Abstract
The use of biomarkers in liver transplantation (LT) may be useful for the noninvasive assessment of T-cell-mediated rejection (TCMR). This study aimed to: (1) determine if changes in circulating microRNAs (miRNAs) in rejector patients reflect those at the tissue level; (2) assess a biomarker score (including miRNAs 155-5p, 181a-5p, and CXCL-10) in an independent cohort and (3) evaluate the score’s utility in detecting other LT-related complications. A prospective cohort of 151 LT recipients was followed during the first year after surgery. Patients with abnormal liver function tests (LFTs) underwent liver biopsy (LB) to rule out TCMR. Plasma samples were collected at predefined time points to analyse miRNAs and chemokines and their ability to predict and diagnose TCMR. To correlate plasma biomarkers with their tissue expression, 27 liver tissue samples (9 with normal LFTs and 18 with TCMR) were selected. Forty patients were diagnosed with TCMR. Circulating miRNAs (155-5p and 122-5p) reflected tissue-level changes in TCMR (p < 0.001, 3.3-fold increase for miR-155 and 2.3 for miR-122). Additionally assessed plasma miRNA 885-5p and chemokines CXCL-9 and CXCL-11, were also significantly elevated in TCMR but did not improve the performance of our previously biomarker score for TCMR (AUROC 0.94 for prediction, 0.99 for diagnosis). In conclusion, we confirmed in an independent cohort the performance of the score based on miR-155-5p, miR-181a-5p, and CXCL10. The correlation between circulating miRNAs and liver tissue expression underscores their value for predicting and diagnosing TCMR.
Similar content being viewed by others
Introduction
Liver transplantation (LT) is the treatment of choice for patients with end-stage liver disease or hepatocellular carcinoma (HCC) at certain stages1,2. Rejection is one of the most common causes of graft dysfunction (GD) during the first months after LT, with an incidence estimated between 15 and 34% in the latest reports3,4. The most frequent form of rejection is T-cell-mediated rejection (TCMR)5. For the diagnosis of TCMR, liver biopsy (LB) remains the gold standard; however, it is an invasive method with known biases6. The manifestations of TCMR in liver function tests (LFTs) are indistinguishable from those of other causes of GD, such as biliary strictures (BS). In this context, the use of biomarkers could be helpful to noninvasively predict and diagnose TCMR. This could be especially impactful during the first weeks and months after LT, when immunosuppression (IS) is often tapered and the incidence of TCMR is highest. Moreover, these biomarkers could help overcome the lack of specificity of abnormal LFTs, allowing for differentiation between TCMR and other causes of GD7.
In transplantation, miRNAs are essential modulators of immunological responses and graft function. Local inflammation associated with allograft rejection is orchestrated by distinct T helper (Th) subsets and regulated by cytokine signaling and miRNA-mediated gene expression8,9. Among these, miRNA-155-5p has been shown to control CD4⁺ T-cell differentiation, promote regulatory T-cell development, and regulate IFN-γ production in T and NK cells, linking its expression to T-cell–mediated immune activation10. Similarly, miRNA-181a-5p modulates T-cell receptor sensitivity and signaling strength, thereby influencing the magnitude of the alloresponse and graft rejection11,12. In contrast, miRNA-122-5p, the most abundant liver-derived miRNA, is predominantly associated with hepatocyte injury and thus serves as an indicator of liver-specific damage rather than immune activation13. Our group identified a plasma miRNA signature (miRNA-155-5p, miRNA-122-5p, and miRNA-181a-5p) useful for prediction and diagnosis of rejection, distinctive from other causes of GD14,15. We confirmed these results and published a logistic regression model for rejection assessment combining the plasma levels of the chemokine CXCL-10 with miRNAs 155-5p and 181a-5p. This score had an area under the ROC curve (AUROC) value of 0.975 and 0.99 for rejection prediction and diagnosis, respectively16. Other scores include the one by Levitsky et al.17,18 incorporating 59 differentially expressed genes before rejection-associated graft injury (AUROC 0.83 for TCMR diagnosis) and the recently published by Schaapman et al.-19 combining magnetic resonance imaging and serum tests (AUROC 0.7).
miRNAs are released into circulation and may reflect ongoing cellular or tissue processes. However, the link between tissue expression and circulating levels is sometimes unclear. For noninvasive biomarkers, it’s important to confirm that changes seen in circulating miRNAs also occur in tissue. This is key to understanding their potential as markers for specific clinical events.
On the other hand, among the plethora of miRNAs, there are few studies in which the same miRNA panel was used, making it difficult to compare results across centres and reach a consensus on which signatures are best. With the goal of identifying a more universal signature, in this study, we propose the evaluation of three new miRNAs (miRNA-483-3p, miRNA-885-5p, and miRNA-194-5p) potentially related with rejection. miRNA-483-3p, miRNA-885-5p, and miRNA-194-5p, have been associated with metabolic stress, hepatocellular injury, and inflammatory conditions, and were therefore explored as potential complementary biomarkers. The expression of miRNA-885-5p is significantly elevated in patients with cirrhosis, HCC, or hepatitis B and C20,21. miRNA-483-3p is significantly upregulated in diabetes and cardiovascular diseases and has been shown to induce cellular apoptosis and lipotoxicity across various cell types22. Shaked et al.23 demonstrated that a signature combining miRNA-483-3p and miRNA-885-5p could be used to diagnose and predict rejection with high accuracy. On the other hand, miRNA-194-5p, like miRNA-122-5p, is specifically expressed in hepatocytes24 and is considered a serum biomarker for liver injury25,26.
In LT, studies assessing chemokines as biomarkers for the risk of rejection are more limited. Our group recently demonstrated that patients with TCMR had significantly higher plasma CXCL-10 levels (AUROC = 0.962)16. Like CXCL-10, CXCL-9 and CXCL-11 may contribute to rejection risk assessment because of their important roles in lymphocyte recruitment to sites of injury and inflammation27,28,29,30,31. Although they act through the same receptor, these chemokines exhibit distinct temporal and affinity profiles: CXCL-9 is typically upregulated during the early stages of endothelial activation, whereas CXCL-11 shows the highest affinity for CXCR3 and may reflect more sustained Th1-driven immune responses. Furthermore, studies by Raschzok et al.26 and Kim et al. 27 suggest the potential of monitoring these chemokines in plasma as predictive biomarkers of rejection in LT. The combined assessment of CXCL-9 and CXCL-11 (two additional ligands of the CXCR3 receptor) with CXCL-10 could better capture the complexity of IFN-γ–mediated immune responses, potentially improving the sensitivity and specificity of the score in diverse clinical contexts. The aims of this study were to 1) investigate whether the miRNA overexpression observed at the circulating level in patients with rejection is reflective of what occurs at the tissue level; 2) evaluate the biomarker score (including CXCL-10 and miRNAs 155-5p and 181a-5p) in a new independent cohort; 3) test the biomarker score in other complications, such as BS and cytomegalovirus (CMV) infections; 4) assess whether the plasma levels of miRNA-483-3p, miRNA-885-5p, and miRNA-194-5p, as well as the plasma production of CXCL-9 and CXCL-11, could provide greater robustness to our previous score.
