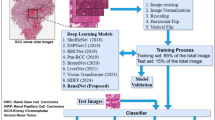
Abstract
Recently, as the number of cancer patients has increased, much research is being conducted for efficient treatment, including the use of artificial intelligence in genitourinary pathology. Recent research has focused largely on the classification of renal cell carcinoma subtypes. Nonetheless, the broader categorization of renal tissue into non-neoplastic normal tissue, benign tumor and malignant tumor remains understudied. This gap in research can primarily be attributed to the limited availability of extensive datasets including benign tumor and normal tissue in addition to specific type of renal cell carcinoma, which hampers the ability to conduct comprehensive studies in these broader categories. This research introduces a model aimed at classifying renal tissue into three primary categories: normal (non-neoplastic), benign tumor, and malignant tumor. Utilizing digital pathology while slide images (WSIs) from nephrectomy specimens of 2,535 patients from multiple institutions, the model provides a foundational approach for distinguishing these key tissue types. The study utilized a dataset of 12,223 WSIs comprising 1,300 WSIs of normal tissue, 700 WSIs of benign tumors, and 10,223 WSIs of malignant tumors. Employing the ResNet-18 architecture and a Multiple Instance Learning approach, the model demonstrated high accuracy, with F1-scores of 0.934 (CI: 0.933–0.934) for normal tissue, 0.684 (CI: 0.682–0.687) for benign tumors, and 0.878 (CI: 0.877–0.879) for malignant tumors. The overall performance was also notable, achieving a weighted average F1-score of 0.879 (CI: 0.879–0.880) and a weighted average area under the receiver operating characteristic curve of 0.969 (CI: 0.969–0.969). This model significantly aids in the swift and accurate diagnosis of renal tissue, encompassing non-neoplastic normal tissue, benign tumor, and malignant tumor.
Similar content being viewed by others
Introduction
In recent years, the United States has witnessed a growing incidence of renal cancer, attributed to factors such as aging, obesity, and the prevalence of metabolic syndrome1. In 2019, approximately 73,820 new cases were identified, accounting for 4.2% of all cancer cases nationwide2. The accurate and swift diagnosis of renal tumors, recognizing the broad spectrum of their histological presentations, is vital for making informed treatment decisions3. Consequently, artificial intelligence (AI) technologies have risen to prominence, playing a crucial role in refining the precision of diagnoses, aiding in the prediction of prognoses, and reducing the time required for diagnosis4,5,6,7,8. In genitourinary pathology, the diagnostic evaluation of renal cell carcinoma (RCC) involves multiple critical factors that guide treatment and prognosis. Pathologists not only classify tissues as benign, malignant, or non-neoplastic, but also focus on essential details such as histological subtype, grade, and stage9,10. Identifying the RCC subtype—such as clear cell, papillary, or chromophobe—is fundamental, as each subtype exhibits distinct clinical behaviors and therapeutic responses, making precise classification vital for patient management10. Tumor grade, assessed through the WHO/ISUP grading system, further aids in understanding tumor aggressiveness, with higher grades (G3/G4) typically indicating more aggressive disease9,10. Staging, based on the TNM classification, provides additional insights into tumor progression, including the size of the tumor, lymph node involvement, and the presence of metastasis11. For instance, tumor invasion into perinephric fat or the renal vein can significantly impact treatment strategies and prognosis. Furthermore, the evaluation of histological features such as lymphovascular invasion, tumor necrosis, and sarcomatoid or rhabdoid differentiation, as outlined in the College of American Pathologists (CAP) protocol, is crucial9. These features provide deeper insights into the tumor’s behavior and further refine clinical decision-making, ensuring personalized and effective treatment approaches. This diagnostic complexity illustrates that pathologists’ tasks extend well beyond a simple classification of tissue types and into nuanced decisions that affect staging, treatment strategies, and patient prognosis. However, inherent challenges remain in analyzing complex histopathological images, often compounded by the availability of pathologists12,13,14. Leveraging AI as a diagnostic aid can mitigate the impact of pathologist shortages by transforming complex histopathological image data into quantitative metrics—such as cell size, shape, and density—thereby enhancing the objectivity, precision, and accuracy of renal tumor diagnoses12,13,15,16,17.
