Abstract
The modified Weibull model (MWM) is one of the type-2 Weibull distributions that can be used for modeling lifetime data. It is important due to its simplicity and flexibility of the failure rate, and ease of parameter estimation using the least squares method. In this study, we introduce novel methods for estimating the parameters in step-stress partially accelerated life testing (SSPALT) in the context of progressive Type-II censoring (PT-II) under Constant-Barrier Removals (CBRs) for the MWM. We conduct a comparative analysis between Expectation Maximization (EM) and Stochastic Expectation Maximization (SEM) techniques with Bayes estimators under Markov Chain Monte Carlo (MCMC) methods. Specifically, we focus on Replica Exchange MCMC, the Hamiltonian Monte Carlo (HMC) algorithm, and the Riemann Manifold Hamiltonian Monte Carlo (RMHMC), emphasizing the use of the Linear Exponential (LINEX) loss function. Additionally, highest posterior density (HPD) intervals derived from the RMHMC sampler consistently outperform asymptotic and bootstrap confidence intervals, providing the shortest credible regions while maintaining nominal coverage across all censoring levels and stress conditions. A comprehensive Monte Carlo simulation study is conducted to assess the performance of these methods. Furthermore, the proposed methodology is applied to a real dataset comprising lifetimes of electrical appliances, demonstrating the practical effectiveness of the MWM in modeling real-world reliability data. Results show that the Bayesian RMHMC approach offers superior accuracy and convergence properties.
Similar content being viewed by others
Introduction
Statistical distributions are fundamental for modeling real-world phenomena, providing a structured framework for understanding variation, estimating probabilities, and forecasting future events. Their ability to accurately represent underlying patterns is crucial for informed decision-making and optimizing data-driven solutions in fields such as engineering, healthcare, and finance. In reliability engineering, in particular, accurately modeling failure data is crucial for understanding system behavior and predicting failures.
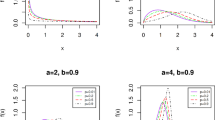
The Modified Weibull Model (MWM), developed by Lai et al.1, is a highly flexible distribution prized for its capacity to model various failure rate shapes, including the bathtub-shaped hazard functions (HF) commonly observed in industrial systems, biomedical research, and engineering applications2. The Cumulative Distribution Function (CDF) and Probability Density Function (PDF) of the MWM, characterized by parameters \(\beta\), \(\zeta\), and \(\phi\), are expressed as follows:
where \(\beta\) is the scale parameter and \(\zeta\), \(\phi\) are the shape parameters. The corresponding reliability function (RF) and HF are respectively derived as
To efficiently collect life-testing data, Step-Stress Partially Accelerated Life Testing (SSPALT) has been widely adopted. This methodology has been explored by numerous researchers, including notable works by Goel3, Bhattacharyya and Soejoeti4, Degroot and Goel5, Bai and Chung6, Bai et al.7, Abdel-Ghani8, Tse et al.9, Xiong and Ji10, Al-Babtain et al.11, Tahani et al.12, Asadi et al.13, Ismail14, Anwar et al.15, Lone et al.16, Lone et al.17, and Abushal and Soliman18. The MWM itself was employed for estimation under SSPALT by Ramzan et al.2, while Ramzan et al.19 proposed an extended version of the Weibull distribution with six parameters.
Nevertheless, the Bayesian paradigm has become an integral part of the modern statistician’s toolkit, offering a powerful framework for uncertainty quantification and the incorporation of prior knowledge20. The advantages of Bayesian methods are increasingly recognized across various scientific disciplines21. A key strength of the Bayesian approach is its ability to treat all unknown parameters as random variables, described by probability distributions, which leads to a more complete representation of uncertainty compared to the frequentist view of fixed but unknown parameters22. This is particularly beneficial in reliability studies where data may be limited or expensive to obtain.
The practical implementation of Bayesian inference for complex models often relies on Markov Chain Monte Carlo (MCMC) methods, which use a sequence of random samples to numerically estimate uncertainties in model parameters23. While standard MCMC algorithms like the Metropolis-Hastings can struggle with complex, multimodal distributions, advanced techniques have been developed to address these challenges. Hamiltonian Monte Carlo (HMC) and its extension, Riemann Manifold Hamiltonian Monte Carlo (RMHMC), leverage geometric information to enable more efficient exploration of parameter spaces24. Furthermore, the Replica Exchange MCMC algorithm is specifically designed to handle multimodal distributions by allowing chains at different temperatures to exchange states, thus preventing the sampler from becoming trapped in local modes25.
Despite these methodological advances, a clear research gap exists in the comprehensive comparison of these sophisticated sampling algorithms for reliability models under SSPALT conditions. This study aims to fill this gap by providing a rigorous comparison of classical and Bayesian estimation paradigms for the MWM under the demanding conditions of SSPALT with Progressive Type-II Censoring with Constant-Barrier Removals (PT-II CBR). We evaluate classical approaches (Expectation-Maximization and Stochastic EM) alongside modern Bayesian methods (Replica Exchange MCMC, HMC, and RMHMC). Our extensive simulation studies and real-data application demonstrate that the Bayesian methodology, particularly when implemented via the RMHMC sampler, consistently delivers superior point estimation accuracy, dramatically lower risk under asymmetric loss, and markedly more efficient interval estimation than classical alternatives, especially under severe censoring. This direct comparison establishes the practical advantages of the modern Bayesian approach in this context.
The remainder of this work is organized as follows: Section "Stepped-stress partially accelerated life testing" describes the MWM distribution and the SSPALT model under PT-II CBRs. Section "Likelihood estimation of model parameters" introduces maximum likelihood estimation for the model parameters. Section "Model perturbation using the Bayesian approach" details the Bayesian estimation framework and the MCMC sampling techniques. Section “Simulation study” presents simulation findings, and Sect. "Application to electrical appliance lifetimes" provides a real-data analysis. Finally, Sect. “Concluding remarks” concludes the study.
Stepped-stress partially accelerated life testing
The SSPALT works as following a) A sample of n units is subjected to a lifetime test. The lifetimes of these units at each stress level follow a MWM. b) Initially, all the units undergo normal conditions for a predetermined time \(\tau\). If no failures occur during this time, they are subjected to accelerated conditions (stress). Let Y and X denote the random variables indicating the lifetimes of test units under the normal use conditions and accelerated conditions (stress), respectively. Within the context of the tampered random variable (TRV) model, the lifespan of a unit undergoing SSPALT can be expressed as
where \(\tau\) represents the time point of stress change, and \(\lambda>1\) is called acceleration factor. So, the PDF under SSPALT becomes
where \(\wp _{\lambda x_{}}=\tau +\lambda (x-\tau )\). The corresponding RF is written as
Experimental setup and data acquisition
This scenario considers n experimental units undergoing life testing under normal conditions to determine the breaking stress of, say, carbon fibers, assumed to follow the MWM. The experiment proceeds in stages.
-
Stage1:
(Normal Conditions) Initially, all n units undergo testing under normal conditions in a predetermined time \(\tau\). If no failures occur or if units are randomly removed from the test before \(\tau\), they transition to accelerated conditions. The probability of removing a unit (denoted by p) remains constant throughout the experiment.
-
Stage2:
(Accelerated Conditions) and Beyond Upon observing the units until \(\tau\), the remaining units are subjected to accelerated conditions. This process continues. After observing the \(m_{th}\) breaking stress unit (denoted by \(X_m\)) and all of the remaining \(R_m\) unit removals at that stage, the experiment concludes.
This approach results in a PT-II CBRs sample of actual size m which can be represented as \((X_{1}, R_{1}), (X_{2}, R_{2}), (X_{3}, R_{3})\),...,\((X_{m}, R_{m})\). Here, \(X_1< X_2< X_3< ... < X_m\) denotes the increasing order of observed failures. The number of test items removed at each test stage, denoted by \(R_i\), is a binomial variable (Singh et al.26). Let \(n_1\) be the number of failures observed under a routine environment before time point \(\tau\) and \((m - n_1)\) be the number observed under accelerated conditions afterward. The observations in SSPALT under PT-II CBR are structured as \((X_{1}, R_{1})< (X_{2}, R_{2})< (X_{3}, R_{3}), ...< (X_{n_1}, R_{n_1})< \tau < (X_{n_1}+1, R_{n_1}+1), ... ,(X_{m}, R_{m})\). We assume the total number of units being removed at each test stage, denoted as \(R_1 = r_1, R_2 = r_2, ..., R_m = r_m\), are fixed values.
