Abstract
Transfer RNA-derived small RNAs (tsRNAs) play crucial regulatory roles in tumour biology; however, their potential as biomarkers for colorectal cancer (CRC) remains underexplored. Plasma samples from 123 patients with CRC and 79 healthy controls (HCs) were collected for this study. Exosomes were extracted from plasma, validated, and tRF-3004a levels were detected using quantitative real-time polymerase chain reaction (qRT-PCR). The correlation between plasma-derived exosomal tRF-3004a expression levels and clinicopathological parameters was analysed using the chi-square test. Receiver operating characteristic (ROC) curve analysis was performed to evaluate the diagnostic performance of plasma-derived exosomal tRF-3004a. The results showed that compared with HCs, plasma-derived exosomal tRF-3004a was significantly elevated in patients with CRC and decreased after surgery. Moreover, high tRF-3004a expression was significantly associated with lymph node metastasis, tumour node-metastasis staging, carcinoembryonic antigen (CEA) levels, and nerve/vascular invasion in patients with CRC. ROC analysis revealed that plasma-derived exosomal tRF-3004a demonstrated promising diagnostic utility for CRC, with an area under the curve (AUC) of 0.819 (sensitivity, 0.691; specificity, 0.861). The combination of CEA and carbohydrate antigen 19 − 9 (CA19-9) levels increased the AUC to 0.867. The results of this study demonstrate that plasma-derived exosomal tRF-3004a may serve as a novel diagnostic biomarker for CRC.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is a prevalent malignancy worldwide, with a progressively increasing incidence in younger ages1. Moreover, CRC often manifests covertly, with approximately 30–40% of patients presenting with distant metastasis upon diagnosis, thereby missing the optimal treatment window2. Consequently, early detection and timely treatment of CRC are pivotal for halting tumour progression and enhancing patient prognosis. Although colonoscopy remains the preferred diagnostic method for CRC, its invasiveness, high cost, and poor patient compliance contribute to low screening rates and hinder the prompt identification of early stage cases3,4. Furthermore, commonly used CRC screening markers such as carcinoembryonic antigen (CEA) and carbohydrate antigen 19 − 9 (CA19-9) exhibit limited sensitivity and specificity, leading to misdiagnoses or missed diagnoses5,6. Therefore, a simple non-invasive biomarker to facilitate early CRC diagnosis is urgently needed.
Liquid biopsy, a novel concept that has emerged in recent years, differs from traditional tissue biopsy by detecting circulating tumour DNA (ctDNA), circulating tumour cells (CTCs), and exosomes in body fluids, such as blood, saliva, and urine7. This non-invasive, real-time technology provides comprehensive information about tumours, making it crucial for early cancer screening8,9. Exosomes are extracellular vesicles secreted by various cells, with diameters ranging from 30 to 150 nm. They carry diverse constituents, including DNA, RNA, and lipids, and are abundant in body fluids. Owing to their involvement in numerous physiological and pathological processes, exosomes are effective biomarkers for cancers10,11.
With the emergence of the post-genomic era, non-coding RNAs (ncRNAs) have gained increasing attention in humans. However, this research has primarily focused on long non-coding RNAs, microRNAs (miRNAs), and circular RNAs12,13. In recent years, transfer RNA (tRNA)-derived small RNAs (tsRNAs), previously considered random degradation products of tRNAs, have garnered attention owing to advancements in high-throughput RNA sequencing technology14. tsRNAs are a class of small ncRNA fragments cleaved at specific sites within pre-tRNAs or mature tRNAs. Depending on the cleavage site, tsRNAs can be categorised into two types: tRNA halves (tiRNAs) and tRNA-derived fragments (tRFs)15. tsRNAs play diverse roles in epigenetic, transcriptional, and translational regulation. Consequently, tsRNA mutations, modifications, and dysregulation contribute to various human diseases, including cancer development16. The tsRNA tRF3008A inhibits CRC cell proliferation and migration by targeting FOXK1 and regulating the Wnt pathway17. Furthermore, tsRNAs can be encapsulated within exosomes and are widely distributed throughout the body, where they play a crucial role in intercellular communication18. Dysregulation of plasma-derived exosomal tsRNAs promotes the progression of cancers, such as non-small cell lung cancer, gastric carcinoma, and hepatocellular carcinoma, and could serve as a novel liquid biopsy marker19,20,21. However, the expression and function of plasma-derived exosomal tsRNAs in CRC remain poorly understood.
