Abstract
Serotonin (5-hydroxytryptamine, 5-HT) has central roles enabling learning and memory, particularly via serotonin receptor 2A (5-HT2AR) signaling. Drosophila Fragile X syndrome model (dfmr1 null mutant) studies reveal impaired learning and memory, which may reflect serotonergic signaling deficits. Here, we use classical olfactory T-maze conditioning to assess behavior, combined with imaging to assess 5-HT and 5-HT2AR levels within the underlying Mushroom Body (MB) brain circuitry. Null dfmr1 mutants exhibit learning and memory deficits that are corrected by elevating 5-HT signaling via 1) overexpression of the serotonin biosynthetic enzyme tryptophan hydroxylase (Trhn) or 2) knockdown of the serotonin reuptake transporter (SERT). Direct comparisons reveal both Trhn and SERT manipulations equally restore learning and memory in dfmr1 null mutants. 5-HT2AR levels in the MB circuit are reduced relative to controls in dfmr1 mutants, and 5-HT2AR RNAi phenocopies dfmr1 null behavioral deficits, suggesting these phenotypes are primarily caused by the loss of 5-HT2AR signaling. Consistently, 5-HT2AR overexpression in dfmr1 nulls restores normal learning and memory compared to controls. These findings suggest loss of 5-HT2AR signaling causes learning and memory deficits in this Fragile X syndrome model, and that rectifying this signaling impairment can restore learning and memory, providing a framework for serotonergic intervention strategies.
Similar content being viewed by others
Introduction
Fragile X syndrome (FXS) is the leading monogenetic intellectual disability (ID)1,2, caused predominantly by epigenetic loss of the Fragile X Messenger Ribonucleoprotein (FMRP)3,4. Like human FXS patients, the well-established Drosophila FXS disease model5,6 displays impairments in learning and memory7,8, with defects in underlying brain Mushroom Body (MB) learning/memory circuitry9,10 and dysfunction in multiple characterized molecular mechanisms11,12,13. This multifaceted FXS disease state may reflect truly independent FMRP functions, or could be interconnected via common foundational signaling nodes14. One intriguing possibility is that fundamental neuromodulator systems might compensate for impaired FXS learning/memory function15. Systematic brain proteomic analyses employing the Drosophila FXS model identified altered monoamine signaling pathways16 utilizing serotonin (5-hydroxytryptamine,5-HT17,18. We hypothesized transgenic manipulation of serotonergic signaling could prove an effective compensatory method for the treatment of learning and memory behavioral impairments in the Fragile X syndrome disease condition.
Serotonin/5-HT is a key regulator of mammalian learning and memory19,20,21. Likewise, Drosophila serotonergic signaling regulates learning and memory, including courtship and aversive olfactory conditioning behaviors22,23. The MB circuit underlying olfactory learning and memory24,25 receives serotonergic input and expresses conserved components of the serotonergic pathway, including 5-HT biosynthetic enzyme tryptophan hydroxylase (Trhn) and the serotonin transporter (SERT)26,27,28,29. Importantly, Trhn produces serotonin in neurons and glia in Drosophila30. Serotonin levels are upregulated in both FXS patients and disease models16,18, although this is likely compensatory as selective serotonin reuptake inhibitor (SSRI) use improves FXS symptoms31,32. The serotonergic 5-HT2A receptor (5-HT2AR) helps mediate mammalian learning and memory21,33. Drosophila 5-HT2ARs in neurons and glia function in olfactory circuitry34, but olfactory learning/memory roles have not been tested. Nothing is known about 5-HT2AR signaling in the Drosophila FXS model35, although 5-HT2AR signaling is altered in the mouse FXS model36, with 5-HT2AR proposed as a potential therapeutic target17,18. We hypothesized 5-HT2AR signaling regulates normal learning and memory, and may be impaired in the FXS condition.
Here, we employ the Drosophila FXS model (dfmr1 null mutants) to systematically investigate the contributions of serotonergic signaling to learning and memory, using classic aversive olfactory conditioning coupled to MB imaging of 5-HT and 5-HT2ARs. We designed experiments to probe serotonin signaling at three levels: serotonin synthesis (Trhn), serotonin uptake (SERT), and serotonin receptor (5-HT2AR) involvement. First, we manipulated serotonin/5-HT synthesis and reuptake using two cell-targeted transgenic methods: Trhn biosynthetic enzyme overexpression (OE) and SERT knockdown (RNAi). These complementary approaches test whether elevating serotonin alters learning and memory, and could compensate for dfmr1 null mutant deficits. Second, 5-HT2A receptors were similarly manipulated with cell-targeted 5-HT2AR OE and RNAi. By manipulating the 5-HT2AR levels, we aimed to determine whether 5-HT2A receptor signaling is necessary and sufficient for normal learning/memory behavior, or could improve outcomes in the FXS model. We used classical olfactory conditioning to evaluate the effects of these serotonergic manipulations on associative learning and memory, paired to MB confocal imaging to quantify 5-HT and 5-HT2AR levels. This combinatory approach allows us to link neuromodulatory ligand-receptor circuit signaling to cognitive behavioral output. Our goal was to determine whether targeted serotonergic interventions could restore learning and memory in the Drosophila FXS genetic disease model and to determine whether 5-HT2A receptor signaling contributes to learning and memory rescue in this FXS model.
Methods
Drosophila genetics
All stocks fed standard Drosophila food were reared at 25 °C in humidified incubators on a 12:12-h light/dark cycle. Adults of both sexes staged to 7–9 days post-eclosion (dpe) were used in all studies. All stocks were backcrossed to the w1118 background, which was used as the genetic background control. For the Fragile X syndrome (FXS) disease model, the dfmr1Δ50M excision deletion null allele6,37 was used in combination with multiple Gal4 drivers and UAS responder lines. The Gal4 driver lines were: ubiquitous daughterless UH1-Gal4 (RRID: BDSC 55850)38, neuron-specific elav-Gal4 (RRID: BDSC 8765)39, and glia-specific repo-Gal4 (RRID: BDSC 7415)40. The UAS responder lines were: a wildtype overexpression (OE) UAS-Trhn line (RRID: BDSC 27638)41, UAS-SERT RNAi (RRID: VDRC 11346)42, UAS-5HT2AROE (RRID: BDSC 4830), and UAS-5HT2AR RNAi (RRID: VDRC 31882)43. Transgenic combinations were generated using standard Drosophila genetic techniques. The control lines used were: (1) genetic background w1118 (RRID: BDSC 3605), (2) w1118,TrhnOE/ + ; UH1-Gal4/ + , (3) w1118; SERT RNAi/+; UH1-Gal4/+, (4) w1118; 5HT2AR RNAi/+; UH1-Gal4/+, and (5) w1118; dfmr1Δ50M; UH1-Gal4/ + . Experimental and control studies were always conducted in parallel for all behavioral and imaging analyses.
Behavioral analyses
Aversive olfactory conditioning was used to assess associative learning and memory, as previously described8,44. Briefly,In every trial, ~ 100 adults (7–9 days dpe) were tested for odorant conditioning behavior using a T-maze training and testing apparatus (Fig. 1A). All behavioral studies were performed in mixed sex cohorts, with the number of animals per replicate reported in Supplementary Table 1. 16 independent biological replicates were tested for each genotype and condition, with ~ 100 combined males and females per replicate (Table S1). The odorant conditioned stimulus (CS+) was paired to electric shock in a copper grid training tube, followed by exposure to the unconditioned odorant (CS⁻; Fig. 1B). Odorant solutions (9 μL in 8 mL mineral oil) were prepared fresh for each trial, balanced for equal aversiveness. In each individual training phase (12 trains of shock-training), a 60 s exposure to the CS⁺ (alternately 3-octanol [OCT] or 4-methylcyclohexanol [MCH]) was paired with electrical shock (1.5-s 80 V trains at 5-s intervals), then a 60 s air rest period, followed by another 60 s exposure to the CS⁻ odorant (opposing OCT or MCH) without electrical shock44. Immediately following training, animals were transferred within the T-maze using a central elevator (Fig. 1A,B) to test immediate odorant-shock association (defined as “learning”). Following memory training, animals were returned to food vials for a 24-h rest period and then re-introduced into the same T-maze (Fig. 1A,B) to test maintained odorant-shock association (defined as “memory”). For both trials, a 2 min period allowed choice between tubes containing CS⁺ and CS⁻ odorants. Tubes were closed, animals anesthetized using CO2, and then performance index (PI) was calculated blind to all genotypes using the formula: PI = CS- tube number—CS+ tube number/total animal number. The T-maze direction and odorants (OCT or MCH) used as CS+ versus CS- were reversed between every experiment to control for any bias.
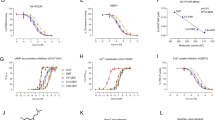
Learning and memory regulation via Mushroom Body serotonin signaling. (A) Aversive olfactory conditioning apparatus used for Pavlovian learning/memory assays. The setup includes red light illumination, humidity control, vacuum airflow odorant delivery, copper-grid shock tube with electrical stimulator (training), a vertical elevator for animal delivery, and a T-maze with two choice tubes (testing). Animals are trained with odorant cues paired to electric shock in the shock tube, and then scored in the T-maze test tubes. (B) Schematic flowchart of the learning and memory assays. The conditioning consists of alternating exposures to CS + odor paired with 12 shocks (80 V, 1.5 s duration, 5 s interval), air interval, and then the CS- odor without shock, followed by either immediate or delayed (24 h) T-maze testing. Across independent trials, OCT and MCH were assigned as CS⁺ and CS⁻ in alternative experiments. Performance index (PI = (CS- – CS+)/(CS- + CS+)) reports learning/memory abilities. (C) Quantification of learning (left) and memory (right) PIs in 5 genotypes shown: genetic background control (w1118); global UH1-Gal4 serotonin transporter (SERT) and serotonin receptor 2A (5HT2AR) RNAi; FXS disease model (dfmr1 null mutant) alone and with UH1-Gal4/ + (dfmr1 control). Data show individual trials (n = 16/genotype), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For learning (left), there is a significant effect of genotype (F(4,75) = 16.54), with no significant difference between w1118 and SERT RNAi (p = 0.9999), while 5HT2AR RNAi (p = 2.588 × 10–4) and dfmr1 null mutants (p = 1.924 × 10–6) show reduced performance, with the dfmr1 transgenic control not significantly different (p = 0.9999). For memory, there is a significant effect of genotype (F(4, 75) = 36.21), with no significant difference between w1118 and SERT RNAi (p = 0.9759), whereas 5HT2AR RNAi (p = 6.347 × 10–3) and dfmr1 null mutants (p = 3.0 × 10–11) show decreased performance. Again, dfmr1 null and dfmr1 control are not significantly different (p = 0.9999). The significance is indicated as p > 0.05 (not significant; ns), p < 0.01 (**), p < 0.001 (***), and p < 0.0001 (****). (D) Drosophila brain confocal images labeled with anti-Trio to reveal the Mushroom Body. Left: Whole-brain at 40 × magnification shows Mushroom Body (MB) and optic lobes (OL). Right: Higher magnification at 63 × showing MB α′, β′, and γ lobes. Fluorescence intensity is shown on a 16-color LUT scale. (E) Serotonergic innervation of the MB in an anatomical reconstruction from FlyWire codex. Dorsal paired medial neurons (DPM, green) provide serotonergic input onto MB Kenyon cells (KC, magenta; small subset shown for clarity).