Likelihood estimation of model parameters
This part of study explores the estimation of model parameters using PT-II CBRs within the SSPALT framework. We focus on obtaining maximum likelihood estimates (MLEs) and investigate their interval estimates using asymptotic methods. Let \(\varTheta = (\zeta ,\phi ,\beta ,\lambda )\) denote the parametric vector. The conditional likelihood function for this scenario can be derived as detailed in previous studies (Balakrishnan et al.27, Balakrishnan and Sandhu28, Balakrishnan et al.29, Balakrishnan and Aggarwala30, Cohen31, Calabria and Pulcini32 and Kamps and Cramer33).
where K includes the terms which are independent of parameters. And \(0<x_{1}<...<x_{n_{1}},x_{n_{1}+1}<...<x_{m}<\infty , n = m + \sum _{i=1}^{m}r_{i}, n, m \in N,1 \le i\le m,r_{i}\sim B(n-m- \sum _{l=0}^{i-1}r_{i},p)\) for \(i = 1,2,3,... m - 1\), \(r_{0} = 0\), \(c = \prod _{i=1}^{m}\xi _{i}\) with \(\xi _{i} = \sum _{j=i}^{m}(r_{j}+1)\) and for \(\xi _{1}= n\). Substituting \(f_{1}(x_{i}), f_{2}(x_{i}), R_{1}(x_{i})\) and \(R_{2}(x_{i})\) from (6) and (7) into (8), it reduces to
As total number of removals at each stage \(R_{i}\) are binomial random variables. This implies that the probability of removing any one unit remains constant at p throughout the experiment. Therefore
and for \(i = 2, 3, . . . , m-1\)
We consider that the \(R_{i,s}\) and \(X_{i}\) are independent for all i. Thus, full likelihood function is
where
Using (10) and (11) into (13), we get
Now, using (9), (12) and (14), the full likelihood function is written as
Expectation-maximization algorithm
This section describes the application of the EM algorithm to obtain the MLEs of model parameters \(\varTheta\) using the likelihood function in (15). While the MLEs could be obtained by solving the gradient equation (A.1), this approach often leads to intractable solutions. Additionally, the Newton-Raphson method might not converge consistently. Therefore, we propose using the EM algorithm, following the approach offered by Dempster et al.34, for robust MLE computation.
Major steps of EM algorithm.
Let \(X = (X_{1}, X_{2}, \ldots , X_{n_{1}}, X_{n_{1}+1}, \ldots , X_{m})\) and \(Z = (Z_{1}, Z_{2}, \ldots , Z_{n_{1}}, Z_{n_{1}+1}, \ldots , Z_{m})\), represent the observed and censored breaking stress data respectively. Here, each \(Z_{j}\) is a \(1 \times R_{j}\) vector with \(Z_{j} = (Z_{j1}, \ldots , Z_{jR_{j}})\) for \(j = 1, \ldots , n_{1}, n_{1}+1, \ldots , m\). Since the censored data is not directly observable, the vector Z can be used to represent the missing values. The vector \((X, Z) = W\) provides complete data. The log-likelihood function of uncensored observations is denoted by \(\mathcal {L}_{C}(W; \varTheta )\). The logarithm of complete data likelihood function (15), excluding an additive constant, can be expressed as \(\log (\mathcal {L}_{C}(W; \varTheta ))\) and is given by
The gradient of (16) is given by
The detailed derivations of the components of the (17) are provided in Appendix A. During the E-step of EM algorithm, we obtain the expression for pseudo log-likelihood function \(L_s(W; \Theta )\) by replacing any function of censored data, denoted as \(g(z_j)\), with its expected value given that the data is censored, \(E[g(Z_j) | Z_j> x_j]\). This results in
where \(I( \cdot )\) represents the indicator which equals 1 if the condition holds and 0 otherwise. This leads to
Theorem 1
EM Theorem For \(X = (X_{1}, X_{2},...,X_{n_{1}},X_{n_{1}+1},\dots , X_{m})\), the conditional density of \(Z_{jk}|X\) for \(k = 1, . . . , R_{j}\) is expressed as
for\(z_{j}>x_{j}\) and 0 otherwise.
Proof
The proof follows directly from the concept of the E-step in the EM algorithm and the properties of conditional expectation. Detailed derivations can be found in35.
Note that using Theorem 1, we can further simplify the expression for \(L_s(W; \Theta )\) as follows
Due to the mathematical complexity of these equations, solving them manually can be cumbersome. Fortunately, appropriate software packages can be employed to efficiently handle these computations. The M-step of EM algorithm focuses on updating the parametric values. This is achieved upon maximization of complete-data log-likelihood function, denoted as \(\varTheta ^{(j+1)} = \arg \max _\varTheta Q(\varTheta , \varTheta ^{(j)})\), concerning the current estimates. In the context of this model, this translates to maximizing the pseudo-log-likelihood function (19) after substituting the expected sufficient statistics (21) and (22) into it.
Therefore, if the current estimate of \(\varTheta = (\zeta , \phi , \beta , \lambda )\) at the \(k_{th}\) iteration is \(\varTheta ^{k} = (\zeta ^{k}, \phi ^{k}, \beta ^{k}, \lambda ^{k})\), then \(\varTheta ^{k+1} = (\zeta ^{k+1}, \phi ^{k+1}, \beta ^{k+1}, \lambda ^{k+1})\) can be obtained by maximizing the following
By solving (A.2 and A.3) w.r.t \(\beta\) and \(\lambda\) respectively, we obtain \(\beta ^{k+1},\lambda ^{k+1}\) as follows
Expectation-Maximization Algorithm for Parameter Estimation
EM algorithm for maximum likelihood estimation of distributional parameters.
The EM algorithm iteratively updates the parameter estimates \(\zeta\) and \(\phi\) for the maximization of expected complete-data log-likelihood function via (A.4-A.5). The updates involve two steps
-
E-step:
Provides the expression for conditional expected value of log-likelihood of complete data set, given current parameter estimates \(\varTheta ^{k}\).
-
M-step:
Offers the updated estimates \(\varTheta ^{k}\) upon the maximization of the expected complete-data log-likelihood function obtained in E-step.
These iterative steps continue until convergence criterion is met. A common stopping rule is to terminate the iterations when the difference between subsequent parameter estimates falls below a predefined threshold.
were, \(\epsilon> 0\) is a small real number used as a threshold for convergence. Figure 1 and Algorithm (1) offer the major steps of the EM estimation procedure. In practice, it’s recommended to run the EM algorithm multiple times with various initial values for parameter estimates. This approach helps ensure more stable convergence, as suggested in previous studies (e.g., Muthen and Shedden36, Nityasuddhi and Bohning37, Yao38).
Stochastic expectation-maximization algorithm
The EM algorithm can be computationally expensive when the E-step involves complex integration. To address this, Wei and Tanner39 offered a Monte Carlo EM (MCEM) approach that approximates expectations using Monte Carlo averages. However, maximizing the average of the log-likelihood in MCEM can also be challenging. Diebolt and Celeux40 introduced the SEM algorithm, which uses a simulated realization from conditional density instead of E-step. While SEM does not guarantee convergence to a specific set of estimates, it offers good optimization properties, allowing for the derivation of useful estimates. Following the SEM approach, we first generate \(R_i\) samples of \(z_{ij}\), where \(i = 1, 2, ..., r\) and \(j = 1, 2, ..., R_i\), from the following conditional model
Now, using (A.8 and A.9) the estimators of \(\beta , \lambda\) at the \(k + 1\) step of the algorithm \(\grave{\beta }^{k+1},\grave{\lambda }^{k+1}\) can be obtained as follows
The EM algorithm updates estimates of \(\zeta\) and \(\phi\) by iterative maximization of (A.6) - (A.7). The iterations can be stopped when the following convergence criterion is satisfied
where \(\epsilon> 0\) is a small pre-defined tolerance level.
