Abstract
WD40 repeat proteins play crucial roles in various biological processes, including stress responses, development, and signaling. In this study, we identified and characterized 142 WD40 genes (CsWDR) in cucumber (Cucumis sativus), using the Hidden Markov Model to detect WD40 domains. These genes exhibited considerable variation in protein length and domain structure, with some containing additional functional domains such as UTPB, zinc finger, and F-box. Phylogenetic analysis revealed seven distinct clusters of CsWDR proteins, suggesting functional divergence, including roles in stress tolerance. Gene Ontology analysis highlighted their involvement in metabolic processes, stress responses, and protein binding, with most localized to the nucleus. Expression profiling revealed distinct patterns across tissues and developmental stages, with significant responses to temperature and light conditions. Notably, the CsWDR36 gene, associated with parthenocarpy in cucumber, was further investigated. Two transcripts, CsWDR36-1 and CsWDR36-2, were identified, differing in the number of WD40 motifs. Real-time PCR showed higher expression of both genes in flowers, especially male flowers, with increased expression following flowering, suggesting a role in early fruit development. These findings provide insights into the functional diversity of CsWDR genes and their potential roles in cucumber development and stress response.
Similar content being viewed by others
Introduction
WD-repeat (WDR) proteins form a highly diverse superfamily of regulatory proteins1. Structurally, the WD40 domain is characterized by multiple WD40 repeats, each consisting of 44–60 amino acid residues. These repeats typically contain a glycine-histidine (GH) dipeptide near the N-terminus and a tryptophan-aspartate (WD) dipeptide at the C-terminus2. The repeats form four-stranded antiparallel β-sheets that fold into a characteristic β-propeller structure3. The diversity of WD40 structures suggests they have evolved through intragenic duplication and recombination over evolutionary time4. It is believed that repeats within the same protein are more similar to one another than to repeats from different proteins, indicating distinct evolutionary timescales. WD40 proteins with lower self-similarity are thought to have originated earlier in evolutionary history5.
WD40 domain-containing proteins are widespread in eukaryotes but rare in prokaryotes, with few examples found in bacteria6. This domain ranks among the ten most abundant domains in eukaryotic genomes5. WD40 proteins serve as scaffolding molecules, facilitating protein–protein interactions and influencing various cellular processes. In plants, WD40 proteins play essential roles in growth, development, and cell cycle regulation7,8,9,10,11. They are also crucial in anthocyanin biosynthesis12,13,14, defense responses, and adaptation to both biotic and abiotic stresses15,16,17.
Genome-wide characterization of the WDR gene family has been conducted in several plant species, including Arabidopsis1,19, rice20, wheat21, and tomato22. Cucumber, an economically important crop, also serves as a model system for studying sex determination23,24. It offers several advantages, such as a small genome (~ 367 Mb, 2n = 2x = 14), a short life cycle, and cross-pollination, with a relatively narrow genetic background. The availability of its genome sequence has greatly facilitated structural and functional genomics research, with potential applications in crop improvement and food security.
Parthenocarpy is one of the economically important traits in cucumber, It has the potential to increase yield, especially under unfavorable conditions. Parthenocarpic fruits are often firmer and fleshier than their seeded ones25. Although parthenocarpic cucumber has been widely used in commercial production for a long time, its genetic basis is not well understood. It is a complex trait which can be influenced by environmental, physiological, and genetic factors. There were many studies on parthenocarpy in cucumber, which mainly in the positioning of genes related to parthenocarpic traits as well as the functional analysis of genes associated with related hormone biosynthesis or signaling pathways, whereas reports on genes that directly regulate parthenocarpic traits was less, A comprehensive analysis of all the studies revealed that WD40 has been mentioned several times in relation to parthenocarpic. It could be play a crucial role of hormonal regulation and transcription factors6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27, which was one of the parthenocapic candidate genes related to phytohormone28. While WD40 repeat proteins may influence parthenocarpy indirectly through the regulation of key hormones such as gibberellins29, they are more likely involved as components of broader regulatory networks that govern complex developmental processes30. The genome-wide characterization of the WD40 gene family in cucumber offer valuable insights into the molecular mechanisms underlying these pathways and may further clarify the potential role of WD40 proteins in regulating parthenocarpy in this species.
Materials and methods
Identification and characterization of WD40 domain proteins in cucumber genome using hidden markov model and database searches
The Hidden Markov Model (HMM) profile of the WD40 domain (PF00400), retrieved from Pfam v27.0 (http://Pfam.sanger.ac.uk/), was used to search the cucumber genome in the PHYTOZOME v8.0 (www.phytozome.net/) and Cucurbit Genomics databases. An E-value threshold of less than 10 was applied for the search. Hits with E-values below 1.0 were selected, and redundant sequences were removed using Clustal v2.17. Each unique sequence was manually verified for the presence of conserved WD40 domains using SMART (http://smart.embl-heidelberg.de/) 31 and Pfam32 tools.
Physical mapping and chromosomal localization of CsWDR genes in cucumber
Physical mapping of the CsWDR genes in cucumber was performed by conducting BLASTP searches against the Cucurbit Genomics Database. The identified genes were then mapped onto the seven cucumber chromosomes, arranged by their physical position from the short-arm telomere to the long-arm telomere. The final map was generated using JoinMap 4.033.
