Abstract
The gallbladder is a small, hollow organ positioned beneath the liver, primarily responsible for temporarily storing bile. Bile is a fluid formed by the liver that helps digestion. There are various types of gallbladder disease. Early diagnosis and identification are crucial for effective treatment of gallbladder disorders. Poor medical outcomes and improved patient symptoms may result from errors or delays in diagnosis. Numerous symptoms and signs, particularly those associated with gallbladder disorder, might be blurred. As a result, medical specialists must understand and interpret ultrasound images. Considering that ultrasound imaging for diagnosis is labour- and time-consuming, it might be difficult to support financially. Deep learning (DL) is an effective model that could help early identify gallbladder disease using ultrasound (US) images. In this paper, a Hybrid Deep Learning Model with Feature Engineering for the Accurate Diagnosis of Gallbladder Disease Types (HDLMFE-ADGDT) approach is proposed. The primary purpose of the HDLMFE-ADGDT approach is to develop an effective DL-based method for classifying GB disease categories. As an initial step, the HDLMFE-ADGDT technique employs a Non-Local Means (NLM) filter to remove noise and enhance the overall image quality. For feature extraction, the Squeeze-and-Excitation Capsule Network (SE-CapsNet) is employed. At last, the hybrid convolutional neural network with bidirectional long short-term memory (CNN-BiLSTM) is implemented for gallbladder disease diagnosis. A comprehensive set of experiments is conducted to validate the performance of the HDLMFE-ADGDT model on the Gallbladder diseases dataset. The empirical results demonstrate that the HDLMFE-ADGDT model outperformed existing methodologies, achieving 99.09% accuracy, 95.83% precision, 95.87% sensitivity, and 99.49% specificity.
Similar content being viewed by others
Introduction
Gallbladder (GB) illness is a common condition that requires early and accurate analysis. The GB is an intraperitoneal organ located on the lower surface of the liver. The GB is near the transverse colon, pancreas, and duodenum1. Structurally, the GB is divided into three parts: the fundus and body, the neck, and the infundibulum. Bile is a fluid secreted by the liver that aids digestion2. Cholelithiasis (gallstones) is a common GB illness that affects around 10–15% of adults. Nine pathological problems are revealed among the prevalent GB disorders currently. This comprises polyps and cholesterol crystals, GB perforation, GB adenomyomatosis, GB wall thickening, cholecystitis, gallstones, carcinoma, gangrenous cholecystitis, and retroperitoneal and intra-abdominal problems3. It is essential to identify the GB disease type and its severity in the initial phase to prevent or reduce its spread4. To accurately diagnose GB disease, ultrasound (US) is often used as an imaging modality, a strong, universal screening and diagnostic tool for radiologists and doctors. Next to screening, a precise diagnosis is needed to recognise a suitable treatment plan5.
Usually, diagnostic information is gathered from the patient’s past medical tests. Symptoms and indications may be unclear, specifically in GB diseases. Hence, US imaging must be understood and interpreted by highly trained medical service providers6. Considering that diagnosis via US image is labour-intensive and expensive, and that accessing this service in remote locations is challenging. With the rapid development of medical science and technology over the past few decades, artificial intelligence (AI) models can significantly reduce human effort7. In US imaging, the AI’s DL domain is an instructive medical tomography model that helps in the timely detection of GB disease8. To help medical professionals detect and classify various GB pathologic differences, DL methodologies have demonstrated outstanding ability to analyse medical images9. By automating the analysis procedure and leveraging large datasets and advanced models, this method can improve the precision of GB disease diagnosis. Furthermore, DL models can assist specialists in the initial diagnosis, treatment, and detection of illnesses, thereby providing effective techniques for medical analysis10.
In this study, a Hybrid DL Model with Feature Engineering for the Accurate Diagnosis of Gallbladder Disease Types (HDLMFE-ADGDT) model is proposed. The main contributions of this paper are summarised below:
-
The HDLMFE-ADGDT technique aims to develop an effective DL-based method for classifying GB disease categories.
-
The non-local means (NLM) filter is initially applied to enhance image quality.
-
The Squeeze-and-Excitation Capsule Network (SE-CapsNet) is employed as the feature extractor.
-
The hybrid CNN-BiLSTM technique is finally implemented for gallbladder disease classification.
-
A comprehensive set of experiments was conducted to validate the performance of the HDLMFE-ADGDT model.