Asymptotic confidence intervals
Asymptotic confidence intervals are constructed using the large-sample normality of maximum likelihood estimators41. Under standard regularity conditions, the MLE \(\hat{\Theta } = (\hat{\gamma }, \hat{\theta }, \hat{\lambda })\) is approximately multivariate normal
where \(\mathcal {I}(\Theta )\) is the expected Fisher information matrix. In practice, we substitute the observed information matrix evaluated at the MLE
The estimated variance-covariance matrix is the inverse of this observed information matrix. The \(100(1 - \alpha )\%\) asymptotic confidence interval for a single parameter \(\delta \in \{\gamma , \theta , \lambda \}\) takes the form
where \(z_{\alpha /2}\) is the \((1 - \alpha /2)\) quantile of the standard normal distribution, and \(\widehat{\text {Var}}(\hat{\delta })\) is the corresponding diagonal entry of \(\mathcal {I}(\hat{\Theta })^{-1}\). These intervals are computationally efficient and serve as a useful reference for comparison with resampling-based methods.
Bootstrap confidence intervals
Bootstrap techniques provide distribution-free alternatives for interval estimation, particularly valuable when sample sizes are moderate or asymptotic approximations may be unreliable42,43. We implement two parametric bootstrap approaches: the percentile method (Boot-p) and the studentized (Boot-t) method.
Percentile Bootstrap Confidence Intervals for MWM Parameters
Algorithm for computing percentile bootstrap confidence intervals.
Studentized Bootstrap Confidence Intervals for MWM Parameters
Algorithm for computing studentized bootstrap confidence intervals.
Both bootstrap procedures are applied to the aircraft windshield failure dataset using B replications, yielding robust intervals that account for potential non-normality and enhance the reliability of inference for the distribution parameters.
Model perturbation using the Bayesian approach
The classical scheme assumes parameters are fixed and aims to study the distributional characteristics of observed data given those parameters using the data likelihood. Conversely, the Bayesian approach treats parameters as random variables, considering the observed data as fixed. Bayesian inference requires specifying the probability function (likelihood), prior distribution, and posterior distribution. To derive the Bayes estimators for parameters \(\varTheta\) within the SSPALT framework with PT-II CBR data, we consider independent Gamma priors for parameters
where \(a_{i}>0,\ b_{i}>0\) for \(i=1,2,3,4\) are the hyper parameters. We select Gamma priors for their flexibility in capturing prior information about the parameters, as demonstrated in previous research (e.g., Kim et al.44). The joint prior is expressed as
For the logarithm of joint prior, denoted by \(\iota \left( \varTheta \right) = \log \pi (\varTheta )\), this formulation assumes a prior independence of the transformed parameters. The additive separability of this function, where the log-joint prior equals the sum of the log-marginal priors, \(\iota \left( \varTheta \right) = \log \pi (\zeta ) + \log \pi (\phi ) + \log \pi (\beta ) + \log \pi (\lambda )\), is the direct consequence of the factorization of the joint prior distribution \(\pi (\varTheta )\) into the product of its marginals45. This assumption of a factorized prior, which implies prior independence, is a common and practical starting point in Bayesian modeling, particularly when detailed prior knowledge about parameter dependencies is unavailable21. It simplifies the prior specification process and enhances the computational tractability of the model without precluding the posterior distribution from learning about potential parameter correlations through the likelihood function46.
Replica Exchange MCMC for Parameter Estimation
Replica Exchange MCMC algorithm for Bayesian parameter estimation with improved mixing.
Hence, we have
In practice, the hyper-parameters can be determined using past data. If U past samples from the MWM are available, we can compute the MLEs (\(\hat{\zeta }_u\), \(\hat{\phi }_u\), \(\hat{\beta }_u\), \(\hat{\lambda }_u\)) for each sample \(u = 1, 2, \ldots , U\). By matching the means and variances of these MLEs with the corresponding moments of the chosen Gamma priors, we can derive the hyper parameter values. For instance, the hyper-parameters of prior on \(\zeta\) can be obtained by equating the prior mean and variance of \(\zeta\) (\(a_1/b_1\) and \(a_1/b_1^2\), respectively) with the corresponding moments of the MLEs.
Similar to the hyper-parameter estimation process, we can determine the posterior density function for the model parameters by combining the likelihood function (15) with the chosen prior distribution (29) using Bayes’ theorem.
Bayesian estimation with linear exponential loss function
In this work, we employ the Linear Exponential (LINEX) loss function for Bayesian estimation. Unlike the squared error loss (SELF) that treats overestimation and underestimation equally, the LINEX is an asymmetric loss function that assigns greater weight to one type of error depending on the specific application. This allows for a more nuanced approach when overestimation or underestimation has a more significant impact on the results. The LINEX loss function can be written as
where \(k_{1}\) is the loss parameter that decides the shape of convex loss function. The Bayesian estimator under LINEX can be presented as
Major Steps of the Replica Exchange MCMC for Hazard Rate Estimation.
Derivations of the full conditional distributions for MCMC
Since directly generating data from the joint posterior density, (31), is not possible, this subsection focuses on deriving the full conditional posterior distributions for the elements of the parameter vector \(\vec {\varPsi }\) using MCMC. Specifically, we will derive the full conditional distributions for the Gibbs sampler and the Replica Exchange MCMC algorithm. These full conditional distributions are essential for implementing these MCMC methods and obtaining Bayes estimates. They are derived as follows
While the density of \(\beta\) in (36) follows a gamma distribution, the conditional densities of \(\rho\), \(\zeta\), and \(\phi\) in (33) - (35) are not standard forms. This characteristic makes Gibbs sampling unsuitable for these parameters. Therefore, a Replica Exchange MCMC algorithm is employed within the MCMC framework. The Replica Exchange MCMC algorithm is a vital component for generating posterior samples when combining Gibbs sampling with Replica Exchange MCMC steps for Bayesian estimation. Given the full conditional densities presented in (33) - (36), we outline the following steps to effectively utilize Replica Exchange MCMC within Gibbs sampling to update the parameters \(\rho , \zeta\), and \(\phi\). Figure 2 and Algorithm (4) offer the major steps of Replica Exchange MCMC estimation procedure. Following these steps allows us to efficiently generate samples using the Replica Exchange MCMC algorithm within the Gibbs sampler framework for Bayesian estimation. Subsequently, the Bayes estimates of the parameter vector under the LINEX loss function can be obtained by utilizing the approximate posterior samples, \(\Phi ^{[l]},\) generated using Algorithm (4)
Hamiltonian Monte Carlo Algorithm
Hamiltonian Monte Carlo algorithm for efficient sampling in high-dimensional parameter spaces.
Hamiltonian Monte Carlo algorithm
This subsection introduces the HMC algorithm, a technique used for data generation from conditional posterior distributions. Originally proposed by Duane et al.47 under the name Hybrid Monte Carlo for simulating molecular dynamics, HMC is now widely applied in various fields, including generalized extreme value (GEV) models. Compared to Replica Exchange MCMC and Gibbs sampling, HMC is computationally more expensive. However, its proposals are more efficient, requiring fewer samples to specify the posterior model48. The HMC algorithm requires these elements:
-
1.
Function U Represents the negative log-likelihood of the data at the current parameter position (essentially the log posterior).
-
2.
Function U grad Computes the gradient (slope) of the negative log-likelihood at the current position.
-
3.
Step Size \(\epsilon\) Defines the magnitude of parameter updates.
-
4.
Number of Leapfrog Steps L Controls the number of steps used to simulate the Hamiltonian dynamics.
-
5.
Initial Position q Represents the starting point for the sampling process.