AI has been already increasingly researched as a diagnostic aid in renal cell carcinoma classification for pathologists. Lu et al. (2022) utilized a federated learning framework to classify malignant kidney tumor subtypes, achieving F1-scores between 0.88 and 0.984. Wu et al. (2021) reported accuracies around 93% in distinguishing normal and malignant renal tissues using whole slide images (WSIs)5. Haeyeh et al. (2022) and Tabibu et al. employed ResNet architectures to classify renal cancer subtypes, achieving high accuracy and AUC scores above 0.956,7. Chanchal et al. developed a model to classify clear cell RCC grades with 90.14% accuracy8.While these studies show promising results using AI as a diagnostic tool, key challenges remain, such as small dataset sizes performed in single institution, narrow diagnostic focus, and ambiguity in model inference criteria.
In a national project with the National Information Society Agency (NIA), we collected a comprehensive kidney dataset by collecting extensive data from multiple tertiary care hospitals. This dataset includes high-quality labeling masks created by board-certified pathologists and labelers, ensuring superior quantity and quality compared to existing datasets. Our innovative research approach of developing a general-purpose model that includes various subcategories is founded on solving mentioned above problems through model training using this robust dataset. To our knowledge, this study is the first to utilize this nationwide project dataset. In response, our study develops a ResNet-18-based classification model that utilizes a comprehensive kidney dataset, encompassing benign and malignant tumors as well as non-neoplastic renal tissue, and applies the multiple instance learning (MIL) method in instance level. This approach is designed to address some of the limitations observed in previous research by offering a more generalized solution for renal tissue classification, supported by quantitative performance metrics.
Methods
Data collection
This study was conducted with the approval of the Institutional Review Board (IRB) of Seoul St. Mary’s Hospital, and the need for informed consent was waived due to the retrospective nature of the study (Approval Number: KC23RNDI0458). All experimental protocols were performed in accordance with the relevant guidelines and regulations of the Declaration of Helsinki. The data used in this research comprises kidney whole slide images (WSIs) collected from 2,535 patients who underwent either partial or radical nephrectomy for renal masses, or biopsy due to suspected kidney cancer, across five university hospitals. The dataset comprises a male-to-female ratio of 6:4, including 1,300 WSIs of normal (non-neoplastic) tissue, 700 WSIs of benign tumors, and 10,223 WSIs of malignant tumors. All pathology slides were digitally scanned at 20× magnification, using a Hamamatsu NanoZoomer S360 Digital Slide Scanner (Hamamatsu Photonics, Hamamatsu, Japan), Aperio GT450 (Leica Biosystems, Deer Park, IL, USA), and Pannoramic 250 Flash III (3DHISTECH, Budapest, Hungary). The data was assembled in 2023 as part of the Artificial Intelligence Learning Data Construction Support Project organized by the National Information Society Agency in Korea.
The collected raw WSIs were labeled as normal (non-neoplastic), benign tumor, and malignant tumor regions through the cloud-based MeDIAuto platform (Urbandatalab, Seoul, Korea) (Fig. 1). This process was performed directly by the labeler and confirmed by a pathologists. The pathologists who participated in the inspection have 26 and 11 years of experience, respectively, and have at least 10 years of sufficient field experience. For ambiguous cases, such as papillary adenoma and papillary carcinoma, where there may be differing opinions on collection criteria, a consensus was reached among the participating pathologists, and consistent standards were applied throughout the review process. The labeled images were divided into training, validation, and test sets at a ratio of 8:1:1, forming datasets of 9,723, 1,250, and 1,250 WSIs, respectively. The composition of the constructed dataset resulted in the training set containing 886 normal, 370 benign, and 8,467 malignant WSIs, the validation set consisted of 113 normal, 38 benign, and 1,099 malignant WSIs and the test set was 301 normal, 292 benign and 657 malignant WSIs. Between 2 and 8 WSIs were collected per patient, with careful attention to ensure that WSIs from the same patient were not used across the training, validation, and test sets. From the collected WSIs, we extracted and utilized 111,754 benign tiles, 255,646 normal tiles, and 267,033 malignant tiles. Although the number of normal WSIs was fewer, we could extract a sufficient number of normal tiles from the WSIs of both malignant and benign cases to ensure an adequate representation of normal tissue. Normal tissue shows minimal variation across cases and remains largely uniform in its histological features, making it morphologically similar between different patients. In contrast, malignant tumors exhibit significant variability, both across and within the various histologic subtypes of renal cell carcinoma. This diverse morphology in malignant cases requires a larger number of images to capture the full range of variability accurately. Therefore, the increased number of malignant WSIs reflects the need to accommodate the morphological heterogeneity of malignant renal tumors. In addition, the number of detailed diagnostic items for each grade and the nuclear grade according to WHO/ISUP standards for clear cell RCC and Papillary RCC are shown in the table and figure below (Table 1; Fig. 2).
