Abstract
Acanthocephalan parasites are often overlooked in many areas of research, and satellitome and cytogenetic analyzes are no exception. The species of the genus Acanthocephalus are known for their very small chromosomes with ambiguous morphology, which makes karyotyping difficult. In this study, we performed the first satellitome analysis of three Acanthocephalus species to identify species- and chromosome-specific satellites that could serve as cytogenetic markers. RepeatExplorer2 revealed a remarkably high number of species-specific repeats, with a predominance of satellite DNAs, alongside variations in repetitive content between sexes. Five satellites in A. anguillae, two in A. lucii and six in A. ranae were successfully mapped to chromosomes using FISH. Each satellite showed a clustered hybridization signal at specific chromosomal locations, which allowed us to create a schematic representation of the distribution of satellites for each species. These newly identified satellites proved to be useful chromosomal markers for the accurate identification of homologous chromosome pairs. No FISH-positive signals were observed on the supernumerary chromosomes of A. anguillae and A. lucii, supporting the hypothesis that these chromosomes have recent origin.
Similar content being viewed by others
Introduction
Repetitive sequences are an important organizational component of eukaryotic genomes and comprise several classes, with the predominating being satellite DNA (satDNA) and transposable elements (TEs)1,2. Satellite DNA is classically characterized as non-coding, highly abundant repetitive DNA consisting of short, multiple repeated sequences. It is usually found in heterochromatin regions, mostly in pericentromeric, subtelomeric and interstitial regions, which are part of vital structures such as centromeres and telomeres3. Despite their association with critical chromosomal structures, satDNA exhibit highly dynamic behavior and can vary considerably even between closely related species. Therefore, these sequences are often genus- or species-specific and reflect short-term evolutionary changes4,5.
Acanthocephala is a group of endoparasites characterized by the possession of a retractable proboscis armed with hooks. These parasites are commonly found in livestock and birds, but also in marine and freshwater fauna and aquaculture. Despite their worldwide occurrence and their veterinary, medical and economic importance, many scientific areas, including repeatome analysis, remain overlooked or under-researched. To date, only one study has provided preliminary insights into the repetitive content of Acanthocephala. This study focused on the first nuclear genome and transcriptome assemblies of Pomphorhynchus laevis6. However, cytogenetic approaches were lacking in this study and information on repetitive elements was limited. Understanding the repetitive parts of the genomes and their distribution on the chromosomes could be of great value for phylogenetic studies and karyotyping process. Currently, the number of satellitome studies on various plant and animal species, including cytogenetic approaches, is increasing7,8,9,10,11,12,13,14. This increase is mainly due to the recent development of rapid and cost-effective whole genome sequencing in conjunction with bioinformatics tools that facilitate the identification and characterization of repetitive elements. Among these tools, RepeatExplorer2 (RE)15 stands out by offering several advantages: (i) use of low-throughput Illumina sequence data, (ii) no need for genome assembly, and (iii) no reference genome required. However, TAREAN/RepeatExplorer2 shows some limitations since the analysis sensitivity and accuracy are limited by low-complexity sequences masking similarities and the focus on top clusters that overlook less abundant or variable repeats15. Additionally, the number of reads analyzed is constrained by hardware resources like RAM, especially for large genomes or comparative studies. Despite this growing trend in satellite studies using cytogenetic approaches, acanthocephalans have been neglected. To date, only four species have been analyzed for the number and distribution of 18S rRNA and/or histone H3 genes, with different species-specific patterns found on chromosomes16,17,18.
Here we present the first study in which RE15,19 was used to perform satellitome analyses within the genus Acanthocephalus. We performed a comparative analysis of three species common in Slovakia, one amphibian (A. ranae) and two fish parasites (A. anguillae and A. lucii). These species are known to have very small chromosomes, which makes the pairing of homologous chromosomes challenging. The aim was to study the satellitome of these species and to identify species- and chromosome-specific satDNA families that could potentially serve as new cytogenetic markers. Satellites serve as effective chromosomal markers for the determination of intra- and interspecific genome diversity as well as for taxonomic and phylogenetic studies9,11,20,21. Our comparative analysis revealed a considerable number of species-specific satellites, and the most suitable repeats were subsequently mapped to chromosomes by fluorescence in situ hybridization.
Results
RepeatExplorer2 cluster analysis
We first performed separate analyzes for each species and sex to ensure optimal sensitivity in their characterization, followed by a comparative analysis to examine species-specific satellite repeats. The separate analyzes revealed that both the proportion of reads in top clusters (clusters representing more than 0.01% of the genome) and the number of identified repeat elements were increased in A. lucii compared to the other two species. In addition, RE showed remarkable differences between the sexes in terms of repeat element content in the analyzed species, where male genomes had a higher number of identified repeat elements than female genomes (Table 1). The differences were also observed in the genomic abundance of repetitive elements in the individual species. The highest prevalence of repeats above 0.9% in the genome was found in A. anguillae, namely 12 repeats. Moreover, the repeat of the highest abundance, 7.7% in female and 9.5% in male, was also found in A. anguillae, a proportion not reached in the other species studied. The species A. lucii contained nine repeats above 0.9% genomic abundance, with the highest abundant repeat being 2.1% in female and 4.8% in male. In contrast, A. ranae had no repetitive elements above 0.9% in its genome, with the top cluster having an abundance of 0.86% in female and 0.89% in male. In A. anguillae and A. lucii, the most prevalent cluster was annotated as satDNA. In A. ranae, however, the most important cluster was either annotated as Helitrons or remained unannotated. Overall, the majority of repeats identified by RE were annotated as satDNA in all species, with a higher proportion observed in male genomes (Table 1).
The comparative analysis of studied species was carried out with a total of 2,100,000 input reads. After RE clustering, 1,491,992 reads were grouped into 209,486 clusters, with 40% of the reads in the top clusters. The comparison of repeat content between the species is presented in Fig. 1. TAREAN analysis yielded a list of 134 putative satellite repeats. Of these, only nine clusters represented shared satDNAs across all three studied species, and only three of these contained more than 200 reads per species. In contrast, a considerable number of the found putative satellites represented species-specific satDNA: 29 in A. anguillae, 41 in A. lucii and 43 in A. ranae. Using a threshold of a minimum of 700 reads within repeat clusters, we identified 11 species-specific satellite repeats in the genome of A. anguillae, 16 in A. lucii and 28 in A. ranae. The threshold ensured that we focused on repeats with a higher representation and filtered out less significant or noise-related repeats.