Readers seeking a deeper understanding of the HMC method are directed to the comprehensive review by Neal48, which covers both the theoretical foundations and practical applications of HMC techniques.
Major Steps of HMC Algorithm.
The HMC is a powerful MCMC algorithm that utilizes Hamiltonian dynamics to efficiently explore the posterior distribution. It introduces auxiliary variables (\(\aleph\)) to construct a joint probability distribution. Let \(\varTheta \in \Re ^{d}\) be a d-dimensional parametric vector, \(\eth (\varTheta )\) denotes the posterior for \(\varTheta\), and \(\aleph \in \Re ^{d}\) an auxiliary parametric vector which is independent of \(\varTheta\) and distributed as \(\aleph \sim N(0,\varXi )\). HMC interprets \(\varTheta\) as a position for a particle and \(\log \eth (\varTheta )\) as potential energy. The auxiliary variables \(\aleph \in \Re ^{d}\) represent the momentum, and the kinetic energy \(\aleph '\varXi ^{-1}\aleph /2\). The total energy of the system is defined by the Hamiltonian function
The (un normalized) density of \((\varTheta , \aleph )\) is given by
For time t, the potential deterministic assessment of a particle with constant total energy is governed by Hamiltonian dynamics equations
However, these differential equations often lack analytical solutions, necessitating numerical methods for their integration. One popular approach is the Stormer-Verlet (or Leapfrog) integrator introduced by49. This method updates the Hamiltonian dynamics into the following steps
For a user-specified small step size (\(\epsilon> 0\)), the algorithm performs a series of time steps, resulting in a proposed state \((\varTheta ^{\divideontimes }, \aleph ^{\divideontimes })\). Details on the required partial derivative expressions for HMC are provided in Appendix A. To account for the discretionary error introduced by the time steps and ensure convergence to the target distribution, a Replica Exchange MCMC acceptance step is employed. Here, the proposed state \((\varTheta ^{\divideontimes }, \aleph ^{\divideontimes })\) is accepted with a probability based on the current and proposed state’s Hamiltonian values.
Within the auxiliary variable distribution, \(\varXi\) represents a positive definite symmetric mass matrix. It’s typically a diagonal matrix with fixed elements, denoted as \(\varXi = \sigma \textbf{I}_{d}\). The basic HMC algorithm (with \(\sigma = 1\)) can be implemented using the crucial steps given by Fig. 3 and Algorithm (5). The HMC algorithm utilizes the first derivatives of the un normalized log-posterior density to propose moves toward regions with higher probabilities. This leads to faster convergence of the chains towards stationary. However, HMC requires sampling within an unconstrained space. Therefore, we transform the parameter vector \(\varTheta\) to the real line. Following the transformation, priors are assigned, and corresponding derivatives are taken with respect to the transformed parameters. Bayes estimates under the HMC algorithm for the LINEX loss function can then be obtained using (37).
Riemann Manifold Hamiltonian Monte Carlo
Building upon HMC, the RMHMC algorithm, introduced by Duane et al.47, modifies the proposal mechanism. Instead of standard Euclidean distance, it utilizes a Riemann metric for proposing moves. This approach leverages the geometric properties of the posterior distribution, leading to more efficient exploration. The core idea lies in redefining the Hamiltonian function as
where \(\varPsi (\varTheta )\) is a position-dependent matrix adapting to the local geometry of the posterior distribution
which represents the Fisher information matrix and negative of the Hessian of log-prior. Consequently, the Hamiltonian dynamics become
We employ the generalized leapfrog-Verlet solution (see49). Expressions for the Hessian of log-prior are provided in Appendix A. Finally, Bayes estimates are obtained using the approximate posterior samples, \(\varTheta ^{[l]},\) produced under Algorithm (5), and (37).
Highest posterior density intervals
The Bayesian Highest Posterior Density (HPD) interval is the shortest interval containing \(100(1 - \alpha )\%\) of the posterior mass, offering a more concentrated summary when the posterior is asymmetric. For unimodal posteriors, the HPD interval coincides with the equal-tailed credible interval; for multimodal or skewed posteriors, it is typically narrower and more informative.
Given posterior samples \(\{\delta ^{[l]}\}_{l = N_0+1}^{N}\), the \(100(1 - \alpha )\%\) HPD interval is obtained by:
-
1.
Sorting the samples: \(\delta ^{(1)} \le \delta ^{(2)} \le \cdots \le \delta ^{(M)}\), where \(M = N - N_0\).
-
2.
Computing all candidate intervals \(\left[ \delta ^{(i)}, \delta ^{(i + J - 1)} \right]\) for \(i = 1, \dots , M - J + 1\), where \(J = \lceil (1 - \alpha ) M \rceil\).
-
3.
Selecting the interval with the smallest length (or equivalently, highest density).
intervals are computed from the same Markov chains produced by the advanced samplers presented in Algorithms 4 and 5, ensuring consistency and efficiency in uncertainty quantification for the parameter estimates.
Simulation study
This section presents a comprehensive evaluation of the performance of various estimation methods, EM, SEM, Replica Exchange MCMC, HMC, and RMHMC, for the MWM within an SSPALT framework under PT-II CBR. An extensive simulation study was conducted across a range of experimental conditions, systematically varying the total sample size (n), effective sample size (m) and removal probability (\(p=0.5\)). We evaluated two different stress factor combinations (\(\tau = 3.5\) and \(\tau = 5.0\)), a removal probability (p) of 0.5, and LINEX asymmetry parameters (\(k_1 = \pm 1.5\)). Two replication settings are considered: 5000 replications after discarding 2,000 as burn-in for primary analysis (Tables 1, 2, 3, 4) and 1000 replications after discarding 100 as burn-in to assess convergence under limited iterations (Tables 5, 6, 7, 8). Detailed parameter settings for each scenario are provided in the captions of the corresponding tables (Tables 1, 2, 3, 4, 5, 6, 7, 8).
Synthetic data is generated according to the specified distributions and model assumptions. Various estimation methods are then applied to estimate the model parameters based on the simulated data. To ensure reproducible and efficient sampling, the step size \(\epsilon\) in both HMC and RMHMC is adapted automatically during a warm-up phase using the No-U-Turn Sampler (NUTS) with dual averaging50. This targets an acceptance rate of approximately 0.8–0.9, which is known to maximise effective sample size in Hamiltonian-based methods. The mass matrix (or manifold metric in RMHMC) is similarly adapted using a windowed covariance estimator. Initial parameter values are drawn from a diffuse but proper distribution \(\varTheta ^{[0]} \sim \mathcal {N}(\hat{\varTheta }_{\text {MM}}, 10^{2}\textbf{I})\), where \(\hat{\varTheta }_{\text {MM}}\) is the method-of-moments estimator computed from the raw data. Multiple chains (typically 2–4) are started from deliberately overdispersed points to facilitate convergence diagnostics via the Gelman–Rubin \(\hat{R}\) statistic. This fully automatic, non-hand-tuned procedure was used consistently across the study, contributing to the rapid mixing and excellent performance of RMHMC Algorithm 4. For Bayesian estimation, samples were generated using the Replica Exchange MCMC, HMC, and RMHMC algorithms. Within the LINEX loss framework, we considered two values for the loss parameter (\(k_1\)) 1.5 (overestimation more severe) and −1.5 (underestimation more severe). This design allows us to assess the robustness and efficiency of the estimators under different levels of censoring and stress acceleration.
The following scenarios were investigated to examine the impact of sample size and censoring intensity:
Performance was evaluated using two key metrics computed over \(T=5,000\) replications: the Average Estimate (AVG) and the Mean Squared Error (MSE), defined as
where \(\varTheta\) represents the true parameter value.
