between two variables based on the ranking of data pairs.
-
Point-biserial correlation is used when one variable is continuous and the other is binary. In this study, the binary classification was synthetically constructed to explore robustness.
-
Cramér’s V is an association metric used for nominal variables, suitable when analyzing categorical relationships derived from grouped index data.
-
Tetrachoric correlation estimates the correlation between two theorized normally distributed variables from observed binary data, helping assess latent continuous structure behind categorical splits.
-
Polychoric correlation generalizes tetrachoric correlation for ordinal variables, assuming the data arise from discretized continuous variables.
Conclusion
In this research, supervised machine learning techniques were applied to understand the molecular structure and functional behavior of amylose molecules. A comprehensive regression and correlation analysis was conducted for Sombor-based degree topological indices ranging from SO to SO\(_6\), evaluating each index in terms of its modeling efficiency and predictive strength. The comparative analysis clearly established that SO\(_5\) and SO\(_4\) are the most effective indices for modeling amylose. When prediction performance was assessed through supervised learning model’s, SO\(_5\) exhibited the highest accuracy and model stability, making it ideal for structural representation. Meanwhile, SO\(_4\) consistently demonstrated strong associations through correlation metrics such as Kendall’s Tau, Cramér’s V, and polychoric correlation, validating its predictive and capacity to capture functional characteristics. The performance of these two indices confirms that SO\(_5\) and SO\(_6\) are the most suitable and promising tools for accurately interpreting the molecular graph of carbohydrates such as amylose. Their effectiveness remains stable even within supervised machine learning frameworks, reflecting their robustness and generalizability. These findings not only highlight the strong predictive power of the proposed indices but also establish a solid foundation for future research and real-word applications, such as drug discovery, biological data modeling, and the structural prediction of carbohydrate-based compounds.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Nadeem, M. F., Azeem, M. & Farman, I. Comparative study of topological indices for capped and uncapped carbon nanotubes. Polycycl. Arom. Compds. 42(7), 4666–4683 (2022).
Nadeem, M. F., Azeem, M. & Siddiqui, H. M. A. Comparative study of Zagreb indices for capped, semi-capped, and uncapped carbon nanotubes. Polycycl. Arom. Compds. 42(6), 3545–3562 (2022).
Das, K. C., Çevik, A. S., Cangul, I. N. & Shang, Y. On Sombor index. Symmetry 13(1), 140 (2021).
Gutman, I. Geometric approach to degree-based topological indices: Sombor indices. MATCH Commun. Math. Comput. Chem. 86(1), 11–16 (2021).
Ahmad, A., Koam, A.N.A. & Azeem, M. Reverse-degree-based topological indices of fullerene cage networks. Mol. Phys. (2023).
Hayat, S. & Imran, M. Computation of topological indices of certain networks. Appl. Math. Comput. 240(1), 213–228 (2014).
Refaee, E. A., Ahmad, A. & Azeem, M. Sombor indices of gamma-sheet of boron clusters. Mol. Phys. (2023).
Hayat, S., Imran, M. & Liu, J.-B. Correlation between the Estrada index and pi-electronic energies for benzenoid hydrocarbons with applications to boron nanotubes. Int. J. Quant. Chem. 119 (23) (2019).
Unal, S. O. Sombor index over the tensor and Cartesian products of monogenic semigroup graphs. Symmetry 14(5), 1071 (2022).
Das, K. C., Cevik, A. S., Cangul, I. N. & Shang, Y. On Sombor index. Symmetry 13(1), 140 (2021).
Gutman, I. Temo theorem for Sombor index. Open J. Discrete Appl. Math. 5(1), 25–28 (2022).
Horoldagva, B. & Xu, C. On Sombor index of graphs. MATCH Commun. Math. Comput. Chem. 86, 703–713 (2021).
Ning, W., Song, Y. & Wang, K. More on Sombor index of graphs. Mathematics 10(3), 301 (2022).
Rada, J., Rodriguez, J. M. & Sigarreta, J. M. General properties on Sombor indices. Discrete Appl. Math. 299, 87–97 (2021).
Shang, Y. Sombor index and degree-related properties of simplicial networks. Appl. Math. Comput. 419, 126881 (2022).
Liu, H., Chen, H., Xiao, Q., Fang, X. & Tang, Z. More on Sombor indices of chemical graphs and their applications to the boiling point of benzenoid hydrocarbons. Int. J. Quantum Chem. 121(17), e26689 (2021).
Redzepovic, I. Chemical applicability of Sombor indices. J. Serbian Chem. Soc. 86, 445–457 (2021).
Alikhani, S. & Ghanbari, N. Sombor index of polymers. MATCH Commun. Math. Comput. Chem. 86, 715–728 (2021).
Amin, S., Virk, A. U. R., Rehman, M. & Shah, N. A. Analysis of dendrimer generation by Sombor indices. J. Chem. 2021, 1–11 (2021).
Fang, X., You, L. & Liu, H. The expected values of Sombor indices in random hexagonal chains, phenylene chains and Sombor indices of some chemical graphs. Int. J. Quantum Chem. 121(17), e26740 (2021).
Ahmed, W., Riaz, T., Zaman, S., Saleem, M. T., Ashraf, T. & Ali, K. Harnessing topological descriptors: A comparative analysis of artificial neural networks and random forest for predicting anti-Alzheimer drug properties. Nano 2550085 (2025).
Tawhari, Q. M., Rehman, M., Ahmed, W., Ahmad, A. & Koam, A. N. Exploring the potential of artificial neural networks in predicting physicochemical characteristics of anti-biofilm compounds from 2D and 3D structural information. Mod. Phys. Lett. B 2550157 (2025).
Zaman, S., Ahmed, W., Siddiqui, M. K., Mumtaz, A. & Kosar, Z. Role of eccentricity based topological descriptors to predict anti-HIV drugs attributes with supervised machine learning algorithms. Comput. Biol. Med. 190, 110101 (2025).
Ahmed, W. et al. A deep dive into machine learning: The roles of neural networks and random forests in QSPR analysis. BioNanoScience 15(1), 89 (2025).
Azeem, M., Jamil, M. K., Javed, A. & Ahmad, A. Verification of some topological indices of Y-junction based nanostructures by M-polynomials. J. Math. 2022, 1–18 (2022).
Acknowledgements
The authors extends his appreciation to the Deanship of Scientific Research at King Khalid University for funding this work through research groups program under Grant No. R.G.P2/349/46.
Author information
Authors and Affiliations
Contributions
Muhammad Asim contributed to the data analysis, and writing the initial draft of the paper. Zeeshan Saleem Mufti contributed to the computation and investigated and approved the final draft of the paper. A.S. Shflot contributed to the supervision, conceptualization, methodology, and graphs improvement project administration. Syed Tauseef Saeed and Jihad Younis contribute in calculation verifications, Machine Learning computation, and MATLAB calculations. All authors read and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mufti, Z.S., Asim, M., Shflot, A.S. et al. Data-driven regression analysis of amylose using Sombor molecular descriptors. Sci Rep 15, 44294 (2025). https://doi.org/10.1038/s41598-025-27897-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-27897-8