Homology modeling and 3D structural prediction of CsWDR proteins in cucumber
To model the three-dimensional (3D) structure of CsWDR proteins, we performed a BLASTP search of all CsWDR proteins against the Protein Data Bank34 to identify suitable templates with known 3D structures. Homology modeling of the proteins was conducted using Phyre2 (http://www.sbg.bio.ic.ac.uk/phyre2) in ‘intensive’ mode.
Phylogenetic analysis and functional annotation of CsWDR proteins in cucumber
The amino acid sequences of CsWDR proteins were aligned using MEGA535 with Clustal W36, using a gap open penalty of 10 and a gap extension penalty of 0.1. An unrooted phylogenetic tree was constructed using the neighbor-joining method37, with bootstrap analysis (1000 replicates). Functional annotation of the CsWDR proteins was performed using the WEGO tool38 (http://wego.genomics.org.cn/cgi-bin/wego/index.pl), which maps BLASTP results to retrieve the corresponding GO terms.
Gene expression analysis based on RNA-Seq data
In this study, RNA-based data analyses were conducted to explore the family-level expression patterns of the CsWDR genes in cucumber, focusing on both developmental stages (root, stem, leaf, tendril, male flower, female flower, and young fruit) and biotic stress responses. Transcriptome data from the genome-wide gene expression atlas of cucumber were downloaded from the Cucurbit Genomics Database (http://cucurbitgenomics.org/v2/) to analyze the spatial and temporal expression profiles of the CsWDR genes during development. To prepare the data for clustering analysis, absolute FPKM values were normalized by dividing them by the mean of all values, followed by a log2 transformation of the ratios. Genes with FPKM (Fragments Per Kilobase per Million mapped reads) values: < 5 were considered low expression, 5–20 as moderate expression, and > 20 as high expression. The expression data were then hierarchically clustered based on Euclidean distance using average linkage in MeV 4.5.
Plant material: parthenocarpic and non-parthenocarpic cucumber lines
Two cucumber lines, EC1 (parthenocarpic) and 8419 s-1 (non-parthenocarpic), were grown in plastic greenhouses under a 12-h photoperiod with mean daily temperatures of 29 °C (day) and 17 °C (night) at the Jiangpu Experiment Farm, Nanjing Agricultural University, during the fall of 2015. Plants were spaced 30 cm apart in rows, with 80 cm between rows. Female flowers were isolated one day prior to anthesis to prevent pollination. Flowers from EC1 were used for sampling, while 8419 s-1 flowers were pollinated the following day. Five tissues (roots, stems, leaves, male and female flowers) were collected from 10-week-old plants, along with ovary samples from EC1 (parthenocarpic) and pollinated 8419 s-1 at stages − 3 days post-anthesis (dpa), − 2 dpa, − 1 dpa, 0 dpa, 1 dpa, 2 dpa, and 3 dpa for qRT-PCR analysis. In addition, 8419 s-1 seedlings were treated with various hormones. Three-week-old seedlings were treated by spraying their leaves with NAA (5, 10, 50 μM), 6-BA (10 μM), GA3 (10 μM), ABA (10 μM), or ethephon (10 μM). Samples were collected at 0, 3, 6, and 9 h post-treatment, frozen in liquid nitrogen, and stored at -80 °C.
RNA extraction, cDNA synthesis, and primers of CsWDR36 in cucumber
Total RNA was extracted using Trizol reagent (Invitrogen) following the manufacturer’s instructions. RNA samples were treated with RNase-free DNase I to eliminate DNA contamination. First-strand cDNA synthesis was performed using 2 μg of total RNA with a cDNA Synthesis Kit (Fermentas). The CsWDR36 gene was amplified using gene-specific primers (Table S1). PCR conditions were as follows: 98 °C for 1 min, followed by 35 cycles of 98 °C for 15 s, 56 °C for 15 s, and 68 °C for 2 min 30 s, with a final extension at 68 °C for 5 min.
qRT-PCR analysis of CsWDR36 in cucumber
Ovary samples from EC1 and 8419 s-1 were collected at various developmental stages for qRT-PCR analysis. Each sample consisted of 20 ovaries ground in liquid nitrogen. Total RNA was extracted using Trizol (Invitrogen), treated with RNase-free DNase I, and converted to cDNA using 2 μg of total RNA (Fermentas). Quantitative real-time PCR was conducted using the SYBR Premix Ex Taq™ Kit (TAKARA) on a Bio-Rad CFX96 system. The PCR program included denaturation at 95 °C for 30 s, followed by 40 cycles of 95 °C for 5 s and 60 °C for 30 s. The actin gene (GenBank ID: AB010922) served as the internal control. The relative expression levels of each gene were normalized to actin and calculated using the 2−△△Ct method. All reactions were performed in triplicate. The primers used for qRT-PCR are listed in Table S1.
