Abstract
Hydrothermal vents are ecosystems inhabited by a highly specialized fauna. To date, more than 30 gastropod species have been recorded from vent fields along the Central and Southeast Indian Ridge and all of them are assumed to be vent-endemic. During the INDEX project, 701 representatives of the genus Anatoma (Mollusca: Vetigastropoda) were sampled from six abyssal hydrothermal vent fields. Traditional morphology and COI barcoding of Hoffman et al. (Eur J Taxon 826:135–162, 2022) were combined with 2b-RAD sequencing to investigate the anatomid community structure and connectivity between the different vent fields. Consequently, 2b-RAD sequencing supported the primary species hypothesis (based on morphology) for 125 individuals of the recently described taxa A. discapex, A. declivis, A. laevapex and A. paucisculpta. We assigned 22 additional specimens to species with 2b-RAD sequencing and updated the community analyses that confirmed the pattern of expanding populations. Population structure and FST values indicated high connectivity along the six sampled vent fields for the three most abundant species. High levels of gene flow are suggested, pointing to high dispersal potential of the target species along the study area. However, low levels of heterozygosity revealed a small gene pool and therefore an increased vulnerability towards environmental change. Our results demonstrate that 2b-RAD sequencing, in combination with other molecular methods, can accurately characterise macrobenthic mollusc communities. Sequencing technology is an essential tool for ongoing monitoring. Furthermore, we highlight that the inferred molecular and ecological patterns provide valuable insights into hydrothermal vent ecosystems, which are crucial for the successful conservation of these ecosystems.
Similar content being viewed by others
Introduction
A recent agreement among members of the United Nations has resulted in the formulation of the High Seas Treaty to protect biodiversity in international waters. The objective is to ensure the protection of at least 30% of international waters by 2030 2. This new agreement demonstrates that the marine environment and its inhabitants are susceptible to adverse human impacts from commercial fishing, shipping, pollution and climate change. The deep seabed is also becoming increasingly economically interesting to the gas, oil and mineral industries. Moreover, future human activities will affect the deep-sea habitats of manganese nodules and massive sulphides at hydrothermal vent systems3,4,5. It is important to study the present fauna and its unique position in these ecosystems to define mitigation measures for detrimental industrial activities and to facilitate nature conservation.
To ensure the protection of hydrothermal ecosystems under potential human intervention, besides documenting the distribution of the local fauna, their dispersal potential and genetic connectivity must also be considered. Specific environmental conditions facilitate or hinder dispersal at hydrothermal vents, including biotic (larval longevity, feeding mode, physiology and behaviour) and abiotic factors (circulation, water column density, oxygen levels, hydrothermal plume geochemistry)6. However, the most widely recognized driver is the duration of the planktonic stage7. Although, a longer larval stage may facilitate a higher potential for dispersal6, it does not necessarily result in an elevated recruitment or survival rate of species at hydrothermal vents. For instance, prolonged larval residence in currents increases the likelihood of being transported off-axis and therefore missing the species’ intended habitat8.
Species connectivity at hydrothermal vent fields has been studied in the Indian Ocean ridge system, along the Central Indian Ridge (CIR), Southwest Indian Ridge (SWIR) and Carlsberg Ridge (CR). Studies on genetic connectivity are only lacking on the Southeast Indian Ridge (SEIR). Previous connectivity studies focusing on gastropods (Alviniconcha spp., Chrysomallon squamiferum) and decapods (Rimicaris kairei and Austinograea rodriguezensis) revealed no genetic differentiation along the CIR, suggesting a high dispersion ability of the species examined9,10,11,12. In contrast, genetic isolation is observed between the CIR and the SWIR populations11,13,14, suggesting that transform faults are the main barrier between these populations13. Recent studies have analysed populations from the northernmost part of the Indian Ridge system, the CR, and concluded that these populations are significantly different from CIR populations for Neoplepas marisindica, Chrysomallon squamiferum, Bathymodiolus septemdierum and Hesiolyra heteropoda14. They suggest that the Indian Ocean vents should be treated as three provinces for conservation purposes.
To evaluate the feasibility and impacts of deep-sea massive sulphide mining, the Indian Ocean Exploration (INDEX) project conducts biodiversity inventories and connectivity studies of hydrothermal vent ecosystems along the CIR and SEIR. As part of the INDEX project, Hoffman et al.1 identified six species of the genus Anatoma (family Anatomidae) from abyssal hydrothermal vent environments by mitochondrial DNA (COI, Cytochrome oxidase subunit I) analysis and described four new species based on morphological characteristics: Anatoma discapex, A. declivis, A. laevapex and A. paucisculpta. The remaining two species are evidently distinct yet remain undescribed since they are represented by a single individual each. The species A. paucisculpta forms a sister group to the undescribed species Anatoma sp. Lau (GenBank accession number: AB365210)15 from hydrothermal vents in the Lau Basin, Pacific Ocean1. Moreover, some specimens in the study remain unidentified due to the damage of the shells and the failure of barcoding COI, which is a consequence of the fixation and age of the samples.
In the present study, we aim to investigate the genetic connectivity of these anatomid species identified from Hoffman et al.1 across six sampled hydrothermal vent fields along the CIR and SEIR using both genetic barcode and genome-wide data. The question is of particular importance with respect to potential future deep-sea mining in the study area. We use 2b-RAD (restriction-site associated DNA with type IIB restriction endonucleases) sequencing to analyse the Anatoma species with regard to the former species delimitation and to identify additional specimens. Furthermore, we compare the genomic dataset with the previously published COI data, examine population structure and genetic diversity. In addition, we present the distribution of the genus Anatoma along the CIR and SEIR.
Materials and methods
Sampling and sample treatment
The material used in this study was sampled in the Indian Ocean along the CIR and SEIR in six hydrothermal vent areas (Fig. 1) during three cruises of the INDEX project. Samples were collected using the Canadian ROV ROPOS, mainly by rock picking or suction sampling close to the vent fields.
This study is based on the data published in Hoffman et al.1, where a total of 701 anatomids were handpicked from the samples. A subset of 169 specimens was chosen for COI barcoding, resulting in 95 high-quality sequences and six Molecular Operational Taxonomic Units (MOTUs). Subsequently, the species were morphologically studied, confirming the molecular identification, thereby resulting in the description of four new species. The analytical methodology used for COI barcoding is given in Hoffman et al.1. A comprehensive specimen list and sampling details are available in the supplementary information of this study (Supplementary Table S1).
The study of anatomids was impeded by two factors: the loss of the shell during DNA extraction and the age and fixation of the specimen material. Morphological identification of some specimens was almost exclusively based on low resolution microscope images, which introduced a considerable degree of uncertainty. In addition, differences in the way specimens were handled during the three cruises resulted in varying success rates for COI barcoding1, with 53.2% of the specimens from cruise INDEX2015 remaining without barcode. Specimens from INDEX2015 were left in the sediment for one year after fixation in 96% undenatured ethanol. They stayed at room temperature and may have been warmed by transport under tropical conditions (temperatures can reach 30–60 °C in the transport container16,17). In comparison, the samples from INDEX2018 and INDEX2019 were processed immediately on board or cooled to a temperature of at least − 20 °C until they were processed. In addition, the rolling and ethanol exchange steps recommended by Riehl et al.18 were performed. The combination of highly concentrated ethanol and a lower storage temperature is useful to reduce DNA degradation over time19,20,21. All collected specimens were stored at -20 °C.
