Abstract
Major depressive disorder (MDD) and myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) frequently occur together; yet their causal relationship remains unclear. To investigate the potential genetic causal link between these conditions, we conducted a two-sample Mendelian randomization (MR) analysis. Summary data from Genome-Wide Association Studies (GWAS) for MDD were sourced from the UK Biobank and the Psychiatric Genomics Consortium, while GWAS data for ME/CFS were retrieved from the UK Biobank. Inverse-variance weighting (IVW), the MR-Egger method, and weighted median, simple and weighted modes were used to perform the MR analysis. In addition, Cochrane’s Q-test was used to detect heterogeneity among the MR results. Horizontal pleiotropy was detected using the MR-Egger intercept and the MR pleiotropy residual sum and outlier (MR-PRESSO) tests. Leave-one-out analysis was performed to investigate the sensitivity of the association between MDD and ME/CFS. The results of the MR analysis revealed no causal relationship between MDD and ME/CFS. The pleiotropy test revealed that causality bias was improbable, and no evidence of heterogeneity was found among the genetic variants. Finally, the leave-one-out test confirmed the stability and robustness of our findings.
Similar content being viewed by others
Introduction
Major depressive disorder (MDD) is the most common cause of disability, affecting approximately 16% of the global population1. According to a report published by the World Health Organization, MDD is projected to be the leading cause of disability worldwide by 20302. The chronic and debilitating nature of MDD not only complicates the management of many chronic conditions but can also worsen their symptoms and associated functional impairments3.
Myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS) is a chronic condition of unknown origin that can be debilitating. It typically manifests with a diverse array of symptoms, including post-exertional malaise, chronic pain, and cognitive impairment4. Nonetheless, the etiology and pathogenesis of ME/CFS are poorly understood and are potentially attributable to a complex interplay of genetic, epidemiological, and environmental factors. Individuals with ME/CFS report a worse quality of life than those with other diseases, including cancer5. In contemporary society, a significant portion of population experiences a sub-health state of chronic fatigue, and the incidence of ME/CFS, a widespread disease that endangers human health, increases every year6. In recent years, the causal relationship between MDD and ME/CFS has been a subject of extensive investigation. For instance, a study employing various samples indicated that MDD serves as an independent risk factor in CFS samples7. Although ME/CFS and MDD frequently co-occur, their association has not been systematically studied due to potential biases, including confounding factors or reverse causality. Consequently, it remains unknown whether MDD plays a causal role in the development of ME/CFS, and there is a lack of underlying genetic evidence to support a causal relationship.
The Mendelian randomization (MR) study design adheres to Mendel’s principles of inheritance, utilizing genetic variations strongly correlated with exposure as instrumental variables (IVs) to explore the causal relationship between exposure and outcome. In this study, we applied two-sample MR summary statistics from a large-scale genome-wide association study (GWAS) of MDD and ME/CFS to investigate their potential causal relationship.
Methods
Data source
Associations between single-nucleotide polymorphisms (SNPs) and MDD were estimated using data from the largest published GWAS meta-analysis of European ancestry to date8. The meta-analysis encompassed total of 807,553 individuals (246,363 individuals with MDD and 561,190 controls of major European ancestry) from the UK Biobank Study, 23andMe, and the Psychiatric Genomics Consortium (PGC) Working Group on MDD. Pooled statistics for all assessed genetic variants in the three GWASs were publicly available from the UK Biobank and PGC only; therefore, we incorporated the comprehensive pooled statistics for both cohorts of the UK Biobank and PGC cohorts, as furnished by Howard et al. (170,756 cases, 329,443 controls), all participants in these studies belonged to European ancestry, and informed consent was obtained from each participant8.