Results
Genome-wide identification and characterization of WD40 repeat (WDR) genes in cucumber
A systematic analysis of the cucumber genome was conducted to identify WD40 repeat (WDR) genes. The Hidden Markov Model (HMM) profile of the WD40 domain (PF00400) was used as a query to search the cucumber genome via BLASTP, with an E-value threshold of ≤ 10. This search resulted in the identification of 161 putative WD40 genes, with redundant transcripts of the same gene removed. Further validation using SMART and Pfam tools confirmed that 142 genes were part of the CsWDR family, each containing one or more WD40 domains. Of these, 131 CsWDR genes had been previously identified by Li (2014), while 10 were newly discovered in this study. The CsWDR genes exhibited considerable variation in protein length, ranging from 190 amino acids (Csa3M611390.1) to 2,973 amino acids (Csa3M124950.1), and the number of WD40 domains ranged from 1 to 9 (e.g., Csa3M687770.1). Detailed information on these 142 genes, including chromosomal locations, amino acid lengths, and the number of WD40 domains, is provided in Table S2.
Chromosomal distribution of CsWRD genes
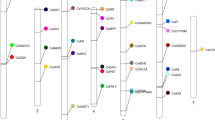
The chromosomal distribution of the 142 CsWDR genes was mapped using BLASTN, revealing an uneven distribution across cucumber chromosomes 1 to 7 (Fig. 1). Chromosome 3 had the highest number of CsWDR genes (32 genes, 22.5%), followed by chromosomes 1 (28 genes, 19.8%) and 4, 6 (19 genes, 13.4% each). Chromosomes 2 and 5 each contained 16 genes (11.3%), while chromosome 7 had the fewest (12 genes, 8.4%). Specific regions on certain chromosomes exhibited higher concentrations of CsWDR genes, particularly in the upper and lower arms of chromosome 4.
Chromosomal distribution of the WD40 repeat protein family in the cucumber genome. The diagram illustrates the chromosomal localization of genes encoding WD40 repeat proteins across the seven cucumber chromosomes (Chr1–Chr7). Each chromosome is represented as a vertical bar, with the positions of WD40 repeat protein genes marked along its length. The distribution provides insight into the genomic organization and potential clustering of WD40 genes within the cucumber genome, which may reflect evolutionary patterns or functional specializations.
Classification and domain diversity of CsWDR proteins in cucumber: insights into functional specialization
The amino acid sequences of the 142 WD40 domains were aligned using Clustal X 2.1. Based on domain structure and sequence alignment, the CsWDR proteins were categorized into 12 subfamilies (Fig. 2, Table S3). Approximately 99 CsWDR proteins, containing only the WD40 domain, were grouped into subfamily A. Other proteins, which contained additional functional domains, were classified into different subfamilies: Subfamily B: 5 members with a UTPB domain. Subfamily C: 4 members with a zinc finger domain. Subfamily D: 4 members with a Coatomer WD-associated region (COPI). Subfamily E: 2 members with a Katanin_con80 domain. Subfamily F: 3 members with a LisH domain. Subfamily G: 2 members with a protein S_TKc domain. Subfamily H: 2 members with a CAF1C_H4-bd domain. Subfamily I: 2 members with an NLE (NUC) domain. Subfamily J: 2 members with F-BOX and U-BOX domains. Subfamily K: 1 member with a DUF3337 domain. Subfamily L: 15 members with various domains (e.g., SFM, ATG16, Sof1, Cyclophilin). This diversity in domain composition suggests functional divergence and potential specialization of the proteins.
Structural representation of representative CsWDR proteins in cucumber (Cucumis sativus L.) from each subfamily: Annotations of conserved domains, motifs, and gene structures for functional and evolutionary classification.
Phylogenetic analysis and functional inference of CsWDR proteins in cucumber
A phylogenetic tree of the 142 CsWDR proteins was constructed using the neighbor-joining (NJ) method, with 12 additional WDR genes from other plants included for comparative analysis (Fig. 3, Table S4). The tree revealed seven distinct clusters (I to VII), with Cluster VII further subdivided into VIIa and VIIb. Bootstrap analysis indicated high reliability for most groupings. Based on gene function in other species, potential roles for each cluster were inferred:
Phylogenetic tree of the CsWDR proteins from cucumber and Arabidopsis: All of the protein sequences were aligned using ClustalX1.83, and the phylogenetic tree was constructed in using a Neighbor-Joining (NJ) method with MEGA5.0 sofware. The boot-strap value was 1,000 replications.
Cluster I contains genes involved in cell division, such as AtYAO, with potential roles in pollen and embryo sac development. Cluster II includes genes like AtWDR55.1, which are likely involved in microtubule organization and polar nuclei fusion. Cluster III contains genes associated with abiotic stress tolerance, such as TaWD40D. Cluster V includes genes like At1g65030, which may influence seed weight and plant size. Cluster VIIa includes StAN11 and NtTTG2, suggesting involvement in anthocyanin biosynthesis and auxin signaling. Cluster VIIb contains genes such as AtARCA and AtRACK1, indicating a role in drought stress and ABA signaling.
Gene ontology enrichment analysis of CsWDR genes related to cellular and developmental processes
Gene Ontology (GO) slim analysis using WEGO categorized the 142 CsWDR proteins into 14 biological process categories. Most CsWDR genes (85.21%) were associated with primary metabolic processes, followed by cellular metabolic processes (70.42%). Around 47.18% of CsWDR proteins were involved in response to stimulus, with notable inclusion of CsWDR36 (Csa2M070230.1). A significant proportion (41.55%) was also implicated in developmental processes, including reproductive process and growth. Additionally, 33 CsWDR proteins were associated with biological regulation, with CsWDR36 also included in this category.