Six sampled vent areas in the Central Indian Ocean. Map is showing the mid ocean ridges: Central, Southwest and Southeast Indian Ridge. Map by QGIS using data from BGR (Bundesanstalt für Geowissenschaften und Rohstoffe), Hannover.
DNA extraction
Of the subset of 169 specimens of Anatoma used for COI barcoding, a total of 138 specimens was analysed by means of 2b-RAD sequencing: of which 45 specimens being sampled during INDEX2015, 24 specimens during INDEX2018 and 69 specimens during INDEX2019 (Supplementary Table S1). We used the E.Z.N.A.® Mollusc DNA Kit (Omega Bio-tek, Inc., Norcross, GA, USA) to obtain high quality DNA. Specimens were photographed, and the shell opened to allow the enzymes to reach the tissue. DNA was extracted according to the manufacturer’s protocol and the entire specimen, including the shell, was used to obtain the highest possible DNA content.
DNA was measured on the Qubit Fluorometer, using the dsDNA HS (High Sensitivity) Assay Kit (Invitrogen-ThermoFisher Scientific, MA, USA). The amount of DNA required from each specimen was calculated to normalise the concentration to 150 ng DNA in 4.525 µl H2O. Therefore, the calculated amount of DNA from each sample was placed on a heat block at 60 °C for 2 h to evaporate the water. If the DNA concentration was too low to measure, the total amount was used.
2b-RAD library construction and sequencing
We prepared the 2b-RAD libraries by following the approach developed by Wang et al.22. DNA from each sample was digested by adding 0.5 µl of the enzyme BcgI (New England Biolabs, Ipswich, MA, USA), 0.6 µl of 10x NEBuffer 3.1 (New England Biolabs), 4.7125 µl of H2O and 0.1875 µl 320 µM SAM (S-adenosylmethionine; New England Biolabs) for 1 h at 37 °C and 20 min at 65 °C. The digested DNA was then ligated in a 26 µl total volume reaction consisting of 0.5 µl 10 nM ATP (New England Biolabs), 1 µl T4 DNA ligase (New England Biolabs), 2 µl 10 x T4 Buffer (New England Biolabs), 14.5 µl H2O and 1 µl Adapter R (specific adapters 2–5), 1 µl Adapter F (non-specific adapter) (Adapter information Supplementary Table S2). Finally, 6 µl of digestion product was added and the products were placed on the heat block for 2 h at 25 °C and 20 min at 65 °C.
Each amplification consisted of 8 µl DNA template, 1 µl specific index primer, 0.5 µl each primer (Pri IC1-P5 and Pri IC1-P7) (primer information Supplementary Table S2), 10 µl 2x Phusion Green Hot Start II High Fidelity PCR Master Mix (ThermoFisher Scientific, MA, USA). Cycling conditions of the 2-step PCR were 98 °C for 1 min, 98 °C for 10 s and 72 °C for 15 s (40 cycles), 72 °C for 5 min. The samples were applied to a 2% agarose gel to check the amplification and the target length of the fragments. Then 2 µl of the PCR product were pooled to a new tube (8 specimens per tube). Depending on the strength of the band from previous step, a higher amount of product was used for the pooling (up to 8 µl). To obtain the target products, bands were separated on a 4% agarose gel and further extracted using the Monarch DNA Gel Extraction Kit (New England Biolabs). As a final step, we measured the concentration of each product using the Qubit™ ds DNA Assay Kit and pooled the products for single end sequencing.
The sequenced library consisted of 138 individuals (this study, see Supplementary Table S1) and 103 individuals (another study, another two species) and was first tested on Illumina Miseq using a NanoKit (Illumina) (1 million reads) to check the quality of the runs. Final sequencing was performed on a NextSeq 500 (120 million reads) at the Carl von Ossietzky Universität Oldenburg, Oldenburg, Germany. Raw reads from the Illumina sequencing were deposited in the European Nucleotide Archive (ENA) at EMBL-EBI (accession number: PRJEB63999) https://www.ebi.ac.uk/ena/browser/view/PRJEB63999.
Genotype calling and filtering
Raw sequencing reads were processed using a custom bash script 2bRADpp downloaded from https://github.com/pmartinezarbizu/2bRADpp. The script uses bbmap for adapter trimming. Reads were oriented in forward direction, PCR duplicates were removed, and reads were demultiplexed by internal barcode.
The remaining data from 125 specimens were processed between and within species, although only the species A. declivis, A. discapex and A. laevapex had enough representatives to be analysed further. Each set of samples (among species, within A. declivis, A. discapex and A. laevapex) was analysed separately using STACKS software version 2.62 23,24. STACKS is a pipeline for building loci from short-read sequences. The algorithm of STACKS reconstructs ‘stacks’ from identical reads from each sample (-m), then either merges them with others to form a single polymorphic locus or keeps them as separate monomorphic loci depending on the number of nucleotide mismatches (-M). We applied the following parameters: -m 8 -M 2 -N 4.
Different scenarios were calculated by POPULATIONS program (within STACKS software version 2.62) to check the influence of the estimated number of populations, as well as the minimum percentage of a read within a population. The two parameters p (minimum number of populations a locus must be present in to process a locus) and r (minimum percentage of individuals in a population required to process a locus for that population) were used for this purpose. The calculated scenarios are: p = 1 and r = 0.1, p = 1 and r = 0.7, p = 2 and r = 0.1, p = 2 and r = 0.7, p = 3 and r = 0.1, p = 3 and r = 0.7. For the between-species analysis, the best approach was to apply parameters for p = 1, r = 0.1 (DS_INMAC_RAD01), and for within-species analysis the more restrictive approach of p = 2, r = 0.1 was used to observe potential differences under strict adjustment (DS_INMAC_RAD02-04). The resulting loci and variant sites of all scenarios can be checked in the supplementary information (Supplementary Table S3).
Analysis of population structure
The scenarios calculated by POPULATIONS program within STACKS (DS_INMAC_RAD01-04) were further analysed by using the STRUCTURE 2.3.4 25 program. STRUCTURE analyses differences in the distribution of genetic variants amongst populations with a Bayesian iterative algorithm by placing samples into groups (or clusters) whose members share similar patterns of variation. The following parameters were used in the analyses: “admixture model” (assuming that each individual has ancestry from one or more of K genetically distinct sources), correlated allele frequencies, and a burn-in period of 100,000 iterations and 200,000 sampling iterations (following the default settings of Pritchard et al.26 ). Analyses were repeated three times for each cluster (k) with a range of 1 to 10 between species and 1 to 5 within species. In addition, the online tool CLUMPAK (Clustering Markov Packager Across K)27 was employed for the visualisation of the STRUCTURE plots.
We applied two approaches to calculate the most probable number of clusters K (based on DS_INMAC_RAD01-04, files produced with POPULATIONS): Evanno’s method28 using STRUCTURE HARVESTER software version 0.6.94 29 and DAPC (Discriminant Analysis of Principal Components) using the package adegenet version 2.1.5 30,31 in R version 4.2.232 within R STUDIO 2022.12.0 33.
STRUCTURE HARVESTER is a web-based program designed to collate results generated by the program STRUCTURE. It offers a rapid method of assessing and visualising likelihood values across a range of K values and hundreds of iterations, thereby facilitating the identification of the optimal number of genetic groups that align with the dataset. Furthermore, STRUCTURE HARVESTER is capable of reformatting data for utilisation in downstream programs, such as CLUMPAK29.
