Abstract
Bromoform-rich extracts of Asparagopsis taxiformis represent a promising sustainable strategy for mitigating methane emissions in ruminants. Accurate quantification of bromoform is essential to ensure both efficacy and safety. Although gas chromatography-mass spectrometry (GC-MS) offers high accuracy, it is time-consuming, resource-intensive, and requires significant chemical reagents. This study pioneers the use of Fourier-transform near-infrared (FT-NIR), Fourier-transform mid-infrared (FT-MIR), and their data fusion (FT-MIR-NIR) combined with a recursive weighted partial least squares (rPLS) variable selection algorithm for rapid, non-destructive quantification of bromoform in seaweed extracts, validated against GC-MS. The partial least squares regression (PLSR) models employing rPLS based on FT-MIR spectra (R2CV = 0.95, RMSECV = 3.59 ppm (µL/L)) and fused FT-MIR-NIR spectra (R2CV = 0.94, RMSECV = 3.90 ppm) demonstrated robust predictive performance for bromoform quantification, though slightly lower than GC-MS (R2CV = 0.99, RMSECV = 3.21 ppm). This work establishes a high-throughput, accurate, and efficient alternative method for bromoform analysis, aligning with green chemistry principles by reducing chemical usage and enhancing sustainability. The approach holds significant potential for applications in animal nutrition and industrial feed processing. Future studies should further explore this method for bromoform quantification in other seaweed extracts and for high-throughput screening in animal feed industrial processes.
Similar content being viewed by others
Introduction
A major issue that the agri-food sector is currently facing relates to the increased emissions of greenhouse gases (GHGs) caused by traditional agricultural production systems guided also by the increased food demands by a growing human population1. Amongst all GHGs, methane has a global warming potential 28 times greater than that of carbon dioxide, and ruminant livestock contribute have been estimated to contribute to a quarter of global methane emissions2. In order to address this challenge, several animal nutrition strategies have been implemented, including feeding ruminants various anti-methanogenic agents (such as bromoform, tannins, saponins, flavonoids and chemical compounds such as 3-nitrooxypropanol (3-NOP) and ethyl-3-NOP,) which could potentially result in a reduction or inhibition of methane emissions3. Among them, the use of red seaweeds (mainly Asparagopsis taxiformis and Asparagopsis armata) have been described as one of the most promising feeding strategy, attributing these anti-methanogenic effects to the presence of bromoform (CHBr3) in some of these species4. Kinley et al.5 proved that the addition of Asparagopsis taxiformis to ruminants feed improved the productivity of the livestock while reducing methane emissions, and no bromoform residues were detected in the animals’ meat, fat, organs or faeces. However, other studies in lactating dairy cows revealed that traces of bromoform (about 35% of the EU drinking water limit) were detected in the milk from animals supplemented high amounts of seaweed (333 g seaweed DW/day)6. Therefore, when using A. taxiformis in ruminant feeds, careful control of bromoform dosage is required from a safety perspective, including bromoform analysis, to prevent risks such as insufficient methane inhibition or bromoform carryover due to inconsistent seaweed intake7,8. New research in animal feed is focusing on the generation and application of bromoform-rich seaweed extracts instead of whole seaweed to improve the delivery efficiency and stability of the active ingredients, and thus, there is a need for accurate methods of analysis of bromoform in extracts9.
Currently, due to the volatility of bromoform, its quantitative analysis is usually performed on a gas chromatography-mass spectrometry (GC-MS) systems10. Although GC-MS is effective and reliable and can be considered as a standard method for these determinations, the analyses (sample preparation and analytical run) can be time-consuming and involve the use of high amounts of chemicals. In recent years, driven by the principles of green chemistry, laboratory work is moving towards more efficient use of sustainable biomass, use of water or agro-solvents instead of organic solvents, reduction or recovery of energy, efficient use of by-products, simpler experimental and testing processes, and extracts that are free of pollutants, solvent residues, and biodegradable11,12. Therefore, there is a need to develop accurate and non-destructive methods to quantify bromoform concentrations in seaweed extracts that could be comparable to GC-MS, while using less amount of chemicals operating easily and fast, producing little waste, and requiring lower energy inputs.
In recent years, with the rapid development of spectroscopy techniques, some emerging detection methods have attracted the attention of researchers, especially Fourier-transform near infrared (FT-NIR) and Fourier-transform mid infrared (FT-MIR). Both methods can obtain the overall chemical characteristics of the sample based on the vibration of the functional groups and chemical bonds of the molecules, and they have also been described as fast, clean and non-destructive qualitative analysis methods13,14. FT-MIR and FT-NIR have been used to analyse the polyphenol content, protein content in seaweed biomass, and the purity of fucoidan from brown macroalgae extracts15,16,17,18. However, in previous literature, researchers mainly focused on the determination of the content of chemical components in biomass or the purity of complex mixtures. To our knowledge there is currently no method using FT-NIR and/or FT-MIR for the prediction of bromoform concentration, and there is also no exploration of bromoform quantification based on data fusion of FT-NIR and FT-MIR spectral data. As interest in bromoform extracts grows, researching the novel analytical capabilities of FT-NIR and/or FT-MIR for bromoform quantification can be extremely valuable to shorten the time of the analysis performed by GC-MS while maintaining the accuracy and reducing the chemical consumption of currently available techniques.
This study aims to develop and validate rapid, environmentally friendly analytical methods based on FT-MIR, FT-NIR and FT-MIR-NIR fused spectroscopy as an alternative to GC-MS for the quantitative determination of bromoform in seaweed extracts. The potential advantages of these spectroscopic techniques compared to GC-MS have been also evaluated in terms of accuracy of the methods together with other aspects of green chemistry, including non-destructive, reduction of chemical consumption, analysis time and energy inputs. This research seeks to provide a reliable, green analytical method to support the efficient development and quality control of bromoform-rich seaweed extracts, contributing the application of bromoform extracts in sustainable animal nutrition.
Materials and methods
Biological Raw material and chemical reagents
Red seaweed A. taxiformis was provided by European Freeze Dry Ltd. (Preston, UK). HPLC grade methanol was purchased from Thermo Fisher Scientific (Dublin, Ireland). Bromoform standards (contains 2-methyl-2-butene as stabilizer, 99%) and other chemical reagents were of analytical grade and purchased from Sigma-Aldrich, Inc. (Wicklow, Ireland).