Literature survey on gallbladder disease diagnosis
Alazwari et al.11 presented an automatic GB Cancer Detection utilising an AGTO with TL (GBCD-AGTOTL) method on UI. This method analyses the US image for GB cancer occurrence using a DL approach. This method uses the Inception module to extract features from the pre-processed image. Hasan et al.12 introduced a new DL model to precisely classify GB Cancer (GBC) into normal, malignant, and benign types utilising US images. The presented method improves image quality and delineates GB wall boundaries using state-of-the-art image processing methods, such as median filtering (MF) and CLAHE. Madan et al.13 focused on the GBC detection problem from US images. That problem poses specific challenges for existing Deep Neural Network (DNN) models due to lower image quality caused by textures, noise, and viewpoint differences. Addressing those challenges requires precise localisation performance from the DNN to recognise the discriminating features for downstream malignancy estimation. The authors examine ViT-Adapter for the GBC recognition issues. It is observed that ViT-Adapter primarily relies on a primitive CNN-based spatial prior model to incorporate localisation information via cross-attention. Meng et al.14 proposed a combined framework leveraging medical features, radiomics, and DL-enabled contrast-enhanced CT imaging to predict the presence of GBC in patients. A medical radiomics and prognostic signature was generated using ML models based on the best medical features or handcrafted radiomics features.
Yin et al.15 intended to investigate whether CT-aided radiomic features of doubtful GB lesions examined through ML models can effectively distinguish benign GB disorder from GBC. Furthermore, the additional value of ML algorithms to radiological CT scan interpretation was evaluated. Basu et al.16 focused on GBC recognition employing only image-level labels. This is because of lower inter-class variance, higher intra-class variance, and lower availability of training data. Dou et al.17 planned to assess the utility of dried serum Fourier-transform infrared (FTIR) spectroscopy integrated with ML architectures for distinguishing patients. The difference between normal people and GBC serum is utilised by linear discriminant analysis (LDA) and PCA. Chang et al.18 developed a GBC LDCT image denoising system. Capability to process diverse amounts of GBC LDCT images with significant variances in artefacts and noise distribution. This research developed a noise-level approximation subnetwork as a codec structure, and the decoding part is used to estimate the noise level of GBC LDCT images. ANN achieves its objective of processing information by altering the connections between many internal nodes. Chen et al.19 proposed an Enhanced Selective Kernel Convolution (ESKConv) method integrated into the U-Net with deep supervision to adaptively capture robust tumour features. Huang et al.20 presented a k-means Mask Transformer enhanced U-Net (kMaXU) technique that incorporates Convolutional Neural Network (CNN) and Transformer features, multi-scale kMaX blocks, and cross-contrastive cluster assignment for more accurate segmentation.
Research design and methodology
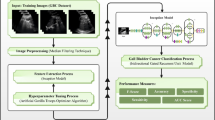
In this study, an HDLMFE-ADGDT approach is proposed. The primary purpose of the HDLMFE-ADGDT approach is to develop an effective method for classifying GB disease categories using the DL model. To accomplish this, the HDLMFE-ADGDT model has noise elimination, feature extraction, and a hybrid classification process. Figure 1 depicts the entire workflow of the HDLMFE-ADGDT model.
Entire workflow of the HDLMFE-ADGDT model.
Data processing and data collection
The performance of the HDLMFE-ADGDT technique is evaluated on the Gallbladder diseases dataset21,22. This dataset comprises 10,692 images across 9 class labels, as shown in Table 1 below. Figure 2 illustrates the sample image.
Sample images.
Gallstones: They are small cholesterol, bile salts, and calcium crystals that form in the gallbladder. The primary bile constituent is bilirubin, which is a by-product of red blood cell breakdown.
Intra-abdominal and retroperitoneal problems: These problems are anatomical areas located within the abdominopelvic cavity. It is visible on MRI or CT in an axial or transverse section. Ultrasound can detect challenging gallbladder cancers, including those with lymph node involvement.
Cholecystitis: It is a dangerous condition characterised by rapid cholecystitis. It can cause a hazardous perforation. It mostly takes place in the wounded with a parallel immune system depression, diabetes, and cardiac illnesses.
Gangrenous Cholecystitis: The GB becomes inflamed and irritated, leading to cholecystitis. Due to stones, the bile duct may become obstructed. These lead to bile trapping, which quickly traps illness-causing microorganisms.
Perforation: Once the GB wall loses its integrity, the risk of GB perforation increases among the most critical and main difficulties of the inflamed GB, particularly in Diabetics.
Polyps and Cholesterol Crystals: Lipid buildup in macrophages lining the GB contributes to this type of GB disease. Cholesterol polyps are benign, whereas endothelial polyps larger than 10 mm may progress to cancer.