In contrast, the multivariate statistical approach of DAPC partitions the variance in the sample into a between-group and within-group component, thereby optimising discrimination between groups. The genetic data was initially transformed using a PCA (principal component analysis), and clusters are subsequently identified by a DA (discriminant analysis). The DAPC analysis was performed as suggested by Miller et al.34 using two different attempts. For the de novo approach, we used the find.clusters function of DAPC to infer the most likely number of clusters. The optimal number of PCs retained was N/3; where N = number of samples as recommended in the manual. The BIC (Bayesian Information Criterion) was calculated and the optimal number of populations with the lowest BIC value was identified. As a second approach, an a priori DAPC analysis was performed using the expected number of clusters (either based on the expected number of species and/or the previously calculated cluster K by Evanno’s method).
Results of the STACKS pipeline (structure files) were used to test hypotheses for analyses of molecular variance (AMOVA). The default method (ade4) was selected to perform an AMOVA by the package poppr35 version 2.9.3 in R based on 999 iterations of the three species dataset (DS_INMAC_RAD02, DS_INMAC_RAD03, DS_INMAC_RAD04) to test whether genetic variation was greater (1) between the populations of each vent field, (2) between the samples within one population, or (3) within individuals. For the within individual variance, poppr splits genotypes into haplotypes. Default settings were used.
In addition, the program STRUCTURE calculates the inferred ancestry. Based on this inferred ancestry matrix (of the dataset DS_INMAC_RAD01) we performed an NMDS (Nonmetric Multidimensional Scaling) by using the metaMDS function in the vegan36 package version 2.6.2 in R.
The obtained datasets (DS_INMAC_RAD01-04) can be accessed through the Senckenberg Metadata Portal https://dataportal.senckenberg.de/dataset/318754d6-e802-4cb2-a8e5-7f3a4d68af0d.
Species delimitation
We applied a Bayes Factor Delimitation (*with genomic data; BFD*)37 to differentiate species by testing alternative hypotheses of species boundaries based on 2b-RADseq data. The hypotheses were tested against the base scenario (a), which is the current taxonomy as proposed by Hoffman et al.1. The following alternative species delimitation models were employed: b) (A. discapex) (A. paucisculpta) (Anatoma sp. 1 DZMB_2021_0095) (A. declivis and A. laevapex), based on the observation of similarities in their shells, anterior soft parts and radulae1 and c) (A. laevapex) (A. paucisculpta) (Anatoma sp. 1 DZMB_2021_0095) (A. declivis and A. discapex), are similar, as evidenced by their genetic similarity according to the mitochondrial COI data. To ascertain whether the result would differ if fewer sites were excluded, the analysis was repeated with the exclusion of species represented by a limited number of specimens (A. paucisculpta and Anatoma sp. 1 DZMB_2021_0095).
The data (DS_INMAC_RAD01) produced by POPULATIONS program within STACKS (VCF file) were initially transformed into an XML file by the BEAUti tool, a component of the BEAST 2.6.738 software package. The mutation rates were set to u = 1 and v = 1, and the coalescence rate (population size parameter with one value for each node in the tree) was sampled. This was done while u represented the instantaneous rate of mutating from the ‘0’ allele to the ‘1’ allele and v represented the instantaneous rate of mutating from the ‘1’ allele to the ‘0’ allele. In case of SNP data where the ‘0’ and ‘1’ alleles are arbitrarily assigned from the data, uncoupling these rates is typically not a useful approach39. A Γ-distributed prior was employed for the θ parameter (α = 2 and β = 200). To accommodate for uncertainty in the λ parameter (λ refers to the speciation rate in the Yule model), a Γ-distributed hyperprior was applied to this parameter.
Once we established the base scenario in an XML file, we had to edit the file manually for each of the four hypotheses in order to enable the implementation of a path sampling (or stepping stone) analysis. Subsequently, path sampling was performed by utilising the model-selection package, version 1.5.3 in BEAST 2.6.738. A total of 48 steps (1,000 MCMC steps, 0 pre-burnin steps) were employed to estimate marginal likelihoods and species trees for each of the four hypotheses (a, b, c and d). The hypotheses were then ranked in accordance with their estimation of marginal likelihood, and Bayes factors (BF) were calculated to identify the optimal hypothesis for species delimitation, as BF = 2 (ln L1 – L0). L0 and L1 represent the estimated marginal likelihoods of the two models under comparison. To assess the significance of the BF, the estimates were employed in compliance with the methodology proposed by Kass and Raftery40: 0 < BF < 6 is positive evidence, 6 < BF < 10 is strong support and BF > 10 is decisive.
Furthermore, the XML file was analysed with the SNAPP (SNP and AFLP Package for Phylogenetic analysis) plugin version 1.5.6 implemented in the software BEAST 2 38 for estimating species trees41.
Population genetic metrics
For the three species datasets (DS_INMAC_RAD02-04), the POPULATIONS software in the STACKS pipeline was used to obtain the total number of alleles, number of variant loci (variations of a locus), number of private alleles (found only once in a set of populations), observed heterozygosity (HO, observed differing alleles within one gene of one individual), expected heterozygosity (HE) under Hardy-Weinberg equilibrium, nucleotide diversity (π), fixation index (FIS ) and population differentiation (FST). In addition, isolation-by-distance (Mantel test) based on PhiST (from an AMOVA) was calculated by GENODIVE version 3.06 42.
The genomic datasets were compared with the published mitochondrial COI data (95 sequences) of Hoffman et al.1, accessible via the following link: http://www.https://doi.org/10.5883/DS-INMAC03. The COI data was analysed by using the DnaSP6 43,44 software by estimating parameters for populations with sample size of n ≥ 4 including: gene (u), haplotype (h) and nucleotide (π) diversities45, Fu’s Fs46 and Tajima’s D45.
We estimated minimum spanning networks47 to visualise the relationships among the six sampled species based on the published COI dataset by PopART (Population Analysis with Reticulate Trees) (http://popart.otago.ac.nz). We included all sequences from reference libraries in the haplotype network: Anatoma euglypta (GenBank accession number: AY923934) from the Antarctic Basin48, A. pseudoequatoria (MW278816) from the western Pacific Basin, Anatoma sp. Lau (AB365210) from the Pacific Lau Basin and Anatoma sp. Izu (AB365211) from the northern Pacific Izu Basin15. The COI data (95 sequences) published in Hoffman et al.1 were updated according to the results of this study and can be downloaded via BOLD (http://www.https://doi.org/10.5883/DS-INMAC03).
All data conversions of this study were performed by PGD Spider version 2.1.1.5 49. Distribution map was created by ggOceanMaps49 version 2.2.0 in R. Figures were graphically adjusted using Adobe® Photoshop® 25.11.0 software.
Results
Raw data filtering
Sequences with average sequencing quality Q < 30 were filtered. All demultiplexed, raw data reads (.fasta) belonging to 138 specimens were checked using the FASTQC High Throughput Sequence Report software (version 0.11.9) and 13 individuals with low number of reads (< 15,000 reads), low number of loci (< 1,500 loci) or abnormal GC content (GC < 50 and > 58) were excluded from further analysis. A total of 125 individuals of Anatoma were successfully analysed using 2b-RAD sequencing from six vent fields (VF1, Gauss, VF2, VF3, VF4 and VF5) (Fig. 1) for five different species published by Hoffman et al.1: A. declivis, A. discapex, A. laevapex, A. paucisculpta and Anatoma sp. 1 DZMB_2021_0095. Filtration steps to include loci found in at least 10% of the individuals of one population resulted in 23,856 loci, of which were 12,254 variant (Supplementary Table S3).
