Abstract
Environmental DNA (eDNA) is increasingly employed for biodiversity monitoring, yet its accuracy and consistency relative to conventional fish surveys remain under debate. Here, we evaluated fish assemblages at 14 sites across six major rivers in Liaoning Province, China, using four primer sets (12 S rRNA, COI, MiFish, and a high-variation MiFish) coupled with high-throughput sequencing, and compared results with capture-based surveys. eDNA detected 211 species across 17 orders, 71 families, and 146 genera—substantially exceeding traditional methods—but failed to identify nine species recorded morphologically and could not provide ecological attributes such as body size or population structure. Species richness and composition varied between tributary and mainstem sites, reflecting hydrological and habitat gradients. Detection efficiency also differed among primers, underscoring the importance of multi-primer strategies. Overall, eDNA provides a sensitive and efficient complement to conventional surveys, but its limitations in taxonomic resolution and ecological inference highlight the need for integration of molecular and conventional morphological methods to achieve comprehensive fish diversity assessments.
Similar content being viewed by others
Introduction
Freshwater biodiversity is experiencing steep declines due to pollution, habitat alteration, overexploitation and biological invasions1,2. As central components of riverine ecosystems, freshwater fishes contribute to nutrient cycling and food-web dynamics and are widely used as indicators of ecosystem condition3. Yet monitoring these changes remains challenging: conventional surveys (e.g., gill-netting, electrofishing, visual identification) yield direct biological information but are invasive, labour-intensive, costly and reliant on taxonomic expertise. These constraints motivate complementary, low-impact approaches4.
eDNA metabarcoding offers such a complement5. First used to detect invasive American bullfrogs (Rana catesbeiana) in fresh waters6, eDNA has since been widely applied to survey aquatic biodiversity across rivers, lakes and marine systems7,8. In China, eDNA applications are rapidly expanding, including riverine assessments in Beijing (Chaobai and Beiyun Rivers) and the Danjiang River, as well as river–lake networks and estuaries (Pearl River)9, collectively demonstrating feasibility across diverse hydrological settings10,11.
Compared with traditional surveys, eDNA can detect species without capture, reduce ecological disturbance and often improve sensitivity for rare or elusive taxa12,13. However, performance depends on primer choice, environmental conditions and reference-database completeness14, and false positives can arise from DNA persistence or contamination15. Thus, strengths and weaknesses must be weighed in context, and eDNA is best viewed as a valuable but non-standalone tool for comprehensive monitoring16.
Here we compare eDNA metabarcoding with conventional surveys across six rivers in Liaoning Province, China. We targeted two mitochondrial markers—12S rRNA and cytochrome c oxidase subunit I (COI)—and used widely adopted primer sets for 12S rRNA (MiFish-U and a high-variation MiFish set) and validated mini-barcode primers for COI, then contrasted these results with cage-fishing data to evaluate detection sensitivity and taxonomic coverage. By analysing concordance and divergence between approaches, we propose an integrated framework that leverages the complementarity of molecular and traditional methods to optimize fish biodiversity monitoring in riverine ecosystems.
Materials and methods
Study area and sampling locations
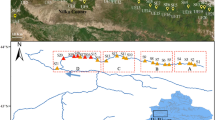
Based on hydrography, channel morphology, and confluence structure of the Liao River Basin17, we established 14 riverine sampling sites in September 2024 (autumn). Sites were distributed as follows: Xiaoling River (A1–A3), Liao River mainstem (A4–A6, A8, A9, A11), Dayang River (A7), Qing River mouth (A10), Taizi River (A12–A13), and Yingna River (A14). The spatial arrangement captured longitudinal and tributary gradients as well as the estuarine transition (Fig. 1).
Spatial distribution map of the study area in the Liao River Basin and location map of sampling sites, where A1, A2 and A3 are Xiaoling River, A4, A5, A6, A8, A9 and A11 are Liao River, A7 is Dayang River, A10 is Qing River, A12 and A13 are Taizi River, A14 is Yingna River.