Adenomyomatosis: A disease categorised by irregular mucosal epithelial hypertrophy that leads to Luschka’s crypts. This disorder has three kinds: segmental, generalised (diffuse), and fundal (localised).
Carcinoma: GB cancer is a very uncommon tumour. It rises in individuals around 70 years of age. Nearly every individual with this cancer has gallstones in the GBs. Carcinoma arises from the tissue lining the interior organs, including the GB, liver, and kidney.
Gallbladder wall thickening: It usually results from chronic GB inflammation. Once it is treated, a localised wall thickening progresses due to persistent irritation from sharp stones.
Noise reduction through NLM filter
As a first step, the HDLMFE-ADGDT technique utilises NLM-filter-based noise reduction to enhance image quality23. This is used to improve image quality, whereas the NLM filter is particularly effective at preserving fine structural details. The model excels at removing speckle and Gaussian noise, which are common in ultrasound images. NLM considers the redundancy of identical patches across images, thereby achieving superior edge and texture preservation. This is considered significant, where subtle features can highlight the existence of disease. Furthermore, blurring risk is mitigated using NLM, thereby providing cleaner input for subsequent feature extraction and classification. Thus, this technique is considered ideal over conventional denoising models due to its robustness and its ability to maintain high-frequency details.
The NLM model is also presented to improve denoising performance. It is an excellent method for image denoising that exploits the redundancy inherent in natural images. In detail, for each pixel in the denoised image, calculate the weighted average of the related patches from the complete imagery. In contrast, the weights are determined by the similarities between patches. Combining NLM in this work aims to achieve better denoising while preserving key image features.
Provided a noisy image \(\:v\), the value of denoised \(\:u\left(i\right)\) at the\(\:\:ith\) pixel was calculated as the weighted average of each pixel \(\:j\). The weight assigned to each pixel \(\:j\) is based on the similarities between the neighbourhoods of pixels \(\:i\) and \(\:j\). The NLM model is stated as shown:
Whereas \(\:u\left(i\right)\) denotes the pixel’s denoised value \(\:i,\) \(\:\varOmega\:\) denotes the group of each pixel in the images, and \(\:w(i,j)\) represents the weight, which measures the similarity between the neighbourhoods \(\:of\:pixels\:i\) and \(\:j.\) \(\:C\left(i\right)\) denotes the normalisation feature guaranteeing that the weights account for 1, and \(\:v\left(j\right)\) represents the noisy image value at \(\:the\:j\)th pixel\(\:.\).
The weight \(\:w(i,j)\) was calculated below:
Here, \(\:h\) stands for the filtering parameter, which controls the exponential function decay, and \(\:B\left(i\right)\) means:
Now, \(\:R\left(i\right)\subseteq\:\varOmega\:\) and symbolises the square area at pixel \(\:i,\) \(\:\left|R\right(i\left)\right|\left|R\right(p\left)\right|\) indicates pixel counts in the region \(\:R.\).
Feature extractor using SE-capsNet
For feature extraction, the SE-CapsNet model is employed24. DL applications in the detection field have achieved remarkable research advances; particularly, the capsule network considers relationships among features, and this model shows advantages when used on smaller datasets.
This model is considered effective as it efficiently captures inter-feature dependencies and spatial hierarchies in medical images. The technique also incorporates capsule networks that preserve the orientation, pose, and spatial relationships of crucial image structures. Furthermore, SE-CapsNet primarily extracts features via convolutional layers, unlike conventional CNNs such as ResNet or EfficientNet. The inclusion of Squeeze-and-Excitation (SE) blocks further improves the model’s ability to recalibrate feature maps. The technique also accentuates the most informative features while suppressing irrelevant ones. Also, this incorporation is specifically advantageous for ultrasound images of the gallbladder, where subtle textural discrepancies and spatial arrangements are key for accurate disease detection. The method also provides richer, more discriminative feature representations than conventional CNNs, thereby improving the reliability of downstream classification tasks. This study incorporates a channel attention mechanism, the SE block, into the CapsNet to create the SE-CapsNet, which comprises the succeeding four layers.
1) Convolutional Layer.
It is a simpler convolutional layer tailored for removing local attributes, utilising \(\:3\text{x}3\) convolution kernels with a step size of 1 and the ReLU activation function.
2) SE Layer.