Species delimitation and assignment
The result of both cluster analyses, DAPC and STRUCTURE HARVESTER, indicated that K = 3 (Supplementary Figures S1 and S2) is the most probable number of clusters K. However, a clear assignment of the specimens to five different clusters was evident in the STRUCTURE barplot (Fig. 2a), which supported the morphological identification and delimitation of species by COI barcoding. The accuracy of the cluster analyses was likely constrained by the underrepresentation of A. paucisculpta and Anatoma sp. 1 DZMB_2021_0095. Furthermore, an additional 22 specimens for which COI identification was unsuccessful due to DNA degradation were successfully sequenced by 2b-RAD and could consequently be assigned to species.
The species delimitation based on morphology and COI can also be observed in the clustering on the NMDS plot. Five clusters were identified using the genetic ancestry, inferred from the genomic data (Fig. 2b). Two specimens exhibited a signal indicative of potential hybridisation. Both were unambiguously attributed to the species A. declivis and A. discapex through morphology and COI barcoding.
In addition, path sampling based on the 2b-RAD data provided further confirmation of the current taxonomy, with full support. The species trees inferred using SNAPP within the software BEAST 2 38 revealed A. discapex and A. declivis as the most closely related species, for both applied approaches comprising 3115 sites (Supplementary Table S4 and Fig. 3a) and 61 sites (both dataset DS_INMAC_RAD01; Supplementary Table S5 and Fig. 3b).
Results of population structure analysis between the species. (a) STRUCTURE assignment based on an admixture model with correlated allele frequencies. The plot indicates the most likely estimated number of clusters K = 3 calculated by STRUCTURE HARVESTER and according to the (de novo) cluster analysis by the package adegenet30,31 version 2.1.5 in R . Also highlighted is the expected number of clusters K = 5 according to Hoffman et al.1. The barplot was visualized using CLUMPAK. (b) Nonmetric Multidimensional Scaling (NMDS) plot based on the inferred ancestry matrix calculated by STRUCTURE showing the five different species (stress = 0.0868), generated by package vegan36 version 2.6.2 in R.
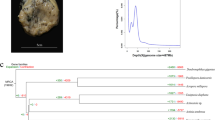
Species tree for the Anatoma species estimated with (a) 3115 sites (including the three most abundant species A. declivis, A. discapex and A. laevapex); and (b) 61 sites (including all five species) by using the 2b-RADs dataset and the current taxonomy model (RunA) that separates the species according to the taxonomy done by Hoffman et al.1. Posterior probabilities are shown on branches.
Anatoma species distribution
Figure 4 illustrates the updated species distribution within the genus Anatoma across the six vent fields. Maximum diversity of the four species was observed in Vent Fields 1, 4 and 5, while Gauss, Vent Field 2 and 3 exhibited three species (Supplementary Table S6). All four species were observed in both the CIR and SEIR. Anatoma discapex was documented in five vent fields, A. declivis in all six vent fields, A. laevapex in five vent fields, and A. paucisculpta in only three vent fields. It is evident that the most prevalent species, A. declivis, is the most extensively represented.
Genetic differentiation and population genetic structure
We measured the intraspecific difference for A. declivis, A. discapex and A. laevapex by analysing the RAD data. Anatoma paucisculpta and Anatoma sp. 1 DZMB_2021_0095 were excluded from the analysis due to the insufficient number of representatives (three individuals of A. paucisculpta; one individual of Anatoma sp. 1 DZMB_2021_0095).
A total of 48 individuals from four vents were studied within the species of A. declivis, (Gauss = 4, VF2 = 11, VF4 = 22, VF5 = 11). Following the application of the STACKS analysis, 5,064 loci and 5,148 variant sites were identified as remaining (Supplementary Table S3). The cluster analysis conducted using the STRUCTURE HARVESTER software revealed that the optimum number of clusters K for A. declivis was three and DAPC indicated K = 1 (Fig. 5a, Supplementary Figure S3). No evidence for genetic differentiation was observed. The FST values calculated by STACKS ranged from 0.0077 to 0.0122 and were therefore not statistically significant (Fig. 5b). No species differentiation was observed in the four vent fields in the CIR and SEIR, which corroborated the results of the cluster analysis. For the less distant vent fields, such as Vent Field 4 and 5 (FST = 0.008) and Vent Field 2 and Gauss (FST = 0.009) the FST values were comparatively lower.
We examined 50 individuals of the species A. discapex from four vents (VF1 = 22, Gauss = 1, VF2 = 7, VF3 = 20). For A. discapex 4,829 loci and 4,617 variant sites were retrieved (Supplementary Table S3), resulting in the optimal number of clusters K = 3 for STRUCTURE HARVESTER and K = 1 for DAPC (Fig. 5a, Supplementary Figure S3). This indicated no evidence of differentiation, as demonstrated by non-significant FST values ranging from 0.006 to 0.014 (Fig. 5b).
The study of A. laevapex encompassed 23 individuals from three distinct vent fields (Gauss = 18, VF2 = 1, VF5 = 4). The cluster analysis included 4,283 loci and 2,207 variant sites (Supplementary Table S3), and yielded the most probable number of clusters K = 3 (STRUCTURE HARVESTER) and K = 1 (DAPC; Fig. 5a, Supplementary Figure S3). The STRUCTURE analysis suggested a genetic differentiation for the population at Vent Field 5; however, the FST values were not significant (Fig. 5b).
AMOVA showed that the genetic variation is (slightly) lower among samples within populations (− 16.8 − 0.43 %) than among populations (0.4 − 4.1 %), while most of the genetic variation is within individuals (98.5 − 112.7 %) (Table 1). AMOVA statistics among populations resulted in significant p-values, indicating the presence of population structure (Table 1).
Results for the population analysis of A. declivis, A. discapex and A. laevapex. (a) Cluster analysis of the STRUCTURE calculation (based on an admixture model with correlated allele frequency). The most likely estimated number of clusters K was calculated by STRUCTURE HARVESTER (red) and according to the (de novo) cluster analysis by the package adegenet (blue). The barplot was produced using CLUMPAK. (b) Pairwise FST matrix was constructed using the sampled vent fields. No pairwise FST values were significant.
Population metrics
Genetic distance (PhiST) of the 2b-RAD data (DS_INMAC_RAD02-04) was tested against geographic distance, and a positive correlation was identified. However, the Mantel test for isolation-by-distance yielded a non-significant result (A. declivis: r2 = 0.609, p = 0.269; A. discapex: r2 = 0.115, p = 0.603; A. laevapex: r2 = 0.237, p = 0.666).
The genomic RAD data revealed an overall low heterozygosity for the three species studied across all vent populations and the metapopulations. The FIS value was represented by values close to 0 (Table 2). Additionally, the nucleotide diversity was observed to be low, ranging from 0.00399 to 0.00644 for all loci (variant and fixed). These findings were confirmed by the low nucleotide diversity observed in the COI data, which ranged between π = 0.00473 for A. discapex (combined dataset) and π = 0.00481 for A. declivis at Gauss (Table 3).
The haplotype diversity calculated based on the COI data was high for A. discapex (VF3; h = 0.961), its combined dataset (VF2, Gauss, VF3; h = 0.931) and in A. declivis (Gauss; h = 1.000 and the combined set VF1, VF2, Gauss, VF4, VF5; h = 0.829) (Table 3).