The pooled data for ME/CFS were derived from the GWAS, and we included a study conducted by the UK Biobank. Each GWAS involved in this research has obtained ethical approval from its respective institution9. ME/CFS data were selected from this pooled data for 462,933 individuals (2,076 cases, 460,857 controls). ME/CFS was confirmed based on patient-reported diagnoses. We used SNPs as the IVs, MDD as the exposure variable, and ME/CFS as the outcome variable. Data from the ME/CFS studies can be accessed at: https://gwas.mrcieu.ac.uk/datasets/ukb-b-8961/.
Selection of IVs
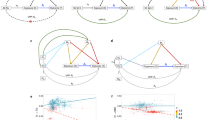
In our MR analyses, we utilized SNPs as the IVs to examine the causal relationships between exposures and outcomes. The selection of these IVs in this study adhered to stringent criteria: (1)the SNP was exhibited a robust association with MDD; (2) no significant association was observed between the SNP and ME/CFS; (3) the SNP only affected ME/CFS through MDD, with no other mediating pathways. First, SNPs significant associations to MDD were identified at a genome-wide level (P < 5 × 10[-8]). Subsequently, linkage disequilibrium parameters (clump_kb = 10,000 and clump_r2 < 0.001) were established to ensure the independence of IVs used for MDD. Lastly, the IVs associated with ME/CFS confounders were exclued, confounders were identified via the PhenoScanner V2 database, and SNPs associated with confounders such as mononucleosis were removed10. The correlation hypothesis was further validated by employing an F-statistic threshold of SNP > 10 to filter out weak instrumental variable shifts; the F-calculation method used in this study was F = R2/(1- R2)/(n-k-1/k), where R2 is the exposure variance for each IV explanation (R2 = 2 × (1-EAF) × EAF × beta2, where EAF is the effect allele frequency). The detailed methodology is depicted in Fig. 1.
Selection of instrumental variables. In this figure, SNPs exhibiting a strong association with MDD are considered as instrumental variables (IVs) (correlation assumptions). These selected SNPs can only influence the outcome through exposure (exclusive assumptions), and all IVs should not affect the outcome directly or through confounding factors (independence assumptions).
Statistical analysis
In this study, we conducted two sample MR analyses utilizing inverse-variance weighting (IVW), the MR-Egger approach, and the weighted median, simple and weighted modes to infer the genetic association between MDD and MC/CFS. IVW was combined with the MR effect estimation of each SNP to derive a composite weighted estimation of potential causal effects. The IVW approach is considered the most reliable when there is no pleiotropy present in the genetic instruments, and thus it was selected as the primary method for MR analysis Heterogeneity among SNP estimates was assessed using Cochran’s Q test and further evaluated for heterogeneity based on the symmetry of the funnel plot. To detect potential horizontal pleiotropy and examine the consistency of the associations, MR-Egger11 and Mendelian randomized pleiotropic residuals and outliers (MR-PRESSO)12 methods were applied. Finally, to confirm the dependability and robustness of the causality assessment results, a leave-one-out sensitivity analysis was conducted to evaluate the impact of removing each single SNP on the results individually. This approach enabled us to evaluate the reliability and robustness of the MR analysis results.
Ethics approval and consent to participate
Our investigation adhered to the STROBE-MR guidelines for the presentation of findings. All data included in this manuscript were sourced from published literature and publicly accessible GWAS. Each GWAS utilized in our research received ethical approval from the respective institution, which included obtaining informed consent from participants. Additionally, the data sources referenced in the paper are accessible to the public.
Results
The result of the IVs selection
Upon application of rigorous criteria to eliminate linkage disequilibrium (R2 < 0.001, with a physical window of 10,000 kb), we identified 69 SNPs that demonstrated a robust association with MDD from the exposure factor dataset. These SNPs satisfied the genome-wide significance threshold (P < 5 × 10[-8] and F > 10). Subsequent to querying the PhenoScanner database to assess their potential association with ME/CFS risk factors, we found that all 69 SNPs remained eligible for inclusion. Following a meticulous reconciliation of exposure and outcome data, a subset of 57 SNPs, which exhibited the strongest associations with MDD and had the highest levels of statistical confidence, were selected as IVs for the subsequent MR analysis. The selected IVs included SNPs with the most robust associations, for a comprehensive list of SNPs and their associated data, please refer to Supplementary Material 1.