In this study, we observed elevated levels of plasma-derived exosomal tRF-3004a in patients with CRC compared with healthy controls (HCs). Moreover, receiver operating characteristic (ROC) analysis demonstrated that tRF-3004a expression exhibited superior diagnostic efficacy compared to CEA and CA19-9 levels. The combination of these biomarkers further enhanced diagnostic accuracy. These findings suggest that plasma-derived exosomal tRF-3004a is a potential diagnostic biomarker for CRC.
Results
Basic information on tRF-3004a
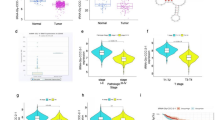
We analysed tRF and tiRNA sequence data from CRC and adjacent normal tissues using the tRF explorer database (https://trfexplorer.cloud/), with thresholds of |log2FC|>1.5 and P < 0.05. The analysis revealed significant upregulation of a novel CRC-associated tsRNA, tRF-3004a, in CRC tissues (log2FC = 2.75, P < 0.05) (Fig. 1A and B). tsRNAs are fragments generated through specific cleavage sites on tRNAs and can be classified as tiRNAs and tRFs based on these cleavage patterns. Among these, tiRNAs consist of 5’-tiRNA and 3’-tiRNA, while tRFs include tRF-1, tRF-3, tRF-5, tRF-2, and i-tRF (Fig. 1C). Using databases such as tRFdb (http://genome.bioch.virginia.edu/trfdb/) and GtRNAdb (http://gtrnadb.ucsc.edu/), we observed that tRF-3004a was a 17-nt fragment originating from tRNA-Gln-CTG-1-1 (TCTCGGTGGAACCTCCA) (Fig. 1D and E). Secondary analysis by Lu et al.22. revealed that plasma-derived exosomal tRF-3004a was upregulated in patients with CRC (log2FC = 3.16, P < 0.05). Therefore, we speculated that plasma-derived exosomal tRF-3004a could serve as a potential liquid biopsy biomarker for CRC.
Features of tRF-3004a. (A) Volcano plots of tsRNA expression profiles in CRC and adjacent normal tissues based on the tRF explorer database. (B) Expression levels of tRF-3004a in CRC and normal tissues based on the tRF explorer database. (C) Biogenesis and classification of tsRNAs. (D) tRF-3004a originates from tRNA-Gln-CTG-1-1. (E) tRNA-Gln-CTG-1-1 information in the GtRNAdb database. CRC, colorectal cancer; tsRNA, tRNA-derived small RNA; tRF, tRNA-derived fragment; GtRNAdb, Genomic tRNA database.****P<0.0001.
Methodological evaluation of plasma-derived Exosomal tRF-3004a
Plasma exosomes were collected and isolated from patients with CRC and HCs using a precipitation reagent. Transmission electron microscopy (TEM) revealed cup-shaped structures 30–150 nm in diameter, consistent with typical exosome size and morphology (Fig. 2A). Nanoscale flow cytometry revealed a monodispersed particle population with a peak diameter of approximately 80 nm (Fig. 2B). Western blot analysis confirmed the presence of exosome-specific marker proteins (CD63 and TSG101) and the absence of the non-exosomal protein calnexin (Fig. 2C). To investigate the potential of tRF-3004a as a novel diagnostic biomarker for CRC, we comprehensively evaluated detection methods. The assay demonstrated specificity with smooth amplification curves and single-peak melting curves (Fig. 2D and E). Sanger sequencing confirmed the presence of the complete tRF-3004a sequence in quantitative real-time polymerase chain reaction (qRT-PCR) products (Fig. 2F).