The neutrality and population expansion tests yielded negative Tajima’s D values (Table 3), which were found to be statistically significant (p-value < 0.05) for the populations of A. declivis (metapopulation and VF4), A. discapex (VF3) and for A. laevapex (metapopulation and Gauss). The negative D values indicated an excess of rare nucleotides thus an expansion of the populations. This hypothesis was supported by the negative Fu’s Fs values (Table 3).
The number of observed haplotypes varied from 1 to 19 (Table 3; Fig. 6). Based on the haplotype network (Fig. 6) a distinct separation between the six species with a mutation rate of 25 to 56 substitutions was demonstrated. The minimum intra-specific difference was observed between A. declivis and A. discapex (25 substitutions). The corresponding mean intra-specific difference was notably smaller than the inter-specific difference. These two species were therefore conclusively separated.
Results of the statistical haplotype network analysis conducted using the PopART program on the COI dataset, including all Anatoma species from GenBank. The analysis is based on an alignment of 658 bp. The colours correspond to the different sampling sites and the size of each circle is proportional to the number of individuals (see legend). Hatch marks indicate single substitutions. Mutations (n > 10) are presented as numbers. The triangles in the legend refer to hydrothermal vents at the sampling locality. The high degree of genetic similarity between A. paucisculpta and Anatoma sp. Lau is indicated with a red circle.
Discussion
RAD sequencing - suitable method for species delimitation
By assigning additional specimens to species this study demonstrates that 2b-RAD sequencing is a more suitable method for degraded DNA51,52 compared to the COI marker. The cytochrome c oxidase I (COI) gene, which is around 650 basepairs (bp) in length and located within the mitochondrial genome, is a useful DNA barcode to provide signals of population history over short periods of time due to its relatively high mutation rate53. In addition, to gain insights into the population over longer time frames, it is necessary to examine the nuclear DNA. The assessment of the entire nuclear genome is expensive, especially when comparing several specimens from different populations. By contrast, 2b-RAD sequencing is a whole-genome sequencing method that examines multiple loci through the analysis of short fragments of 32–35 bp.
High similarity to other species from hydrothermal vent fields – are the species vent-endemic?
Based on COI analyses, our studied anatomids were genetically similar to Anatoma sp. Lau, a specimen obtained from hydrothermal vent fields in the Lau Basin (South Pacific Ocean) at a depth of 1817 m15 (Fig. 6). The distance between the Lau Basin and the Rodriguez Triple Junction (RTJ) is approximately 11,200 km. The three other species for which a COI barcode was available exhibited a greater genetic distance from the species under this study (Fig. 6): Anatoma euglypta (Pelseneer, 1903) from Pine Island Bay, Amundsen Sea, Antarctica48 (approx. 4,700 km from the RTJ), A. pseudoequatoria (Kay, 1979) from reef and shore in Hawaii (unpublished, approx. 15,600 km from the RTJ) and Anatoma sp. Izu from intertidal zones Izu, Shizuoka, Japan15 (approx. 10,200 km from the RTJ). It can therefore be concluded that A. euglypta and Anatoma sp. Izu are geographically less distant from our species, which indicates that anatomids inhabiting vent ecosystems may be genetically closer to one another than non-vent species.
The INDEX project encompasses the study of the seafloor at hydrothermal vent fields and their surrounding habitats. It offers a distinctive opportunity for annual sampling with an ROV, which produced a substantial amount of data, including video footage, seafloor maps, and information on the water masses. In addition, a total of 20,000 macrofauna specimens were collected from hydrothermal vents and surrounding non-vent seafloors. Therefore, we can conclude, that the anatomid species analysed in this study were exclusively sampled in the immediate vicinity of vents (active and inactive hydrothermal vent areas) and not even one in non-vent habitats (personal observation). This leads us to the assumption that the anatomids are only associated with hydrothermal vents.
The combination of the occurrence of our sampled anatomids exclusively in chemosynthetically active environments and the high similarity to the specimen Anatoma sp. Lau leads us to the hypothesis, that our species are probably vent-endemic. This term is used to describe species that are restricted to an ecosystem rather than to a specific location54. The feeding traits of our species may also enhance the likelihood of endemism, as they feed most probably on the present bacterial mats1.
Anatoma populations indicate geneflow along the CIR and SEIR
2b-RAD sequencing was employed to evaluate the population structure of the three most prevalent anatomid species along the CIR and SEIR, A. declivis, A. discapex and A. laevapex. The results demonstrated that these species exhibited identical population patterns, characterised by panmictic populations and the absence of genetic differentiation across the entire ridge system (Table 1; Figs. 5 and 6). Although the AMOVA results indicated weak but significant population structure among populations (Table 1), this contrasted with non-significant FST values and non-significant Mantel test for isolation-by-distance. Highly connected populations are the result of most of the connectivity studies along the CIR9,10,11,12,55. This study underscores the well-connected vent populations along the entire ridge system, despite the patchy and dispersed nature of their habitats.
Furthermore, it indicates that species of Anatoma are capable of dispersing between vent fields over a distance of approximately 800 km, which can be explained by the present water masses. The sampled vent fields in this study are influenced by the Circumpolar Deep Water (CDW), which ranges in depth from 2,000 to 2,500 m (see Fig. 6 in Harms et al.56). The Indian Deep Water (IDW), with a flow direction from north to south and a depth range of 2,000–1,250 m, is present above the CDW. In addition, a complex sequence of water masses is known to exist, characterised by a variety of current directions and properties56. If larvae can enter the currents of these different water masses, the studied species may disperse in any direction.
Moreover, an extended duration in a planktonic phase will serve to enhance dispersion. Many vent invertebrate species exhibit lecithotrophic larval development, which is characterised by a dependency on energy reserves stored in the yolk. Nevertheless, this results in a particular larval phase57. Alternatively, there may be a lecithotrophic larval stage for at least part of their development, followed by a plankton-based diet in subsequent stages8. In contrast, it has been proposed that planktotrophic larvae have considerable potential for dispersal6. The gastropod (genera suggested: Lepetodrilus & Phymorhynchus) and bivalve larvae (genus suggested: Bathymodiolus) at Solitaire and Onnuri Vent Field on the CIR were sampled in the upper layers of the water column (0–200 m), indicating that these larvae may disperse approximately 2,000 m above the vents58. The larval shells of the sampled Anatoma indicate either a direct development with large (and hence few) eggs or a short lecithotrophic phase1. To the best of our knowledge, anatomid larvae have not yet been sampled in plankton samples.
It seems reasonable to posit that transform faults may act as an impassable barrier for veliger larvae of numerous gastropods, given that these larvae are inclined to remain in the proximity of the seabed59. An example of low gastropod species connectivity in the Indian Ocean is Chrysomallon squamiferum, which is endemic to the central Indian Ocean vent fields11. Significant genetic differentiation is observed between the southern SWIR and CIR, as well as between the CIR and CR13,14. This differentiation can likely be attributed to transform faults. Furthermore, differentiation is observed between the southern SWIR and CIR for two additional species (Neoplepas marisindica and Bathymodiolus septemdierum), and between the CIR and CR for Neoplepas marisindica, Chrysomallon squamiferum, Bathymodiolus septemdierum and Hesiolyra heteropoda14. To determine whether these “differentiation-zones” are present within Anatoma, further sampling on the CR and SWIR is required.