Results of mendelian randomization analysis
Results of MR
The IVW approach is the primary method for MR, as it yields the most accurate estimates of causality. The analysis did not reveal a statistically significant association between MDD and ME/CFS. The estimated odds ratio (OR) was 0.9992, with a 95% confidence interval (CI) ranging from 0.9974 to 1.0010, and the corresponding P-value was 0.3743. Additional MR methods, including the MR-Egger regression (P = 0.1854; OR [95% CI] = 1.0036 [0.9983–1.0089]), the weighted median approach (P = 0.6289; OR [95% CI] = 1.0006 [0.9982–1.0030]), the simple mode method (P = 0.5481; OR [95% CI] = 1.0016 [0.9963–1.0070]), and the weighted mode method (P = 0.6548; OR [95% CI] = 1.0008 [0.9972–1.0045]), yielded consistent findings, furnishing no evidence for a genetic causal link between MDD and ME/CFS (refer to Fig. 2A and B for graphical representations).
Heterogeneity test
To assess the presence of heterogeneity among the selected instrumental variables, we conducted Cochrane’s Q test. The results indicated no significant heterogeneity for both the MR-Egger regression (Cochran’s Q = 62.0144, P = 0.1185) and the IVW method (Cochran’s Q = 65.7700, P = 0.0799). Furthermore, symmetry of the funnel plot (Fig. 2C) indicated the absence of publication bias or other forms of small-study effects influencing the results.
Pleiotropy test
We assessed for potential pleiotropic effects using the MR-Egger and MR-PRESSO tests. The MR-Egger intercept did not deviate significantly from zero (P = 0.3785), indicating no horizontal pleiotropy. Similarly, the MR-PRESSO test did not identify any outliers, reinforcing the robustness of our findings against violations of the MR assumption.
Sensitivity analysis
To assess the stability of our MR results, we performed a leave-one-out analysis, iteratively removing each SNP one at a time. This sensitivity analysis revealed that the exclusion of individual SNPs had a negligible impact on the overall causal estimate, with the ORs remaining largely unchanged. The stability of the results across multiple MR methods and sensitivity analyses (Fig. 2D) provides confidence in the reliability of our findings, suggesting that the absence of a causal relationship between MDD and ME/CFS is not an artifact of the analytical approach or the selection of instrumental variables.
The association between MDD and ME/CFS. (A) Forest plot of causal effects of MDD on ME/CFS. (B) Scatter plot of causal effects of MDD on ME/CFS, the slope of the line represents the causal effect of each method. (C) The funnel plot showed that the SNPs were symmetric, indicating that there was no heterogeneity in the association. (D) The leave-one-out test showed that the result was not affected by study a single influential SNP, so this association was stable.
Discussion
We utilized a two-sample MR design to rigorously assess the causal relationship between MDD and the occurrence of ME/CFS. Our analysis revealed no definitive evidence supporting a genetic predisposition to MDD as a causal factor for ME/CFS, with findings remaining consistent across various sensitivity analyses. This provides novel perspectives into understanding the relationship between them.
Previous studies have reported associations between MDD and ME/CFS. Specifically, patients with ME/CFS exhibit a higher prevalence of MDD and anxiety disorders prior to diagnosis compared to control participants, and the administration of certain antidepressant medications increasing the risk of a subsequent diagnosis of ME/CFS13. These findings indicate an association between MDD and the development of ME/CFS. However, these findings cannot rule out the possibility of confounding factors and reverse causation, making it unclear whether MDD directly causes ME/CFS. This study utilized the MR approach to effectively address the potential effects of confounding factors and reverse causation by using genetic variants associated with the MDD gene as instrumental variables. The results revealed no causal genetic relationship between MDD and ME/CFS, implying that MDD is not a direct genetic cause of ME/CFS. This finding contrasts with previous studies and highlights the advantages of the MR approach in causal inference.