Methodological evaluation of tRF-3004a. (A) Representative TEM image revealing the characteristic cup-shaped morphology of exosomes. Scale bar: 200 nm. (B) Exosome size distribution as determined using nanoscale flow cytometry. (C) Western blot analysis of exosome-specific protein markers (CD63 and TSG101) and the non-exosome marker calnexin. (D, E) Amplification and melting curves of tRF-3004a. (F) Confirmation of the qRT-PCR product of tRF-3004a using Sanger sequencing. TEM, transmission electron microscopy; qRT-PCR, quantitative real-time polymerase chain reaction; TSG101, tumour susceptibility gene 101.
Plasma-derived Exosomal tRF-3004a levels in patients with CRC and their correlation with clinical pathological parameters
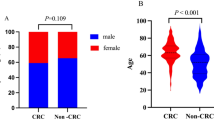
We assessed expression levels of plasma-derived exosomal tRF-3004a in plasma samples from a cohort of patients with CRC (n = 123) and HCs (n = 79). qRT-PCR analysis revealed significantly higher tRF-3004a expression levels in patients than in HCs (Fig. 3A). Subsequently, we investigated the correlation between tRF-3004a expression and the clinicopathological parameters in patients with CRC. Based on the median expression value of tRF-3004a, we categorised the 123 patients with CRC into high- (> 2.282, n = 62) and low- (≤ 2.282, n = 61) expression groups. We observed a significant positive association between tRF-3004a expression and lymph node metastasis (P = 0.008), tumour node-metastasis (TNM) stage (P = 0.005), nerve/vascular invasion (P < 0.001), and CEA levels (P = 0.036) but not with sex, age, tumour size, colon or rectal location, T stage, or CA19-9 levels (Table 1). These findings highlight the upregulation of plasma-derived exosomal tRF-3004a in patients with CRC and its association with disease progression.
Subsequently, we stratified patients based on clinicopathological parameters to further analyse the differential expression levels of tRF-3004a in each subgroup. We observed significantly higher expression levels of plasma-derived exosomal tRF-3004a in patients with CRC with lymph node metastasis than in those without lymph node metastasis (Fig. 3B). Moreover, patients with stage III–IV CRC exhibited significantly elevated expression levels of tRF-3004a compared with those in patients with stage I–II disease (Fig. 3C). Additionally, tRF-3004a levels were increased in patients with CRC with nerve/vascular invasion (Fig. 3D). Furthermore, postoperative plasma samples from 20 patients with CRC showed significantly reduced tRF-3004a levels after surgery, which ultimately returned to levels with no significant difference from those in HCs (P = 0.3016) (Fig. 3E and F). These findings suggest that plasma-derived exosomal tRF-3004a is a promising biomarker for CRC diagnosis.
Levels of plasma-derived exosomal tRF-3004a in patients with CRC. (A) Plasma-derived exosomal tRF-3004a expression levels in patients with CRC (n = 123) and HCs (n = 79). (B) Patients with CRC with (n = 55) and without lymph node metastasis (n = 68). (C) Patients with stage I–II (n = 67) and III–IV (n = 56) CRC. (D) Patients with CRC with (n = 62) and without nerve/vascular invasion (n = 61). (E) Before and after surgery in patients with CRC (n = 20). (F) Comparison of postoperative expression levels of plasma-derived exosomal tRF-3004a in patients with CRC compared with HCs. CRC, colorectal cancer; HCs, healthy controls; **P < 0.01; ***P < 0.001; ****P < 0.0001; ns, not significant; P-values were calculated using t-test, Mann–Whitney U test, and Wilcoxon signed-rank test..