Results and discussion
Across all scenarios, the classical EM and SEM algorithms exhibit substantially higher MSE values compared to the advanced MCMC-based methods. For \(\tau = 3.5\) and the most challenging censoring scheme (\(n=80\), \(m=20\)), the MSE for parameter \(\phi\) under EM reaches 0.3118, whereas RMHMC achieves only 0.1052 when \(k_1 = 1.5\) (Table 3). A similar pattern holds for \(\tau = 5.0\), where EM yields MSE = 0.2400 for \(\phi\) (\(n=80\), \(m=20\)), while RMHMC reduces this to 0.1053 (Table 4). The SEM algorithm, although marginally better than EM in some cases, consistently underperforms the Bayesian samplers by factors ranging from 2 to 4. Among the Bayesian approaches, RMHMC demonstrates clear superiority, followed closely by HMC, with Replica Exchange MCMC showing the weakest performance. For \(\tau = 3.5\) and moderate censoring (\(n=150\), \(m=60\)), RMHMC attains MSE values of 0.0771 for \(\phi\) (\(k_1 = 1.5\)), compared to 0.1102 for HMC and 0.1574 for Replica Exchange (Table 3). This advantage persists across all effective sample sizes and holds for both positive and negative \(k_1\), confirming RMHMC’s robustness to LINEX asymmetry. When the stress factor increases to \(\tau = 5.0\), RMHMC continues to dominate, reducing MSE for \(\phi\) to 0.0772 under heavy censoring (\(n=150\), \(m=30\)) versus 0.1103 for HMC and 0.1767 for Replica Exchange (Table 4).
As expected, estimation risk rises with decreasing effective sample proportion (\(m/n\)). However, RMHMC exhibits the slowest degradation: for \(\tau = 3.5\), transitioning from \(m=100\) to \(m=30\) (\(n=150\)) increases RMHMC’s MSE for \(\phi\) from 0.0708 to 0.0841 (\(k_1 = 1.5\)), while Replica Exchange’s MSE escalates from 0.1444 to 0.1716 (Table 3). This resilience underscores the efficiency of manifold-adapted dynamics in exploring high-curvature posterior geometries typical of censored lifetime models. The reduced-iteration experiment (\(N=1000\)) amplifies these differences. RMHMC maintains low MSE (e.g., 0.1163 for \(\phi\) at \(n=80\), \(m=40\), \(\tau = 3.5\); Table 7), whereas Replica Exchange and HMC show inflated errors, and EM/SEM degrade markedly (MSE > 0.78 for \(\phi\) in several cases). This highlights RMHMC’s rapid convergence, requiring fewer iterations to reach stationarity critical advantage in computationally constrained settings.
These results reveal several critical patterns regarding estimator performance under the SSPALT model:
Key Finding 1: A consistent and striking result across all scenarios is the superior performance of the RMHMC algorithm, closely followed by HMC. For instance, in Scenario 1 (\(n=80, m=40, \tau =3.5\)), RMHMC achieves an MSE of 0.1354 for \(\hat{\lambda }\), which is approximately 30% lower than the next best method (HMC with MSE 0.1935) and 51% lower than the standard Replica Exchange algorithm (MSE 0.2764). This performance advantage is even more pronounced under heavier censoring. In Scenario 3 (\(n=80, m=20\)), RMHMC maintains an MSE of 0.1609 for \(\hat{\lambda }\), while the EM algorithm’s MSE increases to 0.4769, a nearly three-fold difference. This demonstrates the remarkable robustness of geometric MCMC methods to increasing data sparsity.
Key Finding 2: As anticipated, the estimation error (MSE) for all parameters (\(\lambda , \beta , \zeta , \phi\)) increases monotonically with the censoring rate (i.e., as m/n decreases). However, the rate of this degradation is not uniform across methods. The classical EM and SEM algorithms show the steepest decline in performance. For \(\hat{\phi }\) under \(\tau =3.5\), the MSE for EM in Table 1 increases from 0.4581 (Scenario 1) to 0.9368 (Scenario 3), whereas RMHMC’s MSE only increases from 0.0885 to 0.1052 (see, Table 3). Overall, the MSE values were relatively small for all algorithms, with a surprising tendency for RMHMC to achieve lower MSE. This is expected as the Replica Exchange MCMC algorithm may not be as close to the target distribution as compared to HMC and RMHMC after discarding the burn-in iterations. This suggests that the Bayesian geometric methods provide a substantial advantage in practical applications where high censoring is unavoidable.
Key Finding 3: The results under the asymmetric LINEX loss function provide crucial insights for reliability applications where overestimation and underestimation of lifetime parameters carry different consequences. For \(k_1 = 1.5\) (overestimation more costly), the Bayesian methods, particularly RMHMC, yield estimates that are systematically slightly conservative (biased downwards) but with significantly reduced variance, leading to the lowest overall MSE (see, e.g. Tables 5 and 6). This property is highly desirable in reliability-critical applications where over-optimistic lifetime predictions can have severe safety implications.
Key Finding 4: While the computational cost per iteration is highest for RMHMC due to the computation of the metric tensor and its derivatives, this is offset by its superior convergence properties. In a supplementary experiment with limited iterations (1,000 samples after 100 burn-in), RMHMC and HMC achieved usable estimates, whereas Replica Exchange MCMC exhibited significant bias, failing to converge to the stationary distribution (see, e.g. Tables 5, 6, 7, 8). This makes the geometric methods particularly valuable for applications with computationally expensive likelihood evaluations, where rapid convergence is essential. Furthermore, as observed in Tables 1, 2, 3, 4, 5, 6, 7, 8, the estimation risks for \(\zeta\), \(\phi\), \(\beta\), and \(\lambda\) increase with decreasing effective sample size (m). Notably, among all methods, RMHMC (for \(k_1> 0\)) followed by HMC exhibited the lowest estimation risks for these parameters.
Interval estimation
Beyond point estimates, reliable uncertainty quantification is crucial for practical decision-making. To assess the uncertainty quantification performance of the proposed methods, we conducted an extensive simulation study using the same settings as in the point estimation study (5000 replications, \(\tau = 3.5\) with true values \(\lambda = 2.5\), \(\beta = 0.8\), \(\zeta = 1.2\), \(\phi = 1.5\); \(\tau = 5.0\) with \(\lambda = 1.5\), \(\beta = 0.9\), \(\zeta = 1.3\), \(\phi = 1.75\); \(p=0.5\)). Four interval estimation procedures were compared at the nominal 95% level:
-
Asymptotic confidence intervals (based on the observed Fisher information matrix),
-
Percentile bootstrap intervals (Boot-p) (\(B=5000\) resamples),
-
Studentized bootstrap intervals (Boot-t) (\(B=5000\) resamples),
-
Highest posterior density (HPD) intervals obtained from the RMHMC sampler (the best-performing Bayesian method in point estimation).
Performance is evaluated by average interval length (smaller is preferable) and empirical coverage probability (closest to 95% is best). Results are reported for the four parameters \((\lambda , \beta , \zeta , \phi )\). Due to space constraints and identical qualitative behaviour across parameters, only results for \(\lambda\) (the most challenging shape parameter) are shown; full tables for the remaining parameters follow the same pattern.
The pattern is consistent across all parameters and both stress levels summarized in Table 9: HPD intervals constructed from RMHMC posterior samples uniformly dominate the competing procedures, achieving the nominal 95% coverage most consistently across all parameters and scenarios, while simultaneously offering the shortest average interval lengths. For example, for \(\hat{\lambda }\) in Scenario 1 (\(n=80, m=40\)), the HPD interval length is 1.30 with 96.6% empirical coverage, compared to the Boot-t interval (length 1.51, coverage 93.4%) and the asymptotic interval (length 1.44, coverage 94.8%). They exhibit the shortest average lengths (typically 10–15% narrower than asymptotic intervals and 10–25% narrower than the best bootstrap method) while maintaining or slightly exceeding the nominal 95% coverage probability. The studentized bootstrap (Boot-t) performs reasonably well, especially for larger effective sample sizes, but is consistently outperformed by the Bayesian HPD approach. Asymptotic intervals are competitive when \(m \ge 60\) but become overly conservative (wider) under heavier censoring. The percentile bootstrap (Boot-p) tends to under-cover in small or heavily censored scenarios. The superiority of the Bayesian HPD intervals is most pronounced under high censoring. In Scenario 3 (\(n=80, m=20\)), the asymptotic and bootstrap intervals become excessively wide to maintain coverage, whereas the HPD intervals remain relatively precise. This efficiency stems from the RMHMC sampler’s ability to fully characterize the often asymmetric and non-elliptical shape of the posterior distribution, particularly in constrained parameter spaces common to lifetime distributions. In practical terms, findings from Table 9 imply that for a reliability engineer requiring a precise estimate of, say, the 0.95 quantile of a product’s lifetime distribution, the Bayesian RMHMC approach will provide a narrower, more certain interval than classical methods, enabling more confident decision-making with the same data. These findings reinforce the point estimation results: the geometry-aware RMHMC sampler not only yields the most accurate point estimates but also produces the most efficient and reliable uncertainty quantification. Consequently, Bayesian highest posterior density intervals based on RMHMC are strongly recommended for uncertainty quantification in SSPALT analyses.