These SE blocks are easier to apply; they could expand the method’s feature-removal capacity, and they are beneficial for classification tasks that involve dual processes: squeeze-and-excitation. The objective of this process is to get the global features of the provided channels. \(\:{u}_{C}\) denotes \(\:the\:Cth\) feature mapping output by the convolutional layer. Channel-wise statistics \(\:{Z}_{C}\) are obtained using global average pooling. The excitation procedure corresponds to the learning procedure for channel weights. In contrast, \(\:\sigma\:\) signifies the activation function of the sigmoid, \(\:\delta\:\) indicates the function of ReLU, and \(\:{W}_{2}\) and \(\:{W}_{1}\) are the dimensionality‐increasing and ‐reducing activities, respectively. During the excitation procedure, one may rapidly learn a non-linear interface amongst channels and, finally, obtain the learned channel weights \(\:s\). Lastly, during scaling, the learned channel weights are multiplied by the new feature mapping to produce the SE block’s attention output.
3) Primarycaps Layer.
Afterwards, the block of SE captures all feature maps using its consistent attention weights as the PrimaryCap’s input. This layer differs from a typical convolutional layer. Based on its description, this layer might later receive capsules, also called vectors, which can store additional information.
4) Digitcaps Layer.
It is applied for storing non- and Ponzi capsules. The vector symbolised the last output. The capsule network applies the squashing function. While maintaining the vector path, the resultant vector length is used as the likelihood of an object’s existence.
Whereas \(\:{W}_{ij}\) represents the weighted matrix that captures the relation between capsule \(\:j\) and capsule \(\:i\), \(\:{\widehat{u}}_{j|i}\) denotes the prediction that the \(\:ith\) low-level capsule is activated by the \(\:jth\) high-level capsule. \(\:{c}_{ij}\) denotes the coupling coefficient gained over dynamic routing. The output \(\:{v}_{j}\) was determined by the outcome of the last squashing function.
CNN-BiGRU hybrid architecture for gallbladder disease classification
At last, the hybrid CNN-BiLSTM is employed for gallbladder disease classification25. This method effectually captures both spatial and sequential dependencies. The CNN is effective at extracting local and high-level spatial features, while the BiLSTM efficiently models temporal relationships and contextual dependencies across sequential feature maps. The bidirectional processing of the BiLSTM model ensures that data from both preceding and succeeding features contribute to classification, thereby improving diagnostic accuracy. The hybrid methodology also provides a more robust and comprehensive understanding of intrinsic ultrasound patterns, resulting in enhanced gallbladder disease detection performance compared to conventional techniques. Figure 3 characterises the framework of the CNN-BiLSTM technique.
Architecture of CNN-BiLSTM.
The hybrid CNN-BiLSTM approach is developed to leverage the strengths of both structures. The CNN serves as a local feature extractor, providing a data representation of the input to the Bi-LSTM. It is mainly advantageous due to the higher data dimensionality. The Bi-LSTM sees captures longer-range dependencies within the removed features to improve classification performance. This hybrid classification contains a concatenation, a pooling, an input, a convolutional, and a Bi-LSTM layer, accompanied by dense layers with L2 regularisation, dropout, flattening, classification, and batch normalisation (BN). These layers are further discussed below:
Input layer
The presented model’s input layer gets the pre-processed data sequence. The resultant output is provided to the following layer.
Convolutional layer
To improve forecasting precision, a 1D convolutional layer is applied to remove local characteristics. It uses convolutional filters to capture local dependencies and extract features from the input data. The ReLU activation function was employed to improve the network’s learning dynamics and reduce the number of iterations required for convergence. Strides are then applied to shift the filter effectively through the input data. The convolutional layer’s output is passed to the next layer.
Pooling layer
After the previous layer, a max-pooling layer is used to remove local features. It is attained by capturing the maximal data from the identified window rate from the input feature mapping. The value characterises the further important feature inside that area. This layer’s output is then delivered to the Bi-LSTM layer.
BiLSTM layer
After utilising the CNN layer to remove spatial characteristics from the input, the Bi-LSTM layer is applied to capture longer-range dependencies and relations within the data. It includes dual LSTM cells: LSTM cells process the input order in both forward and reverse directions. It utilises a gating mechanism with three gates: a forget gate, an output gate, and an input gate. The input gate controls the sum of novel inputs restored from the cell’s layer. The forget gate manages the total of the former layer of the cell that is forgotten. The output gate leads the number of layers of the cell that output to the following time step. Provided the input order \(\:\mathcal{X}\), the LSTM element’s output at every time step (\(\:t\)) is formulated in Eqs. (10)-(15).