Graph comparing the repeat content of three Acanthocephalus species studied. The bar plot shows the size (number of reads) in the individual top clusters. The color of the rectangles represents the RepeatExplorer2 annotation of each cluster, and the size is in proportion to the abundance of the repeat in each genome. A high number of species-specific repeats is evident. Species codes: AA—A. anguillae, AL—A. lucii, AR—A. ranae.
Divergence of satDNA is inversely related to amplification and homogenization processes and directly related to mutation rate22,23. Focusing on the species specific satDNAs landscapes, we found that their profile indicates a recent expansion on their genomes as the main peak of divergence have a value of 0 (see 3 examples on Supplementary Fig. S1). In addition, the repeatome analysis of the investigated species revealed a different proportion of ribosomal DNA (2.15% in A. anguillae, 1.03% in A. lucii and 0.92% in A. ranae). The genome proportion was calculated as the ratio of reads specific to individual repeat type to the total reads used for clustering analyses.
Physical mapping by FISH
The karyotypes of the species A. anguillae and A. lucii were determined in our recently published studies17,18; 2n = 7/8 (male/female); n = 1 m + 1 m-sm + 1a + 1a (X) in the former and 2n = 7/8; n = 1 m + 2sm + 1a (X) in the latter. While an old study determined the chromosome characteristics of A. ranae as 2n = 8; n = 4 m, with heterogametic (XY) sex chromosome system24, in our study we did not observe the Y chromosome, as will be discussed later. Here, we performed single- and two-color FISH and combined the satDNA probes to accurately identify the localization and satDNA-carrying chromosomes. All mapped satellites showed a clustered pattern at specific chromosomal locations. Hybridization signals were detected in the telo-subtelomeric, pericentromeric, centromeric and interstitial chromosomal regions. Based on the morphology of the chromosomes and the distribution of the satellites, the homologous pairs were identified and a scheme was drawn up showing the distribution of the newly identified species-specific markers (Fig. 2).
Schematic representation of the species-specific satellites on the chromosomes of: (a) Acanthocephalus anguillae; (b) Acanthocephalus lucii; (c) Acanthocephalus ranae.
Acanthocephalus anguillae
Of the 11 species-specific satDNA repeats identified, only five were successfully mapped to chromosomes, namely AAc5, AAc25, AAc31, AAc72 and AAc85 (Fig. 3). The largest metacentric chromosome pair No. 1 carried four of the identified repeats. Two satellites, AAc25 and AAc85, were located close to each other in the pericentromeric region of this chromosome, as revealed by two-color FISH (Fig. 3a–c). While the specific localization in the nuclei of metaphase and diplotene was indistinct (Fig. 3a,c), the particular locus was most clearly visible in the nuclei of pachytene. Both repeats were located on the long arms of the chromosome, with AAc85 closer to the centromere region, which was highlighted with DAPI (Fig. 3b). The next repeat detected on chromosome pair No.1 was AAc72. Distinct hybridization signals were observed on the short arms in the telomeric region at the metaphase and diplotene stages (Fig. 3d–f). Satellite AAc5 signals were detected on both the long and short arms of chromosome pairs No.1 and 2 in the interstitial regions near the centromere. The distribution of these repeats on the chromosomes was well observed in the nuclei of metaphase and diplotene (Fig. 3g–i). Finally, the satellite repeat AAc31 was detected on the X chromosome. The repeat clusters were located in a subtelomeric position (Fig. 3j–l). No positive FISH signals were observed on the third pair of chromosomes nor on the B chromosomes (Fig. 3a,d,g,l).
FISH mapping of repeats specific to Acanthocephalus anguillae. (a–c) AAc25 (red) and AAc85 (green) probes on: (a) female metaphase chromosome with one B chromosome; (b) male pachytene nucleus; (c) male diplotene nucleus. (d–f) AAc72 probe on: (d) male metaphase chromosome with one B chromosome; (e,f) male diplotene nucleus. (g–i) AAc5 probe on: (g) male metaphase chromosome with B chromosome; (h,i) male diplotene nucleus. (j–l) AAc31 probe on: (j) male metaphase chromosome; (k) male pachytene nucleus; (l) male diplotene nucleus with one B chromosome. Chromosomes were counterstained with DAPI. Asterisks indicate X chromosomes and arrows indicate B chromosomes. Scale bar = 5 µm.
During our study, we found a similar distribution pattern between the new species-specific satellite AAc5 and the 18S rDNA mapped in our previous work17. After two-color FISH, consistent localization was observed on the first two pairs of chromosomes with chromosome No.1 showed an additional signal of AAc5 on the short arms (Fig. 4a–c).
Mutual chromosomal location of selected satDNAs detected by two-color FISH on the chromosomes of Acanthocephalus anguillae (a–c) and A. ranae (d–i). (a–c) 18S rDNA (green) and AAc5 satellite (red) on male metaphase chromosomes of Acanthocephalus anguillae. (d–i) ARc42 satellite (green) and ARc27 satellite (red) on female metaphase chromosomes of A. ranae. (g–i) ARc51 satellite (green) and ARc39 satellite (red) on male diplotene chromosomes of A. ranae. Chromosomes were counterstained with DAPI. Asterisks indicate X chromosomes. Scale bar = 5 µm.
Acanthocephalus lucii
The comparative analysis identified 16 species-specific satDNAs, of which only two (ALc1, ALc98) were successfully mapped by FISH. The hybridization signals of the ALc1 repeat were restricted to the centromeric regions of all chromosomes (Fig. 5a–c), with the exception of the B chromosomes (Fig. 5b,c). The second repeat successfully mapped by FISH, ALc98, was clustered at three loci. The clusters were located in the telomeric region on the long arms of chromosome pairs No. 1, 2 and 3 (Fig. 5d–f). No positive FISH signals were found on the X chromosome or the B chromosome (Fig. 5d,f).
Satellite clusters detected by FISH in chromosomes of Acanthocephalus lucii. (a–c) ALc1 on: (a) male metaphase chromosomes; (b) male pachytene nucleus with one B chromosomes; (c) male diplotene nucleus with one B chromosome. (d–f) ALc98 on: (d) male metaphase chromosomes with one B chromosome; (e,f) male diplotene nucleus without and with one B chromosome. Chromosomes were counterstained with DAPI. Asterisks indicate X chromosomes and arrows indicate B chromosomes. Scale bar = 5 µm.