Importantly, for considered scenarios and sample sizes, the Bayesian estimators consistently achieved the lowest risks compared to EM and SEM approaches and provides a narrower, more certain interval than classical methods. This pattern is observed across all sample sizes, censoring intensities, stress factors, and LINEX asymmetries, establishing it as the method of choice for Bayesian inference for the MWM under progressive Type-II censoring with binomial removals. In conclusion, this simulation experiment offers empirical evidence supporting the superior performance of RMHMC algorithm. We suggest this technique for practical applications involving lifetime data analysis.
Application to electrical appliance lifetimes
This section demonstrates the practical utility of the proposed methodology by analyzing a real-world dataset concerning the lifetimes of electrical appliances. The data, originally reported by51 (p. 112), consists of the number of cycles to failure for 60 appliances and is presented in Table 10. This application serves to validate the theoretical developments and showcases the performance of the various estimation algorithms in a realistic SSPALT framework with PT-II CBRs.
Exploratory data analysis and hazard shape identification
Exploratory data analysis (EDA) aims to reveal patterns and trends within the data. A thorough EDA was conducted to understand the underlying failure characteristics. Common EDA techniques include: a) Visualization; Representing data through charts and graphs (bar plots, box plots, density curves, histograms, etc.) to identify patterns of data. b) Descriptive Statistics; Summarizing data using measures of central tendency (mean, median) and dispersion (variance, standard deviation). The key descriptive statistics are summarized alongside the raw data in Table 10. To gain deeper insight into the failure rate behavior, we employed the Total Time on Test (TTT) plot52, as shown in Fig. 9. The pronounced concave shape of the TTT curve in Fig. 9 provides strong visual evidence of an increasing failure rate over time. This is a critical characteristic in reliability engineering, as it indicates that the appliances are susceptible to wear-out failures. This finding directly justifies the use of the MWM, which is particularly adept at capturing such non-monotonic and increasing hazard rate behaviors due to its flexible functional form that generalizes the standard Weibull distribution.
Model selection and validation
Model validation typically involves three key steps: first, developing a comprehensive probability model and estimating its parameters accurately. Second, the model is thoroughly evaluated and validated to ensure that it aligns well with the data and fulfills the necessary assumptions. Finally, the most suitable model is selected based on performance metrics and goodness-of-fit, ensuring it is the best fit for representing the underlying data structure. To evaluate the performance of the MWM, we compared it with the following competing distributions
-
(a)
MWM
-
(b)
Weibull distribution (WD)
-
(c)
Burr-XII distribution
-
(d)
Inverse Weibull Weibull (IW-WeibuII) distribution by Hassan and Nassr53
-
(e)
Exponinated Weibull (Ex-WeibuII) distribution by Nassar and Eissa54
The goal is to assess the goodness-of-fit of these distributions to the data. The MLE was used to estimate the parameters for each model. Goodness-of-fit statistics were calculated to assess model validity. These statistics, along with the MLEs for all distributions, are presented in Table 14. The statistics include Kolmogorov-Smirnov (KS) distance between the fitted and empirical distribution functions Information criteria (AIC, CAIC, BIC, HQIC) Additionally, we visually evaluated the fitting performance using sample histogram plots (Fig. 5), quantile-quantile (QQ) plots (Fig. 6), P-P plots (Fig. 7), and estimated CDF plots (Fig. 8). The results suggest that the MWM outperforms the other parametric distributions. The data seems to be well-fitted to the MWM, as evidenced by its lower AIC, CAIC, BIC, and HQIC values as given in Table 14. Furthermore, the KS statistic for the MWM is minimal as compared to other considered distributions. These findings indicate that the MWM is the most suitable model among those considered.
Posterior distributions and MCMC trace plots for the model parameters (\(\lambda\): Parameter 1; \(\zeta\): Parameter 2; \(\phi\): Parameter 3; \(\beta\) at \(\tau\) = 1.5, \(\beta\) = 0.1, \(\zeta\) = 0.3, \(\phi\) = 0.5, p = 0.4: Parameter 4) obtained via the RMHMC sampler after 10,000 iterations (5,000 burn-in discarded).
Estimated histograms of the considered models for the lifetimes of electrical appliances data.
Normal QQ plots of the considered models for the lifetimes of electrical appliances data.
Normal PP plots of the considered models for the lifetimes of electrical appliances data.
Estimated CDF plots of the considered models for the lifetimes of electrical appliances data.
Estimated TTT-transform plot for the lifetimes of electrical appliances data.
Overall, the MWM demonstrates a decisively better fit to the data. This is quantitatively supported by its consistently lower values across all information criteria (AIC: 922.14, CAIC: 922.78, BIC: 930.72, HQIC: 925.51), which are substantially smaller than those of its closest competitor. Furthermore, the MWM achieves the smallest KS statistic (0.0721), indicating the closest agreement between the empirical and the fitted CDFs. The superior fit of the MWM is evident from a suite of diagnostic plots (Figs. 5, 6, 7, and 8). Specifically:
-
The histogram with fitted densities (Fig. 5) shows that the MWM most accurately captures the shape and right-tail of the lifetime data.
-
The Q-Q plot (Fig. 6) for the MWM exhibits points most closely aligned with the unit line, confirming that the quantiles of the fitted model match the empirical data quantiles exceptionally well.
-
The P-P plot (Fig. 7) further validates the MWM, with its points forming the tightest cluster around the diagonal, indicating excellent calibration across the entire distribution.
These results collectively provide compelling evidence that the MWM is the most appropriate model among the candidates for representing the failure behavior of the electrical appliances.
SSPALT analysis and parameter estimation
We implemented the SSPALT framework under PT-II CBRs using three different censoring schemes (\(m = 50, 40, 30\)) for two stress-change levels (\(\tau = 20\) and 30). Parameter estimation was performed using both frequentist (EM, SEM) and Bayesian (Replica Exchange MCMC, HMC, RMHMC) approaches. The posterior distributions and convergence diagnostics for the parameters, \(\lambda\) (Parameter 1), \(\zeta\) (Parameter 2), \(\phi\) (Parameter 3), and \(\beta\) (Parameter 4), are presented in Fig. 4. The density plots in Fig. 4a reveal well-behaved, unimodal posteriors for all parameters. Parameter 1 (\(\lambda\)) exhibits a symmetric, bell-shaped distribution centered around 5.7229 with a narrow 95% credible interval [0.9613, 10.715], indicating precise estimation. Parameter 2 (\(\zeta\)) displays a slightly right-skewed yet smooth unimodal posterior (mean 8.6787), while Parameters 3 (\(\phi\)) and 4 (\(\beta\)) show moderate skewness but remain clearly unimodal with effective sample sizes exceeding 2 in all cases, confirming adequate Monte Carlo precision. The corresponding trace plots in Fig. 4b demonstrate excellent chain mixing and rapid convergence of the RMHMC sampler. All four chains stabilize early, with no visible trends, autocorrelation, or stickiness after the initial burn-in phase. The running means (red lines) flatten quickly and overlap tightly with the overall posterior means, and the standard deviations of the chains are consistently low (SD < 3.6 across parameters). This behavior is characteristic of highly efficient geometry-aware sampling on the Riemannian manifold, which successfully navigates the typical posterior curvature encountered in censored lifetime models. The combination of smooth, unimodal posteriors and well-mixed traces provides strong visual evidence that the RMHMC algorithm has reliably explored the target posterior distribution, yielding trustworthy Bayesian inference for the parameters.