Sample collection and processing
eDNA water sampling and laboratory processing
At each site, 15 L of surface water (0–30 cm) was collected using a sterile water sampler and kept on ice (~ 4 °C) during transport. Before each sampling event, all reusable equipment was disinfected with 10% sodium hypochlorite and rinsed thoroughly with deionized water; single-use consumables were employed whenever possible to minimize contamination risk18. In the laboratory, water from the same site was composited and split into three technical replicates (4 L each); the remaining 3 L was discarded following common practice19,Luo et al., 2015; Xu et al., 2016). Replicates were filtered within 24 h through 0.45 µm mixed-cellulose-ester membranes (Whatman, UK) using a vacuum manifold. When turbidity or suspended sediment was high, samples were pre-filtered in the field through sterile medical gauze20,21. Filtration units were decontaminated between samples as above. We processed field/lab blanks (2 L molecular-grade water) alongside samples to monitor exogenous DNA. Filters were stored at − 80 °C until extraction.
Total DNA was extracted from filters using the PowerWater DNA Isolation Kit (Qiagen Genomic-Tips) per the manufacturer’s protocol. DNA quality was screened by 1% agarose gel electrophoresis22,23. Each replicate was extracted independently, and unused filters served as extraction blanks. DNA was stored at − 20 °C prior to PCR.
Conventional capture-based sampling and specimen processing
To generate a morphology-based benchmark, we conducted standardized capture-based surveys at the same site. At each site we deployed one 6 m ground cage (0.5 m × 0.5 m; 1.5 cm mesh) for a 24 h soak (start: late afternoon; retrieval: next day), positioned along near-bank habitat with reduced current. This effort (net length and soak time) was standardized across sites. Where hydrodynamic conditions permitted, we additionally used set/hang nets or dip (plunge) nets to supplement species detection; ancillary catches were recorded as presence-only and were not used in quantitative comparisons. Specimens were identified in situ whenever possible; individuals requiring lab confirmation were preserved on ice or in 5–10% neutral buffered formalin to prevent tissue degradation.
Taxonomic identification
eDNA technical identification
To increase coverage of fish diversity, we targeted two mitochondrial markers—12S rRNA and cytochrome c oxidase subunit I (COI)—with validated primer sets. For 12S rRNA, we used the MiFish-U primers and a high-variation MiFish set; we also included a generic fish 12S rRNA primer pair to cross-validate detections. For COI, we used a commonly adopted fish mini-barcode primer pair. PCRs (25 µL) contained 12.5 µL 2 × Taq Master Mix, 1 µL each primer (10 µM), 2 µL DNA template, and 8.5–9.5 µL nuclease-free water; cycling followed primer-specific recommendations. Each extract was amplified in triplicate PCRs with PCR negatives included. Amplicons were visualized by agarose gel, pooled per replicate, gel-purified, and libraries were prepared with Illumina TruSeq kits. Sequencing used MiSeq v3 2 × 300 bp paired-end chemistry. The details referto·Figure. S1.and Table. S1.
Morphology-based identification of captured fish
Fish were identified to the lowest possible taxon using standard keys(Sofia D. et al., 2025); when necessary, vouchers were confirmed in the laboratory by detailed morphological examination (meristics, morphometrics) with body length and mass recorded. Diet (stomach contents), gonadal development, and age structure (as feasible) were assessed to obtain ecological information. Taxonomy followed FishBase (https://www.fishbase.de/) and Species and Distribution of Inland Fishes of China24.
Bioinformatics and statistical analyses
Raw sequences obtained from high-throughput sequencing were subjected to quality control, filtering, splicing, and clustering to generate species abundance data. Operational taxonomic units (OTUs) were clustered at a similarity threshold of ≥ 97%25 Representative OTU sequences were annotated using the National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/) and MitoFish (http://mitofish.aori.u-tokyo.ac.jp/) databases(version 4.09, accessed on 8 February 2025). To ensure accuracy, taxonomic assignments were cross-validated against distribution records of aquatic organisms in typical watersheds of Liaoning, and non-fish sequences were manually excluded. Based on the OTU clustering results, fish species composition and α-diversity indices were calculated. Statistical analyses and data visualizations, including Venn diagrams, bar plots, and heatmaps, were performed using R (v.4.4.1)26,27 with the packages ggplot228 and vegan29.
Results
Identification of fish species composition based on eDNA technology
Using eDNA to analyse water samples from typical rivers in Liaoning Province, we detected 211 fish species spanning 17 orders, 71 families, and 146 genera, with Cyprinidae contributing the largest number of taxa. As shown in Fig. 2, primers (a) and (c) recovered more taxa—particularly at the genus and species levels—than primers (b) and (d), indicating broader coverage and higher resolution at lower taxonomic ranks. By contrast, primers (b) and (d) yielded fewer assignments overall, consistent with narrower coverage and potential taxonomic bias.