Acanthocephalus ranae
Six of the 28 satellites were successfully mapped on the chromosomes of A. ranae, namely ARc27, ARc39, ARc42, ARc49, ARc51 and ARc58 (Fig. 6). Hybridization signals were found on all chromosomes except chromosome pair No. 3. The largest chromosome pair No. 1 showed two satellites, ARc42 (Fig. 6a–c) and ARc51 (Fig. 6d–f), both located on the long chromosome arms. ARc42 was located in the centromeric region as detected in metaphase stages (Fig. 6a), and clear co-localization with DAPI-positive centromeric bands was visible in pachytene nuclei (Fig. 6b). The ARc51 repeat clusters were located interstitially on the long arms of the largest chromosome pair (Fig. 6d–f) and were mainly visible in the diplotene stages (Fig. 6f). Two repeats, Arc27 and Arc39, were detected on the second pair of chromosomes (Fig. 6g–l). The satellite repeat ARc27 was observed in the pericentromeric region (Fig. 6g–i), while hybridization signals from the ARc39 probe were detected interstitially on the long arms (Fig. 6j–l). The centromeric position of repeat ARc27 was confirmed by the presence of a DAPI-positive band at the same position of chromosome pair No. 2 (Fig. 6h,i). The interstitial position of ARc39 was most evident in diplotene nuclei, where the centromere highlighted with DAPI was most visible (Fig. 6l). The specific chromosomes carrying these satDNA sequences were identified by two-color FISH combining ARc27, ARc42 and ARc39, ARc51 as probes (Fig. 4d–i). The last two mapped repeats (ARc49 and ARc58) were found on the sex chromosomes (Fig. 6m–o). Two-color FISH with ARc49 and ARc58 probes revealed their pericentromeric localization, with the ARc49 cluster on the short arms in the secondary constriction site and ARc58 on the long arms near the centromere (Fig. 6m–o). The particular position of these repeats was most evident during the diplotene stages (Fig. 6o).
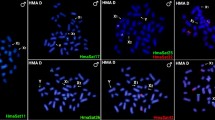
Physical mapping of different satDNAs in Acanthocephalus ranae. (a–c) ARc42 on male chromosomes: (a) metaphase I; (b) pachytene nucleus; (c) diplotene nucleus. (d–f) ARc51 on male chromosomes: (d) metaphase I; (e) pachytene nucleus; (f) diplotene nucleus. (g–i) ARc27 on male chromosomes: (g) metaphase I; (h) pachytene nucleus; (i) diplotene nucleus. (j–l) ARc39 on male chromosomes: (j) metaphase I; (k) pachytene nucleus; (l) diplotene nucleus. (m–o) Two-color FISH with ARc49 (red) and ARc58 (green) on female chromosomes: (m) metaphase I; (n) pachytene nucleus; (o) diplotene nucleus. Chromosomes were counterstained with DAPI. Asterisks indicate X chromosomes. Scale bar = 5 µm.
Discussion
Satellitome analysis
Studies on repetitive DNA, satDNAs and transposable elements (TEs), are of great interest to better understand the structure and dynamics of the genome. SatDNAs are one of the most abundant elements in the genome and their characterization based on low-coverage sequencing data has been published in the last few years for several animal9,10,12,13,14 but mostly for plant species8,11,25,26,27. Knowledge about the repeatome of parasitic species is generally limited. The data originate exclusively from genome projects that are primarily address the proportion of repetitive elements within specific species. Among tapeworms, species from the orders Cyclophyllidea and Diphyllobothriidea have been studied [Table 1 in28]. Notably, cyclophyllidean species have smaller genomes with fewer repetitive elements than diphyllobothridean species, with transposable elements predominating28. In trematodes, genome sequencing projects have shown that repetitive elements are generally substantial. The genome of Schistosoma mansoni contains about 45% repetitive elements, Fasciola hepatica 35% and Clonorchis sinensis about 30% [Supplementary Table S2 in29]. As for the parasites from the Acanthocephala group, as mentioned in the introduction, there is a study that marginally deals with the repetitive content. The first genome assembly by Mauer et al.6 revealed that repetitive elements make up a significant proportion (63%) of the draft genome of P. laevis. However, there are currently no data on the organization of satDNA repeat units in parasite species. Our study provides the first such analysis in parasites on three species of the genus Acanthocephalus.
RepeatExplorer2 analysis of A. anguillae, A. lucii and A. ranae revealed several interesting differences between the species and sexes examined. The majority of the identified repeats were annotated as satDNA, with A. anguillae having the highest proportion (13.34% of analyzed reads in female, 15.75% in male), followed by A. lucii (9.87% in female, 13.75% in male) and A. ranae (8.77% in female, 10.27% in male). The increased proportion of satDNA in A. anguillae correlates with our previous results obtained after DAPI staining which detected AT-rich heterochromatin. In A. anguillae17, distinct blocks were found in the centromeric regions of all chromosomes and also in the pericentromeric region on the long arms of chromosome pair No. 2, whereas in A. lucii18 only weak centromeric heterochromatin bands were found and in A. ranae, barely perceptible blocks were found. In addition, the analysis revealed differences in the proportion of reads annotated as rDNAs, with the predominant proportion of aligned reads also originating from the A. anguillae genome (2.15% in A. anguillae, 1.03% in A. lucii and 0.92% in A. ranae). This situation could be related to the different number of rDNA loci in the karyotypes of A. anguillae (three) and A. lucii (one)17,18. Finally, A. anguillae showed a significant dominance of a specific cluster annotated as satDNA, which comprised about 9% of the reads. A similar dominance of one repeat class was also observed in the club-legged grasshopper Gomphocerus sibiricus30. Shah et al.30 proposed that the dominance of this repeat class may indicate a recent dispersal within this species or in recent ancestor, as an ancient expansion would likely have caused the repeat sequences to diverge by mutation, resulting in them assembling into multiple clusters. The significant interspecific differences in satDNAs between the studied species indicate a high satDNA diversity in the genomes of Acanthocephalus species. SatDNA sequences can differ in a number of characteristics, e.g. in nucleotide composition, distribution and abundance in the genomes of even closely related species. It is assumed that related species share an ancestral set of conserved satDNAs families, the “library”, each of which may undergo different amplification in the species5,10,12,31,32,33. TEs can play an important role in the spread of satDNAs throughout the genome. Once a satDNA family is spread throughout the genome, the sequences undergo a process known as “intragenomic concerted evolution”. This leads to a more efficient homogenization of repeats within species than between species and also of repeats located in the same array/chromosome than between different ones34,35. New variants arising from mutations in single units are extended to the remaining repeats by various mechanisms such as unequal crossing-over, gene conversion or transposition, which together with gene expression contribute to the homogenization of satDNAs. Consequently, the newly expanded variants are fixed in a population/species through sexual reproduction36,37.