Patients and Methods
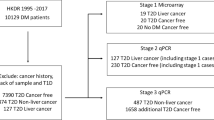
An observational, prospective cohort study was conducted at the Liver Transplant Unit of Hospital Clínic Barcelona. Biomarker analysis and immunosuppressive monitoring were performed at the Pharmacology and Toxicology Laboratory (CIBERehd; Project PI19/00,378 ISCIII, co-funded by the EU). Consecutive candidates were enrolled from September 2020 to January 2024, excluding combined liver-kidney transplants and those without consent. The patients were enrolled while on the LT waiting list and followed for the first year after surgery. Only patients who completed the follow-up period and attended at least 50% of the scheduled study visits were included in the analysis to ensure data completeness. This criterion does not indicate that 50% of visits were missed but rather served as a threshold for inclusion (Fig. 1S).
IS was tailored to pre-LT status. Patients with Child‒Pugh A cirrhosis or low renal risk received corticosteroids and tacrolimus (TAC) within 24 h post-LT (target trough: 8–10 ng/mL). Those with Child‒Pugh B/C cirrhosis or with acute liver failure received a single dose of basiliximab 20 mg induction, followed by corticosteroids, mycophenolate mofetil (MMF, 2000 mg/day), and TAC from day 5 (target: 5–8 ng/mL). Corticosteroids were tapered and discontinued by 6 months, and MMF was reduced to 1500 mg/day after 1 month after LT.
Follow-up and graft dysfunction (GD) episodes.
Participants were followed and underwent clinical and laboratory surveillance, including LFTs, pharmacokinetic monitoring, outpatient visits, and additional plasma collection for biomarker analysis. Assessments occurred at weeks 1 and 2 and at months 1, 2, 3, 6, 9, and 12. GD was diagnosed when patients had abnormal LFTs, defined as aspartate aminotransferase (AST), alanine aminotransferase (ALT), or bilirubin serum levels more than two times the upper limit of normal during routine follow-up laboratory surveillance, or if these parameters did not decrease within the first two weeks after LT. Additionally, abdominal ultrasound was performed to exclude vascular or biliary complications in GD cases. Patients with GD underwent liver biopsy (LB) to rule out rejection, evaluated by an expert pathologist using Banff criteria34. TCMR severity was graded by the Rejection Activity Index (RAI), and other biopsy-confirmed causes of GD were recorded35. Biliary strictures (BS), a frequent event that may lead to abnormal LFTs, were also documented. This event was recorded during the study visit prior to BS treatment. Finally, all patients were monitored for CMV viral load in plasma at every visit during the first 6 months. CMV infection was defined as CMV DNA > 1000 copies/mL.
Pharmacokinetic monitoring
The TAC trough concentrations (TAC-C0) were analysed during the study visits. Whole-blood TAC-C0 were measured via the Tacrolimus-CMIA-Architect assay (Abbott, Wiesbaden, Germany) according to the manufacturer’s instructions. The laboratory’s participation in the United Kingdom External Analytical Quality Assessment Service ensured compliance with LGC Standard Proficiency Testing.
Plasmatic chemokine measurements
Plasma levels of CXCL-9, CXCL-10, and CXCL-11 were analyzed using blood samples collected in EDTA tubes before the morning treatment dose during clinical visits and biopsies. The samples were centrifuged within 2 h at 3000 rpm for 10 min, and the plasma was stored at -70 °C for later analysis. Chemokine concentrations were measured by ELISA (Quantikine kits, R&D Systems) following the manufacturer’s instructions. The detection limits were 3.84 pg/mL for CXCL-9, 1.67 pg/mL for CXCL-10, and 13.9 pg/mL for CXCL-11.
Plasmatic miRNA analysis
Plasma expression of miR-155-5p, miR-122-5p, miR-181a-5p, miR-194-5p, miR-483-3p, and miR-885-5p was measured during clinical visits, pharmacokinetic assessments, and biopsies using qPCR (LightCycler 480, Roche). Blood was collected in EDTA-K3 tubes before IS administration or any treatment changes during rejection episodes. Plasma was separated by centrifugation 3000 rpm, 10 min, within 2h and stored at − 70 °C.
Plasma expression was analysed as previously described by our group8. Total RNA was purified from 200 uL of patient plasma according to the manufacturer’s instructions (miRCURY™ RNA Isolation Kits – Biofluids ref. #300,112 from QIAGEN (Netherlands)). Briefly, plasma components were lysed with the provided Lysis Solution, and proteins were precipitated with the provided Protein Precipitation Solution. Isopropanol was added to the collected supernatant, and the solution was loaded into the column. The solution was washed with Wash Solutions 1 and 2, the RNA was eluted with RNase-free water, and the concentration and quantity of the total RNA were measured at 260 nm and 280 nm (A260/A280) using a NanoDrop device (NanoDrop Technologies). Hemolysis contamination was tested using a spectrophotometer and measuring the oxyhemoglobin absorbance at 414 nm. An OD scan was performed from approximately 200 to 700 nm, and distinguishing absorbance peaks at 414 nm were used to disqualify hemolysis samples. Total RNA was reverse transcribed into cDNA following the manufacturer’s instructions (miRCURY LNA™ Universal RT ref #203,301 from QIAGEN). cDNA served as a template for miRNA RT‐qPCR amplification with locked nucleic acid (LNA) primers and SYBR Green master mix. The following specific LNA PCR primer sets, all from QIAGEN (Netherlands), were used (Table 1S): hsa-miRNA-155-5p LNA™ PCR primer set, UniRT (ref.# YP00204308); hsa-miRNA-181a- 5p LNA™ PCR primer set, UniRT (ref. # YP00206081); hsa-miRNA-122- 5p LNA™ PCR primer set, UniRT (ref, # YP00205664); hsa-miRNA-194-5p LNA™ PCR primer set, UniRT (ref.# YP00204080); hsa-miRNA-483-3p LNA™ PCR primer set, UniRT (ref.# YP00204012); hsa-miRNA-885-5p LNA™ PCR primer set, UniRT (ref.# YP00204473); hsa-miRNA-103a-3p, LNA™ PCR primer set, UniRT (ref. YP00204063); hsa-miRNA-191-5p, LNA™ PCR primer set, UniRT (ref. # YP00204306) and UniSp6 LNA™ PCR primer set, UniRT (ref. YP00203954). To monitor the cDNA synthesis reaction, the synthetic spike-in UniSP6 (ref. #339,340) was used for signs of inhibition (prior to the reverse transcription reaction, we added 1 μl of synthetic spike-in (108 copies/μl) per 20 ng sample RNA). PCRs were performed using a Light Cycler 480 instrument. The amplification profile was denatured at 95 °C for 10 min followed by 45 amplification cycles of 95 °C for 10 s and 60 °C for 1 min. At the end of the PCR cycles, melting curve analyses were performed. Negative controls with 1 µg of MS2 carrier RNA as a mock template from the reverse transcription reaction were produced and profiled similarly to the samples. The amplification curves were analyzed using Roche LC Software for determination of Cq by the second derivative method. Average Cq values were normalized to the stably expressed reference miR-103a-3p and miR-191-5p, following the manufacturer’s instructions. First, the Cq values for all samples were determined; the average Cq of miRNA-191-5p + miRNA-103-3p was calculated, and the ∆Cq was calculated as the difference in Cq values between the miRNA target and the reference control (Cq average of miR-191-5p + miR-103-5p). Relative expression levels of target miRNAs were then evaluated within a sample according to the formula 2^(-∆Cq), where high values corresponded to higher expression.