Generation of seaweed extracts containing bromoform
Bromoform extracts (n = 35) were generated using ultrasound-assisted extraction. Briefly, 3 g of red seaweed samples were mixed with 60 mL of pure methanol. The solvent of choice was based on previous results reported by Romanazzi et al.8 and Paul et al.19 that proposed methanol as the most efficient solvent for the extraction of halogenated alkanes compared with dichloromethane, ethanol, hexane, and chloroform. The mixtures were extracted using off/on time of 1 s in an ultrasound bath (UP400S, Hielscher Ultrasonics GmbH, Teltow, Germany) at 24 kHz at variable ultrasonic power (80–400 W, 20%−100% power) and extraction times (3–30 min). Following ultrasonic treatments the mixtures were centrifuged (4000 g, 5 min) at room temperature using a Rotina 38 R (Hettich, Tuttlingen, Germany), and the bromoform-rich supernatants were collected. Bromoform extracts were prepared on the day of analysis, stored in sealed vials at 4 °C in the dark to minimize volatilization and degradation, and analysed on the same day.
Bromoform determination via GC-MS
Bromoform content was quantified by GC-MS. Specifically, headspace sampling (HS) and gas chromatography (GC) parameters were modified from Szelewski20. For mass spectrometry (MS), the methods were adapted from Pérez Pavón et al.21 for the selection of internal standards naphthalene and bromoform. 1 mL of samples were diluted in 9 mL of methanol. 0.2 mL of bromoform standard (0–75 ppm (µL/L)) or sample solutions were mixed with 0.2 mL of 10 ppm naphthalene internal standard solution and 1.6 mL of water. The quantification was performed in a GC-MS (gas chromatograph: Clarus® 690, Perkin Elmer; mass spectrometer: Clarus® SQ 8 C, Perkin Elmer, Waltham, MA, USA) equipped with a headspace trap (TurboMatrix 40, Perkin Elmer, MA, USA), and an Agilent VF-200ms silica capillary column (30 m × 0.32 mm id, 1 μm film thickness). For the headspace trap process, 2 mL of mixtures were transferred to a 20 mL vial and sealed, the vials were equilibrated (70 °C, 10 min) and pressured to 15 psi for 1 min. The vapour phase was then extracted (1 cycle, 1.6 min decay) and transferred to the trap which was set to range from 40 to 280 °C, dry purge for 7 min and hold for 6 min. Desorption of the trap to the GC column was performed within 0.5 min. The column oven temperature was initially set to 30 ℃ for 0.30 min and then, the temperature increased at a rate of 10 ℃/min until reaching 220 ℃, while the run time was 19.3 min. The carrier gas was helium 99.996% purified using an oxygen, moisture and hydrocarbon gas filter (Agilent gas purifier, CP17973) and was set to 9 psi giving a column flow of 1.60 ml/min and velocity of 26.8 cm/s during the initial column conditions of 30℃. The MS inlet line temperature was 250 ℃, source temperature was 180 ℃ and multiplier was 1600 V. Scan mode was initially performed to determine retention times of bromoform and naphthalene. The scan range was 35 to 350 Da, scan time was 0.1 s and interscan delay was 0.06 s. For the quantification of bromoform and naphthalene, Selected Ion Recording (SIR) method was set up to their specific fragment peaks, respectively, with a dwelling time of 0.05s. The quantitative ion of bromoform and naphthalene was at m/z 172.8 and 128.2, respectively. The bromoform content was calculated based on the ratio of its peak area to that of the internal standard peak, which was then converted to content by using the standard curve (R2 = 0.9988).
Spectra acquisition
FT-NIR
The FT-NIR spectral acquisition was performed as described by Kasemsumran et al.22. Spectra were acquired using a FT-NIR spectrophotometer equipped with an InGaAs detector (MPA II, Bruker, Berlin, Germany). Briefly, 1 mL of samples or standards (0–100 ppm) were transferred into a glass vial with a cap after extraction and stored at 4 °C before measurements. At ambient temperature, each sample was measured with 64 scans over the wavelength range of 11,500–4000 cm− 1 at 8 cm− 1 resolution. Duplicate measurements were carried out for each standard or extracted sample.
FT-MIR
FT-MIR spectra were acquired as described by Li et al.23 with slight modification. The analyses were performed using a FT-IR spectrometer (INVENIO S, Bruker, Berlin, Germany) equipped with a HATR (horizontal attenuated total reflection) 2 mm ZnSe crystal plate (Pike Technologies, Madison, WI, USA) which is a multiple reflection ATR with 20 internal reflections. The spectra acquisition involved 64 scans with a resolution of 4 cm− 1 in the range of 650 to 4000 cm− 1. The samples were stored at 4 ℃ as described in Sect. 2.2 and tested at room temperature. During the measurements, 2 mL samples/standards were transferred onto the crystal.
Quantitative model development
FT-NIR and FT-MIR spectral data were pre-processed under autoscaling. For quantitative analysis, partial least squares regression (PLSR) models were developed on FT-NIR and FT-MIR pre-treated spectral data (as X-value) with the chemical references of bromoform content obtained using GC-MS as Y-value.
In spectral analysis, various algorithms could be used to establish relationships between spectral data and chemical quantification to determine the concentration of target substances, including recursive weighted partial least squares (rPLS), genetic algorithms (GA), stepwise regression, and uninformative variable elimination (UVE). GA simulates an evolutionary process to optimize wavelength selection and reduce multicollinearity, stepwise regression selects features based on their predictive significance, and UVE eliminates variables based on the statistical significance of regression coefficients24,25,26. In contrast, rPLS was used to iteratively reweights the spectral variables based on the regression coefficients calculated during PLSR modelling27. rPLS not only improves model interpretability and ease of operation but also generally achieves better predictive performance than unweighted models, which significantly improves computational efficiency and enables more efficient wavelength selection27.