The karyotypes of A. anguillae and A. lucii are asymmetric17,18. Their mitotic karyotype shows a diploid set of 2n = 7/8 chromosomes, consisting of one (male) or two (female) acrocentric X chromosomes and two pairs of metacentric and one pair of acrocentric autosomes in A. anguillae and one pair of metacentric and two pairs of submetacentric autosomes in A. lucii. As for the karyotype of A. ranae, the same diploid number but a completely different morphology was observed, represented only by metacentric chromosomes24. As mentioned in the introduction, the distribution of the two chromosomal markers, the major rDNA and the histone H3 genes, was analyzed by FISH mapping. In A. anguillae, three 18S rRNA loci per haploid genome were observed, one on chromosome pair No.1 and two on the second chromosome pair. Mapping of histone H3 genes revealed a scattered distribution on all chromosomes, including the B chromosomes17. In contrast, in A. lucii there was a single locus for both markers, with the rRNA genes on the short arms of the X chromosomes and the histone H3 genes on the long arms in the centromeric region of chromosome pair No. 118. Outside the genus Acanthocephalus, the distribution of major rDNA genes was studied in two species, Pomphorhynchus laevis and P. tereticollis, again revealing a distinct, species-specific pattern was also found16. It is therefore evident that the evolution of the karyotype of acanthocephalan species is accompanied by frequent changes in the distribution of cytogenetic markers, suggesting frequent chromosomal rearrangements. The significant diversity of satDNA in the Acanthocephalus species studied could facilitate these rearrangements of the karyotype. Chromosomal rearrangements may involve highly repetitive DNA sequences that can provide sites for karyotype reshuffling without compromising the integrity of the coding sequences38,39.
Interestingly, a higher number of repeats and reads annotated as satDNA was observed in the male genomes of all species studied (Table 1), which was contrary to our expectations. Acanthocephalan species are typically known for an XX/X0 sex determination mechanism, where females have two X chromosomes and males have only one17. However, in an earlier study, John24 proposed that A. ranae has an XY sex determination mechanism, with males having XY chromosomes and females having XX chromosomes. In our study, we did not observe the Y chromosome, and the sex determination in A. ranae is consistent with that proposed in other acanthocephalan species (XX/X0)[Supplementary Table S1 in 17]. The observed differences in the amount of satDNA between sexes may be attributed to an unequal proportion of satDNA between the X chromosome and the autosomes. If the percentage of satDNA in the X chromosome is higher than in the autosomes, the total percentage of satDNA in the genome would increase in females (with two X chromosomes). Conversely, if the satDNA content in the X chromosome is lower than in the autosomes, the percentage of satDNA in the whole genome would be lower in females compared to males, as is this case. This finding is in congruence with the results of Orosová et al.17, which demonstrated that the amount of heterochromatin in the X chromosome is lower than in the autosomes in A. anguillae. The recombination rate in the sex chromosomes in XX/X0 systems is lower than in autosomes, as there is no recombination in males. This may lead to a slower homogenization process of satellite DNA in the X chromosome, which in turn leads to the formation of specific satDNA in the sex chromosome. The accumulation of repetitive DNAs is therefore a common feature of sex chromosome evolution.
Species-specific chromosome markers
One of the aims of this study was to identify the most abundant species-specific satellite repeats that could potentially serve as novel chromosomal markers for each species. The high sequence homology of certain satDNA clusters allows the development of fluorescently labeled probes to study their distribution using molecular cytogenetic techniques. Species-specific distribution patterns of satellites could be of great value in distinguishing morphologically similar species and in identifying specific chromosomes and homologous chromosome pairs within karyologically challenging species40,41. Acanthocephalus anguillae and A. lucii are species with great morphological similarity and the ability to parasitize the same host species42. From a karyological point of view, they have very small chromosomes with ambiguous morphology, which complicates the process of homologous pairing. The karyotype of A. ranae has not yet been analyzed using modern molecular cytogenetic techniques. In our work, we successfully mapped several species-specific satellites on the chromosomes of each species. Each mapped satellite showed a clustered distribution pattern in a few loci located in the centro/pericentromeric, telo-/subtelomeric regions or intercalary. The inclusion of these new species- and/or chromosome-specific markers and the selection of the correct specific marker set is a valuable tool for the accurate identification of chromosomes and chromosome pairs. The inclusion of these new species- and/or chromosome-specific markers and the selection of the correct specific marker set is a very valuable tool for the accurate identification of chromosomes and chromosome pairs. For A. anguillae, we recommend the use of the markers 18S rDNA, AAc72 and AAc31. The first two chromosome pairs can be distinguished by the distribution of the major rDNA, as they differ in the number of loci17. An additional and more precise identification of chromosome pair No. 1 can be achieved by the marker AAc72. Finally, the satellite AAc31 is specific for the X chromosomes, so that chromosomes without a marker would correspond to pair No. 3. In the case of A. ranae, the chromosomes can be identified using different sets of newly identified markers, each specific to a particular pair of chromosomes. The chromosome pair No. 1 can be identified by the satellites ARc42 or ARc51, the second chromosome pair by the satellites ARc39 or ARc27 and the sex chromosomes by the satellites ARc49 and ARc58. Interestingly, as in A. anguillae, the third chromosome pair lacks any satellite marker. Finally, only two specific markers were successfully mapped in A. lucii. The marker ALc1 is specific for the centromeric region of all chromosomes, and ALc98 is specific for chromosome pairs No. 1, 2 and 3. These markers alone are insufficient for accurate chromosome identification, and the inclusion of recently mapped cytogenetic markers, namely 18S rDNA and histone H3 is necessary18. With these two additional markers, the identification of chromosome pair No. 1 and sex chromosomes can be achieved. In addition to being cytogenetic markers, these satDNAs can also serve as molecular markers. Specific primers can be easily designed based on these satellites, enabling species identification through PCR analysis.