Liver graft miRNA analysis
LBs were obtained either percutaneously or by a transjugular approach. The samples were formalin-fixed and paraffin-embedded (FFPE). We analysed 27 tissue samples obtained from LT recipients (9 with normal liver tests and 18 diagnosed with TCMR). Three freshly cut sections of FFPE tissue, each with a thickness of 5 µm, were used to analyse miRNA expression.
Total RNA, including miRNA, was extracted from FFPE liver tissue using the miRNeasy FFPE Kit (Qiagen). After deparaffinization at 56ºC and proteinase K digestion, samples were incubated at 56 °C and 80 °C. Genomic DNA was removed with DNase, and RNA was purified with RNeasy MiniElute columns, eluted in RNase-free water, and reverse transcribed into cDNA using the miRCURY LNA SYBR Green PCR Kit.
qPCR was performed with the miRCURY LNA SYBR Green PCR Kit (Qiagen). Cq values were obtained using the second derivative method (Roche LC software). ∆Cq was calculated using miRNA-103a-3p and miRNA-191-5p as reference controls, and relative expression was determined by the 2^(-∆Cq) method, with higher values indicating greater expression.
Statistical analyses
Continuous variables are presented as medians with interquartile ranges; categorical variables as absolute frequencies and proportions. Group differences were assessed using the Mann–Whitney U test, and correlations using Spearman’s rho. Pairwise comparisons between groups (free clinical event vs. TCMR, free clinical event vs. CMV, and TCMR vs. CMV) were also performed with the Mann–Whitney U test. A Kruskal–Wallis test followed by Dunn–Bonferroni post hoc correction is used in visits with both events (TCMR and CMV, vs no events). Linear mixed-effects models (LMMs) with a random intercept for each patient were used to evaluate the association between TCMR and biomarker levels over time. A p-value < 0.05 was considered statistically significant.
For the diagnostic analyses, biomarker measurements corresponding to the time of the event were used, whereas for the predictive analyses, biomarker values from prior visits in patients who later developed rejection were also included. The predictive and diagnostic performance of the biomarkers was assessed using AUROC with 95% confidence intervals (CI). Cut-off values were determined by maximizing the Youden index, and sensitivity, specificity, PPV, and NPV were estimated for the biomarkers and LFTs showing significant differences in TCMR according to the LMM analysis. A stepwise logistic regression model was developed to predict TCMR, using as explanatory variables those clinical parameters with significant differences between patients with and without TCMR, and the biomarkers and LFTs with significant differences in the LMM analysis. Analyses were performed with SPSS v23.0 (SPSS Inc., Chicago, IL, USA) and Stata 16.0 (StataCorp LLC, College Station, TX, USA).
Results
Study patients and graft dysfunction episodes
In total, 151 patients were included. Clinical, demographic and laboratory data are summarized in Table 1. The majority of the patients were male (72.9%), with a median age of 59 years at the time of LT. The most common primary disease prior to the procedure was alcohol-related cirrhosis, followed by viral hepatitis. In 58 patients (35.8%), the indication for LT was HCC. Most recipients received organs from brain-dead donors (68.2%). The median cold ischaemia time was 395 min.
Fifty-nine patients (39.1%) met GD criteria and underwent LB. TCMR was identified in 40 patients (26.5%), with three experiencing two episodes, totalling 43 rejection episodes. Most TCMR cases (72%) occurred within the first two weeks post-transplant. In terms of severity, 21 episodes were moderate (RAI 5–6) and 22 were mild (RAI 3–4); no severe cases (RAI ≥ 7) were observed. Comparisons showed a higher rate of HCC as the transplant indication and a lower incidence of CMV infections among patients with TCMR.
In the remaining 16 GD episodes, LB mainly showed inflammation that did not meet TCMR criteria or lobular hepatitis of unknown cause. Other causes of non-rejection GD included hepatitis C and ischemia–reperfusion injury (IRI). Additionally, 26 patients (17.2%) developed BS requiring endoscopic or percutaneous intervention, and 29 patients (19.2%) met the criteria for CMV infection.
Tissue miRNA expression of miR-155-5p, miR-181a-5p and miR-122-5p
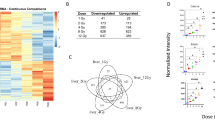
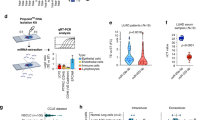
miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p were analysed in LB samples (Fig. 1). The results revealed a significant increase in the expression of miRNA-155-5p (1A) and miRNA-122-5p (1B) in the biopsies with rejection compared with those with normal histology for miRNA-155-5p (p < 0.001, a median value 3.3-fold increase) and miRNA-122-5p (p < 0.001, a median value 2.3-fold increase). For miRNA-181a-5p, despite the tissue expression being 1.3-fold higher in patients with rejection, the difference did not reach statistical significance (p = 0.304) (Fig. 1C).
Tissue expression of miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p. miRNA-155-5p (1A), miRNA-122-5p (1B) and miRNA-181a-5p (1C) were analysed in liver biopsy samples. Box plots showing the tissue expression of miRNAs between patients with rejection (n = 18) (grey boxes) and those without rejection (n = 9) (white boxes).