Immunocytochemistry imaging
Mushroom Body (MB) antibody labeling and confocal imaging was done as previously described5. Briefly,Adults (7–9 dpe) were anesthetized in 70% ethanol for 1 min and then brains were dissected using sharpened forceps (Dumont #5) in 1 × phosphate-buffered saline (PBS; Invitrogen). Brains were fixed for 30 min at room temperature (RT) in 4% paraformaldehyde (PFA; EMS 15714) + 4% sucrose in PBS. The fixed brains were washed 3 × with PBS and then blocked for 1.5 h at RT with 1% bovine serum albumin (BSA; Sigma-Aldrich) + 0.2% Triton X-100 in PBS (PBS-T; Fisher Chemical). Brains were then incubated with primary antibodies + 0.2% BSA in PBS-T at 4 °C overnight. Primary antibodies used: rabbit anti-5-HT (Immunostar, 20,080 1:1,000), rabbit anti-5HT2AR (Abcam, ab140524, 1:100), and mouse anti-Trio (Developmental Studies Hybridoma Bank (DSHB), 9.4A, 1:100). Our previous studies demonstrate anti-5HT2AR antibody specificity with cell-targeted 5HT2AR knockdown and overexpression30. Brains were washed 3 × for 20 min each with PBS-T and then incubated overnight with fluorescently-conjugated secondary antibodies. The secondary antibodies used were: AlexaFluor-488 goat anti-mouse (Invitrogen, A21202, 1:250) and AlexaFluor-555 donkey anti-rabbit (Invitrogen, A31572, 1:250). Brains were washed in PBS-T 3 × for 20 min each, followed by PBS and dH2O 1 × for 20 min each. Brains were mounted in Fluoromount-G (EMS 17984-25) onto glass slides (75 × 25 mm, 0.9 to 1.06 mm; Corning) with a glass coverslip (No. 1.5H, Carl Zeiss). Double-sided adhesive tape (Scotch) was used to raise coverslips over the brains, with clear nail polish (Sally Hansen) used to seal the coverslip edges. All images were collected on a 510 META laser-scanning confocal microscope (Carl Zeiss) using either 40 × or 63 × oil-immersion objectives. All images were collected at 1024 × 1024 resolution with a Z-slice thickness of 0.75 μm. All confocal microscope imaging settings were kept absolutely constant in every experiment. All fluorescent measurement intensities were linear within the confocal range assayed. All the imaging quantifications were conducted blind to genotype and conditions, with the imaging analyses performed using the ImageJ Blind Analysis Tool.
Image quantification
Confocal image measurements were performed on Mushroom Body lobes as previously described5. Briefly,The MB lobes were labeled with anti-Trio (DSHB 9.4A) immunocytochemistry and then the MB was manually outlined to delineate the region of interest (ROI). Fluorescence intensity quantification was conducted within the outlined MB ROI for either anti-5-HT or 5-HT2AR labeling as indicated, with the mean pixel fluorescence intensity values measured using NIH ImageJ as previously described30,34. All quantification measurements were done blind to both the genotype and training conditions using the ImageJ Blind Analysis Tool plug-in. All image acquisition settings, including the laser pinhole, gain, and offset, were held constant within every experiment. All raw images are available in the Harvard Dataverse under the "Kendal Broadie Dataverse” heading.
Statistical analyses
All comparisons were performed using Prism software (GraphPad version 9). For the different experiments, one-way analysis of variance (ANOVA) (Figs. 1, 2, 3, 5, and 6) or two-way ANOVA (Fig. 4) were used to assess changes among genotypes or treatment groups, followed by Tukey’s multiple comparisons tests for multiple group comparisons. Sample sizes are specified in the figure legends. For behavioral assays, each data point represents one trial consisting of ~ 100 animals. For imaging assays, each data point represents one Mushroom Body lobe in a different brain. All the data meet normality and homogeneity of variance ANOVA requirements. Normality tests were conducted for every comparison. All datasets passed these criteria by using D’ Agostino & Pearson tests. Data are presented as mean ± standard error of the mean (SEM). All mean ± SEM numerical values for both behavioral and imaging datasets are provided in Supplementary Table 1, with exact p-values reported. Significance in figures is indicated as p < 0.05 (*), p < 0.01 (**), p < 0.001 (***), and p < 0.0001 (****). Values of p > 0.05 are reported as not significant (ns). The exact p-values for all significant comparisons are also provided in the figure legends.
Tryptophan hydroxylase overexpression rectifies FXS learning/memory. Performance index quantification of learning (A) and memory (B) across five genotypes; genetic background control (w1118; green), FXS model (dfmr1; red), and UAS-tryptophan hydroxylase (Trhn) overexpression in the dfmr1 background driven by UH1- (ubiquitous), elav- (neuronal), and repo- (glial) Gal4 lines (blue). Data show individual trials (n = 16 each), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. Learning performance shows a significant effect of genotype (F(4,75) = 17.93). For learning, a significant impairment occurs between w1118 and dfmr1 (p = 4.662 × 10–7), and a significant improvement between dfmr1 alone and with UH1-Gal4 driven UAS-Trhn (p = 3.367 × 10–9). No significant differences occur in any other comparisons. Memory performance also shows a significant effect of genotype (F(4,75) = 10.58). For memory trials, a significant impairment occurs between w1118 and dfmr1 (p = 3.902 × 10–6), and significant improvement between dfmr1 alone and with UH1-Gal4 driven UAS-Trhn (p = 2.645 × 10–6). No significant differences occur in any other comparisons. (C) Representative Mushroom Body lobe (anti-Trio, green outline) anti-serotonin (5-HT, grey scale) labeling in the same five above genotypes in untrained (top row) and trained (bottom row) conditions. Quantification of MB 5-HT fluorescence intensity from untrained (D) and T-maze trained (E) conditions. Data show individual brains (n = 10–15/condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For the untrained basal condition, the overall effect of genotype is significant (F(4, 47) = 28.50), with a significant elevation occurring between dfmr1 nulls alone versus with UH1-Gal4 driven UAS-Trhn (p = 1.477 × 10–8). No significant differences occur in any other comparisons. For the T-maze trained condition, the overall effect of genotype is also significant (F(4, 45) = 25.75), with a significant increase between dfmr1 nulls alone versus with UH1-Gal4 driven UAS-Trhn (p = 2.207 × 10–7). No significant differences occur in any other comparisons. Significance is indicated as p > 0.05 (not significant; ns) and p < 0.0001 (****).
Serotonin transporter (SERT) knockdown restores FXS learning/memory. Learning (A) and memory (B) performance index quantifications across five genotypes; genetic background control (w1118; green), FXS model (dfmr1; red), and UAS-serotonin transporter (SERT) RNAi in the dfmr1 mutant background driven by UH1- (ubiquitous), elav- (neuronal), and repo- (glial) Gal4 lines (blue). Data show individual trials (n = 16 each), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. Learning performance shows a significant effect of genotype (F(4,75) = 16.40). For learning, a significant impairment occurs between w1118 and dfmr1 (p = 8.993 × 10–6), and a significant improvement between dfmr1 alone vs with UH1-Gal4 driven SERT RNAi (p = 4.388 × 10–7). No significant differences occur in any of the other comparisons. Memory performance also shows a significant effect of genotype (F(4,75) = 17.00). For memory, a significant impairment occurs between w1118 and dfmr1 (p = 6.184 × 10–9), and significant improvement between dfmr1 alone and with UH1-Gal4 driven UAS-SERT RNAi (p = 6.644 × 10–7). No significant differences occur in any other comparisons. (C) Mushroom Body lobe outline (anti-Trio, green) and anti-serotonin (5-HT, grey scale) labeling in the same five genotypes in untrained (top row) and trained (bottom row) conditions. Quantification of the MB 5-HT fluorescence intensity in both the untrained (D) and trained (E) conditions. Data show individual brains (n = 10–15/condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For the untrained condition, the overall effect of genotype is significant (F(4, 45) = 15.62), with a significant elevation occurs between dfmr1 alone and with UH1-Gal4 driven UAS-SERT RNAi (p = 1.285 × 10–4). No significant differences occur in any of the other comparisons. For the trained condition, the overall effect of genotype is also significant (F(4, 48) = 12.19), with a significant increase likewise occurring between dfmr1 alone and with UH1-Gal4 driven UAS-SERT RNAi (p = 1.267 × 10–4). No significant differences occur in any other comparisons. Significance is indicated as p > 0.05 (not significant; ns), p < 0.001 (***), and p < 0.0001 (****).