In this study, rPLS variable selection algorithm iteratively reweights the input variables by standardizing the regression coefficients used to filter out non-relevant spectral wavelengths and select latent variables (LVs) for the model from spectral variables (FT-MIR (2348 variables), FT-NIR (2625 variables), and FT-MIR-NIR (4973 variables)) based on the last iteration or the one that provides the lowest root mean square error of cross validation (RMSECV)27. Statistical parameters were used to describe the model performances, including coefficient of determination for calibration (R2C) and leave-one-out cross-validation (R2CV), which were expected to be close to 1, root mean square error of calibration (RMSEC) and RMSECV, which were expected to be close to 0, and bias of cross-validation (CV bias) that is ideally close 0. A data fusion based on concatenation of both FT-NIR and FT-MIR spectral data was also carried out for PLSR model development. Data analysis of this work was carried out using in-house written scripts in MATLAB 2023a (The Mathworks, Natick, MA, USA).
Results and discussion
GC-MS quantification of bromoform
Table 1 shows the effects of different ultrasound power (20–100%) and time of extraction on the extraction of bromoform from A. taxiformis. The extracts achieved at 60% ultrasonic power and 30 min achieved extracts with the highest concentration of bromoform (76.25 ± 3.53 ppm). When extracting bromoform, choosing the right ultrasonic power and time is particularly important to increase the concentration of the bromoform extract, as excessive cavitation can result in the degradation of the extracted bromoform28. The concentration of bromoform in the extracts generated in this study were lower than other previously reported by Magnusson et al.9. This could be explained by differences in the concentration of bromoform in the original biomass used as well as differences in its processing prior, during and post-extraction. Magnusson et al.9 used an oil immersion method to extract freshly harvested A. taxiformis, while the biomass used in the current study was previously dried, milled and stored prior to bromoform extraction. Future studies are needed in order to clarify pre- and post-extraction processing in order to preserve these volatile compounds, as well as the effect of novel processing methods to enable the generation of extracts with high yields of bromoform. It is worth noting that methanol was used as the extraction solvent in this study due to its common and efficient use in bromoform extraction and quantification. In the future, the application prospects of other green extraction solvents such as ethanol, aqueous mixtures, and deep eutectic solvents in bromoform extraction need to be further explored29.
With respect to the quantification of bromoform from these extracts, GC-MS demonstrated its excellent capability for the quantification of the compounds of interest. These GC-MS measurements served as reference values for subsequent quantitative model development based on FT-MIR, FT-NIR, and data fusion of FT-MIR and FT-NIR spectral data (FT-MIR-NIR) aiming to develop and assess the use of greener analytical techniques. The application of the bromoform quantitative technique established by GC-MS and bromoform standards in the quantitative process of bromoform extracts was established in this research. For GC-MS, when measuring standards the method was really accurate (R2C = 0.9988 and R2CV = 0.9963, see Fig. 1A); with RMSEC and RMSECV of 1.47 ppm and 2.55 ppm, respectively. The measurement of bromoform in the extracts continued to perform well with slightly lower R2C (0.9948) and R2CV (0.9946) as seen Fig. 1B. Moreover, the RMSEC and RMSECV of the bromoform extracts quantitatively measured by GC-MS were 3.15 ppm and 3.21 ppm, respectively.
(A) Scatter plot of bromoform standards and (B) extracts (n = 35 extracts, measured in duplicate) on validated bromoform content determined by GC-MS.
Bromoform calibration models based on FT-MIR spectral data
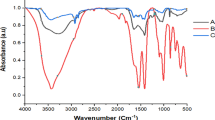
Figure 2A and B present the FT-MIR spectra of the bromoform standard in methanol and the bromoform extracted from A. taxiformis using methanol, respectively. The peak at 675 cm− 1 can be attributed to the stretching vibration of the C-Br bond in bromoform30. The peak at 1023–1038 cm− 1 reflects the C–O stretching vibration, while the broad absorption band observed between 3285 and 3355 cm− 1 corresponds to the O–H stretching vibration of hydroxyl groups of methanol in standards solution31,32. Furthermore, the spectral features in these regions within the A. taxiformis extract likely include contributions from O–H and C–O stretching vibrations of co-extracted compounds, such as polyphenols and polysaccharides, from the seaweed matrix, which is also observed from methanol extracts of the seaweed Sargassum polycystum33. The peak at 1448 cm− 1 and distinct doublet peaks at 2944 cm− 1 and 2831 cm− 1 are representative of C–H bending modes and the asymmetric and symmetric C–H stretching vibrations, which represents the methyl group of both bromoform and methanol34,35. In addition, the weak absorption band at approximately 850 cm− 1 in the extract may be related to the out of plane C–H bending vibrations characteristic of carbohydrates present in its co-extract, which is not observed in bromoform standards solution36.
Furthermore, the FT-MIR spectra of all standards (n = 12) and A. taxiformis methanol extracts (n = 35) were processed for modelling and quantification of bromoform contents. The rPLSvariable selection process was applied to select the FT-MIR wavenumbers by iteratively optimizing their weights27. For the standards, rPLS selected 20 variables from a total of 2348 variables (Fig. 2C), which were subsequently used to develop a PLSR model based on the Semi-Implicit Method for Pressure Linked Equations (SIMPLS) algorithm. The selected FT-MIR wavenumbers included 693.488, 719.173, 1031.67, 1038.81–1040.23.81.23, 1865–1867.85.85, 2851.01–2855.29.01.29, 2922.35, 3300.49, 3329.03–3330.46.03.46, 3336.16–3337.59.16.59, 3928.34–3929.77 cm− 1, which were used to predict bromoform concentration using FT-MIR spectral data of bromoform standards (Fig. 2D). The model was developed based on four LVs and demonstrated robust fitting performance of the calibration (R2C = 0.99, RMSEC = 3.29 ppm) and the leave-one-out cross-validation (R2CV = 0.972, RMSECV = 6.13 ppm) with a systematic bias (cross-validation (CV) bias = −0.568). It can be observed that rPLS variable selection process significantly enhanced the predictive power of FT-NIR. In the model developed based on all the 2348 spectral variables collected, the model performance only achieved R2CV of 0.64 with RMSECV of 21.23 ppm, as shown in Table 2.