Supernumerary “B” chromosomes
Supernumerary chromosomes were observed in the karyotypes of A. anguillae and A. lucii from the Zemplínska Šírava reservoir, a phenomenon already documented in our previous studies17,18. B chromosomes, which are present in addition to the standard A chromosomes, occur in about 15% of eukaryotes43. Typically, they are heterochromatic and rich in repetitive DNA sequences, including satDNAs; several classes of TEs and multigene families44,45,46,47. Conversely, certain species have a low proportion of repetitive DNA and heterochromatin regions on B chromosomes. Few heterochromatin blocks were observed in the fish species Moenkhausia bonita48, and similarly, bioinformatic and cytogenetic analyzes of the South American locust Abracris flavolineata showed that its B chromosome is not enriched with high copy repetitive DNA sequences, only a limited occurrence was detected, and no B-specific satDNA was identified. The authors suggested that the B chromosome in A. flavolineata represents a young element in the early stages of heterochromatization49. Minimal enrichment of AT- or GC-rich heterochromatin on B chromosomes was observed in A. anguillae17 and A. lucii18 after differential staining techniques. FISH revealed the presence of histone H3 genes at several loci on the B chromosomes of A. anguillae, but only faint hybridization signals were detected. In the present work, no satDNAs were mapped on B chromosomes, confirming the low content of repeated sequences in supernumerary chromosomes. In our previous work, we hypothesized that B chromosomes in Acanthocephalus species may have evolved recently in the context of environmental pollution, and the results of the present work appear to confirm our hypothesis. However, it is important to point out that our results concerning B chromosomes are still preliminary. More extensive studies are needed to unravel the mystery of the origin of B chromosomes.
The present study provides a first insight into the satellitome of three endoparasite species. By analyzing the repetitive content, we were able to determine the prevalence of satDNA in their genomes as well as the intersexual and interspecific variability in their abundance. Most importantly, we have identified a high number of species-specific satDNA families, some of which have proven to be valuable cytogenetic markers for the identification of homologous chromosomes. However, it is essential to conduct further and more complex studies on different species to fully understand the functional and structural features of their genomes, especially in species with B chromosomes. In addition, such studies are essential to clarify the pathways of chromosomal and karyotypic reorganization during speciation.
Methods
Ethics
The fish were handled in accordance with the relevant guidelines and regulations (Decree of the Ministry of the Slovak Republic No. 381/2018 Coll. and Act No. 216/2018 Coll. on Fisheries) and we confirm that all methods are reported in accordance with the ARRIVE guidelines50. The fish were brought to the laboratory in aerated tanks with the same water as in the home tank and were killed by severing the spinal cord just behind the head after anesthesia with clove oil–water solution (Sigma-Aldrich). Prior to necropsy, the frogs were euthanized in the chamber with clove oil–water emulsion for anesthesia, approximately 20 drops of clove oil (Sigma-Aldrich) to one liter of water, and their spinal cord was severed to execute one animal. All procedures were reviewed and approved by the Ethics Committee of the Institute of Parasitology of the Slovak Academy of Sciences.
Study site and parasite material
The studied specimens of A. anguillae originated from chub (Squalius cephalus) and the studied specimens of A. lucii were from perch (Perca fluviatilis) caught in March and April 2023 in the Zemplínska Šírava reservoir (ZŠ; 48°47′09.0″N 21°57′20.5″E), a site known for heavy pollution with polychlorinated biphenyls (PCB)51. The fishing was conducted under permit No. 48/2023, issued by the Ministry of Environment of the Slovak Republic. The analyzed specimens of A. ranae were collected from frogs caught in two locations in May and June 2023 with a permit from the Ministry of Environment of the Slovak Republic (No. 519/2022–6.3). A total of 20 frog hosts were caught: 10 Pelophylax ridibundus from Veľký Lel (47°45′35.4 "N 17°56′41.0 "E) and 10 P. esculentus from Rusovce (48°03′26.8 "N 17°09′11.2 "E). The fish and frog hosts were dissected on the spot or transported to the laboratory and processed. Intestines were removed from the abdominal cavity, placed in 0.9% saline solution and examined for presence of acanthocephalans.
A total of 111 parasites were collected: 25 A. anguillae (10 males and 15 females), 48 A. lucii (20 males and 28 females), and 38 A. ranae (21 males and 17 females). All collected parasites were rinsed in 0.9% saline immediately after isolation from the host intestine. The individual species were identified microscopically by counting the hooks and hook rows and by the shape of the proboscis hook roots42. At the same time, several individuals of the collected worms (at least five males and five females from each species) were fixed in 100% ethanol for molecular analysis and the species identification was confirmed by sequencing the PCR-amplified 18S rDNA fragment.
Chromosome preparations
Live worms were placed in a 0.025% colchicine solution for one hour at room temperature (RT, 25 °C), followed by hypotonic treatment with 0.075 M KCl. Whole intact females were placed in the solution for 4–5 h at RT, while the testes were dissected from the males and only these were incubated for 20 min at RT. Finally, tissues were fixed in freshly mixed modified Carnoy’s fixative (methanol/acetic acid = 3:1 v/v) in two changes (15 min) and stored at − 20 °C until further use. Chromosome preparations were made using the “hot plate” technique52. Slides were inspected with phase-contrast optics and those of sufficient quality were dehydrated in an ethanol series (70, 80 and 100%, 1 min each), dried at RT and stored at − 20 °C for further use.
DNA isolation and sequencing
High molecular weight genomic DNA (HMW gDNA) was isolated from females and males of each species using the cetyltrimethylammonium bromide (CTAB) method53. The samples of A. anguillae and A. lucii from which the DNA was isolated contained supernumerary chromosomes. The concentration of the isolated HMW gDNA was quantified using the Qubit 4.0 fluorometer (Invitrogen, Carlsbad, USA) and the quality was assessed using the Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). A sample of optimal quality was selected for each species to be examined. A total of 20 µg of gDNA were used for sequencing at Novogene (HK) Co, Ltd (Hong Kong, China) using the HiSeq 4000 platform. The low-coverage sequencing yielded approximately 1.4 Gb of 150 bp paired-end reads for each species.