In Eqs. (10)-(15), \(\:i,f,0\), and \(\:c\) represent the input, forget, output gates, and cell layer, respectively, with the corresponding time variables \(\:{i}_{t},{f}_{t},{o}_{t}\), and \(\:{c}_{t}\). \(\:b,W,\text{t}\text{a}\text{n}\text{h},\sigma\:\), and \(\:\otimes\:\) symbolise the biases, weighted matrices, hyperbolic, Softmax target function, and element-wise multiplication, correspondingly. The outputs of these dual LSTM networks are then combined to produce the final output of the Bi-LSTM layer, as shown in Eqs. (16)-(18):
Whereas, \(\:{\overleftarrow{h}}_{t}\) and \(\:{\overrightarrow{h}}_{t}\) represent the hidden layer (HL) of the backward and forward Bi-LSTM. The Bi-LSTM layer’s output is then provided to the fatten layer.
Flatten layer
This layer reshapes the multi-dimensional outputs of the Bi-LSTM layer into a 1D vector.
Dense layer
It learns the designs among the diseases and their corresponding objectives, generating additional perceptual representations. To reduce model complexity and avoid overfitting, L2 regularisation is applied. This penalty corresponds to the square of the weight dimensions in the process. Provided the new loss function \(\:\left({L}_{s}\right)\), the L2 loss function \(\:\left({L}_{r}\right)\) is specified by Eq. (19):
Whereas \(\:\lambda\:\) denotes regularisation strength, \(\:x\) characterises the feature counts, and \(\:w\) symbolises feature weights.
Dropout layer
To balance the L2 regularisation method to address overfitting, a dropout layer is introduced. The dropout model prevents overfitting by randomly dropping neurons at the specified rate \(\:{p}_{r}\) during training. This is attained by setting the activations of the chosen Bi-LSTM elements to 0. The output from this layer is then provided to the next layer.
Classification Layer.
It successfully transforms the learned features into specific, proper calculations, which are evaluated alongside the true labels to assess the method’s performance. This layer’s output characterises the possibility of categorising diseases.
Model performance analysis
This article studies the performance of the HDLMFE-ADGDT model on the Gallbladder diseases dataset. The technique is simulated using Python 3.6.5 on a PC with an i5-8600k, 250GB SSD, GeForce 1050Ti 4GB, 16GB RAM, and 1 TB HDD. Parameters include a learning rate of 0.01, ReLU activation, 50 epochs, 0.5 dropout, and a batch size of 5.
80% and 20% of (a-b) confusion matrices and (c-d) curves of PR and ROC.
Figure 4 depicts the classifier results of the HDLMFE-ADGDT method on 80:20. Figure 4a and b portrays the confusion matrices with correct classification and detection of each class. Figure 4c establishes the PR investigation by identifying the maximal outcomes across all classes. At Last, Fig. 4d exemplifies the ROC inspection, yielding higher ROC values for each class.
Table 2; Fig. 5 illustrate the classifier outcomes of the HDLMFE-ADGDT approach at 80:20. The results indicate that the HDLMFE-ADGDT approach correctly identified the samples. Under 80%TRPHE, the HDLMFE-ADGDT method presents an average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:sen{s}_{y}\), \(\:spe{c}_{y}\), and \(\:{F}_{Measure}\) of 98.92%, 95.10%, 95.01%, 99.39%, and 95.05%, respectively. Simultaneously, with 20%TSPHE, the HDLMFE-ADGDT method presents an average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:sen{s}_{y}\), \(\:spe{c}_{y}\), and \(\:{F}_{Measure}\) of 99.09%, 95.83%, 95.87%, 99.49%, and 95.84%, respectively.
Average values of HDLMFE-ADGDT model under 80%:20%.
In Fig. 6, the training (TRAN) \(\:acc{u}_{y}\) and validation (VALD) \(\:acc{u}_{y}\) outcomes of the HDLMFE-ADGDT methodology at 80:20 is shown. The outcomes indicated that the TRAN and VALD \(\:acc{u}_{y}\) values show upward trends, illustrating the potential of the HDLMFE-ADGDT methodology and yielding better results across several iterations. The TRAN and VALD \(\:acc{u}_{y}\) remain close throughout the epochs, validating minimal overfitting and demonstrating the superior performance of the HDLMFE-ADGDT technique.
\(\:Acc{u}_{y}\) curve of HDLMFE-ADGDT technique at 80:20.
Figure 7 displays the TRAN and VALD loss graphs for the HDLMFE-ADGDT model under 80:20. It shows that TRAN and VALD values decrease, indicating the HDLMFE-ADGDT model’s ability to balance trade-offs between data fitting and generality. The persistent drop further confirms the superior performance of the HDLMFE-ADGDT technique and ultimately fine-tunes the predictive outcomes.