Modelling was also explored based on the FT-MIR spectral data of bromoform extracts from A. taxiformis. The model developed based on the whole spectral variables collected (N = 2348) demonstrated that it could not be used to predict the bromoform content in the extract with a R2CV value of 0.5 and a RMSECV value of 12.59 ppm. However, when using rPLS to select variables of FT-MIR spectral data of bromoform extracts samples (n = 35), 22 wavelength variables were selected from a total of 2348 variables (Fig. 2E). Furthermore, the PLSR model was developed based on the rPLS selected spectral variables (N = 22) including 653.534, 657.815, 663.523, 679.219, 683.5, 692.062, 849.024, 1038.81, 2341.59, 3230.57, 3243.41, 3251.97, 3260.54–3261.96.54.96, 3264.82, 3270.53, 3274.81, 3280.51, 3320.47, 3349.01, 3376.12, 3408.94 cm− 1 for bromoform content prediction in the extracts. The model involved 9 LVs and obtained robust calibration and cross-validation performances (i.e., R2C = 0.99, RMSEC = 1.36 ppm; R2CV = 0.95, RMSECV = 3.59 ppm), which demonstrated a significant improvement in predictive ability compared with the performance of the model developed without variable selection. Models developed based on FT-MIR spectral variables selected using rPLS for both bromoform standards’ and extracts’ predictions obtained an excellent and widely accepted threshold for predictive utility, with R2CVs > 0.9. The results indicate the potential of FT-MIR combined with PLSR under rPLS spectral variable selection as a powerful tool for bromoform quantification—even in complex bromoform extracts derived from red seaweed biomass. To our knowledge there are no previous studies using a similar methodology when quantifying bromoform.
(A) FT-MIR spectra of bromoform standards (n = 12); (B) FT-MIR spectra of bromoform extracts (n = 35); (C) wavelength variables (N = 22) with high iteration numbers selected from the FT-MIR spectral data of bromoform standards (n = 12) using rPLS for PLSR modelling; (D) scatter plot and regression fit-line of the PLSR model developed based on selected FT-MIR spectral variables (LVs = 4) of bromoform standards (n = 12) on measured bromoform content vs. cross-validated bromoform content; (E) wavelength variables (N = 22) with high iteration numbers selected from FT-MIR spectral data of bromoform extracts (n = 35) using rPLS for PLSR modelling; (F) scatter plot and regression fit-line of PLSR model developed based on selected FT-MIR spectral variables (LVs = 9) of bromoform extracts (n = 35) on measured bromoform content vs. cross-validated bromoform content.
Bromoform calibration models based on FT-NIR spectral data
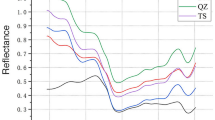
FT-NIR spectra of the bromoform standards (Fig. 3A) and extracts (Fig. 3B) exhibited significant absorbance variations in the 4358–4466 cm− 1 region, which primarily correspond to combination bands involving O-H stretching and C-H stretching vibration of methanol37,38. The peak observed at 4809–4818 cm− 1 is attributed to a combination of O-H bending and C-O stretching vibrations39. The FT-NIR spectra of bromoform-methanol solutions are influenced by the preference of bromoform, owing to its ability to form intermolecular hydrogen bonds with methanol. The intermolecular hydrogen bonding between bromoform and methanol were also observed by Shalit et al.40 using 2D Raman–THz spectroscopy. In the higher wavenumber regions, the band at 5881 cm− 1 is attributed to the first overtone band of C–H stretching, the band at 6326 cm− 1 can be related to the first overtone of O-H stretching, while the peak at 8324 cm− 1 can be attributed to the second overtone of C-H stretching39,41. In addition, the spectral changes in the 4000–5000 cm− 1 region were more significant in the extract samples than in the standards, which was due to the changes in the content of co-extracts as also observed in Lupinus mutabilis Seeds polyphenol extracts42. The presence and variation of functional groups in these co-extracts resulted in the observed spectral variability. Notably, distinct differences between the FT-NIR spectra of the bromoform extracts and standards, there were obvious differences in the region of 4989–5369 cm− 1, which are due to the influence of co-extracts. Okello et al.38 found that the extract of Aspilia africana showed peaks in a similar region and attributed it to the hydroxyl groups of the polyphenols in the extract and their corresponding aromatic ring-related vibrations. This means that our co-extracts are likely polyphenols, as expected for the method of choice for the preparation of the extracts using methanol. Previous studies reported the use of this solvent as the most efficient one for polyphenol compounds from A. taxiformis compared with petroleum ether, chloroform, and ethyl acetate43.
As shown in Table 2, the prediction model developed based on all the collected FT-NIR spectral variables (N = 2625) in the wavelength range of 3997.637–11495.71 cm− 1 had poor predictable ability for bromoform content prediction, with an R2CV value of 0.01 and a RMSECV of 54.49 ppm. After using rPLS, 11 spectral variables including 3997.64–4000.49.64.49, 4326.25, 4354.82, 4434.83, 4503.41, 4523.42, 4860.6, 4874.89, 5149.21–5152.06 cm− 1 were selected for a further PLSR model development (see Fig. 3C). The developed PLSR model significantly improved performance on the prediction of bromoform content in the standard solution samples, which makes the RMSECV from 54.49 ppm (using all spectral variables) to 4.13 ppm (after rPLS spectral variable selection). The calibration and cross-validation results revealed a R2CV of 0.986 (~ 1) with RMSECV of 4.13 ppm; additionally with CV bias of 0.37 (Fig. 3D), which could be classified as robust model.
For the prediction of bromoform content in the seaweed extracts, 34 variables (4020.5–4023.35.5.35, 4043.36–4046.21.36.21, 4117.65, 4154.8, 4206.23, 4297.67–4300.53.67.53, 4309.1, 4394.83, 4420.55, 4437.69–4440.55.69.55, 4480.55–4483.41.55.41, 4503.41, 4511.99, 4537.7–4546.28.7.28, 4783.45, 4797.74, 4806.31, 4817.74–4820.6.74.6, 4837.74, 5649.27–5663.56 cm− 1) were selected using rPLS (Fig. 3E). These selected variables were subsequently used for further PLSR modelling achieving robust prediction performance (Fig. 3F). The PLSR model based on the FT-NIR spectra of bromoform extracts (n = 35) involving 8 latent variables achieved R2CV of 0.88, RMSECV of 5.68 ppm, and CV bias of 0.413, which could be considered as a model with good predictive ability. The potential of FT-NIR spectroscopic technique for bromoform content prediction was well demonstrated by the performances of models developed on the selected FT-NIR spectral variables of both bromoform standards and extracts. The rPLS selected spectral variables were mainly found in the wavelength range of 4000–5000 cm− 1, which were adjacent to the FT-MIR wavelength range. Thus, data fusion of FT-MIR and FT-NIR spectral data in a low level by concatenation could avail the prediction model development with complementary spectral features for bromoform content prediction.