Although the results of this study suggest that MDD is not a genetic predisposition to ME/CFS, there are overlapping symptoms, potential causes, and mechanisms between them. For instance, ME/CFS and MDD frequently co-occur and share symptoms including fatigue, pain, muscle tension, and flu-like discomfort14,15. Both diseases may be associated with the partial activation of immune-inflammatory pathways, particularly involving activated microglial cells, Chaves-Filho et al. demonstrated that shared microglial mechanisms underlie depression and chronic fatigue syndrome and their co-morbidities16. In addition, the hypothalamic-pituitary-adrenal (HPA) axis plays an important role in both ME/CFS and MDD. ME/CFS typically associated with low cortisol levels, whereas MDD is associated with elevated HPA axis activity and cortisol levels17. These similarities suggest that the possibility of a shared pathophysiological mechanism between MDD and ME/CFS, even though the results of the present study indicate that MDD is not a direct genetic cause of ME/CFS, and that the results using IVW, MR-Egger methods, and weighted median, simple, and weighted models are consistent with the absence of a causal relationship between MDD and ME/CFS. We propose that the reason for this result might be related to the differences in symptom attribution patterns between patients with ME/CFS and those with MDD. For instance, Johnson et al. found that symptoms in patients with chronic fatigue syndrome were more likely to be associated with somatic factors, whereas symptoms in patients with depression were more likely to be associated with psychological factors18. This highlights the necessity for ongoing research into the disparities and interconnections between MDD and ME/CFS. In the future, more attention should be paid to identifying the occurrence of environmentally determined MDD and ME/CFS.
A key strength of our study is its MR design, which is the first to utilize a large- scale GWAS to investigate the causal genetic relationship between MDD and ME/CFS, minimizing residual confounding and reverse causality. Our findings suggest that enhanced screening for ME/CFS may not be beneficial in genetically predisposed MDD cases. Although we included the latest large GWAS summary data and analyzed genetic data from large-scale sample populations, there were some limitations in this study, for instance, the inclusion of some non-cancer illness codes and self-reported cases of ME/CFS may have introduced data bias and potential inaccuracies in our estimates. Additionally, all GWAS data were derived from European populations; hence, the generalizability of our results to other global populations is limited. Further clinical trials and advancements in biological information technology are necessary to more thoroughly explore the relationship between MDD and ME/CFS.
Conclusion
Our results confirm that there is no causal relationship between MDD and ME/CFS. While our study does not support a definitive causal link between them, the questions raised in this study are indeed worth exploring in future research to enhance our understanding of these complex disorders.
Data availability
The original contributions presented in the study are included in the article/Supplementary Material, and further inquiries can be directed to the corresponding author.
References
Kessler, R. C. et al. The epidemiology of major depressive disorder: results from the National Comorbidity Survey Replication (NCS-R). JAMA 289(23), 3095–3105. https://doi.org/10.1001/jama.289.23.3095 (2003).
Mathers, C., Boerma, T. & Fat, D. M. The global Burden of Diesease:2004. update. (2008).
Krishnan, V. The molecular neurobiology of depression. Nature. 455, 894–902. https://doi.org/10.1038/nature07455 (2008).
Das, S. et al. Genetic risk factors for ME/CFS identified using combinatorial analysis. J. Transl. Med. 20(1), 598. https://doi.org/10.1186/s12967-022-03815-8 (2022).
Falk Hvidberg, M. et al. The Health-Related Quality of Life for Patients with Myalgic Encephalomyelitis / Chronic Fatigue Syndrome (ME/CFS). PloS One 10(7), e0132421. https://doi.org/10.1371/journal.pone.0132421 (2015).