Data processing and satDNA identification
The quality assessment of the NGS data was checked using FastQC version 0.10.154 and trimming was performed using Trimmomatic version 0.3255. During trimming, Illumina sequencing adapters and reads with low quality were removed and all reads were trimmed to a final length of 105 bp. The quality and accuracy of the trimming was checked again with FastQC. Subsequent analyzes were performed using the RE pipeline15,19, including TAREAN analysis within the Galaxy portal (https://repeatexplorer-elixir.cerit-sc.cz). The first step was to run the pipeline separately for each species and sex. A sample of 500,000 paired reads was analyzed using the default options and the Metazoa version 3.0 database for automatic annotation. We then performed a comparative analysis between males and females of each species. For this analysis, concatenated files containing a random sample of 700,000 reads from each sex were used. Finally, for the comparative analysis between the studied species, the trimmed sequences were tagged with species-specific read name prefixes and suffixes and then concatenated in a single fasta file. This file was analyzed using RE with the default parameters, except for an extended computing time (extra-long). The annotation of the repeats was performed by the automatic annotation of RE using the Metazoa version 3.0 database. Based on the results of RE, we selected several species-specific satellites for each species that were best suited for physical mapping on chromosomes. In addition, the tool RepeatMasker (https://www.repeatmasker.org/) with default settings and the option “-a” to preserve alignment was used to calculate the divergence of each satDNA. As a custom library, we used a concatenation of dimers for those repeats with a repetition unit above 100 bp and at least a concatenation of 200 bp for those satDNAs with a repetition unit below 100 bp. Then we mapped back the custom library to a total of 1,000,000 randomly selected reads from each species. The output of RepeatMasker was used to calculate the divergence for the repeats by applying the Kimura 2-parameter model (K-2p) with the Perl script calcDivergenceFromAlign.pl. Subsequently, the satDNA landscapes were generated using a custom script in RStudio (v 4.1.0) with the ggplot2 package (v 3.4.4)56.
Preparation and labeling of probes and FISH analysis
The consensus sequences of selected satellites were used to design primer sets (Tables 2, 3) using the Geneious Prime software version 2021.1.1 and the Primer3 tool version 4.1.0. SatDNA repeats were amplified in 25 µl PCR mix containing 1 µM of each primer, 10 ng HMW gDNA and 12.5 µl DreamTaq Green PCR Master Mix (2X). The following PCR program was used: initial denaturation at 95 °C for 3 min, followed by 35 cycles of 95 °C for 30 s, from 45 to 60 °C for 30 s (depending on repeat, Table 2) and 72 °C 90 s; the final extension was at 72 °C for 5 min. The results were checked on a 1% standard agarose gel in 1 × TAE buffer. The amplified PCR products of repeats with a genomic proportion below 0.8% were labeled in the PCR reaction with biotin-16-dUTP or digoxigenin-11-dUTP (both Roche Diagnostics, Mannheim, Germany). For repeats with a genomic proportion above/around 1%, the oligonucleotide sets (Table 3) were labeled directly with biotin-16-dUTP or digoxigenin-11-dUTP using terminal transferase (Roche Diagnostics, Mannheim, Germany) following the manufacturer’s protocol.
The labeled PCR products or oligonucleotides were used as probes in single or two-color FISH according to the protocol described in Cabral-de-Mello and Marec57. FISH requires the design of probes that are complementary to the satDNA of interest. The efficiency of this hybridization depends on how well the probes match the satellite sequence, which can be a challenge if the satDNA has significant sequence variation within or between species. The hybridization solutions were mixed at a final concentration of 5 ng/mL in 50% formamide. The denaturation step was performed at 68 °C for 3 min and 30 s, followed by hybridization at 37 °C overnight in a moist chamber. All stringent washes, including 2 × SSC (2 × 5 min), 0.1 × SSC (2 × 5 min), 2 × SSC (1 × 5 min), and WBB (3 × 5 min), were modified to RT. Conditions of FISH experiments ensure about 80–85% of homology between the probe and the target sequence. Biotinylated probes were detected using the Cy3-conjugated streptavidin (Jackson ImmunoRes. Labs. Inc., West Grove, PA, USA) / biotinylated anti-streptavidin (Vector Labs. Inc., Burlingame, CA, USA) system with three rounds of amplification. DIG-labeled probes were detected with anti-Digoxigenin-FITC (Sigma-Aldrich). Slides were counterstained with DAPI in Vectashield Antifade solution (Vector Laboratories, Burlingame, CA, USA). The stained slides were analyzed using an Olympus BX51 fluorescence microscope (Olympus, Hamburg, Germany) equipped with a CCD camera (Olympus DP70) and a LEICA DM 4000 B combined light and fluorescence microscope equipped with a DFC 450 C digital camera. The fluorescence images were pseudo-colored, merged to combine chromosome and probe signals, and optimized for brightness and contrast using Adobe Photoshop version 7.0.
Data availability
DNA sequence data that supports the findings was deposited in Sequence Read Archive (SRA) under the BioProject accession no. ID PRJNA1171452. All other relevant data are within the paper.
References
Biscotti, M. A., Olmo, E. & Heslop-Harrison, J. S. Repetitive DNA in eukaryotic genomes. Chromosome Res. 23, 415–420. https://doi.org/10.1007/s10577-015-9499-z (2015).
Wei, K. H. C. et al. Variable rates of simple satellite gains across the Drosophila phylogeny. Mol. Biol. Evol. 35, 925–941. https://doi.org/10.1093/molbev/msy005 (2018).
Plohl, M., Meštrović, N. & Mravinac, B. Satellite DNA evolution. Genome Dyn. 7, 126–152. https://doi.org/10.1159/000337122 (2012).
Dodsworth, S. et al. Genomic repeat abundances contain phylogenetic signal. Syst. Biol. 64, 112–126. https://doi.org/10.1093/sysbio/syu080 (2015).
Belyayev, A. et al. Natural history of a satellite DNA family: from the ancestral genome component to species-specific sequences, concerted and non-concerted evolution. Int. J. Mol. Med. 20, 1201. https://doi.org/10.3390/ijms20051201 (2019).
Mauer, K. et al. The genome, transcriptome, and proteome of the fish parasite Pomphorhynchus laevis (Acanthocephala). PLoS ONE. 15, e0232973. https://doi.org/10.1371/journal.pone.0232973 (2020).
Pita, S. et al. Comparative repeatome analysis on Triatoma infestans Andean and Non-Andean lineages, main vector of Chagas disease. PLoS ONE. 12, e0181635. https://doi.org/10.1371/journal.pone.0181635 (2017).