Diagnostic significance of plasma-derived Exosomal tRF-3004a in CRC
To assess the diagnostic value of plasma-derived exosomal tRF-3004a for CRC, we conducted an ROC analysis of tRF-3004a, CEA, and CA19-9. The results demonstrated that the expression level of tRF-3004a effectively differentiated patients with CRC from HCs, yielding an area under the curve (AUC) of 0.819 (95% CI 0.763–0.875) (Fig. 4A). This AUC was more than that of CEA at 0.681 (95% CI 0.608–0.754) and CA19-9 at 0.658 (95% CI 0.584–0.732). Moreover, when these biomarkers (tRF-3004a, CEA, and CA19-9) were combined for diagnosis, the AUC reached a peak value of 0.867 (95% CI, 0.819–0.916) (Fig. 4B). The optimal cutoff value for this combination was 0.703, with a sensitivity of 0.707 and specificity of 0.987 (Table 2). Plasma-derived exosomal tRF-3004a represents a novel biomarker for CRC diagnosis, and its integration with other established CRC markers can significantly enhance diagnostic efficiency.
Diagnostic value of plasma-derived exosomal tRF-3004a in patients with CRC. (A) Diagnostic value of plasma-derived exosomal tRF-3004a in patients with CRC. (B) ROC curves for tRF-3004a, CEA, and CA19-9, independently or combined, for differentiating patients with CRC from HCs. ROC, receiver operating characteristic; CEA, carcinoembryonic antigen; CA19-9, carbohydrate antigen 19 − 9.
Downstream prediction of plasma-derived Exosomal tRF-3004a
To investigate the potential molecular mechanisms underlying CRC progression, we used tsRNA target prediction tools (tRFtarget and tRFtar) to identify the putative genes regulated by tRF-3004a. Our analysis revealed 82 common target genes between the databases (Fig. 5A). Subsequent Gene Ontology (GO) functional enrichment analysis revealed that these target genes were implicated in diverse biological processes, including viral processes, cytokine-mediated signaling pathways, transcription regulator complexes, membrane microdomains, transcriptional co-regulator activity, and DNA-binding transcription factor binding (Fig. 5B). Furthermore, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis demonstrated that the identified target genes of tRF-3004a were predominantly enriched in crucial pathways in cancer, apoptosis, and human papillomavirus (Fig. 5C).
Bioinformatic analysis of identified tRF-3004a. (A) Potential target genes. (B) GO enrichment analysis of potential target genes. (C) KEGG pathway analysis of potential target genes23. GO, Gene Ontology; KEGG, Kyoto Encyclopedia of Genes and Genomes.
Discussion
CRC is among the most prevalent malignant tumours worldwide, ranking third in incidence and second in mortality among all malignancies. Therefore, screening and early standardized diagnosis and treatment are critical for improving the prognosis of patients with CRC. Although colonoscopy is considered the gold standard for CRC screening, its associated risks, such as bleeding and perforation, often result in low patient compliance24. Commonly used laboratory tests in clinical practice, including fecal immunochemical tests, CEA, and CA19-9, exhibit significantly inadequate sensitivity and specificity25. Thus, an urgent need exists to identify potential biomarkers and optimise their combinations. Therefore, identifying efficient, convenient, and stable biomarkers is of paramount importance to facilitate early CRC detection and treatment.
Liquid biopsy based on circulating molecular markers has greatly expanded the scope of early CRC diagnosis, enabling the detection of ctDNA, CTC, and exosomes in body fluids such as blood. The advantages of this approach include rapidity, minimal invasiveness, and high patient compliance; moreover, the significance of early tumour diagnosis and prognosis assessment is increasingly recognized26. Exosomes carry various components, including DNA, RNA, and lipids, which are abundant in body fluids and participate in numerous physiological and pathological processes. Therefore, exosomes are promising novel biomarkers for liquid biopsy27,28.
tsRNAs have recently emerged as promising biomarkers. These small RNA fragments, derived from precursors or mature tRNAs, can be stably enriched in body fluids within exosomes. Compared with miRNA-based exosomal biomarkers, tsRNAs have distinct advantages29,30,31. They are generated through tRNA-specific cleavage rather than the canonical miRNA biogenesis pathway, thus conferring greater tissue specificity. Their abundant nucleotide modifications enhance nuclease resistance, leading to superior stability in biofluids, and they display versatile regulatory mechanisms capable of miRNA-like target regulation and direct intervention in translation and epigenetic processes32,33,34. These characteristics suggest that tsRNAs are promising candidates for biomarker development35,36. Cao et al.37 demonstrated an association between tRF-19-PNR8YPJZ expression and lymph node metastasis, as well as clinical staging in patients with pancreatic cancer, suggesting the diagnostic potential of this tsRNA in pancreatic cancer. However, despite the promising prospects of liquid biopsy technology based on tsRNAs, research evaluating the clinical detection and diagnostic value of plasma-derived exosomal tsRNAs is lacking.