(A) FT-NIR spectra of bromoform standards (n = 12); (B) FT-NIR spectra of bromoform extracts (n = 35); (C) wavelength variables (N = 11) with high iteration numbers selected from the FT-NIR spectral data of bromoform standards (n = 12) using rPLS for PLSR modelling; (D) scatter plot and regression fit-line of the PLSR model developed based on selected FT-NIR spectral variables (LVs = 7) of the bromoform standards (n = 12) on measured bromoform content vs. cross-validated bromoform content; (E) the wavelength variables (N = 34) with high iteration numbers selected from FT-NIR data of bromoform extracts (n = 35) using rPLS for PLSR modelling; (F) scatter plot and regression fit-line of the PLSR model developed based on the selected FT-NIR spectral variables (LVs = 8) of the bromoform extracts (n = 35) on measured bromoform content vs. cross-validated bromoform content.
Data fusion FT-MIR-NIR
Spectral data fusion is an emerging spectral data processing method, which can be used in a variety of combinations of different instruments providing synergy and mutual information, and better predictability for data processing44. FT-NIR spectra (4000–12500 cm− 1) can capture the overtones and combined absorption bands of hydrogen-containing groups (C–H, O–H, N–H), which can be fused with FT-MIR spectra (650–4000 cm− 1), while the integrated data after fusion can have the specificity of both FT-NIR and FT-MIR in the PLSR model, enhancing the information richness, accuracy and generalization ability of the model45. Data fusion of FT-MIR and FT-NIR is highly effective for analysing complex chemical compositions, making it widely applicable for geographical traceability of Panax notoginseng44 and detection of adulteration in fruit pulp46 and olive oil47. The data fusion (FT-MIR-NIR) spectra of bromoform standards (Fig. 4A) or extracts (Fig. 4B) covered the wavelength range of 650–12,500 cm− 1, providing complementary spectral features for the prediction of their bromoform contents. As shown in Table 2, The PLSR model developed on the fused FT-MIR-NIR included 4973 spectral variables and it had poor or moderate prediction performance for bromoform standards (R2CV = 0.04) or extracts (R2CV = 0.55), respectively. For spectral feature selection, the spectral variables of the fused spectra were selected using the rPLS variable selection algorithm for both bromoform standards (Fig. 4C) and extracts (Fig. 4E). Figure 4D represents the PLSR model developed based on 19 selected spectral variables (719.173, 1038.81–1040.23.81.23, 1865–1866.43.43, 2851.01–2855.29.01.29, 2920.93–2922.35.93.35, 3300.49, 3336.16, 4460.55, 4491.98–4494.84.98.84, 4797.74, 4823.45–4826.31 cm− 1), selected from the fused FT-MIR-NIR spectra of bromoform standards (n = 12) and included 4 LVs for model development. These data indicated that the model performance for bromoform standards achieved R2CV of 0.96 and RMSECV of 6.66 ppm. For the bromoform standard, FT-MIR, FT-NIR and FT-MIR-NIR are all robust prediction models. However, it can be found that for the bromoform standard, the fusion spectrum did not improve the R2CV when compared with single spectrum (FT-MIR: 0.97 and FT-NIR: 0.99), and the RMSECV of the fusion spectrum was slightly higher than that of the single spectrum (FT-MIR: 6.13 ppm and FT-NIR: 4.13 ppm). For the quantitative analysis of FT-MIR-NIR spectra of bromoform extracts (Fig. 4F), 19 spectral variables (652.107, 656.388, 667.804, 680.646, 690.635, 3233.43, 3251.97, 3261.96, 3270.53, 3274.81, 3323.32, 3367.56, 3374.69, 3421.78, 4209.09, 4503.41, 4543.42, 4766.3, 5657.84 cm− 1) were used (Fig. 4E) to develop the PLSR model shown in Fig. 4F. For those extracts, the fusion spectrum prediction model (R2CV of 0.94 and RMSECV of 3.9 ppm) had better predictive ability compared with FT-NIR (R2CV of 0.88 and RMSECV of 5.69 ppm) and slightly less than FT-MIR (R2CV of 0.95 and RMSECV of 3.59 ppm). Therefore, in this study the bromoform quantitative model developed based on the FT-MIR-NIR fusion spectrum can effectively optimize the bromoform quantitative model based on FT-NIR, and may have a wider application prospect in bromoform extracts from other sources in the future. The observed result could be attributed to the specificity of bromoform, consisted of only three C-Br bonds and one C-H bond. Therefore, FT-MIR spectra contains the most critical predictive information, while FT-NIR provides limited supplementary data and may introduce minor non-informative variance.
(A) FT-MIR-NIR fused spectra of bromoform standards (n = 12); (B) FT-MIR-NIR fused spectra of bromoform extracts (n = 35); (C) wavelength variables (N = 19) selected from FT-MIR-NIR fused data of bromoform standards (n = 12) using rPLS for PLSR modelling; (D) scatter plot and regression fit-line of the PLSR model developed based on selected FT-MIR-NIR spectral variables (LVs = 4) from bromoform standards (n = 12) on measured bromoform content vs. cross-validated bromoform content; (E) wavelength variables (N = 19) selected from FT-MIR-NIR fused data of bromoform extracts (n = 35) using rPLS for PLSR modelling; (F) scatter plot and regression fit-line of the PLSR model developed based on selected FT-MIR-NIR spectral variables (LVs = 8) from bromoform standards (n = 35) on measured bromoform content vs. cross-validated bromoform content.