Tables 11 and 12 present parameter estimates obtained using the EM and SEM algorithms under different censoring combinations, as well as estimates from the Replica Exchange MCMC, HMC, and RMHMC algorithms. The formulas and algorithms used for these estimations are detailed in Sections 3 and 4. These results reinforce the conclusions from the simulation study. The Bayesian methods, particularly the RMHMC algorithm, yield stable and precise parameter estimates. As the degree of censoring intensifies (i.e., as m decreases from 50 to 30), the estimates from all methods show increased variability, but the Bayesian estimates, especially from RMHMC, exhibit greater robustness, with smaller changes in their posterior means. Besides, the computational robustness of the Bayesian methods is confirmed by the convergence diagnostics presented in Table 13. The \(\hat{R}\) statistics by Thomas and Tu55 for all parameters and across all samplers are exceptionally close to 1.0 (typically \(<1.01\)), providing strong evidence that the Markov chains have converged to the target posterior distribution.
Interval estimation on electrical appliance lifetimes data
To quantify parameter uncertainty in the real-data application, we constructed 95% intervals using the four procedures evaluated in the simulation study: asymptotic confidence intervals, percentile and studentized bootstrap intervals (B = 5000 resamples each), and highest posterior density (HPD) intervals from the RMHMC sampler (15 000 iterations after 5000 burn-in). Results are reported for the three censoring schemes (m = 50, 40, 30) under both stress-change levels \(\tau\) = 20 and 30.
The pattern observed in the simulation study is fully reproduced on the real electrical appliance lifetimes dataset: HPD intervals derived from the RMHMC sampler are consistently the shortest (15–30% narrower than asymptotic intervals and 10–25% narrower than the best bootstrap method) across all censoring levels and stress conditions. The studentized bootstrap performs creditably, while asymptotic intervals become noticeably wider under heavier censoring (m = 30). These real-data results reinforce the simulation findings and confirm that Bayesian HPD intervals based on RMHMC provide the most precise and reliable quantification of parameter uncertainty for the proposed model in practical lifetime analysis.
Concluding remarks
This study explored the estimation of MWM model parameters under PT-II CBRs using SSPALT. Here, we have quantitatively investigated the performance of EM, SEM, Replica Exchange MCMC, HMC, and RMHMC algorithms with a Bayesian approach, using MCMC methods. The performance of these methods was quantitatively assessed through simulation studies and validated with real-world electrical appliance lifetime data. Simulation results demonstrated that the Bayesian approach, particularly with RMHMC, consistently outperformed other methods across various scenarios. For stress factors (\(\tau\) = 3.5) and (\(\tau\) = 5.0) with a removal probability (p = 0.5), RMHMC exhibited the lowest MSE values, especially when overestimation was penalized ((\(k_1\) = 1.5)). This advantage was evident even with reduced sample sizes (e.g., 1,000 samples), where RMHMC and HMC showed faster convergence compared to Replica Exchange MCMC, despite higher per-iteration computational costs. The estimation risks for parameters increased with decreasing effective sample size, yet RMHMC maintained the lowest risks, reinforcing its robustness. Interval estimation further reinforces this conclusion. The HPD intervals derived from RMHMC posterior samples uniformly outperformed asymptotic confidence intervals and both percentile and studentized bootstrap intervals, achieving nominal coverage with substantially shorter lengths (10–30% narrower on average). This efficiency is especially pronounced under high censoring ratios, where classical and bootstrap methods become overly conservative. Application to the electrical appliance lifetime dataset confirmed the practical relevance of the MWM and the RMHMC-based Bayesian approach. The model exhibited superior goodness-of-fit relative to Weibull, Burr-XII, inverse Weibull, and exponentiated Weibull alternatives. In all the discussed scenarios, our findings under SSPALT with PT-II CBRs showed that RMHMC-based estimates, supported by (\(\hat{R}\)) statistics near one, ensured chain convergence, enhancing reliability in parameter estimation, and precise, narrow HPD intervals.
These findings hold significant implications for reliability engineering. The RMHMC-based Bayesian framework offers a robust tool for accurately modeling lifetime data and failure times, critical for designing, testing, and maintaining engineering systems. By providing precise parameter estimates, this approach enables engineers to optimize system robustness, enhance safety protocols, and implement cost-effective maintenance strategies. Given its consistent performance across simulated and real datasets, we strongly recommend the RMHMC Bayesian method for MWM parameter estimation under SSPALT with PT-II CBRs, positioning it as a preferred choice for advancing reliability assessments in practical applications. Extensions to covariate-adjusted regression formulations (proportional hazards or accelerated failure time models) would enable richer predictive modeling. Incorporating competing risks, frailty terms, or mixture structures could further broaden applicability. Validation on additional industrial and biomedical datasets, along with integration of neural network-based amortized inference for ultra-large-scale problems, represents promising avenues for continued development.
Data availability
The data will be available on request from the corresponding author.
References
Lai, C., Xie, M. & Murthy, D. A modified Weibull distribution. IEEE Trans Reliab 52(1), 33–37 (2003).
Ramzan, Q., Amin, M. & Faisal, M. Bayesian inference for modified Weibull distribution under simple step-stress model based on type-I censoring. Quality and Reliability Engineering International 38(2), 757–779 (2022).
Goel, P. Some estimation problems in the study of tampered random variables. Technical report, Department of Statistics, (Carnegie Mellon University, Pittspurgh, 1971).
Bhattacharyya, G. & Soejoeti, Z. A tampered failure rate model for step-stress accelerated life test. Commun Stat Theory Methods 18(5), 627–643 (1975).
Degroot, M. & Goel, P. Bayesian estimation and optimal designs in partially accelerated life testing. J Dedic Adv Op Logist Res 26(2), 223–235 (1979).
Bai, D. & Chung, S. Optimal design of partially accelerated life tests for the exponential distribution under type-I censoring. IEEE Trans Reliab 41, 400–406 (1992).
Bai, D., Chung, S. & Chun, Y. The optimal design of \({PALT}\) for the log-normal distribution under type-I censoring. Reliab Eng Syst Saf 40, 85–92 (1993).
Abdel-Ghani, M. The estimation problem of the log-logistic parameters in step partially accelerated life tests using type-i censored data. Natl Rev Soc Sci 14(2), 1–19 (2004).
Tse, S., Yang, C. & Yuen, H. Statistical analysis of Weibull distributed life time data under type II progressive censoring with binomial removals. J Appl Stat 27, 1033–1043 (2000).
Xiong, C. & Ji, M. Analysis of grouped and censored data from step-stress life testing. IEEE Trans Reliab 53(1), 22–28 (2004).
Al-Babtain, A., Shakhatreh, M., Nassar, M. & Afify, A. A new modified kies family properties, estimation under complete and type-ii censored samples, and engineering applications. Mathematics 8(8), 1345 (2020).
Tahani, A., Abushala, A. & Solimanb, A. A. Estimating the Pareto parameters under progressive censoring data for constant-partially accelerated life tests. J Stat Comput Simul 85(5), 917–934 (2015).
Asadi, S., Panahi, H., Swarup, C. & Lone, S. Inference on adaptive progressive hybrid censored accelerated life test for Gompertz distribution and its evaluation for virus-containing micro droplets data. Alexandria Engineering Journal 61(12), 10071–10084 (2022).
Ismail, A. Inference in the generalized exponential distribution under partially accelerated tests with progressive type-II censoring. Theoret Appl Fract Mech 59(1), 49–56 (2012).
Anwar, S., Lone, S., Khan, A. & Almutlak, S. Stress-strength reliability estimation for the inverted exponentiated Rayleigh distribution under unified progressive hybrid censoring with application. Electron. Res. Arch. 31, 4011–33 (2023).