Loss curve of HDLMFE-ADGDT technique at 80:20.
Figure 8 portrays the classifier outcomes of the HDLMFE-ADGDT method at 70:30. Figure 8a and b illustrates the confusion matrices with the correct detection of 9 classes. Figure 8c shows the PR investigation, indicating a better solution for each class. Ultimately, Fig. 8d shows the ROC curves, indicating effective performance with the highest ROC values for each class.
70% and 30% of (a-b) confusion matrices and (c-d) curves of PR and ROC.
Table 3; Fig. 9 present the classifier results of the HDLMFE-ADGDT methodology at 70:30. The results indicate that the HDLMFE-ADGDT methodology correctly identified the instances. With 70%TRPHE, the HDLMFE-ADGDT technique attains an average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:sen{s}_{y}\), \(\:spe{c}_{y}\), and \(\:{F}_{Measure}\) of 98.81%, 94.54%, 94.52%, 99.33%, and 94.51%, respectively. Under 30%TSPHE, the HDLMFE-ADGDT technique attains an average \(\:acc{u}_{y}\), \(\:pre{c}_{n}\), \(\:sen{s}_{y}\), \(\:spe{c}_{y}\), and \(\:{F}_{Measure}\) of 98.68%, 94.00%, 93.96%, 99.25%, and 93.97%, respectively.
Average values of HDLMFE-ADGDT methodology under 70%:30%.
In Fig. 10, the TRAN \(\:acc{u}_{y}\) and VALD \(\:acc{u}_{y}\) results for the HDLMFE-ADGDT technique at 70:30 is displayed. The figure underscored that the TRAN and VALD \(\:acc{u}_{y}\) values exemplify emerging trends, demonstrating the HDLMFE-ADGDT technique’s outstanding performance across iterations. Furthermore, the TRAN and VALD \(\:acc{u}_{y}\) remain close across epochs, indicating minimal overfitting and displaying the enhanced performance of the HDLMFE-ADGDT method.
\(\:Acc{u}_{y}\) curve of HDLMFE-ADGDT methodology under 70:30.
In Fig. 11, the TRAN and VALD loss curves for the HDLMFE-ADGDT approach at 70:30 are shown. It indicates that the TRAN and VALD values reveal declining trends, suggesting the HDLMFE-ADGDT approach’s ability to balance trade-offs between data fitting and generality. The persistent drop, moreover, ensures the model’s advanced performance and tunes the classification outcomes.
Loss curve of HDLMFE-ADGDT methodology under 70:30.
A comparison of the HDLMFE-ADGDT approach with existing techniques is presented in Table 4; Fig. 1219,20,26,27. Under \(\:acc{u}_{y}\), the HDLMFE-ADGDT approach has a higher \(\:acc{u}_{y}\) of 99.09% while the ESKConv, U-Net, kMaXU, Visual Geometry Group 16-layer network (VGG16), InceptionV3, ResNet152, MobileNet, You Only Look Once version8 (YOLOv8), YOLOv2, and Content-Based Image Retrieval (CBIR)-based model methodologies have achieved a lower \(\:acc{u}_{y}\) of 87.09%, 85.97%, 98.85%, 97.78%, 86.50%, 85.30%, 98.35%, 82.79%, 90.80%, and 94.40%, respectively. Similarly, on \(\:pre{c}_{n}\), the HDLMFE-ADGDT approach has a higher \(\:pre{c}_{n}\) of 95.83% while the ESKConv, U-Net, kMaXU, VGG16, InceptionV3, ResNet152, MobileNet, YOLOv8, YOLOv2, and CBIR-based model methodologies have obtained a lower \(\:pre{c}_{n}\) of 90.62%, 94.65%, 90.45%, 93.51%, 89.97%, 93.86%, 89.81%, 92.52%, 92.61%, and 94.46%, correspondingly. Finally, at \(\:\:spe{c}_{y}\), the HDLMFE-ADGDT approach has obtained a higher \(\:\:spe{c}_{y}\) of 99.49% whereas the ESKConv, U-Net, kMaXU, VGG16, InceptionV3, ResNet152, MobileNet, YOLOv8, YOLOv2, and CBIR-based model methodologies have obtained a lower \(\:spe{c}_{y}\) of 94.35%, 91.89%, 98.57%, 96.13%, 93.81%, 91.11%, 98.05%, 98.92%, 98.86%, and 95.19%, correspondingly. Thus, the comparison analysis exhibits higher detection accuracy compared to other methods.