Comparison of the Trhn overexpression and SERT knockdown effects. Quantification of performance index in learning (A) and memory (B) assays by genotype, tested with either Trhn overexpression (OE, blue) or SERT knockdown (RNAi, green). Genotypes include genetic background control (w1118), FXS model (dfmr1), and both UAS transgenes in the dfmr1 null background driven by UH1- (ubiquitous), elav- (neuronal), and repo- (glial) Gal4 lines. Data show individual trials (n = 16 per condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For learning, the two-way ANOVA reveals no significant interaction between genotype and training (F(4,150) = 0.9138), a significant effect of genotype (F(4,150) = 33.16), and no significant effect of Trhn overexpression versus SERT knockdown (F(1,150) = 4.454). Similarly, for memory, the two-way ANOVA reveals no significant interaction between genotype and training (F(4,150) = 0.7009), a significant effect of genotype (F(4,150) = 26.12), and no significant effect of Trhn overexpression versus SERT knockdown (F(1,150) = 1.093). Quantification of basal (C) and trained (D) 5-HT fluorescence intensities in the Mushroom Body across the same genotypes and conditions. Data points show results from individual brains (n = 10–15 per condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For the basal condition, the two-way ANOVA shows no significant interaction between genotype and training (F(4,92) = 9.901), a significant main effect of genotype (F(4,92) = 40.85), and no significant main effect of Trhn overexpression versus SERT RNAi (F(1,92) = 66.85). In the basal condition, significant differences in 5-HT intensity occur between Trhn OE and SERT RNAi with UH1- (p = 4.785 × 10–8), elav- (p = 2.325 × 10–5), and repo- (p = 2.528 × 10–5) Gal4 drivers. For the trained condition, the two-way ANOVA likewise shows no significant interaction (F(4,93) = 10.64), significant effect of genotype (F(4,93) = 38.50), and no significant effect of training (F(1,93) = 81.86). In the trained condition, significant differences also occur between Trhn OE and SERT RNAi with UH1- (p = 5.731 × 10–8), elav- (p = 4.934 × 10–7), and repo- (p = 2.470 × 10–6) drivers. Significance is indicated as p < 0.0001 (****).
Results
The Drosophila FXS model and 5HT 2A R knockdown share learning/memory deficits
Drosophila olfactory conditioning has been used for decades to dissect the genetic mechanisms of learning and memory8,44. Classical aversive conditioning is done in a T-maze with a conditioned stimulus (CS+) odor paired to electric shock training versus an unconditioned stimulus (CS−) odor, and then tests for the ability to discriminate between them (Fig. 1A). Testing can be done either immediately (“learning”) or after a delay (“memory”) to separate these two different behavioral phases (Fig. 1B). Performance index (PI) outcomes are quantified as (CS– − CS+)/(CS– + CS+) for both learning and memory (Fig. 1C). To begin to test links between serotonin signaling and the FXS disease model (dfmr1 null mutants), 5 genotypes were tested: the genetic background control (w1118), a serotonin transporter knockdown (SERT RNAi), a serotonin receptor 2A knockdown (5HT2AR RNAi), dfmr1 null mutants (dfmr1∆50 M), and transgenic UH1-Gal4/ + driver in this background (dfmr1 control). Compared to genetic background controls, UH1-Gal4 driven SERT RNAi has no significant effect on either the immediate learning or maintained memory outcomes (Fig. 1C), suggesting that elevated serotonin (5-HT) levels do not impact baseline behavioral performance. In contrast, 5HT2AR RNAi significantly impairs both immediate learning (p = 2.588 × 10–4) and memory maintenance (p = 6.347 × 10–3), showing that reduced 5HT2AR signaling causes learning and memory deficits (Fig. 1C). Similarly, the dfmr1 null mutants have both significantly reduced learning (p = 1.924 × 10–6) and memory (p = 3.0 × 10–11), which are unaltered with the transgenic Gal4 driver control (Fig. 1C). These findings indicate 5HT2AR and dfmr1 loss similarly impair immediate learning, and both cause strong memory deficits, suggesting a possible role of serotonergic signaling in Drosophila FXS disease model behavioral phenotypes.
The Drosophila central brain Mushroom Body (MB) is the site of learning/memory circuitry (Fig. 1D, left)45. The MB circuit is composed of approximately 2,000 intrinsic MB neurons/hemisphere called Kenyon cells (KCs)46, which extend their axons into discrete MB lobes (α/β, α′/β′, and γ; Fig. 1D, right) to form compartmentalized circuits that mediate associative learning and memory consolidation47. The KC γ neurons primarily mediate odorant associative learning48, the α/β neurons contribute to memory retrieval49,50, and the α′/β′neurons are essential for long-term memory (LTM) storage50. To study the MB circuit underlying the behavioral outcomes, staged brains were visualized with anti-Trio labeling5,51, which reveals the MB lobes in the context of the whole brain (Fig. 1D). High magnification imaging shows the α′, β′, and γ lobes that are the focus of our serotonin pathway studies (Fig. 1D, right). The recently released FlyWire codex allows complete 3D reconstruction to investigate MB serotonergic circuitry52. The serotonergic dorsal paired medial neuron (DPM, green) provides the principal MB serotonergic input (Fig. 1E. DPMs directly innervate MB Kenyon cells (magenta; only a small subset of KCs shown for clarity, providing widespread serotonergic modulation driving learning acquisition and memory consolidation27,53. The FlyWire connectome confirms extensive DPM processes overlapping the MB intrinsic circuitry, with defined synaptic inputs onto MB Kenyon cells, illuminating the spatial organization of serotonergic innervation in relation to this well-defined learning and memory circuit (Fig. 1E). Based on this behavioral and brain MB circuitry framework, we set forth to test the role of serotonergic signaling in Drosophila FXS model learning and memory impairments.
Tryptophan hydroxylase overexpression restores FXS model learning and memory
The rate-limiting serotonin biosynthetic enzyme is tryptophan hydroxylase (Trhn)54. Trhn overexpression elevates 5-HT levels and potentiates serotonergic signaling30,55. To investigate whether increasing serotonin could modulate learning/memory impairments in the Drosophila FXS disease model (dfmr1 null mutants), we overexpressed UAS-Trhn41 using three drivers: ubiquitious (UH1-Gal4), neuronal (elav-Gal4), and glial (repo-Gal4). qPCR testing Trhn overexpression (OE) demonstrates this transgenic method is highly efficacious (UH1-Gal4/w1118 vs. UH1-Gal4; UAS-Trhn ∆∆Cq 14.31 ± 0.108, p = 1.962 × 10–7). All Trhn overexpression genotypes were tested alongside Gal4 driver (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118) as well as the UAS transgene (UAS-Trhn/w1118) controls. Compared to the genetic background control (w1118), dfmr1 null mutants have a significant learning impairment (p = 4.662 × 10–7), which can be restored by UH1-Gal4 driven Trhn overexpression, significantly improving learning relative to the dfmr1 nulls (p = 3.367 × 10–9) and resulting in a performance that shows no significant difference from w1118 controls (p = 0.7789; Fig. 2A). Consistently, neuronal elav-Gal4 driven Trhn overexpression results in a comparable improvement in FXS model learning, and we were surprised that glial overexpression also proved efficacious (Fig. 2A, right). For memory compared to controls, ubiquitous Trhn overexpression in dfmr1 mutants restores performance (p = 3.902 × 10–6) and improves memory over dfmr1 nulls (p = 2.645 × 10–6), with no significant difference left to genetic background controls (p = 0.9999; Fig. 2B). As above, neuronal (elav-Gal4) and glial (repo-Gal4) Trhn overexpression in dfmr1 mutants similarly corrects memory, with no significant differences among the three drivers, albeit with a trend towards an additive effect from neuronal elav-Gal4 and glial repo-Gal4 compared to the ubiquitous UH1-Gal4 (Fig. 2B, right). The 3 Gal4 driver and UAS-Trhn controls showed no behavior differences compared to the w1118 genetic background. These findings suggest elevated serotonin signaling is sufficient to restore learning/memory in the Drosophila FXS model.
To assay serotonin signaling in the underlying Mushroom Body (MB) circuitry, all genotypes were labeled with anti-5-HT30, both under baseline, untrained conditions and following the above training protocol (Fig. 2C). Serotonin levels were imaged by delineating the MB with anti-Trio labeling5. Anti-Trio images used to delineate the MB lobes (green outlines, Fig. 2C) are shown for all genotypes and conditions (Fig. S1). The genetic background control (w1118) and FXS disease model (dfmr1 null mutants) exhibit low basal levels of serotonin, with Trhn overexpression dramatically elevating serotonin (Fig. 2C, top row). T-maze aversive olfactory conditioning training does not detectably alter serotonin in any of the genotypes (Fig. 2C, bottom row). Anti-Trio immunostaining was performed in all trials, but was not displayed for clarity; the anti-trio outline was used to define the MB quantification region. The anti-5-HT fluorescence intensity was quantified within the MB (sum of slices) to quantitatively compare serotonin levels. In untrained conditions, there is no significant difference between control and dfmr1 nulls (p = 0.9927), indicating that basal serotonin levels are not detectably affected by dfmr1 loss alone (Fig. 2D, left). Compared to the dfmr1 mutants, the Trhn overexpression results in a highly significant elevation of serotonin in the MB (UH1-Gal4 driven Trhn overexpression; p = 1.477 × 10–8), with a similar elevation from the other drivers (Fig. 2D, right). In the trained condition, results are similar, with no significant difference between w1118 and dfmr1 nulls (p = 0.9231), compared to a highly significant serotonin elevation arising from UH1-Gal4 driven Trhn overexpression (p = 2.207 × 10–7; Fig. 2E). There is a trend towards an additive effect from elav- and repo-Gal4 versus ubiquitous UH1-Gal4, but no significant differences (Fig. 2D,E). All 3 Gal4 (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118) and UAS transgene (UAS-Trhn/w1118) controls display 5-HT levels indistinguishable from the w1118 background control (Fig. S2). Taken together, these findings suggest elevating brain serotonin levels restores learning and memory to control levels within detectable limits in the Drosophila FXS disease model.