Comparison of model performance
Overall, the three spectroscopic techniques used, FT-MIR, FT-NIR, and data fusion, combined with PLSR modelling and rPLS spectral variable selection algorithms demonstrated high potential for quantitative analysis of bromoform content in the prepared standards and the extracts of A. taxiformis. Table 2 summarized the PLSR model performances of all three techniques for the prediction of bromoform content in both standards and extracts. PLSR models based on all the collected spectral variables of FT-MIR (N = 2348), FT-NIR (N = 2625), and FT-MIR-NIR (N = 4973) generally demonstrated weak performances with R2CVs of 0.01–0.64 and RMSECVs of 21.23–54.49 ppm for bromoform content prediction in standards, and with R2CVs of 0.54–0.55 and RMSECVs of 11.35–12.59 ppm for bromoform prediction in extracts. However, the PLSR models developed based on rPLS selected spectral variables of FT-MIR (N = 20 or 22), FT-NIR (N = 11 or 34), FT-MIR-NIR (N = 19 or 19) for standards or sample demonstrated strong potential for bromoform prediction in both standards and extracts. These models achieved R2CVs of 0.9–0.97, RMSECVs of 4.13–6.66 ppm for bromoform prediction in standards, and R2CVs of 0.88–0.95, RMSECVs of 3.59–5.68 ppm for bromoform prediction in extracts. The rPLS variable selection step had a positive impact on the PLSR model, i.e., decreased RMSECV and increased R2CV.
In conclusion, the rPLS algorithm used in this study played a critical role in selecting the most related spectral bonds from the whole wavelength range for quantitative analysis of bromoform content. In addition, the computation power and time required for modelling was significantly reduced based on the multiple spectral variables selected. Furthermore, it is worthy to noting that the model developed based on the rPLS selected FT-NIR spectral variables (N = 11) obtained overall the best prediction performance (i.e., R2CV = 0.99, RMSECV = 4.13 ppm) on bromoform content in standards. Thus, the high potential of FT-NIR spectroscopic technique for quantification of pure bromoform content in standard was clearly revealed in this study. On the other hand, the model developed based on the selected FT-MIR spectral variables (N = 20) demonstrated significant improved prediction performance (R2CV = 0.95, RMSECV = 3.59 ppm) on bromoform content in extracts.
Compared with the prediction performance of traditional GC-MS (R2CV = 0.9963, RMSECV = 2.55 ppm), FT-NIR spectroscopic technique has demonstrated similar capabilities (R2CV = 0.99, RMSECV = 4.13 ppm) for standards’ prediction; but not sufficient for extracts’ prediction (R2CV = 0.88, RMSECV = 5.68 ppm). On the other hand, FT-MIR and fused FT-MIR-NIR spectral quantitative analysis demonstrated improved bromoform prediction performances (i.e., R2CV = 0.95, RMSECV = 3.59 ppm of FT-MIR; R2CV = 0.94, RMSECV = 3.90 ppm of FT-MIR-NIR). Furthermore, the main parameters when considering using an analytical technique were compared between GC-MS, FT-MIR, FT-NIR, and FT-MIR-NIR and summarised in Table 3. GC-MS is a conventional destructive method for high-precision laboratory analysis, however, it’s also time- and chemical-consuming48. In FT-MIR and FT-NIR techniques, the primary source of energy consumption is the light source, and measurement times are extremely short, resulting in extremely low energy consumption levels of 0.002–0.004 kWh and 0.005–0.01 kWh, respectively. In contrast, GC-MS requires continuous high temperature and vacuum conditions during operation, and due to its longer analysis times, its energy consumption significantly increases, reaching 2.25 kWh, which is hundreds of times more energy than the other two techniques. Furthermore, FT-MIR (2 mL) and FT-NIR (1 mL) analysis generate far less waste than GC-MS (12 mL). As for the bromoform extracts obtained from seaweed, the chemical composition is much more complex than that of the standards, so the predictive abilities of these spectra are reduced compared with that of the standards. FT-MIR spectroscopy is suitable for quantitative speciation analyses of chemical mixtures, whereas FT-NIR spectroscopy contained broad overlapping absorption bands and it was not selective, which was the main reason why these spectra had less predictable abilities compared to those of GC-MS49. However, spectroscopic techniques (FT-MIR, FT-NIR, FT-MIR-NIR) are rapid and non-destructive measurements, which can contribute to the development of efficient, low-cost and highly adaptable preferred quantification strategy for bromoform content determination.
Conclusions
This study pioneered a rapid spectroscopic quantitative method that can be used to replace traditional GC-MS quantification in future studies. To our knowledge this is the first study exploring the potential of FT-MIR, FT-NIR and data fusion (FT-MIR-NIR) combined with PLSR modelling and rPLS spectral variable selection for quantitative analysis of bromoform content in both standard solutions and seaweed extracts. The data fusion of FT-MIR-NIR spectral data (650–12500 cm− 1) significantly improved the information richness and prediction performance of the PLSR model by concatenation of the spectral features of FT-MIR (650–4000 cm− 1) and FT-NIR (4000–12500 cm− 1). FT-NIR had high accuracy in the quantitative analysis of bromoform standards (R2CV = 0.99, RMSECV = 4.13 ppm), while FT-MIR was more suitable for bromoform analysis in extracts (R2CV = 0.95, RMSECV = 3.59 ppm). The investigation in this study will assist on further the commercial application of A. taxiformis as a feed additive, especially the extraction process of bromoform and the prospect of large-scale, accurate, greener, rapid and quantitative application. The limitation of this study is that only methanol was used as the extraction solvent, while future work should explore the application of other green solvents like ethanol, aqueous mixtures, and deep eutectic solvents for bromoform extraction. Furthermore, the matrix-matched calibration effects of different solvents during dilution must be considered in subsequent quantification. In future high-throughput testing, more extracted samples could be included, divided into calibration and validation datasets, and the Kennard-Stone algorithm could be used for more robust model development and applications. Future research can further apply fusion spectroscopy to the rapid and non-destructive quantification of other bioactive substances in seaweed extracts, broadening the future applications of these technologies for an accurate measurement of multiple functional ingredients that could be added as functional ingredients to animal feed.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Halpern, B. S. et al. The environmental footprint of global food production. Nat. Sustain. 5, 1027–1039 (2022).