Statistics on the results of OTU delineation and classification status identification in typical watersheds of the Liao River. Note: Figure (a)-Figure (b) Results of four primers identified by fish mitochondrial 12S rRNA gene amplification, COI, fish mitochondrial 12S rRNA amplification Mifish, and fish mitochondrial 12S rRNA amplification Mifish**, respectively.
Because the four eDNA primer sets are variants of the same molecular approach, primer-wise contrasts are treated as within-eDNA performance comparisons. Any comparison with the capture-based morphological survey is made at the method level, using the pooled eDNA species list (rather than primer-by-primer results), to acknowledge the non-equivalence of four eDNA assays versus a single morphology-based method.Fig. 3 presents the ten dominant fish genera per site, with co-occurring genera shown side by side, totaling 26 genera. Specifically included:Schizothorax、 Acheilognathus、 Rhodeus、 Carassius、 Channa、 Neosalanx、 Rhinogobius、 Glossogobius、 Silurus、 Hypophthalmichthys、 Triplophysa、 Onychostoma、 Odontobutis、 Tor、 Mugil、 Cyprinus、 Macropodus、 Hypomesus、 Oreochromis、 Microphysogobio、 Rhinogobio、 Chanodichthys、 Kuhlia、 Megalobrama、 Larimichthys、 Acipenser、 Opsariichthys。Of these, the genera Channa and Hypophthalmichthys were present in all sites, demonstrating their wide distribution in the region (genera tied for the top ten occurrences have also been included).
Relative Abundance of Fish Dominant Genus Species Between Sampling Sites. Note: The top ten dominant fish genera were selected for this paper, and genera occurring side-by-side were included in the analysis to fully characterize the fish community in the region.
Fish biodiversity
Fish alpha diversity
Figure 4 presents rarefaction curves for the four primer sets (A–D), with sequencing depth on the x-axis and the number of observed features on the y-axis. In all cases, the number of features increased with sequencing depth and approached saturation at higher read numbers. Primer C tended to reach saturation between 60,000 and 80,000 reads, suggesting relatively higher efficiency, whereas primers A and B required 80,000–100,000 reads before leveling off. Primer D consistently produced fewer observed features than the other sets, with curves stabilizing earlier and at lower values. These patterns indicate variation among primers in their ability to recover community diversity, with primer C yielding comparatively higher richness and primer D showing lower sensitivity.
Species alpha diversity index sparse curves for 4 primers.
Figure 5 shows rank–abundance distributions of OTUs for the four primer sets (A–D). All curves exhibited long-tailed patterns, with a few high-abundance OTUs and many rare OTUs. Primers B and C produced broader curves with more OTUs and a more even distribution of abundant and rare taxa, whereas primers A and D showed narrower curves with fewer OTUs and steeper declines in relative abundance. Differences in abundance patterns were evident among primers and across samples.
Clustering curves of species alpha diversity index ranks for the four primers.
Beta biodiversity of fish
Figure 6 shows principal coordinate analysis (PCoA) of fish communities across 14 sites using four primer sets (A–D). The proportion of variance explained by PCo1 and PCo2 differed among primers, ranging from 29.1% + 18.7% for primer C to 65.8% + 11.0% for primer A. Sample distributions also varied: primer A produced more dispersed patterns, indicating greater community differentiation, whereas primers B and C showed clearer grouping of sites. Primer D accounted for 38.3% + 25.5% of the variance and revealed distinct inter-cluster separation. Together, these results indicate that community structure patterns depend on primer choice.
PCoA principal component analysis of the principal coordinates of each point under 4 primer assays.
Figure 7 shows hierarchical clustering of fish communities based on the four primer sets (A–D), using the ten most abundant genera. Each subfigure combines a dendrogram with bar plots of relative abundance. Primer A produced clusters dominated by a few genera, whereas primer B showed more even distributions. Primer C yielded balanced clustering with smaller differences among groups, while primer D formed tighter clusters with greater dominance by a few taxa. Overall, clustering patterns differed among primers, reflecting variation in community structure and dominance of key genera.
Hierarchical clustering analysis of species β-diversity indices for 4 primers.