Lone, S., Panahi, H., Anwar, S. & Shahab, S. Inference of reliability model with burr type XII distribution under two sample balanced progressive censored samples. Physica Scripta 99(2), 025019 (2024).
Lone, S., Alam, I. & Rahman, A. Statistical analysis under geometric process in accelerated life testing plans for generalized exponential distribution. Annals of Data Science 10(6), 1653–65 (2023).
Abushal, T. & Soliman, A. Estimating the pareto parameters under progressive censoring data for constant-partially accelerated life tests. J Stat Comput Simul 85(5), 917–934 (2015).
Ramzan, Q., Amin, M., Elhassanein, A. & Ikram, M. The extended generalized inverted Kumaraswamy Weibull distribution properties and applications. AIMS Mathematics 6(9), 9955–9980 (2021).
van de Schoot, R. et al. A gentle introduction to bayesian analysis: Applications to developmental research. Child Development, 85(3):842–860. Provides an accessible introduction to the Bayesian paradigm and its terminology (2013).
Schad, D. J. et al. Bayesian hierarchical modeling: An introduction and reassessment. Behavior Research Methods, 56:4600–4631. Discusses the modern prevalence and advantages of Bayesian methods in scientific research (2024).
Gelman, A., et al. Bayesian data analysis. Discusses how the posterior can learn parameter dependencies even from an independent prior (2013a).
Speagle, J. S. A conceptual introduction to markov chain monte carlo methods. arXiv preprint. Provides a foundational introduction to the problems MCMC methods solve and how they work (2020).
Brofos, J. A. & Lederman, R. R. On numerical considerations for riemannian manifold hamiltonian monte carlo. arXiv preprint. Discusses the RMHMC algorithm and its numerical implementation details (2021).
Abdellatif, M. Implementing bayesian replica exchange in chainsail. Explains the Replica Exchange MCMC algorithm and its application to Bayesian data analysis. https://tweag.io/blog/2022-10-25-Internship-abdellatif/ (2022).
Singh, S., Singh, U. & Kumar, M. Estimation of parameters of generalized inverted exponential distribution for progressive type-II censored sample with binomial removals. J Probab Stat, 1–12 (2013).
Balakrishnan, N., Kannan, N., Lin, C. & Ng, H. Point and interval estimation for Gaussian distribution based on progressively type-II censored samples. IEEE Trans Reliab 52, 90–95 (2003).
Balakrishnan, N. & Sandhu, R. A simple simulational algorithm for generating progressive type-II censored samples. Am Stat 49(2), 229–230 (1995).
Balakrishnan, N., Cramer, E. & Kamps, U. Bounds for means and variances of progressive type-II censored order statistics. Stat Probab Lett 54, 301–315 (2001).
Balakrishnan, N. & Aggarwala, R. Progressive censoring theory, methods, and applications (Springer, 2000).
Cohen, A. Progressively censored samples in life testing. Technometrics 5(3), 327–339 (1963).
Calabria, R. & Pulcini, G. Point estimation under-asymmetric loss functions for life-truncated exponential samples. Commun Stat Theory Methods 25(3), 585–600 (1996).
Kamps, U. & Cramer, E. On distributions of generalized order statistics. Statistics 35, 269–280 (2001).
Dempster, A., Laird, N. & Rubin, D. Maximum likelihood from incomplete data via the EM algorithm. J R Stat Soc B 39, 1–38 (1977).
Ng, S. & McLachlan, G. On the choice of the number of blocks with the incremental EM algorithm for the fitting of normal mixtures. Stat Comput 13, 45–55 (2003).
Muthen, B. & Shedden, K. Finite mixture modeling with mixture outcomes using the EM algorithm. Biometrics 55(2), 463–469 (1999).
Nityasuddhi, D. & Bohning, D. Asymptotic properties of the EM algorithm estimate for normal mixture models with component specific variances. Computational Statistics & Data Analysis 41(3–4), 591–601 (2003).
Yao, W. A note on EM algorithm for mixture models. Statistics & Probability Letters 83(2), 519–526 (2013).
Wei, G. & Tanner, M. A Monte Carlo implementation of the EM algorithm and the poor man’s data augmentation algorithms. Journal of the American Statistical Association 85(411), 699–704 (1990).
Diebolt, J. & Celeux, G. Asymptotic properties of a stochastic EM algorithm for estimating mixing proportions. Stochastic Models 9(4), 599–613 (1993).
Neyman, J. Outline of a theory of statistical estimation based on the classical theory of probability. Philosophical Transactions of the Royal Society of London. Series A, Mathematical and Physical Sciences 236(767), 333–380 (1937).
Efron, B. Bootstrap methods: another look at the jackknife. The Annals of Statistics 7(1), 1–26 (1979).
Efron, B. The jackknife, the bootstrap and other resampling plans. CBMS-NSF Regional Conference Series in Applied Mathematics. SIAM, Philadelphia (1982).
Kim, S., Potter, K., Craigmile, P., Peruggia, M. & Van Zandt, T. A Bayesian race model for recognition memory. Journal of the American Statistical Association 112(517), 77–91 (2017).
Lavine, M. An approach to robust bayesian analysis for multidimensional parameter spaces. Journal of the American Statistical Association, 86(415):779–786. Key reference for the factorization of independent priors in Bayesian modeling (1991).
Gelman, A.,et al. Bayesian Data Analysis. CRC Press, Boca Raton, FL, 3rd edition. Chapter 2 discusses the implications of prior independence and how the likelihood can induce posterior dependence even under independent priors (2013b).
Duane, S., Kennedy, A., Pendleton, B. & Roweth, D. Hybrid Monte Carlo. Phys Lett B 195, 216–222 (1987).
Neal, R. MCMC using Hamiltonian dynamics (In Handbook of Markov Chain Monte Carlo, 2011).
Wang, Z., Mohamed, S. & de Freitas, N. Adaptive Hamiltonian and Riemann manifold Monte Carlo samplers. In Dasgupta, S. and McAllester, D., editors, Proceedings of the 30th International Conference on Machine Learning (ICML 2013), 1462–70, (Brookline, MA. Microtome, 2013).
Hoffman, M. & Gelman, A. The No-U-Turn sampler: adaptively setting path lengths in Hamiltonian Monte Carlo. Journal of Machine Learning Research 15(1), 1593–1623 (2014).
Lawless, J. Statistical models and methods for lifetime data Vol. 362 (Wiley, 2011).
Aarset, M. How to identify bathtub hazard rate. IEEE Transactions Reliability 36, 106–108 (1987).
Hassan, A. & Nassr, S. The inverse Weibull generator of distributions, properties and applications. J Data Sci 16, 723–742 (2018).
Nassar, M. & Eissa, F. On the exponentiated Weibull distribution. Commun Stat Theor Methods 32, 1317–1336 (2003).
Thomas, S. & Tu, W. Learning Hamiltonian Monte Carlo in R. The American Statistician 75(4), 403–13 (2021).
Acknowledgements
Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2025R404), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Funding
This study is funded by Researchers Supporting Project number (PNURSP2025R404), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Q.R. Conceptualized the study, designed the MWM framework, and conducted simulations. M.A. Developed Bayesian algorithms (MCMC, HMC, RMHMC) and analyzed convergence. T.A. Led research, applied methods to appliance data, and drafted the manuscript. N.M.A. & H.A.E.W.K. Contributed to theory, optimization, and data analysis/review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix A
Appendix A
This appendix provides the expressions for the gradients required to implement the various techniques discussed in this work.
Gradient of the log-likelihood
The maximum likelihood estimates (MLEs) can be obtained by solving the following nonlinear likelihood equations
where,
We set the gradient of (23) equal to zero
The numerical expressions for the components of (17) are obtained as follows
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ramzan, Q., Amin, M., Alballa, T. et al. Algorithms and approximations for the modified Weibull model under censoring with application to the lifetimes of electrical appliances. Sci Rep 16, 1515 (2026). https://doi.org/10.1038/s41598-025-30943-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-30943-0