Serotonin transporter knockdown likewise corrects FXS model learning/memory
To independently test whether increasing serotonin signaling can correct learning and memory deficits in the dfmr1 null mutants—and to distinguish whether serotonin synthesis or reuptake contributes differentially to this restorative effect—we next tested whether inhibiting the serotonin transporter (SERT) could replicate the behavioral and molecular changes compared to Trhn overexpression. SERT inhibition prevents serotonin reuptake to elevate serotonin levels in the brain56. We employed a UAS-SERT RNAi42 in the dfmr1 null mutants using the same three Gal4 drivers above to transgenically control serotonin reuptake in our FXS disease model. All SERT RNAi genotypes were tested beside both Gal4 drivers alone (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118) and UAS-transgene (SERT RNAi/w1118) controls, which all performed at w1118 levels (Fig. 1C). Compared to the genetic background control (w1118), the dfmr1 null mutants again exhibit significant learning (p = 8.993 × 10–6) and memory (p = 6.184 × 10–9) impairments (Fig. 3A,B). Ubiquitous SERT knockdown in the dfmr1 null mutants (UH1-Gal4 driven UAS-SERT RNAi) significantly improves learning performance compared to the dfmr1 null mutants (p = 4.388 × 10–7), restoring behavior that is statistically indistinguishable from the genetic controls (p = 0.9998; Fig. 3A, middle). Likewise, global UH1-driven SERT RNAi significantly improves memory performance compared to the dfmr1 null mutants (p = 6.644 × 10–7), correcting memory performance back to control levels, with no statistical difference remaining compared to the w1118 genetic background animals (p = 0.9960; Fig. 3B, middle). Neuronal (elav-Gal4) and glial (repo-Gal4) SERT knockdown show similar restoration of learning and memory, with no significant differences between any of the three different Gal4 driver lines (Fig. 3A,B). These findings suggest that serotonin transporter inhibition (SERT RNAi) is sufficient to restore both learning and memory performance in the Drosophila FXS disease model.
To test whether SERT knockdown increases brain serotonin levels as expected, we performed anti-5-HT labeling in the basal untrained and T-maze trained conditions in all the genotypes (Fig. 3C). All of the control genotypes (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118, and UAS-SERT RNAi/ w1118) display 5-HT levels comparable to w1118, indicating that any observed changes are specific to SERT knockdown (Fig. S2). With MB anti-Trio labeling, serotonin levels are again very low in genetic background control (w1118) and FXS model (dfmr1 nulls) in both treatment conditions (Fig. 3C). SERT RNAi increases serotonin levels within the MB, but to a much lower extent than the previous Trhn overexpression effects (compare Figs. 2C and 3C). T-maze aversive conditioning training does not detectably alter serotonin compared to baseline in any of the genotypes (Fig. 3C, bottom row). Anti-Trio immunostaining was performed in all trials, with the outline displayed used for quantification. In the anti-Trio defined MB, the anti-5-HT fluorescence intensity was quantified across untrained and trained conditions. In the basal state, there is again no significant differences between w1118 controls and dfmr1 mutants (p = 0.8508), but serotonin levels are significantly elevated by ubiquitous SERT knockdown in dfmr1 mutants (UH1-Gal4 SERT RNAi) compared to dfmr1 alone (p = 1.285 × 10–4; Fig. 3D). There are no significant differences between the three SERT knockdown conditions, suggesting that the serotonin transporter operates in both neurons and glia to limit 5-HT levels in the MB. With T-maze training, there is still no significant difference in MB serotonin levels in the w1118 controls and dfmr1 mutants (p = 0.9839), but UH1-driven SERT RNAi again significantly increases 5-HT levels compared to the dfmr1 nulls (p = 1.267 × 10–4; Fig. 3E). There is a trend for additive effect from elav-Gal4 and repo-Gal4 towards UH1-Gal4, but no significant differences. Taken together, these findings suggest elevating serotonin by two independent methods restores learning/memory in the Drosophila FXS model.
Elevating serotonin levels via Trhn overexpression (increasing 5-HT synthesis) or SERT knockdown (reducing 5-HT reuptake) appears to produce comparable restoration of FXS model learning and memory. To weigh the relative efficacy of these two strategies, we directly compare the UAS-Trhn overexpression (blue) and UAS-SERT RNAi (green) effects on dfmr1 mutants (Fig. 4). Quantification of the learning performance index (PI) reveals that both approaches produce comparable results, with no significant differences between the genetic background controls (w1118), FXS model (dfmr1), or the transgenes driven with any of the three Gal4 drivers (Fig. 4A). Likewise, memory performance was indistinguishable in both manipulations, with no significant differences between the five genotypes with either Trhn overexpression or SERT knockdown (Fig. 4B). These findings suggest that both transgenic strategies effectively restore learning and memory in dfmr1 null mutants, and that neither method yields more benefit. However, 5-HT analyses reveal significant differences in MB serotonin elevation with the two approaches. In the basal, untrained condition, Trhn overexpression promotes significantly greater serotonin levels compared to SERT RNAi under UH1- (p = 4.785 × 10–8), elav- (p = 2.325 × 10–5), and repo- (p = 2.528 × 10–5) Gal4 driver expression (Fig. 4C). For the T-maze trained condition, Trhn overexpression continues to elevate serotonin more than the SERT RNAi counterparts between UH1- (p = 5.731 × 10–8), elav- (p = 4.934 × 10–7), and repo- (p = 2.470 × 10–6) drivers (Fig. 4D). These findings show Trhn overexpression and SERT knockdown both restore behavioral performance, but with distinct magnitudes of serotonin elevation. We next turned to testing the role of serotonin receptors within our FXS disease model.
Serotonin receptor 2A knockdown phenocopies FXS model learning and memory
The behavioral correction of dfmr1 mutants with Trhn overexpression or SERT knockdown suggests increasing serotonin signaling can ameliorate learning and memory deficits in this FXS model. However, the serotonin receptor involved in this mechanism needed to be tested. One candidate is serotonin receptor 2A (5HT2AR)57, which is strongly implicated in learning/memory function21. To test 5HT2AR requirements, we used a UAS-5HT2AR RNAi34,43 driven in the dfmr1 null mutant using the same three Gal4 drivers employed above. All 5-HT2AR RNAi genotypes were tested alongside Gal4 drivers alone (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118) as well as the UAS transgene alone (UAS-5-HT2AR RNAi/ w1118), which all performed at the w1118 level (Fig. 1C; Fig. S3). Compared to w1118 background control, dfmr1 nulls again have significantly lower learning (p = 7.460 × 10–5) and memory (p = 8.599 × 10–5) behavior performance (Fig. 5A,B). In learning, ubiquitous 5HT2AR knockdown with UH1-Gal4 only slightly, albeit significantly, increases the dfmr1 null mutant impairment (p = 0.01548) which, together with the strong 5HT2AR RNAi impact alone in the control background (p = 2.588 × 10–4; Fig. 1C), suggests a large contribution of 5HT2AR loss to the FXS model deficit (Fig. 5A). Global UH1-Gal4 driven 5HT2AR RNAi in dfmr1 nulls remains significantly impaired relative to w1118 (p = 4.6 × 10–10), suggesting a critical requirement in immediate learning. The neuronal elav-Gal4 outcome is not significantly different from UH1-Gal4 (p = 0.9093), but glial repo-Gal4 is significantly different (p = 9.559 × 10–3), showing the 5HT2AR requirement is solely in neurons (Fig. 5A). For memory, ubiquitous 5HT2AR RNAi causes no difference from dfmr1 null mutants alone (p = 0.8586), remaining significantly impaired versus w1118 controls (p = 7.094 × 10–7; Fig. 5B). There are no significant differences between or within the three Gal4 drivers. These findings suggest neuronal 5HT2AR enables learning in concert with dfmr1 function.
5HT2AR knockdown only weakly exacerbates FXS model learning/memory. Performance index quantifications of learning (A) and memory (B) across five genotypes; the genetic background control (w1118; green), FXS model (dfmr1; red), and UAS-5HT2A receptor (5HT2AR) RNAi in the dfmr1 null background driven by UH1- (ubiquitous), elav- (neuronal), and repo- (glial) Gal4 lines (blue). Data show all individual trials (n = 16 each), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. Learning performance shows a significant effect of genotype (F(4,75) = 21.02). For learning, significant differences occur between w1118 and dfmr1 (p = 7.460 × 10–5), w1118 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 null mutant (p = 4.617 × 10–10), and between this condition and repo-Gal4 driven 5HT2AR RNAi in the dfmr1 mutant (p = 9.559 × 10–3). A significant difference also occurs between dfmr1 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 mutant (p = 0.01548). No significant differences occur in the other comparisons. Memory performance also shows a significant effect of genotype (F(4,75) = 14.04). For memory, significant differences occur between w1118 and dfmr1 (p = 8.599 × 10–5), and between w1118 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 mutant (p = 7.094 × 10–7). No other significant differences occur. (C) Representative Mushroom Body lobe (anti-Trio, green outline) anti-serotonin receptor 2A (5HT2AR, grey scale) labeling in the same five genotypes in untrained (top row) and trained (bottom row) conditions. Quantification of MB 5-HT fluorescence intensity from untrained (D) and trained (E) conditions. Data show individual brains (n = 10–15 per condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For the basal untrained condition, the overall effect of genotype is significant (F(4, 53) = 21.92), with a significant difference occurs between w1118 and dfmr1 (p = 7.858 × 10–3), w1118 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 mutant (p = 7.621 × 10–9), and dfmr1 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 mutant (p = 5.732 × 10–3). No significant differences occur in the other comparisons. For the trained condition, the overall effect of genotype is significant (F(4, 52) = 21.58), with a significant 5HT2AR loss occurring between w1118 and dfmr1 null (p = 9.630 × 10–3), w1118 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 null mutant (p = 5.083 × 10–9), and dfmr1 and UH1-Gal4 driven 5HT2AR RNAi in the dfmr1 null mutant (p = 2.454 × 10–3). No other significant differences occur. Significance is indicated as p > 0.05 (not significant, ns), p < 0.05 (*), p < 0.01 (**), and p < 0.0001 (****).