Plasmatic miRNA expression
In the cohort of the present study, the results confirmed the predictive and diagnostic ability of these miRNAs. A significant increase in the plasma levels of miRNA-155-5p (p < 0.000, a median value of 29.6-fold), miRNA-181a-5p (p < 0.001, a median value of 10.6-fold) and miRNA-122-5p (p < 0.001, a median value of 26.3-fold) was detected in patients with rejection (Fig. 2A–C) (Table 2S). When the LMM analysis was applied, adjusting for within-subject variability and accounting for time as a potential confounding factor, the differences remained statistically significant (miRNA-155-5p: p = 0.01 for prediction and p = 0.005 for diagnosis; miRNA-122-5p: p < 0.001 for prediction and p < 0.001 for diagnosis; miRNA-181a-5p: p = 0.002 for prediction and p < 0.001 for diagnosis).
Plasma miRNA expression. Boxplots showing plasma levels of miRNA-155-5p (2A), miRNA-122-5p (2B), miRNA-181a-5p (2C), miRNA-194-5p (2D), miRNA-483-3p (2E), and miRNA-885-5p (2F) at each scheduled follow-up timepoint for TCMR prediction. Data are presented for visits associated with TCMR (black boxes), CMV infection (grey boxes), and no clinical events (white boxes). For clarity, the number of visits contributing to each group at every timepoint is indicated as follow: For clarity, the number of visits contributing to each group at every timepoint is indicated as follow: Week 1: TCMR = 37; CMV = 1; No events = 112 ; Day 15: TCMR = 22; CMV = 2; No events = 126 ; Month 1: TCMR = 12; CMV = 9; No events = 124 ; Month 2: TCMR = 6; CMV = 10; No events = 84 ; Month 3: TCMR = 6; CMV = 3; No events = 136; Month 6: TCMR = 5; CMV = 4; No events = 134; Month 9: TCMR = 4; CMV = 0; No events = 82; Month 12: TCMR = 0; CMV = 1; No events = 130. A value of p < 0.05 was considered statistically significant: * indicates comparisons between rejection vs. free clinical events; ° indicates comparisons between CMV + vs. free clinical events.
Three patients (#26, #30, #63) had two TCMR episodes each. After IS adjustment following the first episode, their plasma miRNA levels normalized, resembling those of rejection-free patients. Prior to the second episode, miRNA expression rose again, reaching levels similar to the first rejection (Fig. 3).
Changes in plasma miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p expression in patients with 2 episodes of TCMR. Posttransplantation changes in the plasma expression of miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p in 3 patients (#26, #30; #63) who experienced two episodes of TCMR. The black arrow indicates the time of the acute rejection (AR) episode.
Furthermore, four patients experienced 2 clinical events during the 1st year of follow-up: one rejection and the other nonrejection (nonspecific inflammation or BS). In the non-rejection event, only miRNA-122-5p expression was increased, although not as high as in TCMR (Fig. 4). Therefore, for this specific miRNA, the presence of unspecific inflammation or BS could be a confounding factor for predicting rejection.
Changes in plasma miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p levels in patients with 2 clinical events during the 1st year of follow-up: one rejection and one nonrejection. Posttransplantation changes in the plasma expression of miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p in 4 patients (#33, #47; #61; #62) who experienced 2 clinical events during the 1st year of follow-up: one rejection and one non-rejection [nonspecific inflammation (UI) and biliary stricture (BS)]. The continuous black arrow indicates the time of the acute rejection (AR) episode, and the discontinuous black arrow indicates the non-rejection event.
Concerning the newly tested miRNAs (194-5p, 483-3p and 885-5p), statistically significant differences were also observed between rejector and nonrejector patients for miRNA-483-5p (0.387 vs. 0.010; p < 0.001, 38.7-fold increase); miRNA-885-5p (0.530 vs. 0.030; p < 0.001, 17.7-fold increase); and miRNA-194-5p (1.060 vs. 0.240; p < 0.001, 4.4-fold increase) (Fig. 2D–F). However, when the LMM analysis was performed, adjusting for individual variability and time, only miRNA-885-5p showed statistically significant differences (p = 0.003 for both prediction and diagnosis of TCMR).
Plasma CXCL-9, CXCL-10 and CXCL-11 production
A significant increase in plasma CXCL-10 production (p < 0.001) was observed in the rejector group (Fig. 5B) (Table 2S). These changes occurred at the time of LB (924 pg/mL vs. 165 pg/mL, p < 0.001); 5.6-fold increase) and before the LB. In agreement with our previous results, patients with CMV infection had significantly higher plasma CXCL-10 concentrations (p < 0.001, 3.5-fold increase).
Posttransplant monitoring of plasma CXCL-9, CXCL-10 and CXCL-11 production. Box plots showing differences in plasma CXCL-9 (A), CXCL-10 (B) and CXCL-11 (C) production among patients with rejection (n = 40) (black boxes), those with CMV + (n = 29) (striped boxes) and those without clinical events (n = 111) (white boxes). A value of p < 0.05 was considered to indicate statistical significance: *patients free of clinical events vs. patients with rejection; †patients free of clinical events vs. CMV + patients.
Among the patients who experienced two episodes of TCMR, the pattern of CXCL-10 plasma production was like that observed with miRNAs; once resolved the TCMR episode, the plasma CXCL-10 levels became like those of rejection-free patients. The production of CXCL-10 again increased prior to the second episode and reached levels similar to those attained during the first episode. Furthermore, patient #30 experienced reactivation of CMV infection after the 2nd rejection episode. As expected, the plasma concentrations of CXCL-10 increased again (Fig. 2S).
The levels of two new chemokines included in this study (CXCL-9 and CXCL-11) at the time of LB were significantly higher in rejector patients (CXCL-9 398 pg/mL vs. 119 pg/mL, p < 0.001, 3.3-fold increase; CXCL-11 197.24 pg/mL vs. 83 pg/mL, p < 0.001, 2.3-fold increase). Like plasma CXCL-10 production, both CXCL-9 (p < 0.001, median 2.9-fold increase) and CXCL-11 (p < 0.001, median 1.1-fold increase) were also significantly increased in patients with active CMV infection (Fig. 5A; 5C).
When the LMM was applied, adjusting for within-subject variability and accounting for time as a potential confounding factor, the differences remained statistically significant for all three cytokines (p < 0.001 for both prediction and diagnosis of TCMR).