In the current study, we observed a significant increase in plasma-derived exosomal tRF-3004a levels among patients with CRC. Moreover, increased expression of tRF-3004a was significantly correlated with various clinicopathological features, including lymph node metastasis, TNM staging, CEA levels, and nerve/vascular invasion in patients with CRC. Additionally, postoperative analysis revealed a substantial reduction in tRF-3004a levels among these patients, indicating its potential as a predictive marker of treatment outcomes.
Subsequent assessment of the diagnostic potential of plasma-derived exosomal tRF-3004a revealed an AUC of 0.819, with sensitivity and specificity values of 0.691 and 0.861, respectively, outperforming commonly used clinical markers such as CEA (AUC: 0.681) and CA19-9 (AUC: 0.658). The combination of CEA and CA19-9 levels with tRF-3004a further improved the AUC to 0.867. These associations with specific clinicopathological features support the promise of plasma-derived exosomal tRF-3004a as a potential biomarker for CRC diagnosis.
An increasing body of evidence has demonstrated the pivotal role of tsRNAs in tumourigenesis and cancer progression through their effects on epigenetic, transcriptional, and translational regulation. For instance, tRF-Glu49 suppresses cervical cancer cell proliferation by targeting fibrinogen-like protein 1, while overexpression of 5’-tRF-GlyGCC inhibits autophagy and promotes breast cancer metastasis38,39. These findings underscore the biological significance of tsRNAs. In this study, we used the GO and KEGG databases to predict the downstream target genes of tRF-3004a, revealing its involvement in transcriptional regulation, apoptosis, and carcinogenesis. Our results provide insight into the underlying mechanism of action of tRF-3004a in CRC progression.
However, this study has certain limitations. First, the single-centre design and relatively small postoperative cohort, coupled with imbalanced sample sizes across certain clinical subgroups (particularly in tumour differentiation grades) and the dichotomization of continuous variables, may have affected the statistical power and generalisability of the findings. Future prospective multicentre studies are needed to ensure adequate recruitment across all differentiation subgroups, thereby enhancing the robustness of the conclusions. Second, the clinical translation of tsRNAs faces substantial challenges, including the need for standardised protocols for exosome isolation and tsRNA detection to ensure reproducibility. Third, mechanistic insights derived from bioinformatic predictions remain preliminary and require further validation through in vitro and in vivo functional experiments to elucidate underlying molecular mechanisms.
The results of this study revealed significantly elevated tRF-3004a expression in patients with CRC, which decreased following surgery. Our assessment of the clinical utility of plasma-derived exosomal tRF-3004a suggests its potential as an innovative liquid biopsy biomarker for real-time monitoring and aiding in the early diagnosis and treatment of CRC.
Conclusion
Plasma-derived exosomal tRF-3004a exhibits potential as a biomarker for early CRC diagnosis, surpassing the diagnostic efficacy of commonly used clinical biomarkers, such as CEA and CA19-9. Furthermore, their combined utilisation enhances diagnostic accuracy.