To test 5HT2AR expression in the MB learning circuit, dual anti-5HT2AR and -Trio co-labeling was done in both untrained and T-maze trained conditions (Fig. 5C). Genetic background controls (w1118) exhibit high 5HT2AR puncta within and around the MB lobes compared to strongly reduced 5HT2AR puncta in the FXS disease model (Fig. 5C, left). Knockdown of 5HT2AR with all Gal4 drivers nearly abolishes detectable 5HT2AR labeling, suggesting interconnected 5HT2AR maintenance in the neurons and glia (Fig. 5C, right). T-maze aversive conditioning training does not detectably alter MB 5HT2AR levels in any of the five genotypes (Fig. 5C, bottom row). Quantification of anti-5HT2AR fluorescence intensity within the MB lobes reveals a significant decrease in the dfmr1 null mutants compared to the w1118 background controls (p = 7.858 × 10–3; Fig. 5D), suggesting the FXS model has lower 5HT2AR levels in the learning circuit. Control genotypes (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118, and UAS-5-HT2AR RNAi/w1118) show 5-HT2AR levels comparable to w1118 alone, confirming a reduction specific to dfmr1 loss. In the untrained condition, ubiquitous 5HT2AR knockdown in the dfmr1 null mutants causes significantly reduced fluorescence intensity compared to both w1118 controls (p = 7.621 × 10–9) and dfmr1 nulls (p = 5.732 × 10–3; Fig. 5D, right). No significant differences occur between the three Gal4 drivers. Following T-maze training, dfmr1 mutants continue to exhibit reduced 5HT2AR level compared to w1118 controls (p = 9.630 × 10–3), consistent with basal conditions (Fig. 5E). Likewise, 5HT2AR levels remain significantly reduced with UH1-driven 5HT2AR RNAi in dfmr1 nulls compared to dfmr1 mutants alone (p = 2.454 × 10–3) and w1118 controls (p = 5.083 × 10–9; Fig. 5E). No significant differences are detected among the Gal4 drivers. These findings show a loss of 5HT2A receptors in dfmr1 mutants, and suggest 5HT2AR overexpression should rectify Drosophila FXS disease model phenotypes.
5HT 2A R overexpression restores learning and memory in the FXS disease model
To test whether elevating serotonin receptor 2A (5HT2AR) signaling can restore learning and memory in dfmr1 nulls, we overexpressed UAS-5HT2AR43. Our hypothesis is that 5HT2AR loss in the FXS disease model results in impaired learning and memory, which should be corrected by potentiating 5-HT signaling either by elevating 5-HT levels (Figs. 2, 3, and 4) or increasing 5HT2AR expression. All 5-HT2AR overexpression genotypes were tested alongside Gal4 driver alone (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118) and UAS-transgene (UAS-5-HT2AR/ w1118) controls, which all performed at w1118 levels. Compared to genetic background controls (w1118), dfmr1 nulls once again show a very significantly impaired learning (p = 4.112 × 10–7) and memory (p = 2.133 × 10–5) performance (Fig. 6A,B). For learning, ubiquitous UAS-5HT2AR overexpression with the UH1-Gal4 driver in the dfmr1 null significantly restores memory performance relative to dfmr1 alone (p = 2.404 × 10–5), to a level not significantly different from the w1118 controls (p = 0.8517), indicating a complete behavioral correction (Fig. 6A). UH1-driven 5HT2AR overexpression results in significantly better performance than the repo-driven receptor (p = 8.799 × 10–3), suggesting a selective 5HT2AR role in neurons. Consistently, there is no significant difference between UH1- and elav-Gal4 5HT2AR overexpression (p = 0.3961), indicating neuronal 5HT2AR function (Fig. 6A). For memory performance, the UH1-driven 5HT2AR overexpression significantly improves behavioral outcomes compared to dfmr1 null mutant alone (p = 2.762 × 10–4), with the restored memory performance not significantly different from the w1118 controls (p = 0.9604), indicating a complete behavioral rectification (Fig. 6B). There is no significant difference between the three Gal4 drivers. Transgenic controls (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118, and UAS-5HT2AR/w1118) show no significant behavioral differences from w1118, confirming rescue is specific to 5-HT2AR overexpression (Fig. S3). These findings indicate 5-HT2AR overexpression can strongly restore both learning and memory performance in the Drosophila FXS disease model.
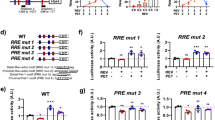
5HT2AR overexpression rectifies FXS model learning/memory performance. Learning (A) and memory (B) performance index quantifications across five genotypes; genetic background control (w1118; green), FXS model (dfmr1; red), and UAS-5HT2A receptor (5HT2AR) overexpression in the dfmr1 background driven by UH1- (ubiquitous), elav- (neuronal), and repo- (glial) Gal4 lines (blue). Data show individual trials (n = 16 each), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. Learning performance shows a significant effect of genotype (F(4,75) = 21.30). For learning, a significant impairment occurs between w1118 and dfmr1 (p = 4.112 × 10–7), improvement compared to dfmr1 with UH1-Gal4 driven UAS-5HT2AR in the dfmr1 null (p = 2.404 × 10–5), and decrease from the UH1- to repo-Gal4 conditions (p = 8.799 × 10–3). Memory performance also shows a significant effect of genotype (F(4,75) = 7.881). For memory, a significant loss again occurs between w1118 and dfmr1 (p = 2.133 × 10–5), which is restored comparing dfmr1 to UH1-Gal4 driven UAS-5HT2AR in the dfmr1 null (p = 2.762 × 10–4). No significant differences occur in the other comparisons. (C) Mushroom Body lobe (anti-Trio, green outline) anti-serotonin receptor 2A (5HT2AR, grey scale) labeling in the same five genotypes in the untrained (top row) and trained (bottom row) conditions. Quantification of the MB 5-HT2AR fluorescence intensity in the untrained (D) and trained (E) conditions. Data show individual brains (n = 10–15 per condition), mean ± SEM, and one-way ANOVA with Tukey’s multiple comparisons tests. For the untrained condition, the overall effect of genotype is significant (F(4, 51) = 16.85), with a significant decrease occurs between w1118 and dfmr1 (p = 3.495 × 10–3), w1118 and UH1-Gal4 driven UAS-5HT2AR in dfmr1 (p = 2.067 × 10–2), and between dfmr1 and UH1-Gal4 driven UAS-5HT2AR in dfmr1 (p = 4.722 × 10–8). For the trained condition, the overall effect of genotype is significant (F(4, 50) = 16.48), with a significant loss between w1118 and dfmr1 (p = 1.423 × 10–2), w1118 and UH1-Gal4 driven UAS-5HT2AR in dfmr1 (p = 6.309 × 10–3), and between dfmr1 and UH1-Gal4 driven UAS-5HT2AR in the dfmr1 null (p = 3.127 × 10–8). No significant differences occur in other comparisons. Significance is indicated as p > 0.05 (not significant, ns), p < 0.05 (*), p < 0.01 (**), p < 0.001 (***), and p < 0.0001 (****).
To test 5HT2AR levels in the MB circuit, brains from all five genotypes were double labeled for anti-Trio and anti-5HT2AR in both untrained and trained conditions (Fig. 6C). For best image clarity, only the MB outline derived from anti-Trio is displayed. Genetic background controls (w1118) show high 5HT2AR puncta compared to many fewer 5HT2AR puncta in the FXS disease model (Fig. 6C, left). 5HT2AR overexpression with all three Gal4 drivers increase the 5HT2AR levels, again suggesting interconnected 5HT2AR expression in MB neurons and glia (Fig. 6C, right). T-maze aversive conditioning training does not detectably alter 5HT2AR levels in the MB circuit of any of the tested genotypes (Fig. 6C, bottom row). Quantification of anti-5HT2AR fluorescence intensity within the MB shows a very significant loss in dfmr1 null mutants compared to the w1118 background controls (p = 3.495 × 10–3; Fig. 6D), again indicating the FXS disease model has reduced 5HT2AR levels in the MB circuit. Control genotypes (UH1-Gal4/w1118, elav-Gal4/w1118, repo-Gal4/w1118, and UAS-5HT2AR/w1118) display 5-HT2AR levels comparable to w1118, indicating the observed increase is specific to 5-HT2AR overexpression. In the basal, untrained condition, UH1-driven 5HT2AR overexpression in the dfmr1 null mutants very significantly elevates MB 5HT2AR levels compared to dfmr1 alone (p = 4.722 × 10–8) and more weakly compared to the w1118 background controls (p = 2.067 × 10–2), confirming successful overexpression (Fig. 6D). There are no significant differences between the Gal4 drivers. Following T-maze training, the same pattern is present with significantly lower MB 5HT2AR levels in dfmr1 mutants compared to matched controls (p = 1.423 × 10–2) and a very significant increase with 5HT2AR overexpression in dfmr1 null mutants, strongly compared to dfmr1 alone (p = 3.127 × 10–8) and more weakly compared to w1118 controls (p = 6.309 × 10–3; Fig. 6E). Taken together, these findings indicate that loss of 5HT2AR signaling causes learning and memory deficits in dfmr1 mutants and that 5HT2AR overexpression can correct these impairments in the Drosophila FXS disease model.
Discussion
Fragile X syndrome (FXS) results from epigenetic loss of the Fragile X Messenger Ribonucleoprotein (FMRP), an mRNA-binding protein tightly regulating the translation of proteins critical for synaptic plasticity58,59. Like FXS patients, mouse and Drosophila FXS models exhibit deficient learning/memory60,61, behaviors known to be modulated by serotonin20,62. Using classical olfactory aversive conditioning44, we find loss of FMRP and 5-HT2AR similarly impairs learning and memory (Fig. 1). Elevating serotonin via overexpression of the biosynthetic enzyme Trhn (Fig. 2) or SERT transporter knockdown (Fig. 3) restores learning and memory performance to control levels within detectable limits in the dfmr1 null mutants. These complementary approaches are equally efficacious in correcting both of the behavioral outcomes, although Trhn overexpression drives much higher serotonin levels in the Mushroom Body (MB) learning/memory circuit (Fig. 4). The Drosophila FXS model shows reduced 5-HT2AR levels, and 5-HT2AR knockdown only weakly exacerbates the dfmr1 null mutant learning impairment, without any worsening of the memory deficit (Fig. 5). Consistently, 5-HT2AR overexpression within our FXS model restores normal learning and memory in dfmr1 null mutants (Fig. 6). Thus, the Drosophila FXS disease model has strongly impaired 5-HT2AR signaling, and elevating either 5-HT or 5-HT2AR levels effectively restores Drosophila FXS disease learning and memory performance. Note all conclusions are based on associative olfactory conditioning and focus specifically on the Mushroom Body circuit, and therefore may not reflect broader FXS processes.