Biomarker performance and score for TCMR prediction and diagnosis
All miRNAs and chemokines showed significantly higher values preceding and during the TCMR episodes. Similarly, both before and during rejection, LFTs also presented significantly higher levels. Nevertheless, the individual biomarker AUROCs for TCMR diagnosis were higher than those of the LFTs (Table 2). These differences were significant when miRNAs 155-5p, 122-5p, and 181a-5p were compared with all the LFTs and even with the remaining miRNAs and chemokines. However, the gap was clearly wider when the predictive performance was compared. All biomarker AUROCs for TCMR prediction were significantly higher than those of the LFTs. miRNAs 155-5p, 122-5p, and 181a-5p showed the best performance for TCMR prediction.
The biomarker score developed in our previous work, based on the expression of miRNAs 155-5p and 181a-5p and plasma CXCL-10 levels, was tested in this cohort. For the diagnosis of TCMR, the score achieved an AUROC of 0.99 (95% CI, 0.98–0.99), with 100% sensitivity, 99.1% specificity, 100% NPV and 79.3% PPV. For TCMR prediction, the AUROC was 0.95 (95% CI, 0.93–0.96), with 69.1% sensitivity, 98.1% specificity, 97.3% NPV and 76% PPV. Additionally, a stepwise logistic regression analysis was repeated in this new cohort to identify the optimal score. The explanatory variables included were those that showed significant differences between patients with and without rejection: the incidence of HCC prior to LT, the incidence of CMV infection, LFTs and the biomarkers (among the latter two, only those with significant differences in the LMM analysis). Consistent with our previous findings, in the present cohort the best combination also included miRNAs 155-5p and 181a-5p, as well as CXCL-10. Given the lack of a validation cohort, a bootstrap analysis with 1000 resamplings was performed for internal validation. The AUROC for rejection diagnosis was 0.992 (bias corrected 95% CI, 0.98–0.99) and 0.94 (bias corrected 95% CI, 0.91–0.97) for prediction. Nevertheless, when the potential relationship between the score values and the severity of rejection was analyzed, no significant differences were observed between patients with mild and moderate rejection.
Lastly, we performed an analysis of each biomarker, as well as of the score based on the levels of miRNA 155-5p, 181a-5p, and CXCL-10 in other post-LT complications such as BS and CMV infections. In the case of BS, no significant elevations were observed in any of the biomarkers. For CMV, the miRNAs showed no differences, whereas the chemokines, as previously mentioned, exhibited significantly elevated levels. In fact, CXCL-9 and CXCL-10 levels were significantly higher in CMV than in TCMR (Table 3S). However, the score was significantly higher in patients with TCMR (0.67, 95% CI, 0.51–0.83) than in those with BS (0.001, 95% CI, 0.001–0.001, p < 0.001) and CMV (0.021, 95% CI, -0.01–0.03). As depicted in Fig. 6, the score identifies rejection episodes but remains unchanged in other events, thus mitigating the potential confounding effect of CMV infection.
Biomarker score values during rejection compared with other complications. Comparison of the biomarker score values between the TCMR episodes and patients without events, BS and CMV infection. All the differences are statistically significant (p > 0.01).
Biomarker performance in patients submitted to LB
A specific analysis was conducted by selecting only patients who met the GD criteria and underwent LB to rule out rejection. All of these patients had abnormal LFTs that defined GD, with no differences between the 43 TCMR episodes and the remaining 16 nonrejection episodes (Table 4S). TAC trough levels were significantly lower in patients with TCMR (5.9 vs. 9.1, P = 0.01). All microRNAs and chemokines had significantly higher values during TCMR episodes than during nonrejection episodes. Specifically, miRNAs 155-5p and 181a-5p identified all TCMR episodes. The values of miRNAs 194-5p, 483-3p, and 885-5p were also significantly higher in rejection but they did not reach the accuracy observed with 155-5p and 181a-5p.
Discussion
Despite the moderate sample size in our study, we have shown that the expression of miRNAs (miRNA-155-5p, miRNA-122-5p, and miRNA-181a-5p) in the circulation of patients with TCMR reflects what is happening at the liver tissue level. For miRNA-181a-5p, although tissue expression was 1.3-fold higher in patients with rejection, the difference did not reach statistical significance, likely because of the small event size. Few groups have established this relationship, and in our opinion, it is critical for the development of more precise diagnostic and prognostic tools that correlate better with clinical outcomes. The differences in the magnitude of increase of miRNA-155-5p and miRNA-181a-5p expression between liver tissue and peripheral circulation may be explained by differences in their cellular origin, release dynamics, and compartmentalization. miRNAs detected in plasma likely originate not only from hepatocytes but also from infiltrating immune cells (e.g., activated T lymphocytes and macrophages), endothelial cells, or other components of the inflammatory microenvironment. Although we did not directly investigate cellular origin in this study, FANTOM5 data36 show that miRNA-155-5p and miRNA-181a-5p are enriched in immune cell subsets (e.g. lymphoid, monocyte lineages), whereas miRNA-122-5p displays high specificity for hepatocytes. These expression pattern suggests that, in the context of rejection, circulating miRNA-155-5p and miRNA-181a-5p may predominantly signal immune activation, while miRNA-122-5p may reflect hepatocyte injury. Circulating miRNAs likely reflect active release from immune or liver cells during injury, whereas tissue levels provide a static snapshot limited to the biopsy site. Moreover, plasma miRNAs may be more stable and accumulate over time, enhancing their detectability and supporting their role as dynamic, non-invasive biomarker. These new findings strengthen the ongoing validation of the composite biomarker score for early TCMR prediction and diagnosis and further support the clinical use of these circulating miRNAs as noninvasive alternatives to liver biopsy. To determine the full potential of miRNAs as biomarkers, a greater understanding of their expression and release is needed. Future studies integrating cell‐specific transcriptomic and single‐cell data would help delineate these contributions more precisely. It is crucial to confirm that the modulation of circulating miRNAs accurately represents the disease state or condition of interest in the tissue.