Muravenko, O. V. et al. Integration of repeatomic and cytogenetic data on satellite DNA for the genome analysis in the genus Salvia (Lamiaceae). Plants. 11, 2244. https://doi.org/10.3390/plants11172244 (2022).
Cuadrado, Á. et al. Contribution of the satellitome to the exceptionally large genome of dinoflagellates: The case of the harmful alga Alexandrium minutum. Harmful algae. 130, 102543. https://doi.org/10.1016/j.hal.2023.102543 (2023).
Mora, P. et al. Making the genome huge: The case of Triatoma delpontei, a Triatominae species with more than 50% of its genome full of satellite DNA. Genes. 14, 371. https://doi.org/10.3390/genes14020371 (2023).
Samatadze, T. E. et al. Genome studies in four species of Calendula L. (Asteraceae) using satellite DNAs as chromosome markers. Plants. 12, 4056. https://doi.org/10.3390/plants12234056 (2023).
Voleníková, A. et al. Fast satellite DNA evolution in Nothobranchius annual killifishes. Chromosome Res. 31, 33. https://doi.org/10.1007/s10577-023-09742-8 (2023).
Cabral-de-Mello, D. C. et al. The spread of satellite DNAs in euchromatin and insights into the multiple sex chromosome evolution in Hemiptera revealed by repeatome analysis of the bug Oxycarenus hyalinipennis. Insect Mol. Biol. 32, 725–737. https://doi.org/10.1111/imb.12868 (2023).
Rico-Porras, J. et al. Heterochromatin is not the only place for satDNAs: The high diversity of satDNAs in the euchromatin of the beetle Chrysolina americana (Coleoptera, Chrysomelidae). Genes. 15, 395. https://doi.org/10.3390/genes15040395 (2024).
Novák, P., Neumann, P. & Macas, J. Global analysis of repetitive DNA from unassembled sequence reads using RepeatExplorer2. Nat. Protoc. 15, 3745–3776. https://doi.org/10.1038/s41596-020-0400-y (2020).
Bombarová, M., Marec, F., Nguyen, P. & Špakulová, M. Divergent location of ribosomal genes in chromosomes of fish thorny-headed worms, Pomphorhynchus laevis and Pomphorhynchus tereticollis (Acanthocephala). Genetica. 131, 141–149. https://doi.org/10.1007/s10709-006-9124-3 (2007).
Orosová, M., Marková, A., Zrzavá, M., Marec, F. & Oros, M. Chromosome analysis and the occurrence of B chromosomes in fish parasite Acanthocephalus anguillae (Palaeacanthocephala: Echinorhynchida). Parasite. 30, 44. https://doi.org/10.1051/parasite/2023045 (2023).
Marková, A., Orosová, M., Marec, F., Barčák, D. & Oros, M. Karyological study of Acanthocephalus lucii (Echinorhynchida): The occurrence of B chromosomes in populations from PCB-polluted waters. Diversity. 16, 140. https://doi.org/10.3390/d16030140 (2024).
Novák, P. et al. TAREAN: a computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res. 45, e111. https://doi.org/10.1093/nar/gkx257 (2017).
Bolsheva, N. L. et al. Characterization of repeated DNA sequences in genomes of blue-flowered flax. BMC Evol. Biol. 19, 49. https://doi.org/10.1186/s12862-019-1375-6 (2019).
Amosova, A. V. et al. Repeatome analyses and satellite DNA chromosome patterns in Deschampsia sukatschewii, D. cespitosa, and D. antarctica (Poaceae). Genes. 13, 762. https://doi.org/10.3390/genes13050762 (2022).
Ruiz-Ruano, F. J., Navarro-Domínguez, B., Camacho, J. P. M. & Garrido-Ramos, M. A. Characterization of the satellitome in lower vascular plants: The case of the endangered fern Vandenboschia speciosa. Ann. Bot. 123, 587–599. https://doi.org/10.1093/aob/mcy192 (2018).
Lorite, P., Carrillo, J. A., Tinaut, A. & Palomeque, T. Evolutionary dynamics of satellite DNA in species of the genus Formica (Hymenoptera, Formicidae). Gene. 332, 159–168. https://doi.org/10.1016/j.gene.2004.02.049 (2004).
John, B. The chromosomes of zooparasites I. Acanthocephalus ranae (Acanthocephala: Echinorhynchidae). Chromosoma. 8, 730–738 (1957).
Pamponét, V. C. C. et al. Low coverage sequencing for repetitive DNA analysis in Passiflora edulis Sims: Citogenomic characterization of transposable elements and satellite DNA. BMC Genom. 20, 262. https://doi.org/10.1186/s12864-019-5576-6 (2019).
Luo, X., Chen, S. & Zhang, Y. PlantRep: A database of plant repetitive elements. Plant Cell Rep. 41, 1163–1166. https://doi.org/10.1007/s00299-021-02817-y (2022).
Argentin, J., Bolser, D., Kersey, P. J. & Flicek, P. Comparative analysis of repeat content in plant genomes, large and small. Front. Plant. Sci. 14, 1103035. https://doi.org/10.3389/fpls.2023.1103035 (2023).
Kamenetzky, L., Maldonado, L. L. & Cucher, M. A. Cestodes in the genomic era. Parasitol. Res. 121, 1077–1089. https://doi.org/10.1007/s00436-021-07346-x (2021).
Soloyeva, A. et al. Transposons-based clonal diversity in trematode involves parts of CR1 (LINE) in eu- and heterochromatin. Genes. 12, 1129. https://doi.org/10.3390/genes12081129 (2021).
Shah, A., Hoffman, J. I. & Schielzeth, H. Comparative analysis of genomic repeat content in Gomphocerine grasshoppers reveals expansion of satellite DNA and helitrons in species with unusually large genomes. Genome Biol. Evol. 12, 1180–1193. https://doi.org/10.1093/gbe/evaa119 (2020).
Meštrović, N., Plohl, M., Mravinac, B. & Ugarković, D. Evolution of satellite DNAs from the genus Palorus-Experimental evidence for the “library” hypothesis. Mol. Biol. Evol. 15, 1062–1068. https://doi.org/10.1093/oxfordjournals.molbev.a026005 (1998).
Meštrović, N., Castagnone-Sereno, P. & Plohl, M. Interplay of selective pressure and stochastic events directs evolution of the MEL172 satellite DNA library in root-knot nematodes. Mol. Biol. Evol. 23, 2316–2325. https://doi.org/10.1093/molbev/msl119 (2006).