Expanding populations reveal small gene pool, hence a vulnerability to mining activities
In addition to the high connectivity among the populations of Anatoma, Tajima’s D and Fu’s Fs calculations on the COI data revealed expanding populations (Table 3). This phenomenon is commonly observed in hydrothermal vent populations60,61,62,63,64. This indicates that the populations may be relatively young sharing a recent common history of bottleneck or founder events, and expansion. This assumption is confirmed by the 2b-RAD data, as the observed heterozygosity (HO) was slightly lower than the expected heterozygosity (HE), suggestive of an earlier population bottleneck.
We observed extremely low heterozygosity (H0 = 0.09155–0.28449; for variant sites) (Table 2), indicating that there is minimal genetic variability. Similarly low values have been documented for the vent species Bathymodiolus platifrons (HO = 0.1480–0.1633)65. In contrast, the values for microsatellite loci for Bathymodiolus manusensis from the Manus Basin were considerably higher (HO = 0.24–0.94)62, with similar results observed by Teixeira et al.66 for Rimicaris exoculata (HO = 0.58–0.68). In the case of Munidopsis lauensis and Chorocaris sp. 2, the calculated HO ranges are larger61.
Coykendall et al.67 establish a correlation between HE of Riftia pachyptila (ranging from approximately HE = 0.1 to 0.5) and tectonic spreading rates. Their findings indicate that lower heterozygosity is associated with faster spreading rates. In our case, however, this theory cannot be confirmed. The CIR is a slow- to intermediate-spreading ridge68, whereas the SEIR is an intermediate- to fast-spreading ridge69. No significant difference was observed in heterozygosity between the CIR and SEIR.
The NMDS plot (Fig. 2b) from the 2b-RAD dataset indicated the potential for hybridisation between A. declivis and A. discapex in two specimens. On the basis of mitochondrial and morphological characteristics they were unambiguously assigned to the respective species as either A. declivis or A. discapex. Hybridisation is a common phenomenon in Gastropoda, and is detected at vent fields within mussel species of Bathymodiolus70,71,72, respectively. Further sampling and analyses will be necessary to support this theory and to estimate the effects that hybridisation will have regarding the formation of new species within Anatoma.
Although connectivity was high for all three species, low heterozygosity indicated a small gene pool and therefore, high sensitivity to environmental change. Serious damage to hydrothermal vent ecosystems is predicted from future deep-sea mining or other anthropogenic impacts73. To ascertain whether the unique vent field fauna can be conserved through the establishment of marine protected areas, further research on genetic connectivity is required to develop a conservation plan also for other species in this ecological niche.
Indian Ocean functions as dispersal corridor for hydrothermal vent species
Several vent-endemic genera occur in Atlantic, Pacific and Indian Ocean, such as Bathymodiolus, Munidopsis and Phymorhynchus74, whereas shared taxa between Atlantic and Pacific Ocean vents are leading to the theory that the Indian Ocean serves as a dispersal conduit between ocean basins74,75. The similarity of A. paucisculpta from Anatoma sp. Lau is supporting this theory. Hoffman et al.1 distinguishes them by four out of six delimitation methods. Haplotype analyses revealed only seven mutations between Anatoma sp. Lau and A. paucisculpta (Fig. 6).
A similar pattern is observed by Hwang et al.76 for the squat lobster Munidopsis lauensis from specimens collected from the Onnuri Vent Field (Indian Ocean), the Brothers Seamount, Manus, and Lau Basins (both southwest Pacific Ocean). The evolution of hydrothermal species from shallow specimens and a dispersion corridor is hypothesized. Besides, their results indicate that the western Pacific population diverged before the Indian Ocean population76.
Species occurrence on small spatial scale suggests sympatric speciation event
In addition, the genetic similarity between A. paucisculpta from Anatoma sp. Lau suggests the possibility of a recently, geographically diverged species. Anatomids in the Indo-Pacific exhibit a wide distribution, with some species extending over 10,000 km77. Only two anatomids were previously reported by Geiger77 to inhabit depths exceeding 2,000 m in the Indian Ocean. However, the recent additions by Hoffman et al.1 suggest that additional species may occupy lower bathyal and abyssal depths.
A broad distribution was evident for A. declivis, A. discapex, A. laevapex and A. paucisculpta along the ridge systems in the Indian Ocean and it is likely that some species were not sampled at certain vents due to either undersampling or unsuitable environmental conditions for the species (environmental filtering). The present study demonstrated the co-existence of six Anatoma species at the small spatial scale, with two to three species observed on a single sampled small rock or piece of chimney, and four to five species sampled within a radius of less than ten metres. The speciation of Anatoma at Indian Ocean vent fields may be attributed to the presence of diverse bacterial mats78,79, which serve as a potential food source for different anatomid species1. The presence of the two single specimens in this study suggests that there are likely numerous additional species of Anatoma at the hydrothermal fields in the Indian Ocean, that have not been sampled due to their occurrence in smaller numbers.
Finally, it must be mentioned, in addition to offering a lot of new insights, this study also presents certain limitations, such as the small number of specimens from some locations. Population genetic studies typically require at least ten individuals per sampling site to ensure accurate results80. Nevertheless, this study makes a significant contribution to the field of connectivity studies along the Indian Ocean ridge system, as there have been no previous studies on population connectivity at the SEIR. The findings of this study indicate that Anatoma populations exhibit no discernible differentiation along the investigated trajectories across the CIR and SEIR. Moreover, this study addresses numerous questions pertaining to the distribution and global connectivity of the Anatomidae. To enhance comprehension of anatomid populations at hydrothermal vents, it is imperative to augment the COI dataset with genetic material from the CR, SWIR, Pacific, and Atlantic vent species, complemented by genomic data. Additionally, the scarcity of the species A. paucisculpta and the two single specimens necessitates further sampling to elucidate their role at Indian Ocean hydrothermal vents.
Conclusions
2b-RAD is a suitable method for species delimitation in addition to the traditional species differentiation by morphology and the molecular identification by COI. Many of the short DNA fragments analysed by 2b-RAD provide superior determinations when the genetic code is degraded. Long-distance connectivity of vent anatomids is indicated by the high similarity between A. paucisculpta and a specimen from a hydrothermal vent in the Pacific Ocean. Furthermore, this study confirms the theory of a dispersal corridor between the western Pacific and the Indian Ocean. Three species A. declivis, A. discapex and A. laevapex show a high genetic similarity over approximately 800 km, revealing a high gene flow and good connectivity between the vent fields. However, low heterozygosity highlights the vulnerability of the present fauna towards catastrophic events due to limited genetic diversity. Finally, this study demonstrates, how 2b-RAD sequencing in combination with other molecular methods can provide important information on population structure and dynamics in hydrothermal vent ecosystems, which is necessary for the conservation and management of these ecosystems with regards to potential future mining.
Data availability
The 2b-RADs raw reads generated during the current study have been deposited in the European Nucleotide Archive (ENA) at EMBL-EBI under accession number PRJEB63999 (https://www.ebi.ac.uk/ena/browser/view/PRJEB63999) and the obtained datasets can be accessed through the Senckenberg Metadata Portal https://dataportal.senckenberg.de/dataset/318754d6-e802-4cb2-a8e5-7f3a4d68af0d. The COI data of Hoffman et al.1 was updated and is available in BOLD (https://doi.org/10.5883/DS-INMAC03).
References
Hoffman, L., Kniesz, K., Martínez Arbizu, P. & Kihara, T. Abyssal vent field habitats along plate margins in the Central Indian Ocean yield new species in the genus Anatoma (Vetigastropoda: Anatomidae). Eur. J. Taxon. 826, 135–162 (2022).