Loan, T. D., Vickers, C. E., Ayliffe, M. & Luo, M. β-Dicarbonyls facilitate engineered microbial bromoform biosynthesis. ACS Synth. Biol. 13, 1492–1497 (2024).
McGurrin, A., Maguire, J., Tiwari, B. K. & Garcia-Vaquero, M. Anti-methanogenic potential of seaweeds and seaweed-derived compounds in ruminant feed: current perspectives, risks and future prospects. J. Anim. Sci. Biotechnol. 14, 1–27 (2023).
Glasson, C. R. K. et al. Benefits and risks of including the bromoform containing seaweed Asparagopsis in feed for the reduction of methane production from ruminants. Algal Res. 64, 102673 (2022).
Kinley, R. D. et al. Mitigating the carbon footprint and improving productivity of ruminant livestock agriculture using a red seaweed. J. Clean. Prod. 259, 120836 (2020).
Muizelaar, W., Groot, M., van Duinkerken, G., Peters, R. & Dijkstra, J. Safety and transfer study: transfer of bromoform present in Asparagopsis taxiformis to milk and urine of lactating dairy cows. Foods 10, 584 (2021).
Vucko, M. J., Magnusson, M., Kinley, R. D., Villart, C. & de Nys, R. The effects of processing on the in vitro antimethanogenic capacity and concentration of secondary metabolites of Asparagopsis taxiformis. J. Appl. Phycol. 29, 1577–1586 (2017).
Romanazzi, D. et al. Rapid analytical method for the quantification of bromoform in the red seaweeds Asparagopsis armata and Asparagopsis taxiformis using gas Chromatography-Mass spectrometry. ACS Agric. Sci. Technol. 1, 436–442 (2021).
Magnusson, M., Vucko, M. J., Neoh, T. L. & de Nys, R. Using oil immersion to deliver a naturally-derived, stable bromoform product from the red seaweed Asparagopsis taxiformis. Algal Res. 51, 102065 (2020).
Hutchings, J. L., Grebneva, Y., Dilmetz, S. J., Pincher, D. W. M. & Hoffmann, P. Analytical methods for the analysis of bromoform in red seaweed Asparagopsis armata and Asparagopsis taxiformis – A review. Algal Res. 79, 103478 (2024).
Sajid, M. & Płotka-Wasylka, J. Green analytical chemistry metrics: A review. Talanta 238, 123046 (2022).
Garcia-Vaquero, M., Sweeney, T., O’Doherty, J. & Rajauria, G. Recent advances in the use of greener extraction technologies for the recovery of valuable bioactive compounds from algae. Recent. Adv. Micro Macroalgal Process. 96–122. https://doi.org/10.1002/9781119542650.ch4 (2021).
Yao, C., Qi, L., Zhong, F., Li, N. & Ma, Y. An integrated chemical characterization based on FT-NIR, GC–MS and LC-MS for the comparative metabolite profiling of wild and cultivated Agarwood. J. Chromatogr. B. 1188, 123056 (2022).
Sakkas, L., Pappas, C. S. & Moatsou, G. FT-MIR analysis of Water-Soluble extracts during the ripening of sheep milk cheese with different phospholipid content. Dairy 2, 530–541 (2021).
Zhao, M., Garcia-Vaquero, M., Przyborska, J., Sivagnanam, S. P. & Tiwari, B. The development of analytical methods for the purity determination of fucoidan extracted from brown seaweed species. Int. J. Biol. Macromol. 173, 90–98 (2021).
Niemi, C. et al. Rapid and accurate determination of protein content in North Atlantic seaweed by NIR and FTIR spectroscopies. Food Chem. 404, 134700 (2023).
Tadmor Shalev, N. et al. NIR spectroscopy and artificial neural network for seaweed protein content assessment in-situ. Comput. Electron. Agric. 201, 107304 (2022).
Cao, X. et al. Near-infrared spectroscopy as a tool to assist sargassum fusiforme quality grading: harvest time discrimination and polyphenol prediction. Postharvest Biol. Technol. 192, 112030 (2022).
Paul, N., de Nys, R. & Steinberg, P. Chemical defence against bacteria in the red Alga Asparagopsis armata: linking structure with function. Mar. Ecol. Prog Ser. 306, 87–101 (2006).
Szelewski, M. Environmental Volatiles using a Headspace-GC/MSD Analyzer. Agilent Technologies, Inc., Application Note, publication 5990–7907 www.agilent.com/chem
Pérez Pavón, J. L., Martín, S. H. & García Pinto, C. Moreno Cordero, B. Headspace-programmed temperature vaporizer-fast gas chromatography-mass spectrometry coupling for the determination of trihalomethanes in water. J. Chromatogr. A. 1194, 103–110 (2008).
Kasemsumran, S. et al. Simultaneous monitoring of the evolution of chemical parameters in the fermentation process of pineapple fruit wine using the liquid probe for Near-Infrared coupled with chemometrics. Foods 11, 377 (2022).
Li, Y. et al. Investigation of enzyme-assisted methods combined with ultrasonication under a controlled alkali pretreatment for agar extraction from gelidium sesquipedale. Food Hydrocoll. 120, 106905 (2021).
Jin, J. & Wang, Q. Evaluation of informative bands used in different PLS regressions for estimating leaf biochemical contents from hyperspectral reflectance. Remote Sens. 11, 197 (2019).
Casian, T. et al. Challenges and opportunities of implementing data fusion in process analytical Technology—A review. Molecules 27, 4846 (2022).
Andries, J. P. M., Heyden, V., Buydens, L. M. C. & Y. & Improved variable reduction in partial least squares modelling by Global-Minimum error Uninformative-Variable elimination. Anal. Chim. Acta. 982, 37–47 (2017).
Rinnan, Å., Andersson, M., Ridder, C. & Engelsen, S. B. Recursive weighted partial least squares (rPLS): an efficient variable selection method using PLS. J. Chemom. 28, 439–447 (2014).
Park, B., Cho, E., Son, Y. & Khim, J. Distribution of electrical energy consumption for the efficient degradation control of THMs mixture in sonophotolytic process. Ultrason. Sonochem. 21, 1982–1987 (2014).