Molecular function analysis revealed that 80.98% of CsWDR proteins were predicted to participate in protein binding, indicating their role in protein–protein interactions. Cellular localization predictions showed that most CsWDR proteins (86.62%) were localized to the cell part and cell, with a significant portion (50.70%) predicted to localize to organelle part (Fig. 4).
Comprehensive gene ontology (GO) distribution analysis of WDR proteins: An insight into (A) biological processes, (B) molecular functions, and (C) cellular components.
Expression profiling and functional grouping of CsWDR genes in development and stress response
Based on tissue expression patterns, genes with FPKM (Fragments Per Kilobase per Million mapped reads) values: < 5 were considered low expression, 5–20 as moderate expression, and > 20 as high expression. We found that 142 genes were classified into four groups (Fig. 5). The first group, comprising 57 genes, shows the lowest expression in male flowers, moderate expression in female flowers, and the highest expression in the ovary. The second group, consisting of 15 genes, displays low expression in both roots and male flowers. The third group, containing 53 genes, shows low expression in tendrils. The fourth group is characterized by high expression in the roots, while the fifth group, containing 16 genes, shows high expression in tendrils and low expression in both the roots and ovary.
Expression profiles of CsWDR genes across various tissues and under abiotic stress conditions as determined by RNA-Seq analysis. (A) fruits at different developmental stages, (B) including roots, stems, leaves, tendrils, female and male organs, and ovaries, (C) under high and low temperature conditions.
Analysis of gene expression in response to temperature and light revealed significant differences between low and high temperatures. The genes were grouped into four categories. The first group, with 45 genes, shows high expression under both high-temperature short-day and high-temperature long-day conditions. The second group, containing 19 genes, is highly expressed under high-temperature short-day conditions but shows low expression under high-temperature long-day conditions. The third group, also with 19 genes, is highly expressed under high-temperature long-day conditions. The fourth group, consisting of 58 genes, shows high expression under low-temperature short-day conditions and low expression under high-temperature long-day conditions.
During fruit development, the 142 genes were again classified into four groups. The first group, containing 54 genes, exhibits high expression during the first week of fruit development, suggesting a potential role in rapid fruit expansion and possibly in cell division. The second group, with 32 genes, shows low expression in the ovary but high expression during the second or third week of fruit development, indicating these genes may be involved in fruit expansion. The third group, comprising 20 genes, shows high expression in the ovary and low expression in the first week of fruit development, likely contributing to fruit set. The fourth group, with 35 genes, shows high expression during the fourth week of fruit development and low expression in the first week, suggesting a role in fruit ripening and coloration.
Characterization and expression analysis of CsWDR36: a candidate gene for parthenocarpy in cucumber
Previously, Csa2M070230 (CsWDR36) is located within a previously reported QTL interval associated with parthenocarpy28, suggesting it may be a positional candidate gene worthy of further investigation. BLAST analysis revealed that Csa2M070230 (CsWDR36) shares a high sequence homology (71.32%) with At4g04940.1 in Arabidopsis thaliana, a member of the transducing family that plays an important role during plant growth and developmental processes. In cucumber, expression profiling further showed that CsWDR36 exhibits low expression levels in tendrils, male flowers, and female flowers, moderate expression in vegetative organs such as roots, stems, and leaves, and the highest expression level in the ovary (Fig. 5), suggesting that this gene may be involved in the regulation of parthenocarpic fruit development in cucumber.
In this study, the gene’s cDNA was cloned, yielding two distinct bands of approximately 2676 bp and 1038 bp, named CsWDR36-1 and CsWDR36-2, respectively (Supplemental Figure S1). Molecular cloning and sequencing revealed that CsWDR36-1 and CsWDR36-2 encode proteins of 891 and 346 amino acids, respectively, with molecular masses of 9.99 kDa and 3.92 kDa. CsWDR36-1 contained nine WD40 motifs, classifying it within the WD40 family.
CsWDR36 was further characterized based on its protein structure. Protein domain analysis showed that CsWDR36-1 contained nine WD40 motifs and one Utp21 domain. Specifically, the nine WD40 motifs were located at positions 82–119, 170–209, 212–262, 256–296, 303–342, 459–502, 505–543, 546–586, and 588–627 in the CsWDR36-1 protein (Fig. 2). In contrast, CsWDR36-2 only contained the Utp21 domain. Previous studies identified two WD40 motifs within the Utp21 domain, suggesting that the difference between CsWDR36-1 and CsWDR36-2 lies in the number of WD40 motifs (Fig. 6). Sequence alignment of the two transcripts confirmed these findings. The existence of multiple transcripts of CsWDR36 suggests that alternative splicing may contribute to its functional diversification.
Schematic representation of the conserved domains in the CsWDR36 proteins. domain architecture of two isoforms of the CsWDR36 protein, CsWDR36-1 and CsWDR36-2. In CsWDR36-1, the protein is depicted with a series of WD40 domains, indicated by the blue hexagonal shapes, followed by a Utp21 domain at the C-terminus, shown in orange. CsWDR36-2 is represented with a single Utp21 domain, colored in a gradient from yellow to red, suggesting a potential functional difference or a simplified structure compared to CsWDR36-1. The absence of WD40 repeats in CsWDR36-2 occurred.