Analysis of fish species differences between sites under four different primers
Figure 8 illustrates the numbers of shared and unique OTUs across 14 sampling sites for the four primer sets (A–D). The number of shared OTUs ranged from 7 (primer D) to 14 (primer B), while the number of unique OTUs varied widely among sites and primers. Site A7 consistently showed the highest number of endemic OTUs across all primers, reaching over 300 for primers B and C, whereas some sites (e.g., A10) yielded few or none. Overall, primer B detected the greatest number of shared OTUs, indicating higher overlap among sites, while primers B and C also revealed strong species uniqueness. Primer D produced the lowest values for both shared and unique OTUs, reflecting weaker detection capacity.
Petal map of common OUT at different sampling points under 4 primers.
Figure 9 shows heatmaps of species composition across 14 sites under the four primer sets (A–D). Abundance patterns varied markedly among sites and primers. Under primer A, certain species were enriched at sites A2 and A3 but scarce elsewhere. Primer B detected species such as Leuciscus idus and Neosalanx taihuensis with strong geographic variation. Primer C revealed higher diversity at sites A7 and A12, with taxa such as Mugil cephalus and Acipenser sinensis concentrated at these locations. Primer D highlighted species enrichment at site A7, while distributions at other sites were more scattered. Overall, the primers differed in their detection of species abundance and clustering patterns, reflecting primer-specific sensitivity to community composition.
Heat map of species composition at different sampling sites under 4 primers.
Comparison of eDNA technology with conventional morphological methods
A total of 19 fish species from 4 orders, 9 families, and 19 genera were captured and identified across the 14 sites using capture-based morphological surveys. The catch was dominated by Cyprinidae (73%), followed by Cobitidae (8.89%), with a third family contributing 5.56%. The most abundant species were white minnow (n = 16) and common carp (n = 10). Species recorded by capture but not recovered by eDNA included Rhynchocypris lagowskii、 Hypophthalmichthys molitrixa、 Sea catfish、 Pelteobagrus fulvidraco、 Hypseleotris swinhonis、 Perccottus gleni、 Cobitis taeniaLinnaeus、 Odontobutis obscurus、 Lophius litulon. representing ~ 4% of the overall species pool. In comparison, eDNA identified a total of 211 fish species across the basin (Fig. 10).
Wayne’s diagram of eDNA methods versus conventional morphological methods methods for investigating species types. Note: The middle section indicates species identified by traditional methods but not by eDNA methods.
Table S2 indicates that nine species recorded using conventional morphological methods surveys were absent from the eDNA results.
Discussions
Species composition of fishes in typical watersheds of Liaoning based on eDNA technology
This basin-wide eDNA survey represents the first multi-marker assessment of fish diversity in the Liao River30, and it yielded a region-wide total of 211 species spanning 17 orders, 71 families, and 146 genera—demonstrating the power of eDNA to generate broad biodiversity inventories and to resolve among-site differences in community composition31. Cyprinidae dominated species richness and relative abundance, aligning with32 and likely driven by the family’s adaptation to fast, cold, montane rivers—conditions that promote widespread occurrence and high local abundance in our samples.The multi-primer strategy (two 12S assays plus COI) revealed clear performance differences, aligning with evidence that MiFish markers generally provide superior species-level resolution in freshwater systems33. Importantly, primer detectability is moderated by eDNA degradation and transport, driven by flow velocity, turbidity, temperature, and microbial activity34. Combining markers therefore maximizes taxonomic recovery and minimizes locus-specific biases35,36
Marked spatial heterogeneity in species richness reflected watershed-scale gradients. Tributaries (A1–A3, A7, A14) supported higher local richness of narrow-niche taxa, whereas mainstem sites hosted broader-tolerance assemblages. Restricted detection of Mugil cephalus and Acipenser sinensis at A7 and A12 further highlights how hydrology, resource availability, and human impacts interact to shape local communities37. These patterns suggest that hydrological connectivity and habitat filtering act as the predominant ecological processes shaping α-diversity, a conclusion that aligns with previous empirical evidence reported by Valentini et al. and Olds et al.38,Olds et al., 2009).
Analysis of fish diversity in typical watersheds of the Liao River based on eDNA technology
Substantial primer effects were evident in both α- and β-diversity metrics. Primers B and C achieved higher richness saturation and more even rank–abundance distributions than primers A and D, emphasizing the significance of marker choice in eDNA-based community profiling39,40,41.