In the present cohort, we confirmed our previous results16 regarding the capacity of plasma miRNA-155-5p, miRNA-122-5p and miRNA-181a-5p expression as well as the plasma production of CXCL-10 to assess the risk of TCMR in the early port-transplant period. Furthermore, we evaluated three new miRNAs (miRNA-483-3p, miRNA-885-5p and miRNA-194-5p), which were selected based on promising findings reported by other transplantation centers, with the aim of identifying a miRNA signature evaluated across studies to enable cross-center validation and ensure comparability of results.14,15,16. All three miRNAs significantly increased before and during rejection episodes, which is in line with previous reports. However, when the LMM analysis was performed, only miRNA-885-5p showed statistically significant differences. Although their predictive capacity was better, their diagnostic performance was similar to that of routine LFTs. One possible explanation may be the timing of the events, mainly during the first weeks after LT. The pathogenic mechanisms involved in this early rejection may differ from those involved in late TCMR, which was predominant in previous studies in which these miRNAs were tested. Additionally, we evaluated two new chemokines, CXCL-9 and CXCL-11 The combined evaluation of CXCL9 and CXCL11 (two additional ligands of the CXCR3 receptor) together with CXCL10 was incorporated into this study to determine whether it could enhance the robustness and discriminatory capacity of the existing biomarker score. We hypothesized that an expanded chemokine profile would more accurately reflect the complexity of IFN-γ–mediated immune responses, enhancing the sensitivity and specificity of the score in diverse clinical settings. Both CXCL9 and CXCL11 exhibited predictive and diagnostic potential for TCMR, both individually and within the LMM analysis. We observed that plasma production also increased when there was an active CMV infection, as with CXCL-10, in line with previous reports in kidney and LT. Combining the newly evaluated biomarkers (miRNA-483-3p, miRNA-885-5p, miRNA-194-5p, CXCL-9 and CXCL-11) with our previously established signature (miRNA-155-5p, miRNA-181a-5p and CXCL-10) did not enhance the robustness of the original score for TCMR risk assessment in the early transplant period. The lack of improvement is likely explained by biological overlap and pathway redundancy within the IFN-γ–driven immune response that characterizes T-cell–mediated rejection. CXCL-9, CXCL-10, and CXCL-11 are co-induced by IFN-γ and signal through the same receptor (CXCR3), resulting in highly correlated expression profiles; thus, adding CXCL-9 and CXCL-11 provided little additional discriminatory value beyond that already captured by CXCL-10. Likewise, some of the newly tested miRNAs (miRNA-483-3p and miRNA-885-5p) participate in inflammatory and metabolic stress pathways that overlap with those regulated by miRNA-155-5p and miRNA-181a-5p during immune activation. Therefore, the inclusion of additional, mechanistically related biomarkers may have introduced redundancy rather than complementarity. These findings reinforce the robustness and reproducibility of the original model and suggest that this biomarker panel may be better suited for clinical translation, offering analytical simplicity and consistency across patient populations.
A common concern among clinicians is the origin of emerging LFT abnormalities in the absence of other diagnostic findings that may explain them. Although the suspicion of rejection is always present, similar disturbances may also be observed in other causes of GD. The predictive accuracy of the miRNAs and chemokines was significantly better than that of LFTs. miRNAs 155-5p, 122-5p, and 181a-5p had the best performance for TCMR diagnosis, with AUROCs of 0.99. Therefore, in a setting where clinicians must make decisions on the basis of routine LFT surveillance, these biomarkers could contribute to an optimized assessment. On the one hand, patients whose miRNA and chemokine values are below the TCMR prediction cut-off may safely minimize their use of IS drugs, given their high NPV. In contrast, patients with elevated LFTs and biomarker values above the TCMR diagnostic cut-off should be considered for LB or an increase in IS. When a LB is performed to confirm rejection, miRNAs 155-5p and 181a-5p may be even more useful. In previous studies9, miRNA-155-5p perfectly identified rejector individuals at the time of diagnostic LB only during the first weeks after LT. In the present cohort, miRNA-155-5p identified all patients with TCMR, regardless of the timing of the event. Furthermore, miRNA-181a-5p showed a perfect correlation with this diagnosis in these patients. Thus, when an LB is considered for TCMR confirmation, these biomarkers could serve a non-invasive role in its diagnosis. Even in patients with indefinite biopsy findings, elevated levels of miRNA-155-5p and -181a-5p may aid in clinical management.
In previous work10, a biomarker score based on miRNAs 155-5p, 181a-5p, and CXCL-10 was developed to predict and diagnose TCMR. The score performed similarly in the current cohort. Regression analysis confirmed these biomarkers as the best predictors of rejection. The score improves accuracy and simplifies interpretation of individual results. In the present study, it was also tested for other complications like CMV infection and bacterial sepsis (BS). Although the three chemokines showed significantly elevated levels during episodes of CMV infection (exceeding those observed during TCMR) their combination with miRNA 155-5p and 181a-5p enabled a clear distinction between rejection and CMV-related infections. This finding supports our hypothesis that a biomarker panel, rather than a single biomarker, may provide greater accuracy for the detection of TCMR. Overall, the score clearly identifies patients with rejection, providing a fast and useful tool for post-transplant monitoring.
Regarding the robustness and feasibility of analytical methods, for the analysis of these biomarkers it is necessary to have robust, reproducible, standardized and interlaboratory validated methodologies. Although newer automated assays for chemokine quantification have been developed, we maintained the use of ELISA across all study cohorts to ensure methodological consistency. The ELISA platform demonstrated high robustness and reproducibility, and its wide availability facilitates implementation in centers interested in applying these biomarkers for graft monitoring. Most laboratories have the capacity and experience to carry out a RT-PCR. These analyses are not very labour-intensive, and the turn-around time is minimal; thus, the results could be available less than 48 h. In the case of the measurement of plasmatic miRNAs RT-PCR is the hallmark method for analysis.
Our study has several limitations: (1) Instead of using an array of miRNAs to screen potential biomarkers of rejection, we selected only a few miRNAs based on existing evidence to simplify interpretation and in consideration of economic constraints. Few groups have the resources for large-scale miRNA screening. Therefore, our research relies on existing studies to select miRNA signatures or focus on their regulatory roles in specific pathways. This approach may obviously leave out other potentially valuable miRNAs. (2) The sample size for miRNA expression analysis in LB tissue from TCMR patients was moderate, however, is within the range commonly reported in similar translational studies evaluating correlations between tissue and circulating miRNA expression37,38. (3) The absence of patients with antibody-mediated rejection did not allow us to evaluate the prognostic capacity of the biomarkers for this clinical event. (4) The score’s performance was shown only in the first months post-transplant and has not been evaluated for late rejection beyond six months. (5) Finally, no protocol biopsies were performed in the present cohort. This limitation prevents ruling out the presence of histological signs of rejection in patients with normal LFTs, thereby excluding the diagnosis of potential subclinical rejection episodes.
Conclusions
A key finding of this study is that the expression of circulating miRNA-155-5p and miRNA-122-5p in patients with TCMR reflect their expression levels and evolution in liver tissue. This correlation suggests that these miRNAs in the bloodstream are consistent with the biological processes occurring in the liver during rejection, supporting their potential as reliable non-invasive biomarkers for TCMR.