Materials and methods
Clinical samples
Blood samples were collected in ethylenediaminetetraacetic acid anticoagulant at the Department of Clinical Laboratory at Xiangya Hospital, Central South University, between January 2023 and October 2024. The sample cohort comprised 123 preoperative patients with CRC, 20 postoperative patients with CRC, and 79 HCs. The diagnosis of CRC was confirmed through tumour tissue pathology reports, and tumour staging was determined according to the TNM system. HCs were selected to match the CRC group in terms of age and sex and had no history of gastrointestinal disorders, malignancies, or chronic systemic diseases, and normal results on routine physical and laboratory examinations. Following collection, venous blood samples were processed within a strict 2 h window. A standardised two-step centrifugation was used: an initial step at 2,000 × g for 10 min to obtain cell-free plasma, followed by a second centrifugation of the supernatant at 10,000 × g for 30 min at 4 °C to remove cellular debris and platelets. The final clarified supernatant was aliquoted and stored at −80 °C for experimental analyses. This study adhered to the principles of the Declaration of Helsinki and was approved by the Ethics Committee of Xiangya Hospital, Central South University (No. 2024091165). Written informed consent was obtained from all the participants.
Exosome isolation
Plasma exosomes were isolated using a precipitation reagent (Yeasen, China) specifically designed for plasma or serum, following the manufacturer’s instructions. In brief, 1 mL of plasma sample was centrifuged at 3000 × g for 10 min at 4℃ to obtain the supernatant, which was then transferred to a new centrifuge tube. The clarified plasma sample was subsequently mixed with four times the volume of phosphate-buffered saline (PBS) and an equal volume of exosome isolation reagent in a clean tube. After incubating the mixture at 4℃ for 2 h, it was centrifuged at 10,000 × g for 1 h, and the resulting supernatant was discarded. The pellet containing exosomes was resuspended in PBS by vortexing. Finally, the enriched exosomes were obtained via centrifugation at 12,000 × g for 2 min at 4℃ for further investigation.
Exosome identification
Morphological examination of the exosomes was performed using TEM. A 10 µL aliquot of plasma-derived exosomes was applied to carbon-coated copper grids and allowed to adsorb for 4 min. Excess liquid was then removed using filter paper, followed by negative staining with 2% phosphotungstic acid for 2 min. After removing residual stain with filter paper, the grids were air-dried at room temperature. Imaging was performed using an HT7700 TEM system (Hitachi Ltd., Japan), with the diameters of 100 individual exosomes measured. The size distribution of the plasma-derived exosomes was analysed using nanoscale flow cytometry. Following instrument calibration and performance verification using standardised nanoparticles, a 10 µL sample was analysed using an N30E nanoparticle analyser (NanoFCM Inc., China). Protein characterisation was performed using western blotting to detect exosome-specific markers and the negative control, calnexin. The antibodies included CD63 (Abclonal, Cat No. A19023), TSG101 (Abcam, Cat No. ab125011), and calnexin (SAB, Cat No. 12186).
RNA extraction and qRT-PCR
Total RNA was extracted from plasma exosomes using a commercial kit (Tiangen, China). cDNA was synthesised using the miRNA 1 st Strand cDNA Synthesis Kit (Stem-loop) (Accurate Biotechnology (Hunan) Co., Ltd., China). Expression levels were quantified using an ABI QuantStudio 5 system (Thermo Fisher Scientific, USA) with a SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biotechnology (Hunan) Co., Ltd., China). Finally, tRF-3004a expression levels were measured relative to cel-miR-39 as an external reference and calculated using the 2−ΔΔCT method. All primers used in this study were synthesised by Accurate Biotechnology (Accurate Biotechnology Co., Ltd.; Hunan, China).
Statistical analysis
Statistical analysis was conducted using IBM SPSS Statistics for Windows, version 23.0, and GraphPad Prism 9.5. Between-group variable comparisons were performed using Student’s t-test, Mann–Whitney U test, and Wilcoxon signed-rank test. The correlation between tRF-3004a expression levels and clinicopathological parameters was analysed using the chi-square test. We assessed the diagnostic performance of plasma-derived exosomal tRF-3004a for CRC by calculating the AUC. Statistical significance was defined as a two-sided P < 0.05.
Data availability
The datasets generated during the current study are available from the corresponding author on reasonable request.
References
Siegel, R. L., Giaquinto, A. N. & Jemal, A. Cancer statistics, 2024. Ca-Cancer J. Clin. 74, 12–49 (2024).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis a single-cell level. Cancer Discov. 12, 134–153 (2022).