Null dfmr1 mutants exhibit significant deficits in classical olfactory conditioning learning and long-term memory (Fig. 1A–C). These results agree with previous studies in the Drosophila FXS model7,8, the mouse FXS model62,63, and in human FXS patients64,65. Serotonergic neuromodulation is a conserved cognitive mechanism in all three cases17,22,66. Among serotonin receptors, 5-HT2AR signaling is recognized as an important modulator of learning acquisition and memory consolidation in mammals21. However, 5-HT2AR roles in learning and memory had not been previously tested in Drosophila. Here, we find that 5-HT2AR knockdown impairs both behaviors (Fig. 1A–C). Elevating 5-HT levels by itself does not enhance performance outcomes, although serotonergic signaling is known to regulate Drosophila learning and memory22. In the Drosophila FXS model, the underlying MB circuit (Fig. 1D–E) shows defects in experience-dependent Kenyon cell structure and function5,11,67 consistent with serotonergic impairments. Fully-mapped brain 3-dimensional neural circuit reconstruction68 reveals MB-innervating serotonergic neurons (Fig. 1E) that are well positioned to control learning and memory behavior in this system. Note this study focuses on a specific brain circuit and behavior, so cannot capture the full complexity of the FXS disease condition, and that further work is needed to clarify precise molecular, cellular, and circuit serotonergic mechanisms underlying broader FXS effects.
Elevating serotonin levels—either by overexpressing the rate-limiting serotonin synthesis enzyme Trhn (Fig. 2) or by knockdown of the serotonin reuptake transporter SERT (Fig. 3)—restores FXS learning and memory performance to the control levels. Global (UH1-Gal4), neural (elav-Gal4), and glial (repo-Gal4) drivers produce comparable correction, which is surprising, but consistent with neurons and glia both using Trhn and SERT to regulate serotonin levels30,34. Similar serotonergic-based effects occur in the mouse FXS model, although direct parallels must be interpreted cautiously, as serotonin receptor distributions and circuit functions differ between models. Specifically, mice with psilocybin can ameliorate cognitive deficits in Fmr1 knockout mice62. SSRI treatments (e.g. sertraline) can likewise improve cognitive function in FXS children69, supporting a conserved role for 5-HT elevation correcting FXS-related impairments. In the Drosophila FXS model, 5-HT levels are not altered in the MB learning/memory circuit, but Trhn overexpression and SERT knockdown elevate 5-HT in dfmr1 nulls (Figs. 2, 3). Basal 5-HT levels are similar in the controls and dfmr1 nulls, but increasing 5-HT levels is sufficient to activate the reduced number of 5-HT2ARs present in the FXS model. Thus, the defect is not a loss of serotonin itself, but rather reduced 5-HT2AR abundance, and the behavioral deficits appear to arise from insufficient receptor availability rather than loss of ligand. Parallel findings in neural and glial drivers agree with Trhn and SERT in both cell types controlling serotonin levels30. Additional Drosophila glial mechanisms (e.g. arylalkylamine N-acetyltransferase 1) can also modulate 5-HT levels70. Mammalian glia also control serotonin homeostasis and signaling71,72. Our results indicate that raising 5-HT to levels very substantially above baseline via neuronal or glial mechanisms restores learning/memory behavior in the Drosophila FXS model, although whether this finding is generalizable remains to be determined.
Comparing these two strategies for increasing serotonin availability shows both restore learning and memory in the FXS model (Fig. 4A,B). This behavioral correction consistency suggests both methods converge on a shared outcome of higher 5-HT ligand levels, rather than targeting separable steps within the serotonergic regulatory pathway73. Our results argue against SERT being a principal site of FMRP action, for example, since reducing SERT does not yield different behavioral outcomes compared to Trhn overexpression. Blocking SERT using SSRIs is reportedly effective in cognitive preclinical trials74,75. Similarly, increasing serotonin via dietary tryptophan supplementation reportedly improves memory in cognitively impaired patients76,77. Elevating 5-HT has been suggested to compensate for 5-HT receptor loss to improve cognitive abilities77. Trhn overexpression is more effective than SERT knockdown in elevating serotonin levels in the MB learning/memory circuit (Fig. 4C,D). The absence of a training-induced 5-HT change indicates behavioral phenotypes do not arise from acute serotonin modulation by conditioning, but rather reflect differences in baseline serotonergic tone that determine the system capacity to support learning and memory (Fig. 4C,D). Although olfactory conditioning T-maze training has no discernible effect on 5-HT levels, neuronal and glial drivers are equally effective, with a trend toward an additive effect occurring with the ubiquitous driver. This result is surprising, but consistent with neurons and glia both using Trhn and SERT for serotonin signaling30,34. Importantly, behavioral changes from elevating 5-HT are observed only in dfmr1 mutants in our studies, indicating that these manipulations act to correct a deficit specific to the FXS model rather than producing a general enhancement of performance. These results prompted us to focus on the downstream 5-HT receptor mechanism in our FXS disease model.
5-HT2AR knockdown impairs learning and memory (Fig. 1), but in dfmr1 nulls has a minimal additional effect on learning and no significant additive effect on memory (Fig. 5A,B). The 5-HT2AR role is within neurons, with no function in glia. These results indicate FXS model behavioral impairments are mediated primarily through lost neuronal 5-HT2AR signaling. Previous Drosophila studies have shown 5-HT2AR signaling regulates locomotion78, metabolic activity79, and synapse pruning30,34, but no prior work assessed roles in learning or memory. However, mouse 5-HT2ARs are involved in synaptic plasticity in mechanisms underlying cognition80,81,82. Human 5-HT2ARs regulate cortical function linked to cognitive impairment across multiple psychiatric conditions21,83. In the Drosophila FXS model, there is a significant 5-HT2AR loss in the MB circuit (Fig. 5C–E). Just like for the 5-HT ligand, T-maze olfactory conditioning training has no discernible effect on the 5-HT2AR levels. Similarly, lack of training-dependent modulation of 5-HT2ARs supports a model of fixed baseline 5-HT2AR level, rather than dynamic regulation during conditioning, determining the efficiency of serotonergic signaling required for associative learning. Both neuronal and glial drivers similarly knockdown 5-HT2ARs, consistent with receptors being present in both cell types30,34. This suggests a regulatory maintenance mechanism within the MB circuit84. Previous studies have shown 5-HT2ARs expressed in the MB circuit53, but this work now connects 5-HT2AR signaling to learning/memory or FXS model behavioral dysfunction.
5-HT2AR overexpression in dfmr1 nulls is sufficient to restore learning and memory in our FXS disease model (Fig. 6A,B), which strongly reinforces the conclusion that serotonergic 5-HT2AR signaling corrects behavioral impairments. 5-HT2ARs act primarily in neurons, but glial signaling also contributes, again suggesting balanced circuit-level modulation of receptor function30. We previously discovered glial 5-HT2AR overexpression allows experience-dependent synaptic connectivity remodeling34, but there are no other studies of 5-HT2AR overexpression in Drosophila. However, 5-HT2AR overexpression in 5-HT2AR knockout mice medial dorsal thalamus restores associative memory, demonstrating that enhancing 5-HT2AR function can directly rescue cognitive deficits in mammals85. Pharmacological activation of 5-HT2ARs using agonists including 2,5-dimethoxy-4-iodoamphetamine (DOI), psilocybin, and LSD has been shown to promote neuroplasticity and ‘cognitive flexibility’, supporting potential applications for treating cognitive impairments86,87,88. In the Drosophila MB learning/memory circuit, 5-HT2AR levels are independently confirmed as reduced in the FXS disease model and also elevated by transgenic overexpression in the dfmr1 null mutants (Fig. 6C–E). 5-HT2AR levels are elevated in all three of the transgenic drivers, with no significant differences comparing the untrained and associatively trained animals. These findings are consistent with a 5-HT2A receptor-limited mechanism, in which the reduced 5-HT2AR level in dfmr1 null mutants constrains serotonergic signaling within the learning/memory circuit, with highly elevated 5-HT levels overcoming the receptor deficit. These abnormally high 5-HT levels could conceivably induce neurotrophic or plasticity-promoting factors to artificially drive correction of learning and memory impairments in our FXS model.
In Fmr1 knockout mice, abnormal 5-HT2A receptor function occurs in cortical circuits36, and pharmacological 5-HT2AR blockade ameliorates learning deficits89. However, serotonergic receptor distribution, downstream signaling, and circuit architectures differ substantially between mice and Drosophila. Thus, while both FXS models reveal 5-HT2AR signaling disrupted by FMRP loss, the consequences and directionality of disruption depend on species-specific circuit contexts. Thus, our results define a 5-HT2AR signaling role in the Drosophila FXS model, and do not contradict the distinct 5-HT receptor phenotypes observed in mice. Besides 5-HT2AR, both 5-HT1A and 5-HT7 receptors are expressed in the Drosophila brain90. 5-HT1AR activation has been shown to rescue mitochondrial malfunction and motor impairments in the Drosophila FXS model91, indicating that multiple 5-HT receptors are affected. Importantly, 5-HT7R signaling plays a role in olfactory associative learning, with MB 5-HT7Rs mediating cAMP-dependent learning92. Our work identifies 5-HT2AR loss in the Drosophila FXS model MB circuit, but other 5-HT receptors could participate in the 5-HT2AR-mediated effect on learning and memory. Since 5-HT ligand levels are unchanged in dfmr1 mutants, and because behavioral restoration aligns precisely with manipulations that compensate for reduced 5-HT2AR signaling, the simplest interpretation is that learning/memory impairments in this aversive conditioning task arise from insufficient 5-HT2AR-mediated signaling. Thus, although 5-HT1A and 5-HT7 receptors likely contribute to serotonergic functions in disease contexts, the combined molecular and behavioral evidence presented here identifies 5-HT2AR as the key receptor mediating FXS-related deficits in MB-dependent learning and memory.
In conclusion, this investigation demonstrates that elevated serotonergic signaling corrects learning and memory impairments in the Drosophila Fragile X syndrome model. By transgenically increasing the 5-HT ligand levels (Trhn overexpression synthesis, SERT knockdown blocking uptake), serotonin is elevated to overcome a 5-HT2AR limitation in the MB learning/memory circuit with learning and memory behavior restored in dfmr1 nulls.