United Nations. Agreement under the United Nations Convention on the Law of the Sea on the Conservation and Sustainable Use of Marine Biological Diversity of Areas Beyond National Jurisdiction. (2023).
Dover, C. et al. Biodiversity loss from deep-sea mining. Nat. Geosci. 10, 464–465 (2017).
Van Dover, C. L. Inactive sulfide ecosystems in the deep sea: a review. Front. Mar. Sci. 6, (2019).
Vadakkepuliyambatta, S. et al. Potential of deep-sea mineral resources for the blue economy. Curr. Sci. 126, 192 (2024).
Mullineaux, L. S. et al. Exploring the ecology of deep-sea hydrothermal vents in a metacommunity framework. Front. Mar. Sci. 5, (2018).
Hilário, A. et al. Estimating dispersal distance in the deep sea: challenges and applications to marine reserves. Front. Mar. Sci. 2, (2015).
Marsh, A. G., Mullineaux, L. S., Young, C. M. & Manahan, D. T. Larval dispersal potential of the tubeworm Riftia pachyptila at deep-sea hydrothermal vents. Nature 411, 77–80 (2001).
Nakamura, K. et al. Discovery of new hydrothermal activity and chemosynthetic fauna on the Central Indian Ridge at 18°–20°S. PLOS ONE. 7, e32965 (2012).
Beedessee, G. et al. High connectivity of animal populations in deep-sea hydrothermal vent fields in the Central Indian Ridge relevant to its geological setting. PLOS ONE. 8, e81570 (2013).
Chen, C., Copley, J. T., Linse, K. & Rogers, A. D. Low connectivity between ‘scaly-foot gastropod’ (Mollusca: Peltospiridae) populations at hydrothermal vents on the Southwest Indian Ridge and the Central Indian Ridge. Org. Divers. Evol. 15, 663–670 (2015).
Breusing, C. et al. Allopatric and sympatric drivers of speciation in Alviniconcha hydrothermal vent snails. Mol. Biol. Evol. 37, 3469–3484 (2020).
Sun, J. et al. Nearest vent, dearest friend: biodiversity of Tiancheng vent field reveals cross-ridge similarities in the Indian Ocean. R Soc. Open. Sci. 7, 200110 (2020).
Zhou, Y. et al. Delineating biogeographic regions in Indian Ocean deep-sea vents and implications for conservation. Divers. Distrib. 28, 2858–2870 (2022).
Kano, Y. Vetigastropod phylogeny and a new concept of Seguenzioidea: independent evolution of copulatory organs in the deep-sea habitats. Zool. Scr. 37, 1–21 (2008).
Leinberger, D. A. Ocean container temperature and humidity study. PreShipment Test. Newsl. (2006). 2nd Quarter/2006.
Csavajda, P. & Böröcz, P. Climate conditions in ISO container shipments from Hungary to South Africa and Asia. Period Polytech. Transp. Eng. 47, (2018).
Riehl, T. et al. Field and laboratory methods for DNA barcoding and molecular-systematic studies on deep-sea isopod crustaceans. Pol. Polar Res. 35, 205–226 (2014).
Post, R. J., Flook, P. K. & Millest, A. L. Methods for the preservation of insects for DNA studies. Biochem. Syst. Ecol. 21, 85–92 (1993).
Carvalho, A. & Vieira, L. Comparison of preservation methods of Atta spp. (Hymenoptera: Formicidae) for RAPD analysis. Soc. Entomológica Bras. 29, 489–496 (2000).
Marquina, D., Buczek, M., Ronquist, F. & Łukasik, P. The effect of ethanol concentration on the morphological and molecular preservation of insects for biodiversity studies. PeerJ 9, e10799 (2021).
Wang, S., Meyer, E., McKay, J. K. & Matz, M. V. 2b-RAD: a simple and flexible method for genome-wide genotyping. Nat. Methods. 9, 808–810 (2012).
Catchen, J. M., Amores, A., Hohenlohe, P., Cresko, W. & Postlethwait, J. H. Stacks: building and genotyping loci De Novo from short-read sequences. G3 GenesGenomesGenetics. 1, 171–182 (2011).
Catchen, J., Hohenlohe, P. A., Bassham, S., Amores, A. & Cresko, W. A. Stacks: an analysis tool set for population genomics. Mol. Ecol. 22, 3124–3140 (2013).
Hubisz, M. J., Falush, D., Stephens, M. & Pritchard, J. K. Inferring weak population structure with the assistance of sample group information. Mol. Ecol. Resour. 9, 1322–1332 (2009).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Earl, D. & Vonholdt, B. Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 1–3 (2012).
Jombart, T. Adegenet: a R package for the multivariate analysis of genetic markers. Bioinforma Oxf. Engl. 24, 1403–1405 (2008).
Jombart, T. & Ahmed, I. Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinforma Oxf. Engl. 27, 3070–3071 (2011).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2022).
Posit Team. RStudio: Integrated Development Environment for R (Posit Software, 2024).
Miller, J. M., Cullingham, C. I. & Peery, R. M. The influence of a priori grouping on inference of genetic clusters: simulation study and literature review of the DAPC method. Heredity 125, 269–280 (2020).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281 (2014).
Oksanen, J. et al. vegan: community ecology package. (2022).
Leache, A., Fujita, M., Minin, V. & Bouckaert, R. Species delimitation using genome-wide SNP data. Syst. Biol. 63, (2014).
Bouckaert, R. et al. BEAST 2.5: an advanced software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 15, e1006650 (2019).
Leaché, A. D. & Bouckaert, R. R. Species Trees and Species Delimitation with SNAPP: A Tutorial and Worked Example. (2018). http://evomics.org/wp-content/uploads/2018/01/BFD-tutorial.pdf
Kass, R. E. & Raftery, A. E. Bayes factors. J. Am. Stat. Assoc. 90, 773–795 (1995).
Bryant, D., Bouckaert, R., Felsenstein, J., Rosenberg, N. A. & RoyChoudhury, A. Inferring species trees directly from biallelic genetic markers: bypassing gene trees in a full coalescent analysis. Mol. Biol. Evol. 29, 1917–1932 (2012).
Meirmans, P. G. GENODIVE version 3.0: easy-to-use software for the analysis of genetic data of diploids and polyploids. (2020).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983).
Fu, Y. X. New statistical tests of neutrality for DNA samples from a population. Genetics 143, 557–570 (1996).
Bandelt, H. J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Geiger, D. L. & Thacker, C. Molecular phylogeny of Vetigastropoda reveals non-monophyletic Scissurellidae, Trochoidea, and Fissurelloidea. Molluscan Res. 25, 47–55 (2005).
Lischer, H. E. L. & Excoffier, L. PGDSpider: an automated data conversion tool for connecting population genetics and genomics programs. Bioinformatics 28, 298–299 (2012).
Vihtakari, M. ggOceanMaps: plot data on oceanographic maps using ’ggplot2’. (2024).
Barbanti, A. et al. Helping decision making for reliable and cost-effective 2b-RAD sequencing and genotyping analyses in non-model species. Mol. Ecol. Resour. 20, 795–806 (2020).
Miller-Crews, I., Matz, M. V. & Hofmann, H. A. A 2b-RAD parentage analysis pipeline for complex and mixed DNA samples. Forensic Sci. Int. Genet. 55, 102590 (2021).
Galtier, N., Nabholz, B., Glémin, S. & Hurst, G. D. D. mitochondrial DNA as a marker of molecular diversity: a reappraisal. Mol. Ecol. 18, 4541–4550 (2009).