Hessel, V. et al. Sustainability of green solvents – review and perspective. Green. Chem. 24, 410–437 (2022).
Dong, M., Zheng, K., Shen, Z. & Liu, C. Light-dependent Br-org production in terrestrial plants under acetaminophen stress and the bromination mechanisms mediated by photosystem. J. Environ. Sci. 153, 275–288 (2025).
Mohan Reddy, Y. et al. Phytochemical profiling of methanolic fruit extract of gardenia latifolia ait. By lc-ms/ms analysis and evaluation of its antioxidant and antimicrobial activity. Plants 10, 1–10 (2021).
Coldea, T. E. et al. Rapid quantitative analysis of ethanol and prediction of methanol content in traditional fruit brandies from Romania, using FTIR spectroscopy and chemometrics. Not Bot. Horti Agrobot Cluj-Napoca. 41, 143 (2013).
Sobuj, M. K. A. et al. Qualitative and quantitative phytochemical analysis of brown seaweed sargassum polycystum collected from Bangladesh with its antioxidant activity determination. Food Chem. Adv. 4, 100565 (2024).
Batley, R. J. et al. Rapid screening of Methane-Reducing compounds for deployment via water with a commercial livestock supplement using in vitro and FTIR-ATR analyses. Methane 3, 437–455 (2024).
Rouhi, H. et al. Point-of-need quantitative detection of trihalomethanes in environmental water samples using a highly sensitive and selective fiber-based preconcentration system. J. Appl. Polym. Sci. 140, 1–10 (2023).
Selmi, A., Khiari, R., Snoussi, A. & Bouzouita, N. Analysis of minerals and heavy metals using ICP-OES and FTIR techniques in two red seaweeds (Gymnogongrus Griffithsiae and Asparagopsis taxiformis) from Tunisia. Biol. Trace Elem. Res. 199, 2342–2350 (2021).
Quintero Arenas, M. A., Meza-Márquez, O. G., Velázquez-Hernández, J. L., Gallardo-Velázquez, T. & Osorio-Revilla, G. Quantification of adulterants in Mezcal by means of FT-MIR and FT-NIR spectroscopy coupled to multivariate analysis. CYTA - J. Food. 18, 229–239 (2020).
Okello, D. et al. Antioxidant Activity, Polyphenolic Content, and FT-NIR Analysis of Different Aspilia africana Medicinal Plant Tissues. Evidence-Based Complement. Altern. Med. 1–11 (2021). (2021).
Zhang, Z. et al. Rapid Geographical Origin Identification and Quality Assessment of Angelicae Sinensis Radix by FT-NIR Spectroscopy. J. Anal. Methods Chem. 1–12 (2021). (2021).
Shalit, A., Mousavi, S. J. & Hamm, P. 2D Raman–THz spectroscopy of binary CHBr 3 –MeOH solvent mixture. J. Phys. Chem. B. 125, 581–586 (2021).
Tahir, H. et al. (ed, E.) HPLC-DAD, GC–MS, UHPLC-Q-TOF/MS2, ICP-MS and FT-NIR combined with the chemometrics explain seasonal variation in Cassia Senna L. leaves from the Coast of red sea in Sudan. South. Afr. J. Bot. 163 20–29 (2023).
Estivi, L. et al. Free phenolic Compounds, antioxidant capacity and FT-NIR survey of debittered Lupinus mutabilis seeds. Processes 10, 1637 (2022).
Neethu, P., Suthindhiran, K. & Jayasri, M. Antioxidant and antiproliferative activity of Asparagopsis taxiformis. Pharmacognosy Res. 9, 238 (2017).
Li, Y., Zhang, J. Y. & Wang, Y. Z. FT-MIR and NIR spectral data fusion: a synergetic strategy for the geographical traceability of Panax Notoginseng. Anal. Bioanal Chem. 410, 91–103 (2018).
Wang, H. et al. NIR and MIR spectral feature information fusion strategy for multivariate quantitative analysis of tobacco components. Chemom Intell. Lab. Syst. 253, 105222 (2024).
Alamar, P. D., Caramês, E. T. S., Poppi, R. J. & Pallone, J. A. L. Detection of fruit pulp adulteration using multivariate analysis: comparison of NIR, MIR and data fusion performance. Food Anal. Methods. 13, 1357–1365 (2020).
Li, Y., Xiong, Y. & Min, S. Data fusion strategy in quantitative analysis of spectroscopy relevant to Olive oil adulteration. Vib. Spectrosc. 101, 20–27 (2019).
Rasekh, M., Karami, H., Kamruzzaman, M., Azizi, V. & Gancarz, M. Impact of different drying approaches on VOCs and chemical composition of mentha spicata L. essential oil: A combined analysis of GC/MS and E-nose with chemometrics methods. Ind. Crops Prod. 206, 117595 (2023).
Kachko, A., van der Ham, L. V., Bardow, A., Vlugt, T. J. H. & Goetheer, E. L. V. Comparison of Raman, NIR, and ATR FTIR spectroscopy as analytical tools for in-line monitoring of CO2 concentration in an amine gas treating process. Int. J. Greenh. Gas Control. 47, 17–24 (2016).
Acknowledgements
The authors acknowledge the funding received from the project BIOVAL funded by the Marine Institute under the umbrella of the Sustainable Blue Economy Partnership (SBEP) (EU Horizon 2020 Research and Innovation Programme).
Author information
Authors and Affiliations
Contributions
W.M. performed investigation, validation, data analysis, visualization, and wrote the original draft; M.Z. developed methodology, performed data analysis, applied software, conducted visualization, and wrote the original draft; J.L. performed validation; C.O’D. provided supervision and reviewed & edited the manuscript; R.C.-M. performed validation; H.S. developed methodology and provided supervision; T.M. conceptualized the study and provided supervision; M.G.-V. developed methodology, conceptualized the study, provided funding and supervision. All participating authors reviewed and edited the manuscript prior to submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Meng, W., Zhao, M., Luo, J. et al. Validation and comparison of GC-MS, FT-MIR, and FT-NIR techniques for rapid bromoform quantification in Asparagopsis taxiformis extracts. Sci Rep 16, 1385 (2026). https://doi.org/10.1038/s41598-025-31263-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31263-z