Ruiz-Ruano, F. J., López-León, M. D., Cabrero, J. & Camacho, J. P. M. High-throughput analysis of the satellitome illuminates satellite DNA Evolution. Sci. Rep. 6, 28333. https://doi.org/10.1038/srep28333 (2016).
Plohl, M., Luchetti, A., Meštrović, N. & Mantovani, B. Satellite DNAs between selfishness and functionality: Structure, genomics and evolution of tandem repeats in centromeric (hetero)chromatin. Gene. 409, 72–82. https://doi.org/10.1016/j.gene.2007.11.013 (2008).
Palomeque, T. & Lorite, P. Satellite DNA in insects: A review. Heredity. 100, 564–573. https://doi.org/10.1038/hdy.2008.24 (2008).
Dover, G. Molecular drive. Trends Genet. 18, 587–589. https://doi.org/10.1016/s0168-9525(02)02789-0 (2002).
Garrido-Ramos, M. A. Satellite DNA: An evolving topic. Genes. 8, 230. https://doi.org/10.3390/genes8090230 (2017).
Hsu, T. C., Pathak, S., Basen, B. M. & Stahl, G. J. Induced Robertsonian fusions and tandem translocations in mammalian cell cultures. Cytogenet. Cell. Genet. 21, 86–98 (1978).
Singer, D. & Downhower, L. Highly repeated DNA of the baboon: organization of sequences homologous to highly repeated DNA of the African green monkey. J. Mol. Biol. 134, 835–842 (1979).
Hasterok, R. & Maluszynska, J. Cytogenetic markers of Brassica napus L. chromosomes. J. Appl. Genet. 41, 1–9 (2000).
Yoshido, A., Bando, H., Yasukochi, Y. & Sahara, K. The Bombyx mori karyotype and the assignment of linkage groups. Genetics. 170, 675–685. https://doi.org/10.1534/genetics.104.040352 (2005).
Lisitsyna, O. Acanthocephala in Fauna of Ukraine (Kiev, 2019).
Camacho, J. P. M. B Chromosomes in The evolution of the genome (ed. Gregory, T. R.) 223–286 (Elsevier Academic Press, 2005).
Hanlon, S. L., Miller, D. E., Eche, S. & Hawley, R. S. Origin, composition, and structure of the supernumerary B chromosome of Drosophila melanogaster. Genetics. 210, 1197–1212. https://doi.org/10.1534/genetics.118.301478 (2018).
Marques, A., Klemme, S. & Houben, A. Evolution of plant B chromosome enriched sequences. Genes. 9, 515. https://doi.org/10.3390/genes9100515 (2018).
Ruiz-Ruano, F. J., Cabrero, J., López-León, M. D., Sánchez, A. & Camacho, J. P. M. Quantitative sequence characterization for repetitive DNA content in the supernumerary chromosome of the migratory locust. Chromosoma. 127, 45–57. https://doi.org/10.1007/s00412-017-0644-7 (2018).
Stornioli, J. H. F. et al. The B chromosomes of Prochilodus lineatus (Teleostei, Characiformes) are highly enriched in satellite DNAs. Cells. 10, 1527. https://doi.org/10.3390/cells10061527 (2021).
Ranucci, L., Fernandes, C. A., Borin-Carvalho, L. A., Martins-Santos, I. C. & de Portela-Castro, A. L. B. Occurrence of euchromatic B chromosomes in natural populations of Moenkhausia bonita and M. forestii (Pisces: Characidae). Neotrop. Ichthyol. 19, e210056. https://doi.org/10.1590/1982-0224-2021-0056 (2021).
Milani, D., Ruiz-Ruano, F. J. & Camacho, J. P. M. Out of patterns, the euchromatic B chromosome of the grasshopper Abracris flavolineata is not enriched in high-copy repeats. Heredity. 127, 475–483. https://doi.org/10.1038/s41437-021-00470-5 (2021).
Percie du Sert, N. et al. The ARRIVE guidelines 2.0: updated guidelines for reporting animal research. PLoS Biol. 14, e3000410. https://doi.org/10.1371/journal.pbio.3000410 (2022).
Šalgovičová, D. & Zmetáková, Z. Polychlorinated biphenyls is in muscle tissue of freshwater fish in east Slovakia. J. Food Nutr. Res. 45, 171–178 (2006).
Orosová, M. & Špakulová, M. Tapeworm chromosomes: Their value in systematics with instructions for cytogenetic study. Folia Parasit. 65, 1. https://doi.org/10.14411/fp.2018.001 (2018).
Winnepenninckx, B., Backeljau, T. & De Wachter, R. Extraction of high molecular weight DNA from molluscs. Trends Genet. 9, 407. https://doi.org/10.1016/0168-9525(93)90102-n (1993).
Andrews, S. FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics. 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Wickham, H. Elegant graphics for data analysis in Ggplot2. (Springer Cham, 2016).
Cabral-de-Mello, D. C. & Marec, F. Universal fluorescence in situ hybridization (FISH) protocol for mapping repetitive DNAs in insects and other arthropods. Mol. Genet. Genom. 296, 513–526. https://doi.org/10.1007/s00438-021-01765-2 (2021).
Acknowledgements
This work was funded by the Scientific Grant Agency of the Ministry of Education, Science, Research and Sport of the Slovak Republic and the Slovak Academy of Sciences (VEGA No. 2/0093/23) and Slovak Academy of Sciences grants for PhD students (APP0452).
Author information
Authors and Affiliations
Contributions
A.M. Methodology, Investigation, Formal analysis, Resources, Writing—Original Draft, Writing—Review & Editing. M.O. Project administration, Methodology, Resources, Conceptualization, Investigation, Funding acquisition, Supervision, Validation, Writing—Review & Editing. P.M. Formal analysis, Writing—Review & Editing. M.B. Resources. P.L. Resources, Writing—Review & Editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Marková, A., Orosová, M., Mora, P. et al. The first insight into Acanthocephalus (Palaeacanthocephala) satellitome: species-specific satellites as potential cytogenetic markers. Sci Rep 15, 2945 (2025). https://doi.org/10.1038/s41598-025-85728-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85728-2
Keywords
This article is cited by
-
Discovery of new chromosomal markers through repeatome analysis of Caryophyllaeus laticeps (Caryophyllidea)
Parasitology Research (2025)