Primer A explained the highest PCoA variance but generated greater among-site dispersion, whereas primers B and C produced clearer, ecologically interpretable clustering. These findings mirror earlier reports that 12S rRNA markers reliably resolve freshwater β-diversity patterns42,43. Despite methodological variation, all primers captured consistent basin structure: tributaries formed distinct clusters from mainstem sites, and Taizi River samples grouped tightly, reflecting shared species pools and hydrological regimes. Thus, ecological signals underpinning community turnover remain robust to primer choice.
Exploring the advantages and disadvantages of eDNA technology over conventional morphological methods methods
eDNA is a powerful tool for ecological research and aquatic biodiversity monitoring, offering several advantages over traditional capture-based surveys44,45. It does not rely on on-site taxonomic expertise, thereby reducing observer bias; it is highly sensitive, allowing detection of elusive or rare species; and it enables efficient processing of large sample volumes, saving time and labor46. In the study, eDNA detected 211 species across the Liao River, substantially more than the traditional method. However, capture-based surveys still identified nine species not recovered by eDNA, highlighting the complementarity of the two approaches47,48. Some discrepancies may be explained by exogenous or “legacy” DNA inputs from upstream transport, aquaculture effluents, or predator feces and carcasses, which can decouple DNA presence from live local populations.
Despite its promise, eDNA currently faces several limitations. False positives primarily stem from sample contamination, laboratory cross-contamination, or bioinformatic errors that misassign sequences(Fonseca, V.G. et al.,2023). Conversely, false negatives frequently occur due to the absence of target taxa from reference databases(Kirtane et al., 2021), preventing species assignment, or because of low eDNA concentration in the environment that falls below the detection limit. These limitations underscore the necessity of stringent laboratory controls and robust bioinformatic filtering(Martin P. et al.,2025). Closely related species remain difficult to discriminate due to high sequence similarity, and eDNA cannot provide critical biological information such as body size, age, sex, biomass, or reproductive status49,50. Moreover, mitochondrial inheritance complicates the detection of hybrids51. High-throughput sequencing itself can introduce erroneous sequences differing from true biological sequences by a few bases52,53,54.
In summary, while eDNA greatly expands species detection, it cannot yet replace capture-based surveys. Instead, the two methods are best viewed as complementary: eDNA provides broad, non-invasive biodiversity screening, while traditional methods deliver essential ecological and demographic data. This complementarity offers practical value for biodiversity monitoring, supporting more robust river health assessments and improving the detection of rare or protected species(Kopp et al., 2023). Continued optimization of primer design, reference databases, and analytical pipelines will further strengthen the role of eDNA in conservation and fisheries management(Kopp et al., 202355;).
Conclusions
While eDNA substantially broadens taxonomic coverage, conventional morphology remains indispensable and cannot be replaced at this stage because it provides vouchered specimens, diagnostic traits, and demographic information and verifies local occurrence. Using this combined approach, we detected 211 fish species spanning 17 orders, 71 families, and 146 genera across six representative rivers in Liaoning Province, whereas conventional surveys alone recorded 19 species from 4 orders, 9 families, and 19 genera. Cyprinidae dominated both datasets, and nine species recovered exclusively by conventional sampling underscore the continuing necessity of morphology-based methods.
Nevertheless, the results clearly demonstrate that eDNA greatly expands the range of taxa detected, improves the discovery of rare and concealed species and provides an efficient and scalable strategy for biodiversity monitoring at the river basin scale. Conventional methods remain essential for validating species presence and obtaining fundamental information on population attributes such as body size, age structure and health status. Therefore, integrating eDNA monitoring with targeted traditional sampling will enhance the comprehensiveness, accuracy and cost efficiency of fish diversity assessments in river ecosystems. This study provides practical guidance for applying eDNA-based monitoring in large river systems and supports scientific decision-making in freshwater conservation, fisheries resource management and ecological restoration in northeastern China and other temperate river basins.
Data availability
All data is stored in the GitHub repository https://github.com/SongShuang408/Comparison-between-eDNA-and-Traditional-Morphological-Methods.
References
Albert, J. S. et al. Scientists’ warning to humanity on the freshwater biodiversity crisis. Ambio 49(5), 865–882. https://doi.org/10.1007/s13280-019-01378-8 (2019).
Almond, R. E. A., Grooten, M., & Peterson, T. Living Planet Report 2020—Bending the curve of biodiversity loss (2020).
R Core Team R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/ (2024).
Albert, J. S. et al. Scientists’ warning to humanity on the freshwater biodiversity crisis. Ambio 50(1), 85–94. https://doi.org/10.1007/s13280-020-01318-8 (2021).