CsWDR36 was initially associated with cucumber parthenocarpy, prompting investigation of its transcription levels using real-time RT-PCR. As shown in Fig. 7, the two genes exhibited distinct expression patterns across cucumber organs, with higher expression levels observed in flowers, particularly in male flowers. However, no significant differences were observed under hormonal treatments (Fig. 8), suggesting that CsWDR36 is not responsive to such stimuli. During early growth and development, the two genes displayed different expression patterns, although their overall trends were similar. Expression of CsWDR36-1 and CsWDR36-2 increased after flowering in EC1, except at 3 days post-anthesis (dpa). In the 8419 variety, expression showed a sharp increase at − 2 dpa, followed by a decrease (Fig. 9). This phenomenon remains unexplained. The observed increase in gene expression after flowering, followed by a return to normal levels, suggests that CsWDR36 may play a role in the early stages of fruit growth and development.
Tissue-specific expression analysis of CsWDR36-1 and CsWDR36-2 in cucumber. (A) Relative expression levels of the CsWDR36-1 gene in various cucumber tissues, including roots, stems, leaves, and reproductive organs (female and male). (B)Relative expression levels of the CsWDR36-2 gene across the same set of cucumber tissues.
Hormone-Induced Expression Analysis of CsWDR36-1 and CsWDR36-2 in Cucumber Tissues. (A) Relative expression levels of the CsWDR36-1 gene in response to various hormone treatments. The treatments include a control (CK), gibberellin (G1), and different concentrations of abscisic acid (ABA), specifically ABA-1, NAA-1, NAA-2, and NAA-3. (B) Relative expression levels of the CsWDR36-2 gene under the same hormone treatments.
Expression Profiles of CsWDR36-1 and CsWDR36-2 across Early Developmental Stages of Cucumber Fruits. (A) Relative expression levels of the CsWDR36-1 gene during the early stages of cucumber fruit development. The expression is measured in two cucumber lines, 84,195–1 and EC1, at various days before and after anthesis (flower opening). The time points are marked as -3, -2, -1, 0, 1, 2, and 3 days after anthesis. (B) Relative expression levels of the CsWDR36-2 gene under the same conditions.
Discussion
Identification and characterization of the cucumber WDR gene family: a revised analysis based on advanced sequencing and precise gene criteria
WD-repeat (WDR) proteins represent a highly diverse superfamily of regulatory proteins, involved in a wide array of biochemical mechanisms and cellular processes. The WDR family has been well-documented in various species, including Arabidopsis thaliana, rice, wheat, and Solanum lycopersicum. In past studies, 191 WDR genes were identified in cucumber through a genome-wide analysis39. However, their study did not delve deeply into the molecular functions, biological pathways, or gene structures of these genes. In contrast, our study identified 142 WDR genes in cucumber, of which 10 were previously unrecognized. Additionally, 59 genes from earlier studies were excluded from our analysis due to the absence of the essential WD40 domain. This exclusion highlights a key difference between our approach and that of previous studies. The discrepancy in the number of identified WDR genes can be attributed to several factors:
Different Parameters for Gene Identification: Variations in the criteria and parameters used to identify genes can result in different outcomes. Unfortunately, the previous study did not provide sufficient details on the parameters they employed, which may have led to the inclusion of genes lacking the canonical WD40 domain. Advances in Sequencing Technologies: The availability of improved cucumber genome data and more accurate annotations, resulting from advances in sequencing technologies, has enabled a more precise determination of gene structures and functions. This study did a more rigorous identification of WDR genes. Therefore, the reduced number of WDR genes identified in our study is a reflection of the more refined methods and enhanced genomic resources used, which offer a more accurate representation of the cucumber WDR family.
Phylogenetic analysis and functional insights of CsWDR36 in cucumber: evolutionary relationships and potential role in stress responses
In this study, we performed a detailed phylogenetic analysis of the 142 cucumber CsWDR proteins to better understand their evolutionary relationships and functional associations. To achieve this, we constructed a phylogenetic tree that incorporated 12 WD40 proteins from other species with known functions, providing a broader context for the analysis. The resulting phylogenetic tree grouped the 154 WDR proteins (142 CsWDR proteins and 12 known functional WD40 proteins) into seven distinct clades.
Previous studies have classified WDR proteins into various subfamilies, albeit using different methods and species. For instance, WDR proteins in cucumber and Arabidopsis were previously classified into 21 subgroups based on domain architecture and phylogenetic relationships39, while studies in foxtail millet40 and Solanum lycopersicum22 identified 5 and 3 subfamilies, respectively. In our study, we observed that several genes with similar functions clustered together, with high confidence scores, such as AtRACK1 and AtARCA, both of which are involved in drought stress responses. This phylogenetic grouping underscores the utility of cross-species ortholog identification, which can help predict gene function based on conserved syntenic relationships.
Among the 142 CsWDR proteins, CsWDR36 was placed within clade III, where it clustered closely with TaWD40D41 and BnSWD1, both of which exhibit a high degree of homology with CsWDR36. Notably, BnSWD1 is a novel WD40 protein in Brassica napus that participates in salt-stress responses, potentially through both abscisic acid (ABA)-dependent and ABA-independent signaling pathways42. Given the homology and functional similarities, it is plausible that CsWDR36 may also play a role in salt-stress responses, potentially through an ABA-independent pathway.