The findings in the present study confirm our previous results regarding the predictive and diagnostic capacity of the score based on plasma miRNAs 155-5p and 181a-5p and CXCL-10 to assess the risk of TCMR in early post-transplant period. Furthermore, miRNA 885-5p, as well as CXCL-9 and CXCL-11, showed these abilities. However, the addition of these new biomarkers did not improve the accuracy of our previously developed score for the assessment of TCMR risk.
The biomarker score allows clear differentiation between TCMR and other complications, including CMV infection and BS. Although these conditions frequently present with similar LFT abnormalities, the score exhibits superior diagnostic performance and may help clinicians in the early, noninvasive management of post-transplant complications.
Since this study was conducted at a single center, the findings should be interpreted with caution regarding their generalizability. To implement these biomarkers in clinical practice, collaborative efforts among multiple transplant centers will be required to advance the validation of this score in multicenter, prospective intervention trials. These studies will be essential to confirm the performance and reproducibility of the biomarker panel across diverse clinical and laboratory settings. The analyses involved in this score are not very labour-intensive, and the turn-around time is minimal, allowing for extensive use in centres dedicated to transplantation.
Data availability
The datasets generated during the current study are not publicly available in order to preserve the privacy of the study participants. If someone wants to request the data from this study please contact with the corresponding author (Dr. M Brunet).
Abbreviations
- LT:
-
Liver Transplantation
- TCMR:
-
T-cell-mediated rejection
- miRNA:
-
MicroRNA
- LFT:
-
Liver function tests
- LB:
-
Liver biopsy
- HCC:
-
Hepatocellular carcinoma
- GD:
-
Graft dysfunction
- BS:
-
Biliary strictures
- IS:
-
Immunosuppression
- AUROC:
-
Area under the receiver operating curve
- CMV:
-
Cytomegalovirus
- TAC:
-
Tacrolimus
- MMF:
-
Mycophenolate mofetil
- AST:
-
Aspartate aminotransferase
- ALT:
-
Alanine aminotransferase
- qPCR:
-
Quantitative real-time PCR
- FFPE:
-
Formalin-fixed paraffin-embedded
- RAI:
-
Rejection activity index
- CI:
-
Confidence interval
- PPV:
-
Positive predictive value
- NPV:
-
Negative predictive value
- LMM:
-
Linear Mixed-effects Model
- HCV:
-
Hepatitis C virus
- HBV:
-
Hepatitis B virus
References
Terrault, N. A., Francoz, C., Berenguer, M., Charlton, M. & Heimbach, J. Liver transplantation 2023: Status report, current and future challenges. Clin Gastroenterol Hepatol. 21(8), 2150–2166. https://doi.org/10.1016/j.cgh.2023.04.005 (2023).
Adam, R. et al. Annual report of the european liver transplant registry (ELTR) - 50-year evolution of liver transplantation. Transpl Int. 31(12), 1293–1317. https://doi.org/10.1111/tri.13358 (2018).
Levitsky, J. et al. Acute rejection increases risk of graft failure and death in recent liver transplant recipients. Clin Gastroenterol Hepatol. 15(4), 584-593.e2. https://doi.org/10.1016/j.cgh.2016.07.035 (2017).
Tang, L. C. Y. et al. Incidence, epidemiology, and outcomes of acute allograft rejection following liver transplantation in Australia. Liver Transpl. 30(10), 1039–1049. https://doi.org/10.1097/LVT.0000000000000375 (2024).
Montano-Loza, A. J., Rodríguez-Perálvarez, M. L., Pageaux, G. P., Sanchez-Fueyo, A. & Feng, S. Liver transplantation immunology: Immunosuppression, rejection, and immunomodulation. J Hepatol. 78(6), 1199–1215. https://doi.org/10.1016/j.jhep.2023.01.030 (2023).
Van Ha, T. G. Liver biopsy in liver transplant recipients. Semin Intervent Radiol. 21(4), 271–274. https://doi.org/10.1055/s-2004-861561 (2004).
Bardhi, E., McDaniels, J., Rousselle, T., Maluf, D. G. & Mas, V. R. Nucleic acid biomarkers to assess graft injury after liver transplantation. JHEP Rep. 4(3), 100439. https://doi.org/10.1016/j.jhepr.2022.100439 (2022).
Banerjee, A., Schambach, F., DeJong, C. S., Hammond, S. M. & Reiner, S. L. Micro-RNA-155 inhibits IFN-gamma signaling in CD4+ T cells. Eur J Immunol. 40(1), 225–231. https://doi.org/10.1002/eji.200939381 (2010).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316(5824), 608–611. https://doi.org/10.1126/science.1139253 (2007).
Trotta, R. et al. miR-155 regulates IFN-γ production in natural killer cells. Blood 119(15), 3478–3485. https://doi.org/10.1182/blood-2011-12-398099 (2012).
Schaffert, S. A. et al. mir-181a-1/b-1 modulates tolerance through opposing activities in selection and peripheral T cell function. J Immunol. 195(4), 1470–1479. https://doi.org/10.4049/jimmunol.1401587 (2015).
Li, Q. J. et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129(1), 147–161. https://doi.org/10.1016/j.cell.2007.03.008 (2007).
Roderburg, C. et al. Elevated miR-122 serum levels are an independent marker of liver injury in inflammatory diseases. Liver Int. 35(4), 1172–1184. https://doi.org/10.1111/liv.12627 (2015).
Millán, O. et al. Monitoring of miR-181a-5p and miR-155-5p plasmatic expression as prognostic biomarkers for acute and subclinical rejection in de novo adult liver transplant recipients. Front Immunol. 24(10), 873. https://doi.org/10.3389/fimmu.2019.00873 (2019).
Ruiz, P. et al. MicroRNAs 155–5p, 122–5p, and 181a–5p identify patients with graft dysfunction due to T cell-mediated rejection after liver transplantation. Liver Transpl. 26(10), 1275–1286. https://doi.org/10.1002/lt.25842 (2020).
Millán, O. et al. A plasmatic score using a miRNA signature and CXCL-10 for accurate prediction and diagnosis of liver allograft rejection. Front Immunol. 30(14), 1196882. https://doi.org/10.3389/fimmu.2023.1196882 (2023).
Levitsky, J. et al. Discovery and validation of a novel blood-based molecular biomarker of rejection following liver transplantation. Am J Transplant. 20(8), 2173–2183. https://doi.org/10.1111/ajt.15953 (2020).