Wang, Y. et al. Application of environmental DNA metabarcoding to identify fish community structure in subtropical rivers. Ecol. Evol. 14(12), e11563. https://doi.org/10.1002/ece3.11563 (2024).
Ficetola, G. F., Miaud, C., Pompanon, F. & Taberlet, P. Species detection using environmental DNA from water samples. Biol. Lett. 4(4), 423–425. https://doi.org/10.1098/rsbl.2008.0118 (2008).
Deiner, K. et al. Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Mol. Ecol. 26(21), 5872–5895. https://doi.org/10.1111/mec.14350 (2017).
Jerde, C. L., Mahon, A. R., Chadderton, W. L. & Lodge, D. M. “Sight-unseen” detection of rare aquatic species using environmental DNA. Conserv. Lett. 4(2), 150–157. https://doi.org/10.1111/j.1755-263X.2010.00158.x (2011).
Yamanaka, H. & Minamoto, T. The use of environmental DNA of fishes as an efficient method of determining habitat connectivity. Ecol. Ind. 62, 147–153. https://doi.org/10.1016/j.ecolind.2015.11.022 (2016).
Gao, H. et al. Fish communities and diversity in river ecosystems on the Qinghai-Tibet Plateau revealed by environmental DNA (eDNA) method. Ecol. Ind. 156, 111185. https://doi.org/10.1016/j.ecolind.2023.111185 (2023).
Valentini, A. et al. Next-generation monitoring of aquatic biodiversity using environmental DNA metabarcoding. Mol. Ecol. 25(4), 929–942. https://doi.org/10.1111/mec.13428 (2016).
Afzali, S. F., Besson, M., Lecellier, G. & Adjeroud, M. Comparing environmental DNA metabarcoding and trawling survey of demersal fish communities. Environ. DNA 2(4), 493–509. https://doi.org/10.1002/edn3.72 (2020).
Hsu, T.-H.T., Chen, W.-J. & Denis, V. Navigating the scales of diversity in subtropical and coastal fish assemblages ascertained by eDNA and visual surveys. Ecol. Ind. 148, 110044. https://doi.org/10.1016/j.ecolind.2023.110044 (2023).
Miya, M. et al. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. R. Soc. Open Sci. https://doi.org/10.1098/rsos.15008 (2015).
Taberlet, P., Coissac, E., Hajibabaei, M. & Rieseberg, L. H. Environmental DNA. Mol. Ecol. 21(8), 1789–1793. https://doi.org/10.1111/j.1365-294X.2012.05542.x (2012).
Carraro, L. Integrating direct observation and environmental DNA data to enhance species distribution models in riverine environments. Eco. Inform. 86, 103056. https://doi.org/10.1016/j.ecoinf.2025.103056 (2025).
Song, F. et al. Change in the sustainability of regional agricultural systems:based on an emergy decomposition analysis. Environ. Res. Lett. 19(10), 104075. https://doi.org/10.1088/1748-9326/ad786b (2024).
Polanco Fernández, A. et al. Comparing environmental DNA metabarcoding and underwater visual census to monitor tropical reef fishes. Environ. DNA 3(1), 142–156. https://doi.org/10.1002/edn3.141 (2021).
Janosik, A. M. & Johnston, C. E. Environmental DNA as an effective tool for detection of imperiled fishes. Environ. Biol. Fishes 98(8), 1889–1893. https://doi.org/10.1007/s10641-015-0405-5 (2015).
Xin, M., Wu, Q., Zhang, Y., Zhang, L. & Zhang, J. Application of environmental DNA metabarcoding in monitoring fish diversity in Liaohe National Park. Front. Mar. Sci. 11, 1403700. https://doi.org/10.3389/fmars.2024.1403700 (2024).
Alberdi, A., Aizpurua, O., Gilbert, M. T. P. & Bohmann, K. Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods Ecol. Evol. 9(1), 134–147. https://doi.org/10.1111/2041-210X.12849 (2018).
Hao, Y., Zhang, A., Liu, J. & Gu, Z. Application of environmental DNA technology in the study of fish resources. Biotechnol. Bull. 34(12), 56 (2018).
Pilliod, D. S., Goldberg, C. S., Arkle, R. S. & Waits, L. P. Estimating occupancy and abundance of stream amphibians using environmental DNA from filtered water samples. Can. J. Fish. Aquat. Sci. 70(8), 1123–1130. https://doi.org/10.1139/cjfas-2013-0047 (2013).