Comparative analysis of HDLMFE-ADGDT approach (a) \(\:Acc{u}_{y}\), (b) \(\:Pre{c}_{n}\), (c) \(\:Sen{s}_{y}\), and (d) \(\:Spe{c}_{y}\).
Conclusion
In this study, an HDLMFE-ADGDT approach is proposed. The aim is to develop an effective method for classifying GB disease categories using a DL model. Initially, NLM filter-based noise reduction is performed to enhance image quality. For the feature extraction process, the SE-CapsNet technique is utilised. Finally, the hybrid CNN-BiLSTM technique is employed for gallbladder disease classification. The comparison study reported an accuracy of 99.09% on the Gallbladder diseases dataset. The limitations include restricted dataset diversity, which may limit the model in generalizing effectually across diverse imaging conditions and patient populace. The model is also sensitive to discrepancies in ultrasound image quality, resulting in potential misclassifications in intrinsic cases. The real-time applicability of the technique is also affected in clinical environments. The reliability of the reported performance is also affected by the absence of large-scale validation and multi-center testing. Future work may concentrate on integrating more diverse datasets, incorporating advanced noise-robust feature learning strategies, and exploring lightweight architectures for faster inference. Furthermore, explainable AI techniques can be included to improve clinical interpretability and decision transparency.
Data availability
The data supporting the findings of this study are openly available in the repository at https://data.mendeley.com/datasets/r6h24d2d3y/1, references [21, 22]. The code used for implementing the proposed model is publicly accessible at https://github.com/zgenz1537-code/GallBladderDiseaseDet.git.
References
Yu, M. H., Kim, Y. J., Park, H. S. & Jung, S. I. Benign gallbladder diseases: Imaging techniques and tips for differentiating with malignant gallbladder diseases. World journal of gastroenterology, 26(22), p.2967. (2020).
García, P. et al. Current and new biomarkers for early detection, prognostic stratification, and management of gallbladder cancer patients. Cancers, 12(12), p.3670. (2020).
de Savornin Lohman, E. A. J. et al. The diagnostic accuracy of CT and MRI for the detection of lymph node metastases in gallbladder cancer: A systematic review and meta-analysis. Eur. J. Radiol. 110, 156–162 (2019).
John, S., Moyana, T., Shabana, W., Walsh, C. & McInnes, M. D. Gallbladder cancer: imaging appearance and pitfalls in diagnosis. Can. Assoc. Radiol. J. 71 (4), 448–458 (2020).
Takahashi, K. et al. Recent advances in endoscopic ultrasound for gallbladder disease diagnosis. Diagnostics, 14(4), p.374. (2024).
Rawla, P., Sunkara, T., Thandra, K. C. & Barsouk, A. Epidemiology of gallbladder cancer. Clin. Experimental Hepatol. 5 (2), 93–102 (2019).
Pickering, O. et al. Prevalence and sonographic detection of gallbladder polyps in a Western European population. J. Surg. Res. 250, 226–231 (2020).
Lopes Vendrami, C., Magnetta, M. J., Mittal, P. K., Moreno, C. C. & Miller, F. H. Gallbladder carcinoma and its differential diagnosis at MRI: what radiologists should know. Radiographics 41 (1), 78–95 (2021).
Kim, H. S., Cho, S. K., Kim, C. S. & Park, J. S. Big data and analysis of risk factors for gallbladder disease in the young generation of Korea. PloS One. 14 (2), e0211480 (2019).
Algamal, Z. Y., Alobaidi, N. N., Hamad, A. A., Alanaz, M. M. & Mustafa, M. Y. Neutrosophic Beta-Lindley distribution: mathematical properties and modeling bladder cancer data. Int. J. Neutrosophic Sci. 23, 186–186 (2024).
Alazwari, S. et al. Automated gall bladder cancer detection using artificial gorilla troops optimiser with transfer learning on ultrasound images. Scientific Reports, 14(1), p.21845. (2024).
Hasan, M. Z., Rony, M. A. H., Chowa, S. S., Bhuiyan, M. R. I. & Moustafa, A. A. GBCHV an advanced deep learning anatomy aware model for accurate classification of gallbladder cancer utilising ultrasound images. Scientific Reports, 15(1), p.7120. (2025).
Madan, C., Gupta, M., Basu, S., Gupta, P. & Arora, C. February. LQ-Adapter: ViT-Adapter with Learnable Queries for Gallbladder Cancer Detection from Ultrasound Images. In 2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV) (pp. 557–567). IEEE. (2025).