Cloning and characterization of two alternative transcripts of CsWDR36 in cucumber: WD40 domain variation and alternative splicing mechanisms
In this study, parthenocarpic cucumber lines (EC1) and non-parthenocarpic lines (8419 s-1) were used to clone the coding sequences (CDS) of CsWDR36. Two different transcripts of CsWDR36 were identified, named CsWDR36-1 and CsWDR36-2 due to the presence of the WD40 domain structure. The key difference between these two transcripts lies in the number of WD40 domains. CsWDR36-2 consists solely of the UTP21 domain, which is part of the UTPB family involved in rRNA precursor modification, processing, and ribosome assembly43. The UTP21 structure is characterized by a C-terminal domain (CTD) containing two WD40 domains (WD1 and WD2). This suggests that the variability between the two transcripts arises from differences in the number of WD40 motifs, potentially due to alternative splicing during translation.
Similar alternative splicing events have been observed in Cucumis melo, Vigna radiata, and Gossypium raimondii, indicating that these CsWDR36 transcripts represent shear isomers. This highlights the flexible and complex regulatory mechanisms governing this gene, reflecting its functional importance. The presence of multiple transcripts also suggests that the gene’s structure is relatively unstable. The most stable and ideal β-sheet configuration is the seven-bladed β-propeller44, and WD40 β-propellers can theoretically contain 4 to 8 WD40 repeatsy45. In our study, CsWDR36-1 contained nine WD40 domains, deviating from the ideal sevenfold β-propeller architecture, which may explain the existence of the two distinct transcripts.
Expression patterns and functional insights of CsWDR36 in cucumber: potential roles in fruit development
Quantitative and semi-quantitative expression analyses revealed that CsWDR36-1 and CsWDR36-2 are predominantly expressed in reproductive organs, especially during early fruit development, with significantly lower expression levels in other organs. Both transcripts show minimal expression before flowering, peak at 1 day post-anthesis (dpa), and subsequently decline. This expression pattern suggests that CsWDR36 plays a role in fruit development, potentially through mechanisms independent of hormonal regulation. The lack of a significant response to hormone treatments further supports the hypothesis that CsWDR36 operates via non-hormonal pathways.
Phylogenetic analysis indicates that CsWDR36 may be involved in salt-stress responses through abscisic acid (ABA)-independent signaling pathways. While the current study confirms that CsWDR36 is not responsive to ABA, further investigations are needed to fully elucidate its role in salt-stress tolerance. Interestingly, an unexpectedly high expression level of both transcripts was observed at -1 dpa, a phenomenon that remains unexplained.
Conclusions
The WD-repeat (WDR) gene family plays a crucial role in regulating various aspects of plant development and responses to both biotic and abiotic stresses. This study enhances our understanding of the CsWDR family by providing detailed insights into the structure and potential functions of CsWDR proteins. Comparative phylogenetic analyses with WDR members from model plants offer valuable perspectives on the functional characterization of the WDR gene family in cucumber and related species. Cloning and expression analyses of CsWDR36 further support its potential involvement in fruit development and stress responses. These findings lay a solid foundation for future research aimed at elucidating the precise roles of CsWDR36 and other WDR family members. Additionally, this work highlights a subset of candidate genes that could be targeted in transgenic approaches to improve agronomic traits and enhance stress tolerance in cucumber and other closely related species.
Data availability
All data generated or analysed during this study are included in this published article.
References
van Nocker, S. & Ludwig, P. The WD-repeat protein superfamily in Arabidopsis: Conservation and divergence in structure and function. BMC Genomics 4, 50. https://doi.org/10.1186/1471-2164-4-50 (2003).
Neer, E. J., Schmidt, C. J., Nambudripad, R. & Smith, T. F. The ancient regulatory-protein family of WD-repeat proteins. Nature 371, 297–300. https://doi.org/10.1038/371812b0 (1994).
Smith, T. F., Gaitatzes, C., Saxena, K. & Neer, E. J. The WD repeat: A common architecture for diverse functions. Trends Biochem. Sci. 24, 181–185. https://doi.org/10.1016/S0968-0004(99)01384-5 (1999).
Andrade, M. A., Perez-Iratxeta, C. & Ponting, C. P. Protein repeats: Structures, functions, and evolution. J. Struct. Biol. 134, 117–131. https://doi.org/10.1006/jsbi.2001.4392 (2001).
Xu, C. & Min, J. Structure and function of WD40 domain proteins. Protein Cell 2, 202–214. https://doi.org/10.1007/s13238-011-1018-1 (2011).
Stirnimann, C. U., Petsalaki, E., Russell, R. B. & Müller, C. W. WD40 proteins propel cellular networks. Trends Biochem. Sci. 35, 565–574. https://doi.org/10.1016/j.tibs.2010.04.003 (2010).
Kim, Y. J. et al. GORI, encoding the WD40 domain protein, is required for pollen tube germination and elongation in rice. Plant J. 105, 1645–1664. https://doi.org/10.1111/TPJ.15139 (2021).