Wolff, T. Composition and endemism of the deep-sea hydrothermal vent fauna. Biol. Mar. 46, 97–104 (2005).
Watanabe, H. K. et al. Phylogeography of hydrothermal vent stalked barnacles: a new species fills a gap in the Indian Ocean ‘dispersal corridor’ hypothesis. R Soc. Open. Sci. 5, 172408 (2018).
Harms, N. et al. Nutrient distribution and nitrogen and oxygen isotopic composition of nitrate in water masses of the subtropical southern Indian Ocean. Biogeosciences 16, 2715–2732 (2019).
Van Dover, C. L. The Ecology of Deep-Sea Hydrothermal Vents (Princeton University Press, 2000).
Kim, M., Kang, J. H. & Kim, D. Holoplanktonic and meroplanktonic larvae in the surface waters of the Onnuri vent field in the Central Indian Ridge. J. Mar. Sci. Eng. 10, 158 (2022).
Mullineaux, L. S. et al. Vertical, lateral and temporal structure in larval distributions at hydrothermal vents. Mar. Ecol. Prog Ser. 293, 1–16 (2005).
Vrijenhoek, R. C. Genetic diversity and connectivity of deep-sea hydrothermal vent metapopulations. Mol. Ecol. 19, 4391–4411 (2010).
Thaler, A. D. et al. Comparative Population structure of two Deep-Sea Hydrothermal-Vent-Associated Decapods (Chorocaris sp. 2 and Munidopsis lauensis) from Southwestern Pacific back-Arc basins. PLOS ONE. 9, e101345 (2014).
Thaler, A. D., Saleu, W., Carlsson, J., Schultz, T. F. & Dover, C. L. V. Population structure of Bathymodiolus manusensis, a deep-sea hydrothermal vent-dependent mussel from Manus Basin, Papua New Guinea. PeerJ 5, e3655 (2017).
Lee, W. K., Kim, S. J., Hou, B. K., Dover, C. L. V. & Ju, S. J. Population genetic differentiation of the hydrothermal vent crab Austinograea Alayseae (Crustacea: Bythograeidae) in the Southwest Pacific Ocean. PLOS ONE. 14, e0215829 (2019).
Yang, C. H. et al. Population genetic differentiation on the hydrothermal vent crabs Xenograpsus testudinatus along depth and geographical gradients in the western Pacific. Diversity 14, 162 (2022).
Xu, T. et al. Population genetic structure of the deep-sea mussel Bathymodiolus platifrons (Bivalvia: Mytilidae) in the Northwest Pacific. Evol. Appl. 11, 1915–1930 (2018).
Teixeira, S., Serrao, E. & Arnaud-Haond, S. Panmixia in a fragmented and unstable environment: the hydrothermal shrimp Rimicaris exoculata disperses extensively along the Mid-atlantic Ridge. PloS One. 7, e38521 (2012).
Coykendall, D. K., Johnson, S. B., Karl, S. A., Lutz, R. A. & Vrijenhoek, R. C. Genetic diversity and demographic instability in Riftia pachyptila tubeworms from eastern Pacific hydrothermal vents. BMC Evol. Biol. 11, 96 (2011).
DeMets, C., Gordon, R. G. & Argus, D. F. Geologically current plate motions. Geophys. J. Int. 181, 1–80 (2010).
Sempéré, J. C. & Cochran, J. R. The Southeast Indian Ridge between 88°E and 118°E: variations in crustal accretion at constant spreading rate. J. Geophys. Res. Solid Earth. 102, 15489–15505 (1997).
Breusing, C., Vrijenhoek, R. C. & Reusch, T. B. H. widespread introgression in deep-sea hydrothermal vent mussels. BMC Evol. Biol. 17, 13 (2017).
Johnson, S. B., Won, Y. J., Harvey, J. B. & Vrijenhoek, R. C. A hybrid zone between Bathymodiolus mussel lineages from eastern Pacific hydrothermal vents. BMC Evol. Biol. 13, 21 (2013).
O’Mullan, G. D., Maas, P. A., Lutz, R. A. & Vrijenhoek, R. C. A hybrid zone between hydrothermal vent mussels (Bivalvia: Mytilidae) from the Mid-atlantic Ridge. Mol. Ecol. 10, 2819–2831 (2001).
van der Most, N., Qian, P. Y., Gao, Y. & Gollner, S. Active hydrothermal vent ecosystems in the Indian Ocean are in need of protection. Front. Mar. Sci. 9, (2023).
Van Dover, C. L. et al. Biogeography and ecological setting of Indian Ocean hydrothermal vents. Science 294, 818–823 (2001).
Hessler, R. R. & Lonsdale, P. F. Biogeography of Mariana Trough hydrothermal vent communities. Deep Sea Res. Part. Oceanogr. Res. Pap. 38, 185–199 (1991).
Hwang, H., Cho, B., Cho, J., Park, B. & Kim, T. New record of hydrothermal vent squat lobster (Munidopsis lauensis) provides evidence of a dispersal corridor between the Pacific and Indian oceans. J. Mar. Sci. Eng. 10, 400 (2022).
Geiger, D. L. Monograph of the Little Slit Shells (Santa Barbara Museum of Natural History, 2012).
Han, Y. et al. Hydrothermal chimneys host habitat-specific microbial communities: analogues for studying the possible impact of mining seafloor massive sulfide deposits. Sci. Rep. 8, 10386 (2018).
Adam-Beyer, N. et al. Microbial ecosystem assessment and hydrogen oxidation potential of newly discovered vent systems from the Central and South-East Indian Ridge. Front. Microbiol. 14, (2023).
Halbritter, D. A., Storer, C. G., Kawahara, A. Y. & Daniels, J. C. Phylogeography and population genetics of pine butterflies: sky islands increase genetic divergence. Ecol. Evol. 9, 13389–13401 (2019).
Acknowledgements
We would like to thank Thomas Kuhn, current Chief Scientist of the INDEX exploration program; Ulrich Schwarz-Schampera, Chief Scientist of the INDEX exploration program during the INDEX2015-INDEX2019 cruises; and the captains and crews of the RV Pelagia and RV Sonne and the ROV team of ROPOS for their assistance during sampling. The maps and samples presented in this study originate from the INDEX exploration project for marine polymetallic sulphides by the BGR (German Federal Institute for Geosciences and Natural Resources) on behalf of the Federal Ministry for Economic Affairs and Climate Action. Exploration activities are carried out in the framework and under the regulations of an exploration license with the International Seabed Authority. We would like to thank our colleagues at the DZMB Wilhelmshaven for their help with the molecular work and data analysis. Sequencing was carried out at Carl von Ossietzky Universität Oldenburg at the Next Seq 500. Special thanks to Sahar Khodami for her assistance with the preparation of the 2b-RAD library, the high-performance computer and by doing the test sequencing. This publication is number 101 of the Senckenberg am Meer Metabarcoding and Molecular Laboratory.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
TCK was the head of the project. TCK and PMA contributed to the conception and design of the study. PMA designed the RADs adapter, barcodes and the bioinformatics pipeline to process the raw reads. KK performed the molecular work, constructed the database, analysed the final data and wrote the first draft of the manuscript. All authors participated in the revision of manuscript and approved the final version for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kniesz, K., Hoffman, L., Martínez Arbizu, P. et al. High genomic connectivity within Anatoma at hydrothermal vents along the Central and Southeast Indian Ridge. Sci Rep 15, 1971 (2025). https://doi.org/10.1038/s41598-025-85507-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85507-z