Chung, J. et al. Recent technologies towards diagnostic and therapeutic applications of Circulating nucleic acids in colorectal cancers. Int J. Mol Sci. 25, 8703 (2024).
Zhang, Z. et al. Identification of faecal extracellular vesicles as novel biomarkers for the non-invasive diagnosis and prognosis of colorectal cancer. J. Extracell. Vesicles. 12, e12300 (2023).
Sun, J. et al. Plasma proteomic and polygenic profiling improve risk stratification and personalized screening for colorectal cancer. Nat. Commun. 15, 8873 (2024).
Huang, Z. et al. Identification and validation of the surface proteins FIBG, PDGF-beta, and TGF-beta on serum extracellular vesicles for non-invasive detection of colorectal cancer, experimental study. Int. J. Surg. 110, 4672–4687 (2024).
Xie, W. et al. Trends in the use of liquid biopsy in oncology. Nat. Rev. Drug Discov. 22, 612–613 (2023).
Loy, C. et al. Liquid biopsy based on Cell-Free DNA and RNA. Annu. Rev. Biomed. Eng. 26, 169–195 (2024).
Cescon, D. W. et al. Circulating tumor DNA and liquid biopsy in oncology. Nat. Cancer. 1, 276–290 (2020).
Zhou, M. et al. Exosome derived from tumor-associated macrophages, biogenesis, functions, and therapeutic implications in human cancers. Biomark. Res. 11, 100 (2023).
Wang, S. et al. Exosomal circrnas as novel cancer biomarkers, challenges and opportunities. Int. J. Biol. Sci. 17, 562–573 (2021).
Antonatos, C. et al. Disentangling the complexity of psoriasis in the post-genome-wide association era. Genes Immun. 24, 236–247 (2023).
Hashemi, M. et al. Emerging roles of CircRNA-miRNA networks in cancer development and therapeutic response. Non-Coding Rna Res. 10, 98–115 (2024).
Wu, D. et al. tRNA modifications and tRNA-derived small RNAs, new insights of tRNA in human disease. Cell. Biol. Toxicol. 40, 76 (2024).
Chen, Y., Shao, Z. & Wu, S. Research progress on the TsRNA biogenesis, function, and application in lung cancer. Non-Coding Rna Res. 10, 63–69 (2024).
Wang, Q. et al. Exploring the role of tRNA-derived small RNAs (tsRNAs) in disease, implications for HIF-1 pathway modulation. J. Mol. Med. 102, 973–985 (2024).
Han, Y. et al. tRF3008A suppresses the progression and metastasis of colorectal cancer by destabilizing FOXK1 in an AGO-dependent manner. J. Exp. Clin. Canc Res. 41, 32 (2022).
Lee, S. et al. Emerging roles of tRNA-derived small RNAs in cancer biology. Exp. Mol. Med. 55, 1293–1304 (2023).
Xiao, L. et al. Disorders and roles of tsRNA, snoRNA, SnRNA and PiRNA in cancer. J. Med. Genet. 59, 623–631 (2022).
Lucien, F. et al. MIBlood-EV: minimal information to enhance the quality and reproducibility of blood extracellular vesicle research. J. Extracell. Vesicles. 12, e12385 (2023).
Wang, J. et al. Plasma tRNA fragments derived from 5’ ends as novel diagnostic biomarkers for early-stage breast cancer. Mol. Ther-Nucl Acids. 21, 954–964 (2020).
Lu, S. et al. A novel tRNA-derived fragment tRF-3022b modulates cell apoptosis and M2 macrophage polarization via binding to cytokines in colorectal cancer. J. Hematol. Oncol. 15, 176 (2022).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res 53, D672–D677 (2025).
Lin, J. S. et al. Screening for colorectal cancer, updated evidence report and systematic review for the US preventive services task force. Jama-J Am. Med. Assoc. 325, 1978–1998 (2021).
Qu, R. et al. Increasing burden of colorectal cancer in China. Lancet Gastroenterol. 7, 700 (2022).