Levitsky, J. et al. Prediction of liver transplant rejection with a biologically relevant gene expression signature. Transplantation 106(5), 1004–1011. https://doi.org/10.1097/TP.0000000000003895 (2022).
Schaapman, J. et al. MRI-serum-based score accurately identifies patients undergoing liver transplant without rejection avoiding the need for liver biopsy: A multisite European study. Liver Transpl. https://doi.org/10.1097/LVT.0000000000000450 (2024).
Gui, J. et al. Serum microRNA characterization identifies miR-885-5p as a potential marker for detecting liver pathologies. Clin Sci (Lond). 120(5), 183–193. https://doi.org/10.1042/CS20100297 (2011).
El-Diwany, R. et al. Acute hepatitis C Virus infection induces consistent changes in circulating MicroRNAs That Are associated with nonlytic hepatocyte release. J Virol. 89(18), 9454–9464. https://doi.org/10.1128/JVI.00955-15 (2015).
Matson, K., Macleod, A., Mehta, N., Sempek, E. & Tang, X. Impacts of MicroRNA-483 on Human Diseases. Noncoding RNA. 9(4), 37. https://doi.org/10.3390/ncrna9040037 (2023).
Shaked, A. et al. An ectopically expressed serum miRNA signature is prognostic, diagnostic, and biologically related to liver allograft rejection. Hepatology 65(1), 269–280. https://doi.org/10.1002/hep.28786 (2017).
Wang, X., He, Y., Mackowiak, B. & Gao, B. MicroRNAs as regulators, biomarkers and therapeutic targets in liver diseases. Gut 70(4), 784–795. https://doi.org/10.1136/gutjnl-2020-322526 (2021).
Farid, W. R. et al. Hepatocyte-derived microRNAs as serum biomarkers of hepatic injury and rejection after liver transplantation. Liver Transpl. 18(3), 290–297. https://doi.org/10.1002/lt.22438 (2012).
Morsiani, C. et al. Circulating miR-122-5p, miR-92a-3p, and miR-18a-5p as potential biomarkers in human liver transplantation follow-up. Int. J. Mol. Sci. 24, 3457. https://doi.org/10.3390/ijms24043457 (2023).
Brunet, M. et al. Barcelona consensus on biomarker-based immunosuppressive drugs management in solid organ transplantation. Ther Drug Monit. 38(Suppl 1), S1-20. https://doi.org/10.1097/FTD.0000000000000287 (2016).
Rabant, M. et al. Early low urinary CXCL9 and CXCL10 might predict immunological quiescence in clinically and histologically stable kidney recipients. Am J Transplant. 16(6), 1868–1881. https://doi.org/10.1111/ajt.13677 (2016).
Hirt-Minkowski, P. et al. Randomized trial to assess the clinical utility of renal allograft monitoring by urine CXCL10 chemokine. J Am Soc Nephrol. 34(8), 1456–1469. https://doi.org/10.1681/ASN.0000000000000160 (2023).
Gandolfini, I. et al. CXCL9 and CXCL10 as biomarkers of kidney graft inflammation across multiple conditions. Clin Transplant. 38(5), e15324. https://doi.org/10.1111/ctr.15324 (2024).
Shino, M. Y. et al. Plasma CXCL9 and CXCL10 at allograft injury predict chronic lung allograft dysfunction. Am J Transplant. 22(9), 2169–2179. https://doi.org/10.1111/ajt.17108 (2022).
Raschzok, N. et al. CD44 and CXCL9 serum protein levels predict the risk of clinically significant allograft rejection after liver transplantation. Liver Transpl. 21(9), 1195–1207. https://doi.org/10.1002/lt.24164 (2015).
Kim, N. et al. Combined detection of serum IL-10, IL-17, and CXCL10 predicts acute rejection following adult liver transplantation. Mol Cells. 39(8), 639–644. https://doi.org/10.14348/molcells.2016.0130 (2016).
Demetris, A. J. et al. Comprehensive update of the Banff working group on liver allograft pathology: Introduction of antibody-mediated rejection. Am J Transplant. 16(10), 2816–2835. https://doi.org/10.1111/ajt.13909 (2016).
Demetris, A. J. et al. Banff schema for grading liver allograft rejection: an international consensus document. Hepatology 25(3), 658–663. https://doi.org/10.1002/hep.510250328 (1997).
https://fantom.gsc.riken.jp/5/suppl/De_Rie_et_al_2017/vis_viewer/#/human
Anglicheau, D. et al. MicroRNA expression profiles predictive of human renal allograft status. Proc Natl Acad Sci U S A. 106(13), 5330–5335. https://doi.org/10.1073/pnas.0813121106 (2009).
Ben-Dov, I. Z. et al. MicroRNA sequence profiles of human kidney allografts with or without tubulointerstitial fibrosis. Transplantation 94(11), 1086–1094. https://doi.org/10.1097/TP.0b013e3182751efd (2012).
Acknowledgements
CIBEREHD is funded by the Instituto de Salud Carlos III (ISCIII), Spain. We thank all the patients who participated in this study and the support received from the Department of Research and Universities of the Generalitat de Catalunya (AGAUR: Expedient code 2021-SGR-01331).
Funding
This study was funded by the Instituto de Salud Carlos III (ISCIII) through the project PI19/00378 and cofounded by the European Union; and by the LETDDNA project of CareDx and Diagnostica Longwood.
Author information
Authors and Affiliations
Contributions
P.R. selected and followed the patients, performed the statistical analysis and drafted the manuscript; O.M. performed the miRNA and chemokine analyses, interpreted the data, performed the statistical analysis and drafted the manuscript (these two authors contributed equally to this work and share first authorship); Y.F performed transplant surgeries and revised the manuscript; J.J revised the manuscript; G.C. followed the patients and revised the manuscript; R.B revised the manuscript; J.C. followed the patients and revised the manuscript; M.B. conceived and designed the study, interpreted the data, and drafted and revised the manuscript. All authors approved the final version of the manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
All the authors declare that they have no competing interests.
Ethics approval and patient consent
This study was performed in accordance with the Declaration of Helsinki and the E6 Good Clinical Practice Standards ICH and fulfilled the criteria established by the Organic Law of Biomedical Research 14/2007 and the Organic Law of Protection of Personal Data 3/2018. The study was approved by the local Clinical Research Ethics Committee (approval number HCB/2019/0258). All patients were informed about the study and provided written informed consent for inclusion.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ruiz, P., Millán, O., Fundora, Y. et al. Plasma microRNAs and chemokines as biomarkers for rejection in liver transplantation with score verification and tissue correlation. Sci Rep 16, 2271 (2026). https://doi.org/10.1038/s41598-025-32120-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32120-9