Yuan, Y., Zhang, J., Wei, F. & Li, X. Study on fish species diversity in the Pingzhai Reservoir based on environmental DNA technology. Fishes 9(10), 382. https://doi.org/10.3390/fishes9100382 (2024).
Carvalho, C. O. et al. Harnessing eDNA metabarcoding to investigate fish community composition and its seasonal changes in the Oslo fjord. Sci. Rep. 14(1), 10154 (2024).
Poyntz-Wright, I. P., Harrison, X. A., Pedersen, S. & Tyler, C. R. Effectiveness of eDNA for monitoring riverine macroinvertebrates. Sci. Total Environ. https://doi.org/10.1016/j.scitotenv.2024.173621 (2024).
Oksanen, J. et al. Vegan: Community Ecology Package. R package version 2.6–4. https://CRAN.R-project.org/package=vegan (2022).
Wang, P. et al. Environmental DNA: An emerging tool in ecological assessment. Bull. Environ. Contam. Toxicol. 103(5), 651–656. https://doi.org/10.1007/s00128-019-02720-z (2019).
Nogueira, H. G. et al. Environmental DNA template variation: Its relevance for species detection and conservation. Freshw. Biol. 70(1), 19–34. https://doi.org/10.1111/fwb.70019 (2025).
Huang, L., Wang, X., Chen, Y., Zhao, P. & Liu, Z. Insights into the process of fish diversity pattern changes and the current status of spatiotemporal dynamics in the Three Gorges Reservoir area using eDNA. Fishes 10(6), 295. https://doi.org/10.3390/fishes10060295 (2025).
Hallam, J., Clare, E. L., Jones, J. I. & Day, J. J. High frequency environmental DNA metabarcoding provides rapid and effective monitoring of fish community dynamics. Environ. DNA 5(6), 1623–1640. https://doi.org/10.1002/edn3.486 (2023).
Xu, N. & Chang, J. Preliminary study on fish species detection in the middle and lower Yangtze River using environmental DNA. J. Hydroecol. 37, 49–55 (2016).
Miya, M., Gotoh, R. O. & Sado, T. MiFish metabarcoding: a high-throughput approach for simultaneous detection of multiple fish species from environmental DNA and other samples. Fish. Sci. 86(6), 939–970. https://doi.org/10.1007/s12562-020-01461-x (2020).
Sun, B. et al. Environmental DNA exploring the distribution of Indo-Pacific humpback dolphins and fish diversity from the Pearl River Estuary China. Marine Pollut. Bull. 216, 118026. https://doi.org/10.1016/j.marpolbul.2025.118026 (2025).
Bessey, C., Neilson, M. E., Pavlova, A., Hogan, Z. S. & Miller, A. D. Maximizing fish detection with eDNA metabarcoding. Environ. DNA 2(4), 493–504. https://doi.org/10.1002/edn3.74 (2020).
Deiner, K., Fronhofer, E. A., Mächler, E., Walser, J. C. & Altermatt, F. Environmental DNA reveals that rivers are conveyer belts of biodiversity information. Nat. Commun. 7, 1–9. https://doi.org/10.1038/ncomms12544 (2016).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. ISBN 978–3–319–24277–4, https://ggplot2.tidyverse.org (2016).
Tzafesta, E. & Shokri, M. The combined negative effect of temperature, UV radiation and salinity on eDNA detection: A global meta-analysis on aquatic ecosystems. Ecol. Ind. 176, 113669. https://doi.org/10.1016/j.ecolind.2025.113669 (2025).
Grey, E. K. et al. Effects of sampling effort on biodiversity patterns estimated from environmental DNA metabarcoding surveys. Sci. Rep. 8, 8843. https://doi.org/10.1038/s41598-018-27048-2 (2018).
Hänfling, B. et al. Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol. Ecol. 25(13), 3101–3119. https://doi.org/10.1111/mec.13660 (2016).
Reid, A. J., Carlson, A. K., Creed, I. F., Eliason, E. J. & Gell, P. A. Johnson, PTJ, Cooke, SJ Emerging threats and persistent conservation challenges for freshwater biodiversity. Biol. Rev. 94(3), 849–873. https://doi.org/10.1111/brv.12480 (2019).