Nikanjam, M., Kato, S. & Kurzrock, R. Liquid biopsy, current technology and clinical applications. J. Hematol. Oncol. 15, 131 (2022).
Min, L. et al. Advanced nanotechnologies for extracellular vesicle-based liquid biopsy. Adv. Sci. (Weinh). 8, e2102789 (2021).
Kalluri, R. & McAndrews, K. M. The role of extracellular vesicles in cancer. Cell 186, 1610–1626 (2023).
Sekar, D., Selvakumar, S. C. & Auxzilia, P. K. Therapeutic nature of MicroRNAs in oral squamous cell carcinoma (OSCC). Oral Oncol. 134, 106106 (2022).
Min, L. et al. Circulating small extracellular vesicle RNA profiling for the detection of T1a stage colorectal cancer and precancerous advanced adenoma. Elife 12, RP88675 (2024).
Min, L. et al. Loss of Circulating Exosomal miR-92b is a novel biomarker of colorectal cancer at early stage. Int. J. Med. Sci. 16, 1231–1237 (2019).
Zhou, M. et al. tRNA-derived small RNAs in human cancers, roles, mechanisms, and clinical application. Mol. Cancer. 23, 76 (2024).
Sekar, D. Implications of MicroRNA 21 and its involvement in the treatment of different type of arthritis. Mol. Cell. Biochem. 476, 941–947 (2021).
Min, L. et al. Evaluation of Circulating small extracellular vesicles derived MiRNAs as biomarkers of early colon cancer, a comparison with plasma total MiRNAs. J. Extracell. Vesicles. 8, 1643670 (2019).
Selvakumar, S. C. et al. The emerging role of microRNA-based therapeutics in the treatment of preeclampsia. Placenta 158, 38–47 (2024).
Weng, Q. et al. Extracellular vesicles-associated tRNA-derived fragments (tRFs), biogenesis, biological functions, and their role as potential biomarkers in human diseases. J. Mol. Med. 100, 679–695 (2022).
Cao, W. et al. Pancreatic stellate cell-derived Exosomal tRF-19-PNR8YPJZ promotes proliferation and mobility of pancreatic cancer through AXIN2. J. Cell. Mol. Med. 27, 2533–2546 (2023).
Wang, Y. et al. tRNA-derived fragment tRF-Glu49 inhibits cell proliferation, migration and invasion in cervical cancer by targeting FGL1. Oncol. Lett. 24, 334 (2022).
Chen, F. et al. 5’-tRF-GlyGCC promotes breast cancer metastasis by increasing fat mass and obesity-associated protein demethylase activity. Int. J. Biol. Macromol. 226, 397–409 (2023).
Acknowledgements
The authors would like to thank KEGG platform for providing their platform and contributors for uploading their meaningful datasets.
Funding
This work was supported by the National Natural Science Foundation of China (82373062), the Natural Science Foundation of Hunan Province (2022JJ30987), Hunan Province Key Research and Development Project (2024JK2107), Hunan Innovative Province Construction Science Popularization Project (2021ZK4221), and the Central South University Innovation-Driven Research Programme (2023CXQD075).
Author information
Authors and Affiliations
Contributions
MLZ: Data curation; methodology; software; writing-original draft. XQY: Data curation; methodology. JZ: Data curation; methodology. ZXH: Data curation. SSF: Data curation. XMX: Data curation. CLO: Conceptualization; funding acquisition; supervision; writing-review and editing. BYZ: Resources; funding acquisition; supervision; writing-review and editing. The author has accepted responsibility for the entire content of this manuscript and approved its submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This study has been reviewed and approved by the Ethics Committee of Xiangya Hospital, Central South University (No. 2024091165). This study was conducted in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Chunlin Ou and Baiyun Zhong have jointly supervised this work.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, M., Yu, X., Zhang, J. et al. Plasma-derived exosomal tRF-3004a as a diagnostic biomarker for colorectal cancer. Sci Rep 15, 45558 (2025). https://doi.org/10.1038/s41598-025-30113-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-30113-2