These findings support a baseline-tone mechanism, with learning and memory capacity determined by sufficient 5-HT2AR levels, not dynamic signaling effects, acting to limit the signaling capacity rather than direct circuit plasticity in response to training. In future work, 5-HT2ARs could be selectively manipulated in defined serotonergic neurons (e.g. DPMs), using transgenic drivers to target the specific cells determining the serotonergic receptor-mediated signaling capacity. Importantly, 5-HT2ARs are downregulated in our FXS model, 5-HT2AR knockdown alone phenocopies dfmr1 null mutant behavioral impairments, and 5-HT2AR knockdown in the FXS disease model only slightly exacerbates learning and memory deficits. These discoveries indicate that 5-HT2AR signaling dysfunction appears causative in FXS disease model behavioral impairments. In future work, systemic use of SSRIs and 5-HT2AR agonists could be attempted to assess whether the pharmacological activation of serotonergic signaling is sufficient to ameliorate learning and/or memory deficits in dfmr1 mutants. Finally, inducing 5-HT2AR overexpression in our FXS disease model elevates receptor levels in the MB learning/memory circuit and corrects behavioral impairments in dfmr1 mutants. These findings suggest serotonergic 5-HT2AR signaling may be a useful means for FXS intervention strategies, pending future investigation in mammalian systems, and could provide a basis for future FXS therapeutic treatments.
Data availability
All original data analyzed during this study are publicly available within the Harvard Dataverse under the “Kendal Broadie Dataverse” heading. [https://dataverse.harvard.edu/dataverse/kendalbroadie].
References
Flavell, J., Franklin, C. & Nestor, P. J. A systematic review of fragile X-associated neuropsychiatric disorders. JNP 35, 110–120 (2023).
Genovese, A. C. & Butler, M. G. Systematic review: Fragile X syndrome across the lifespan with a focus on genetics, neurodevelopmental, behavioral and psychiatric associations. Genes (Basel) 16, 149 (2025).
Pieretti, M. et al. Absence of expression of the FMR-1 gene in Fragile X syndrome. Cell 66, 817–822 (1991).
Verkerk, A. J. et al. Identification of a gene (FMR-1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in Fragile X syndrome. Cell 65, 905–914 (1991).
Sears, J. C. & Broadie, K. Use-dependent, untapped dual kinase signaling localized in brain learning circuitry. J Neurosci 44, e1126232024 (2024).
Zhang, Y. Q. et al. Drosophila fragile X-related gene regulates the MAP1B homolog Futsch to control synaptic structure and function. Cell 107, 591–603 (2001).
Bolduc, F. V., Bell, K., Cox, H., Broadie, K. & Tully, T. Excess protein synthesis is Drosophila Fragile X mutants impairs long-term memory. Nat Neurosci 11(10), 1143–1145 (2008).
Jiang, H. et al. A fully automated Drosophila olfactory classical conditioning and testing system for behavioral learning and memory assessment. J. Neurosci. Methods 261, 62–74 (2016).
Doll, C. A. & Broadie, K. Activity-dependent FMRP requirements in development of the neural circuitry of learning and memory. Development 142, 1346–1356 (2015).
Pan, L., Zhang, Y. Q., Woodruff, E. & Broadie, K. The Drosophila fragile X gene negatively regulates neuronal elaboration and synaptic differentiation. Curr. Biol. 14, 1863–1870 (2004).
Golovin, R. M., Vest, J. & Broadie, K. Neuron-specific FMRP roles in experience-dependent remodeling of olfactory brain innervation during an early-life critical period. J. Neurosci. 41, 1218–1241 (2021).
Leahy, S. N., Song, C., Vita, D. J. & Broadie, K. FMRP activity and control of Csw/SHP2 translation regulate MAPK-dependent synaptic transmission. PLoS Biol 21, e3001969 (2023).
Sears, J. C., Choi, W. J. & Broadie, K. Fragile X Mental Retardation Protein positively regulates PKA anchor Rugose and PKA activity to control actin assembly in learning/memory circuitry. Neurobiol. Dis. 127, 53–64 (2019).
Davis, J. K. & Broadie, K. Multifarious functions of the fragile X mental retardation protein. Trends Genet 33, 703–714 (2017).
Cea-Del Rio, C. A. et al. Disrupted inhibitory plasticity and homeostasis in Fragile X syndrome. Neurobiol Dis 142, 104959 (2020).
Zhang, Y. Q. & Broadie, K. Fathoming Fragile X in fruit flies. Trends Genet 21, 37–45 (2005).
Ciranna, L. & Costa, L. Therapeutic effects of pharmacological modulation of serotonin brain system in human patients and animal models of fragile X syndrome. Int. J. Mol. Sci. 26, 2495 (2025).
Hanson, A. C. & Hagerman, R. J. Serotonin dysregulation in fragile X syndrome: Implications for treatment. Intractable Rare Dis. Res. 3, 110–117 (2014).
Fernandez, S. P. et al. Constitutive and acquired serotonin deficiency alters memory and hippocampal synaptic plasticity. Neuropsychopharmacology 42, 512–523 (2017).
Tortora, F. et al. The role of serotonin in fear learning and memory: A systematic review of human studies. Brain Sci. 13, 1197 (2023).
Zhang, G. & Stackman, R. W. The role of serotonin 5-HT2A receptors in memory and cognition. Front. Pharmacol. 6, 225 (2015).
Johnson, O., Becnel, J. & Nichols, C. D. Serotonin receptor activity is necessary for olfactory learning and memory in Drosophila melanogaster. Neuroscience 192, 372–381 (2011).
Pooryasin, A. & Fiala, A. Identified serotonin-releasing neurons induce behavioral quiescence and suppress mating in Drosophila. J. Neurosci. 35, 12792–12812 (2015).
de Belle, J. S. & Heisenberg, M. Associative odor learning in Drosophila abolished by chemical ablation of mushroom bodies. Science 263, 692–695 (1994).
McGuire, S. E., Le, P. T. & Davis, R. L. The role of Drosophila mushroom body signaling in olfactory memory. Science 293, 1330–1333 (2001).
Coates, K. E. et al. Identified serotonergic modulatory neurons have heterogeneous synaptic connectivity within the olfactory system of Drosophila. J. Neurosci. 37, 7318–7331 (2017).
Lee, P.-T. et al. Serotonin-mushroom body circuit modulating the formation of anesthesia-resistant memory in Drosophila. Proc. Natl. Acad. Sci. U. S. A. 108, 13794–13799 (2011).
Pech, U., Pooryasin, A., Birman, S. & Fiala, A. Localization of the contacts between Kenyon cells and aminergic neurons in the Drosophila melanogaster brain using SplitGFP reconstitution. J. Comp. Neurol. 521, 3992–4026 (2013).
Sitaraman, D., LaFerriere, H., Birman, S. & Zars, T. Serotonin is critical for rewarded olfactory short-term memory in Drosophila. J. Neurogenet. 26, 238–244 (2012).
Miller, V. K. & Broadie, K. Experience-dependent serotonergic signaling in glia regulates targeted synapse elimination. PLoS Biol. 22, e3002822 (2024).
Greiss Hess, L. et al. A randomized, double-blind, placebo-controlled trial of low-dose sertraline in young children with fragile X syndrome. J. Dev. Behav. Pediatr. 37, 619–628 (2016).
Indah Winarni, T. et al. Sertraline may improve language developmental trajectory in young children with fragile x syndrome: A retrospective chart review. Autism Res. Treat. 2012, 104317 (2012).
Zhang, G. et al. Examination of the hippocampal contribution to serotonin 5-HT2A receptor-mediated facilitation of object memory in C57BL/6J mice. Neuropharmacology 109, 332–340 (2016).
Miller, V. K. & Broadie, K. Glia-to-glia serotonin signaling directs MMP-dependent infiltration for experience-dependent synapse pruning. PLoS Biol. 23, e3003524 (2025).
Liu, X., Kumar, V., Tsai, N.-P. & Auerbach, B. D. Hyperexcitability and homeostasis in fragile X syndrome. Front. Mol. Neurosci. 14, 805929 (2021).
Xu, Z. H. et al. Deficits in LTP induction by 5-HT2A receptor antagonist in a mouse model for fragile X syndrome. PLoS ONE 7(10), e48741 (2012).
Song, C. & Broadie, K. Fragile X mental retardation protein coordinates neuron-to-glia communication for clearance of developmentally transient brain neurons. Proc. Natl. Acad. Sci. U. S. A. 120, e2216887120 (2023).
Jafar-Nejad, H., Tien, A.-C., Acar, M. & Bellen, H. J. Senseless and Daughterless confer neuronal identity to epithelial cells in the Drosophila wing margin. Development 133, 1683–1692 (2006).
Sanfilippo, P., Smibert, P., Duan, H. & Lai, E. C. Neural specificity of the RNA-binding protein Elav is achieved by post-transcriptional repression in non-neural tissues. Development 143, 4474–4485 (2016).
Halter, D. A. et al. The homeobox gene repo is required for the differentiation and maintenance of glia function in the embryonic nervous system of Drosophila melanogaster. Development 121, 317–332 (1995).
Zeng, J. et al. Local 5-HT signaling bi-directionally regulates the coincidence time window for associative learning. Neuron 111, 1118-1135.e5 (2023).
Shankar, S. et al. The neuropeptide tachykinin is essential for pheromone detection in a gustatory neural circuit. Elife 4, e06914 (2015).
Gowda, S. B. M., Banu, A., Salim, S., Peker, K. A. & Mohammad, F. Serotonin distinctly controls behavioral states in restrained and freely moving Drosophila. iScience 26, 105886 (2023).
Tully, T. & Quinn, W. G. Classical conditioning and retention in normal and mutant Drosophila melanogaster. J. Comp. Physiol. A 157, 263–277 (1985).
Heisenberg, M. What do the mushroom bodies do for the insect brain? an introduction. Learn. Mem. 5, 1–10 (1998).