Lim, E. J. et al. Systematic review and meta-analysis of the prevalence of chronic fatigue syndrome/myalgic encephalomyelitis (CFS/ME). J. Transl. Med. 18, 100. https://doi.org/10.1186/s12967-020-02269-0 (2020).
Bhui, K. S. et al. Chronic fatigue syndrome in an ethnically diverse population: the influence of psychosocial adversity and physical inactivity. BMC Med. 9, 26. https://doi.org/10.1186/1741-7015-9-26 (2011).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. vol. 22 (3), 343–352. https://doi.org/10.1038/s41593-018-0326-7 (2019).
Non-cancer illness code, self-reported: chronic fatigue syndrome. (2018). Available at: https://gwas.mrcieu.ac.uk/datasets/ukb-b-8961/ (accessed on 20 January 2023).
Palacios, N. et al. Different risk factors distinguish myalgic encephalomyelitis/chronic fatigue syndrome from severe fatigue. Sci. Rep. 13, 2469. https://doi.org/10.1038/s41598-023-29329-x (2023).
Burgess, S., Simon, G. & Thompson Interpreting findings from mendelian randomization using the MR-Egger method. Eur. J. Epidemiol.. 32 (5), 377–389. https://doi.org/10.1007/s10654-017-0255-x (2017).
Verbanck, M. et al. Detection of widespread horizontal pleiotropy in causal relationships inferred from mendelian randomization between complex traits and diseases. Nat. Genet.. 50, 693–698. https://doi.org/10.1038/s41588-018-0099-7 (2018).
Chen, C. et al. Presence of depression and anxiety with distinct patterns of pharmacological treatments before the diagnosis of chronic fatigue syndrome: a population-based study in Taiwan. J. Translational Med. vol. 21 https://doi.org/10.1186/s12967-023-03886-1 (2023).
Maes, M. An intriguing and hitherto unexplained co-occurrence: Depression and chronic fatigue syndrome are manifestations of shared inflammatory, oxidative and nitrosative (IO&NS) pathways. Progress neuro-psychopharmacology Biol. Psychiatry. 35 (3), 784–794. https://doi.org/10.1016/j.pnpbp.2010.06.023 (2011).
Johnson, S. K. et al. Depression in fatiguing illness: comparing patients with chronic fatigue syndrome, multiple sclerosis and depression. J. Affect. Disorders vol. 39 (1), 21–30. https://doi.org/10.1016/0165-0327(96)00015-8 (1996).
Chaves-Filho, M. et al. Shared microglial mechanisms underpinning depression and chronic fatigue syndrome and their comorbidities. Behav. Brain Res. 372, 111975. https://doi.org/10.1016/j.bbr.2019.111975 (2019).
Roberts, A. D. L. et al. Salivary cortisol output before and after cognitive behavioural therapy for chronic fatigue syndrome. J. Affect. Disorders. 115, 1–2. https://doi.org/10.1016/j.jad.2008.09.013 (2009).
Larkin, D. & Colin, R. M. The interface between chronic fatigue syndrome and depression: a psychobiological and neurophysiological conundrum. Clin. Neurophysiol.. 47 (2), 123–129. https://doi.org/10.1016/j.neucli.2017.01.012 (2017).
Funding
This work was supported by the Outstanding Incubation Fund of Heilongjiang University of Chinese Medicine(2019JC03), Key R&D Program of Heilongjiang Province(2022ZX06024) and Key R&D Program of Heilongjiang Province(JD2023SJ41).
Author information
Authors and Affiliations
Contributions
SWJ and ZLW designed the study. The manuscript was written by SWJ. The manuscript was revised by HXL and WMM.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, W., Hou, X., Wu, M. et al. Relationship between major depressive disorder and myalgic encephalomyelitis/chronic fatigue syndrome: a two-sample mendelian randomization study analysis. Sci Rep 15, 1155 (2025). https://doi.org/10.1038/s41598-025-85217-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85217-6