Meng, F. X. et al. Contrast-Enhanced CT-Based deep learning radiomics nomogram for the survival prediction in gallbladder cancer. Acad. Radiol. 31 (6), 2356–2366 (2024).
Yin, Y. et al. Optimal radiological gallbladder lesion characterisation by combining visual assessment with CT-based radiomics. Eur. Radiol. 33 (4), 2725–2734 (2023).
Basu, S., Papanai, A., Gupta, M., Gupta, P. & Arora, C. October. Gall bladder cancer detection from us images with only image level labels. In International Conference on Medical Image Computing and Computer-Assisted Intervention (pp. 206–215). Cham: Springer Nature Switzerland. (2023).
Dou, J. et al. Rapid detection of serological biomarkers in gallbladder carcinoma using fourier transform infrared spectroscopy combined with machine learning. Talanta, 259, p.124457. (2023).
Chang, Y., Wu, Q., Chi, L. & Huo, H. CT manifestations of gallbladder carcinoma based on neural network. Neural Comput. Appl. 35 (3), 2039–2044 (2023).
Chen, G. et al. ESKNet: An enhanced adaptive selection kernel convolution for ultrasound breast tumors segmentation. Expert Systems with Applications, 246, p.123265. (2024).
Huang, C., Wu, Z., Xi, H. & Zhu, J. kMaXU: Medical image segmentation U-Net with k-means Mask Transformer and contrastive cluster assignment. Pattern Recognition, 161, p.111274. (2025).
Turki, A., Obaid, A. M., Bellaaj, H., Ksantini, M. & AlTaee, A. UIdataGB: Multi-Class ultrasound images dataset for gallbladder disease detection. Data in Brief, 54, p.110426. (2024).
Taassori, M. & Vizvári, B. Enhancing Medical Image Denoising: A Hybrid Approach Incorporating Adaptive Kalman Filter and Non-Local Means with Latin Square Optimization. Electronics, 13(13), p.2640. (2024).
Bian, L., Zhang, L., Zhao, K., Wang, H. & Gong, S. Image-based scam detection model using an attention capsule network. IEEE Access. 9, 33654–33665 (2021).
Olatinwo, D., Abu-Mahfouz, A. & Myburgh, H. Mental Disorder Assessment in IoT-Enabled WBAN Systems with Dimensionality Reduction and Deep Learning. Journal of Sensor and Actuator Networks, 14(3), p.49. (2025).
Obaid, A. M. et al. Detection of gallbladder disease types using deep learning: An informative medical method. Diagnostics, 13(10), p.1744. (2023).
Bozdag, A. et al. Detection of Gallbladder Disease Types Using a Feature Engineering-Based Developed CBIR System. Diagnostics, 15(5), p.552. (2025).
Funding
This work was supported by the Institute of Information & Communications Technology Planning & Evaluation (IITP) grant funded by the Korea Government (MSIT) (No. RS-2024-00337489, Development of data drift management technology to overcome performance degradation of AI analysis models).
Author information
Authors and Affiliations
Contributions
Conceptualization, S Jayanthi; Data curation, Inderjeet Kaur, E. Laxmi Lydia and Palamakula Ramesh Babu; Formal analysis, E. Laxmi Lydia and Palamakula Ramesh Babu; Funding acquisition, Cheolhee Yoon and Gyanendra Prasad Joshi; Investigation, Jong-Chul Yoon, Cheolhee Yoon and Gyanendra Prasad Joshi; Methodology, S Jayanthi, Inderjeet Kaur, E. Laxmi Lydia and Palamakula Ramesh Babu; Project administration, Jong-Chul Yoon, Cheolhee Yoon and Gyanendra Prasad Joshi; Resources, Cheolhee Yoon; Software, Inderjeet Kaur and Palamakula Ramesh Babu; Supervision, Jong-Chul Yoon, Cheolhee Yoon and Gyanendra Prasad Joshi; Validation, Jong-Chul Yoon, Cheolhee Yoon and Gyanendra Prasad Joshi; Visualization, S Jayanthi and Inderjeet Kaur; Writing – original draft, S Jayanthi; Writing – review & editing, Gyanendra Prasad Joshi.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jayanthi, S., Kaur, I., Lydia, E.L. et al. Gallbladder disease diagnosis from ultrasound using squeeze-and-excitation capsule network with convolutional bidirectional long short-term memory. Sci Rep 16, 3147 (2026). https://doi.org/10.1038/s41598-025-32978-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-32978-9