Ganguly, I., Heckman, E. L., Litwin-Kumar, A., Clowney, E. J. & Behnia, R. Diversity of visual inputs to Kenyon cells of the Drosophila mushroom body. Nat. Commun. 15, 5698 (2024).
Crittenden, J. R., Skoulakis, E. M., Han, K. A., Kalderon, D. & Davis, R. L. Tripartite mushroom body architecture revealed by antigenic markers. Learn. Mem. 5, 38–51 (1998).
Hancock, C. E. et al. Visualization of learning-induced synaptic plasticity in output neurons of the Drosophila mushroom body γ-lobe. Sci. Rep. 12, 10421 (2022).
Huang, C., Wang, P., Xie, Z., Wang, L. & Zhong, Y. The differential requirement of mushroom body α/β subdivisions in long-term memory retrieval in Drosophila. Protein Cell 4, 512–519 (2013).
Yang, C.-H. et al. Additive expression of consolidated memory through Drosophila mushroom body subsets. PLoS Genet 12, e1006061 (2016).
Lai, Y.-W. et al. Hormone-controlled changes in the differentiation state of post-mitotic neurons. Curr. Biol. 32, 2341-2348.e3 (2022).
Dorkenwald, S. et al. Neuronal wiring diagram of an adult brain. Nature 634, 124–138 (2024).
Lee, W.-P. et al. Serotonin signals modulate mushroom body output neurons for sustaining water-reward long-term memory in Drosophila. Front. Cell Dev. Biol. 9, 755574 (2021).
Kuhn, D. M. & Hasegawa, H. Chapter 12 - Tryptophan hydroxylase and serotonin synthesis regulation. In Handbook of Behavioral Neuroscience (eds Müller, C. P. & Cunningham, K. A.) 239–256 (Elsevier, 2020).
Hu, S. W., Yang, Y. T., Sun, Y., Zhan, Y. P. & Zhu, Y. Serotonin signals overcome loser mentality in Drosophila. iScience 23, 101651 (2020).
Knapp, E. M. et al. Mutation of the Drosophila melanogaster serotonin transporter dSERT impacts sleep, courtship, and feeding behaviors. PLoS Genet 18, e1010289 (2022).
Kim, Y. et al. Pirenperone relieves the symptoms of fragile X syndrome in Fmr1 knockout mice. Sci. Rep. 12, 20966 (2022).
Richter, J. D. & Zhao, X. The molecular biology of FMRP: New insights into fragile X syndrome. Nat. Rev. Neurosci. 22, 209–222 (2021).
Sears, J. C. & Broadie, K. Fragile X mental retardation protein regulates activity-dependent membrane trafficking and trans-synaptic signaling mediating synaptic remodeling. Front. Mol. Neurosci. 10, 440 (2017).
Li, Y.-J. et al. Improvement of learning and memory by elevating brain D-aspartate in a mouse model of fragile X syndrome. Mol. Neurobiol. 60, 6410–6423 (2023).
Sears, J. C. & Broadie, K. FMRP-PKA activity negative feedback regulates RNA binding-dependent fibrillation in brain learning and memory circuitry. Cell Rep. 33, 108266 (2020).
Buzzelli, V. et al. Psilocybin mitigates the cognitive deficits observed in a rat model of fragile X syndrome. Psychopharmacology 240, 137–147 (2023).
Willemsen, R. & Kooy, R. F. Mouse models of fragile X-related disorders. Dis. Model. Mech. 16, dmm049485 (2023).
Bostrom, C. et al. Hippocampal dysfunction and cognitive impairment in fragile-X syndrome. Neurosci. Biobehav. Rev. 68, 563–574 (2016).
Chen, Y.-S. et al. Early 7,8-dihydroxyflavone administration ameliorates synaptic and behavioral deficits in the young FXS animal model by acting on BDNF-TrkB pathway. Mol. Neurobiol. 60, 2539–2552 (2023).
Coray, R. & Quednow, B. B. The role of serotonin in declarative memory: A systematic review of animal and human research. Neurosci. Biobehav. Rev. 139, 104729 (2022).
Doll, C. A. & Broadie, K. Neuron class-specific requirements for fragile X mental retardation protein in critical period development of calcium signaling in learning and memory circuitry. Neurobiol. Dis. 89, 76–87 (2016).
Robinson, J. W. et al. The Drosophila adult brain: Short overview of structure, function, and resources graphical review paper. Curr. Res. Insect Sci. 7, 100113 (2025).
AlOlaby, R. R. et al. Molecular biomarkers predictive of sertraline treatment response in young children with fragile X syndrome. Brain Dev. 39, 483–492 (2017).
Davla, S. et al. AANAT1 functions in astrocytes to regulate sleep homeostasis. Elife 9, e53994 (2020).
Allen, N. J. & Lyons, D. A. Glia as architects of central nervous system formation and function. Science 362, 181–185 (2018).
Lines, J., Corkrum, M., Aguilar, J. & Araque, A. The duality of astrocyte neuromodulation: Astrocytes sense neuromodulators and are neuromodulators. J. Neurochem. 169, e70054 (2025).
Jones, L. A., Sun, E. W., Martin, A. M. & Keating, D. J. The ever-changing roles of serotonin. Int. J. Biochem. Cell Biol. 125, 105776 (2020).
Tahiri, J. et al. Serotonin in depression and Alzheimer’s disease: Focus on SSRI’s beneficial effects. Ageing Res. Rev. 101, 102537 (2024).
Wang, H. et al. Efficacy of selective serotonin reuptake inhibitors-related antidepressants in Alzheimer’s disease: a meta-analysis. Eur. J. Med. Res. 29, 438 (2024).
Jenkins, T. A., Nguyen, J. C. D., Polglaze, K. E. & Bertrand, P. P. Influence of tryptophan and serotonin on mood and cognition with a possible role of the gut-brain axis. Nutrients 8, 56 (2016).
Mohajeri, M. H. et al. Chronic treatment with a tryptophan-rich protein hydrolysate improves emotional processing, mental energy levels and reaction time in middle-aged women. Br. J. Nutr. 113, 350–365 (2015).
Park, J., Kondo, S., Tanimoto, H., Kohsaka, H. & Nose, A. Data-driven analysis of motor activity implicates 5-HT2A neurons in backward locomotion of larval Drosophila. Sci. Rep. 8, 10307 (2018).
Lyu, Y. et al. Drosophila serotonin 2A receptor signaling coordinates central metabolic processes to modulate aging in response to nutrient choice. Elife 10, e59399 (2021).
Berthoux, C., Barre, A., Bockaert, J., Marin, P. & Bécamel, C. Sustained Activation of postsynaptic 5-HT2A receptors gates plasticity at prefrontal cortex synapses. Cereb Cortex 29, 1659–1669 (2019).
Desrochers, S. S. & Nautiyal, K. M. Serotonin 1B receptor effects on response inhibition are independent of inhibitory learning. Neurobiol. Learn. Mem. 187, 107574 (2022).
Grieco, S. F. et al. Psychedelics and neural plasticity: therapeutic implications. J. Neurosci. 42, 8439–8449 (2022).
Luppi, A. I. et al. A role for the serotonin 2A receptor in the expansion and functioning of human transmodal cortex. Brain 147, 56–80 (2024).
Lee, J.-E. et al. The role of glial and neuronal Eph/ephrin signaling in Drosophila mushroom body development and sleep and circadian behavior. Biochem. Biophys. Res. Commun. 720, 150072 (2024).
Barre, A. et al. Presynaptic serotonin 2A receptors modulate thalamocortical plasticity and associative learning. Proc. Natl. Acad. Sci. 113, E1382–E1391 (2016).
Bedford, P. et al. The effect of lysergic acid diethylamide (LSD) on whole-brain functional and effective connectivity. Neuropsychopharmacology 48, 1175–1183 (2023).
Meshkat, S. et al. Impact of psilocybin on cognitive function: A systematic review. Psychiatry Clin. Neurosci. 78, 744–764 (2024).
Šabanović, M. et al. Lasting dynamic effects of the psychedelic 2,5-dimethoxy-4-iodoamphetamine ((±)-DOI) on cognitive flexibility. Mol. Psychiatry 29, 1810–1823 (2024).
Lim, C. S. et al. Pharmacological rescue of Ras signaling, GluA1-dependent synaptic plasticity, and learning deficits in a fragile X model. Genes. Dev. 28(3), 273–289 (2014).
Silva, B., Goles, N. I., Varas, R. & Campusano, J. M. Serotonin receptors expressed in Drosophila mushroom bodies differentially modulate larval locomotion. PLoS ONE 9(2), e89641 (2014).
Vannelli, A., Mariano, V., Bagni, C. & Kanellopoulos, A. K. Activation of the 5-HT1A receptor by eltoprazine restores mitochondrial and motor deficits in a drosophila model of fragile X syndrome. Int. J. Mol. Sci. 25(16), 8787 (2024).
Ganguly, A., Qi, C., Bajaj, J. & Lee, D. Serotonin receptor 5-HT7 in Drosophila mushroom body neurons mediates larval appetitive olfactory learning. Sci. Rep. 10(1), 21267 (2020).
Acknowledgements
We are grateful to the Bloomington Drosophila Stock Center (BDSC; Indiana University, Bloomington, IN, USA) for essential genetic stocks, and to the Developmental Studies Hybridoma Bank (DSHB; University of Iowa, Iowa City, IA, USA) for essential antibodies. We thank Broadie Lab members for extensive and insightful input throughout this study.
Funding
This work is funded by National Institute of Health grants R01 NS131557 and NS13287 to K.B.
Author information
Authors and Affiliations
Contributions
Conceptualization: K.B., V.M.; Methodology: Y.D., V.M., A.M.; Validation: Y.D., A.M.; Formal analyses: Y.D., A.M.; Investigation: Y.D.; Data curation: Y.D.; Visualization: Y.D.; Writing—original draft: Y.D.; Writing—review and editing: K.B., Y.D.; Resources: K.B.; Supervision: V.M., K.B.; Administration: K.B.; Funding: K.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Du, Y., Miller, V.K., Mellies, A.J. et al. Elevated serotonin receptor 2A signaling restores learning and memory in a Fragile X syndrome model. Sci Rep 16, 4450 (2026). https://doi.org/10.1038/s41598-025-34492-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-34492-4