Kumar, S., Jordan, M. C., Datla, R. & Cloutier, S. The LuWD40–1 gene encoding WD repeat protein regulates growth and pollen viability in flax (Linum Usitatissimum L). PLoS ONE 8, e69124. https://doi.org/10.1371/journal.pone.0069124 (2013).
Xu, X. et al. Nucleocytoplasmic trafficking of the arabidopsis WD40 repeat protein XIW1 regulates ABI5 stability and abscisic acid responses. Mol. Plant 12, 1598–1611. https://doi.org/10.1016/j.molp.2019.07.001 (2019).
Zhu, F. et al. A comparative transcriptomics and eQTL approach identifies SlWD40 as a tomato fruit ripening regulator. Plant Physiol. 190, 250–266. https://doi.org/10.1093/plphys/kiac200 (2022).
Gachomo, E. W., Jimenez-Lopez, J. C., Baptiste, L. J. & Kotchoni, S. O. GIGANTUS1 (GTS1), a member of Transducin/WD40 protein superfamily, controls seed germination, growth and biomass accumulation through ribosome-biogenesis protein interactions in Arabidopsis thaliana. BMC Plant. Biol. 14, 37. https://doi.org/10.1186/1471-2229-14-37 (2014).
Tang, P. et al. Genome-wide identification of CaWD40 proteins reveals the involvement of a novel complex (CaAN1-CaDYT1-CaWD40-91) in anthocyanin biosynthesis and genic male sterility in Capsicum annuum. BMC Genomics 25, 851. https://doi.org/10.1186/s12864-024-10681-9 (2024).
Shoeva, O. Y., Mukhanova, M. A., Zakhrabekova, S. & Hansson, M. Ant13 encodes regulatory factor WD40 controlling anthocyanin and proanthocyanidin synthesis in barley (Hordeum vulgare L.). J. Agric. Food Chem. 71, 6967–6977. https://doi.org/10.1021/acs.jafc.2c09051 (2023).
Mol, P. P., Aparna, R. S., Sheeja, T. E. & Deepa, K. Novel bHLH and WD40 transcription factors from turmeric (Curcuma longa L.) as putative regulators of curcumin biosynthesis. J. Plant. Crops. https://doi.org/10.25081/jpc.2021.v49.i1.7057 (2021).
Eryong, C. & Bo, S. OsABT, a rice WD40 domain-containing protein, is involved in abiotic stress tolerance. Rice Sci. 29, 247–256. https://doi.org/10.1016/j.rsci.2021.07.012 (2022).
Tian, G. et al. Allelic variation of TaWD40–4B.1 contributes to drought tolerance by modulating catalase activity in wheat. Nat. Commun. 14, 1200. https://doi.org/10.1038/s41467-023-36901-6 (2023).
Rai, K. K., Singh, S., Rai, R. & Rai, L. C. Functional characterization of two WD40 family proteins, Alr0671 and All2352, from Anabaena PCC 7120 and deciphering their role in abiotic stress management. Plant Mol. Biol. 110, 545–563. https://doi.org/10.1007/s11103-022-01306-4 (2022).
Wu, Z. et al. Genome-wide analysis of WD40 protein family and functional characterization of BvWD40–82 in sugar beet. Front. Plant Sci. https://doi.org/10.3389/fpls.2023.1185440 (2023).
Yang, H. Y. & Zhang, X. Q. WD40 proteins in arabidopsis. Physiol. Mol. Biol. Plants. 44(5), 1025–1033 (2008).
Ouyang, Y., Huang, X., Lu, Z. & Yao, J. Genomic survey, expression profile and co-expression network analysis of OsWD40 family in rice. BMC Genomics 13, 100. https://doi.org/10.1186/1471-2164-13-100 (2012).
Hu, R. et al. Genome-wide identification and analysis of WD40 proteins in wheat (Triticum aestivum L.). BMC Genomics 19, 803. https://doi.org/10.1186/s12864-018-5157-0 (2018).
Zhu, Y. et al. Genome-wide identification, sequence characterization, and protein-protein interaction properties of DDB1 (damaged DNA binding protein-1)-binding WD40-repeat family members in Solanum lycopersicum. Planta 241, 1337–1350. https://doi.org/10.1007/s00425-015-2258-8 (2015).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739. https://doi.org/10.1093/molbev/msr121 (2011).
Li, Z., Pan, J. S., He, H. L., Si, L. T. & Cai, R. Research of sexual inheritance and regulation in cucumber(Cucumis sativus L.). J. Shanghai Jiaotong Univ. (Agric. Sci.) 26(3), 5. https://doi.org/10.3969/j.issn.1671-9964.2008.03.009 (2008).
Bose, S. et al. Characterization of gynoecious-parthenocarpic and monoecious cucumber lines (Cucumis sativus L.) and regression modelling to obtain high yielding and functionally rich genotypes. Hortic. Environ. Biote 65(3), 465–476. https://doi.org/10.1007/s13580-023-00579-y (2024).
Lietzow, C. D., Zhu, H., Pandey, S., Havey, M. J. & Weng, Y. QTL mapping of parthenocarpic fruit set in North American processing cucumber. Theor. Appl. Genet. 129, 2387–2401. https://doi.org/10.1007/s00122-016-2778-z (2016).
Kaur, H. et al. Genome-wide identification and characterization of parthenocarpic fruit set-related gene homologs in cucumber (Cucumis sativus L.). Sci. Rep. 13, 2403. https://doi.org/10.1038/s41598-023-29660-3 (2023).