Dalongeville, A. et al. Benchmarking eleven biodiversity indicators based on environmental DNA. J. Appl. Ecol. 59(8), 2114–2127. https://doi.org/10.1111/1365-2664.14215 (2022).
Valentini, A., Pompanon, F. & Taberlet, P. DNA barcoding for ecologists. Trends Ecol. Evol. 24(2), 110–117. https://doi.org/10.1016/j.tree.2008.09.011 (2009).
Sahu, A. et al. A systematic review on environmental DNA (eDNA) Science: An eco-friendly survey method for conservation and restoration of fragile ecosystems. Ecol. Indicators. 173, 11344. https://doi.org/10.1016/j.ecolind.2025.113441 (2025).
Duarte, S., Santos, A. M., Antunes, C., Sousa, R. & Costa, F. O. Detection of diadromous fish using environmental DNA: prospects for its use in conservation of endangered species occurring in Portugal. Aquat. Conserv. Mar. Freshwat. Ecosyst. 35(8), e70214. https://doi.org/10.1002/aqc.70214 (2025).
Altermatt, F. et al. Blackman, Utilizing aquatic environmental DNA to address global biodiversity targets. Nat. Rev. Biodivers. 1, 332–346. https://doi.org/10.1038/s44358-025-00044-x (2025).
Olds, A. D., Connolly, R. M., Pitt, K. A. & Maxwell, P. S. Habitat connectivity improves reserve performance. Conserv. Lett. 5(1), 56–63. https://doi.org/10.1111/j.1755-263X.2011.00204.x (2012).
Kirtane, A. et al. Quantification of environmental DNA (eDNA) shedding and decay rates for three commercially harvested fish species and comparison between eDNA detection and trawl catches. Environ. DNA 3(6), 1142–1155. https://doi.org/10.1002/edn3.2362021 (2022).
Cristescu, M. E., & Hebert, P. D. N. Uses and Misuses of Environmental DNA in Biodiversity Science and Conservation. In D. J. Futuyma (Ed.), Annual Review of Ecology, Evolution, and Systematics, 49, 209–230 (2018).
Kumar, G., Reaume, A. M., Farrell, E. & Gaither, M. R. Comparing eDNA metabarcoding primers for assessing fish communities in a biodiverse estuary. PLoS ONE 17(6), e0266720. https://doi.org/10.1371/journal.pone.0266720 (2022).
Giles, R. E., Blanc, H., Cann, H. M. & Wallace, D. C. Maternal inheritance of human mitochondrial DNA. Proc Natl Acad Sci USA 77(11), 6715–6719. https://doi.org/10.1073/pnas.77.11.6715 (1980).
Coissac, E., Riaz, T. & Puillandre, N. Bioinformatic challenges for DNA metabarcoding of plants and animals. Mol. Ecol. 21(8), 1834–1847. https://doi.org/10.1111/j.1365-294X.2012.05550.x (2012).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. 081257. https://doi.org/10.1101/081257 %J bioRxiv (2016).
Miya, M. & Nishida, M. Use of mitogenomic information in teleostean molecular phylogenetics: a tree-based exploration under the maximum-parsimony optimality criterion. Mol. Phylogenet. Evol. 17(3), 437–455. https://doi.org/10.1006/mpev.2000.0839 (2000).
Gao, L., Liu, T. & Li, X. Identification and analysis of the endoplasmic reticulum stress hub genes in sepsis-associated ARDS. Sci. Rep. 15, 31007. https://doi.org/10.1038/s41598-025-16644-8 (2025).
Funding
This research was funded by the National Key Research and Development Program of China(2022YFF1301001-03).
Author information
Authors and Affiliations
Contributions
Conceptualisation: Yu, Y.;Shao, M.;Su, F.;Song, S.and Qu, B.Developing methods: Yu, Y.;Shao, M.;Su, F.;Song, S.and Qu, B.Conductingteresearch: Yu, Y.;Shao, M.;Zhou, H.;Su, F.;Guan, P.;Ding, L.;Ma, T.;ZhouB.;Xu, S.;Song, S.and Qu, B.Data analysis, Data interpretation: Yu, Y. and Shao, M.Preparation of figures and tables: Yu, Y.;Shao, M.;Su, F. and Song, S.Writing: all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, Y., Shao, M., Zhou, H. et al. Comparison between eDNA and traditional morphological methods for fish diversity monitoring in rivers. Sci Rep 16, 3098 (2026). https://doi.org/10.1038/s41598-025-32964-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32964-1