Wu, Z. et al. Identification of a stable major-effect QTL (Parth 2.1) controlling parthenocarpy in cucumber and associated candidate gene analysis via whole genome re-sequencing. BMC Plant. Biol. 16, 182. https://doi.org/10.1186/s12870-016-0873-6 (2016).
Jat, G. S., Munshi, A. D., Behera, T. K. & Bharadwaj, C. Inheritance of parthenocarpy in gynoecious cucumber (Cucumis sativus L) cultivar PPC-2. J. Hortic. Sci. 12, 193–197 (2017).
Li, Z., Niu, H. & Guo, Y. Cucumber Sex Determination: Aspects of Gene Interactions. In The Cucumber Genome (eds Pandey, S. et al.) 145–157 (Springer International Publishing, Cham, 2022). https://doi.org/10.1007/978-3-030-88647-9_11.
Ivica, L., Tobias, D. & Peer, B. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40(D1), 302–305. https://doi.org/10.1093/nar/gkr931 (2012).
Clans, P. Pfam: clans, web tools and services. Nucleic. Acids Res. 34(D1), 247–251. https://doi.org/10.1093/nar/gkj149 (2006).
Ooijen, J.W.V., Voorrips, R.E. Software for the calculation of genetic linkage maps. 2001.
John, W., Zukang, F., Li, C., Wang, Y. H. & Berman, H. M. The protein data bank and structural genomics. Nucleic Acids Res. 31(1), 489–491. https://doi.org/10.1093/nar/gkg068 (2003).
Tamura, K. et al. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionarydistance, and maximum parsimony methods. Mol. Biol. Evol. 28(10), 2731–2739. https://doi.org/10.1093/molbev/msr121 (2011).
Yuan, J. et al. MULTICLUSTAL: A systematic method for surveying Clustal W alignment parameters. Bioinformatics 15(15), 862–863. https://doi.org/10.1093/bioinformatics/15.10.862 (1999).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454 (1987).
Ye, J., Wego: a web tool for plotting GO annotations, Nucleic Acids Res. https://doi.org/10.1093/nar/gkl031.
Li, Q. et al. Genome-wide analysis of the WD-repeat protein family in cucumber and Arabidopsis. Mol. Genet. Genomics 289, 103–124. https://doi.org/10.1007/s00438-013-0789-x (2014).
Mishra, A. K., Muthamilarasan, M., Khan, Y., Parida, S. K. & Prasad, M. Genome-wide investigation and expression analyses of WD40 protein family in the model plant foxtail millet (Setaria italica L.). PLoS ONE 9, e86852. https://doi.org/10.1371/journal.pone.0086852 (2014).
Kong, D., Li, M., Dong, Z., Ji, H. & Li, X. Identification of TaWD40D, a wheat WD40 repeat-containing protein that is associated with plant tolerance to abiotic stresses. Plant Cell Rep. 34, 395–410. https://doi.org/10.1007/s00299-014-1717-1 (2015).
Lee, S. et al. A novel WD40 protein, BnSWD1 is involved in salt stress in Brassica napus. Plant Biotechnol. Rep. 4(2), 165–172. https://doi.org/10.1007/s11816-010-0131-6 (2010).
Hunziker, M. et al. UtpA and UtpB chaperone nascent pre-ribosomal RNA and U3 snoRNA to initiate eukaryotic ribosome assembly. Nat. Commun. 7, 12090. https://doi.org/10.1038/ncomms12090 (2016).
Valeyev, N. V., Downing, A. K., Sondek, J. & Deane, C. Electrostatic and functional analysis of the seven-bladed WD beta-propellers. Evol. Bioinform. Online 4, 203–216. https://doi.org/10.4137/EBO.S743 (2008).
Wang, Y., Jiang, F., Zhuo, Z., Wu, X.-H. & Wu, Y.-D. A method for WD40 repeat detection and secondary structure prediction. PLoS ONE 8, e65705. https://doi.org/10.1371/journal.pone.0065705 (2013).
Funding
This research was funded by the Agriculture Project from Zhejiang Provincial Department of Agriculture and Rural Affairs, Expert Workstation Project, Zhejiang Provincial Basic Public Welfare Research Program (Grant No. LTGN23C020002) and the National Natural Science Foundation of China (32260769, 32202521) and the Youth Innovation Fund Project of Inner Mongolia Academy of Agricultural and Husbandry Sciences (2023QNJJN03).
Author information
Authors and Affiliations
Contributions
T.Z. designed the research. T.Z. and X.Y.W. carried out research. Q.W.G. analysed the data. P.P.F. and J.W. wrote the first draft of manuscript. C.S.L and J.L. reviewed and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical declarations
The authors with this declare that the paper is based on our writing except for citations which have been duly acknowledged. We also state that it has not been previously or concurrently submitted for any other journals.
Informed consent
All authors have read and approved the manuscript for publication.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, T., Wang, X., Guo, Q. et al. Genome-wide characterization of WD40 repeat proteins in cucumber reveals their functional roles in stress response and parthenocarpy. Sci Rep 16, 1846 (2026). https://doi.org/10.1038/s41598-025-31502-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-31502-3