Whole slide images of the kidney are annotated with polygons to illustrate three types of labeling: normal, benign tumor, and malignant tumor areas. (a) An image with annotations of only normal kidney tissue. (b) An example with annotations of both normal kidney and benign tumor regions. (c) and (d) Images with polygon annotations of normal kidney and malignant tumor areas, where (c) shows a papillary renal cell carcinoma (RCC) and (d) shows a clear cell RCC.
Cell nuclear grade of clear cell RCC and Papillary RCC for the entire dataset.
Research environment
The experiments in this study were conducted on a system consisting of NVIDIA GeForce RTX 4070 (NVIDIA, California, USA) GPU, Intel® Core™ i7-13700 F (Intel, California, USA) CPU, 32GB RAM, 3 TB storage, and Windows 11 Home operating system. Python (version 3.10.6) was used as the programming language, and Pytorch (version 2.0.1) was applied as the deep learning framework. For image preprocessing, the OpenCV-Python library (version 4.8.1.78) was used to identify and extract specific tissue areas and to divide images into tiles. To evaluate the performance of the developed model, quantitative indicators of classification performance were presented through calculation of the confusion matrix and AUC using the Scikit-learn library (version 1.3.2).
Data preprocessing
To mitigate the impact of noise such as air bubbles, pen marks, background, and tissue folding on learning and inference outcomes18, the Otsu algorithm was employed for noise removal19. By converting images to the Lab space and applying the Otsu Algorithm to the A channel representing red to green, areas exceeding the threshold were treated as tissue, while those below were set to zero, thus segmenting and highlighting tissue in the A channel color.
Due to the use of WSIs obtained from various scanners, each slide may differ in staining density, color, and resolution20. To address this, a color normalization process was implemented as a standardization method. First, a slide image from a reference scanner was selected, and a generative adversarial networks (GAN) model was trained with this image serving as the standard for color normalization of other slides. Color normalization through GANs involves a generator and a discriminator, repeatedly generating new images based on the provided image until the discriminator recognizes the generated image as a real21,22.
Based on the labeled areas, regions of each class were cropped and extracted from the WSIs. Tiling was then performed on the extracted areas by each class to collect tiles for learning by classes, and tiles containing more than 75% background, making them unidentifiable as belonging to a specific class, were excluded. Finally, the selected tiles were resized to a uniform size of 224 pixels by 224 pixels (Fig. 3).
Data preprocessing flowchart from noise removal in top left to tiling in bottom right.
Model and learning method
Among several candidate models, such as GoogleNet and AlexNet, we chose ResNet-18 as the backbone for our study. This decision was guided by its successful application in numerous previous studies, particularly in medical imaging, where it has demonstrated strong performance even with relatively limited datasets. The ResNet-18 model consists of 18 layers and employs skip connections and residual blocks to address the issue of vanishing gradients, which can occur as the model deepens, especially when training on large-scale datasets (Fig. 4).This architecture mitigates the gradient vanishing problem by adding the value from previous layers, enhancing the flow of gradients through the network23. A distinctive feature of our model compared to the original design is the inclusion of a rectified linear unit (ReLU) layer and a Dropout layer in fully connected (FC) layer. This addition enables the model to learn from a broader range of features and reduces overfitting24,25.
Basic ResNet-18 model structure.
To augment the data, we utilized the Albumentations library, applying a variety of transformations such as resizing, brightness adjustment, rotation, and flipping. These transformations increase the diversity of the data, thereby improving the model’s performance and robustness26.
To proceed with MIL learning, we preprocessed slides, then masks are extracted and tiled. Each tile is treated as an instance in MIL learning and is trained by our Resnet-18 model. In the inference, the tiles extracted from the slide are used to construct a bag of instances containing tiles from whole slides, and the inference of the instances in the bag is predicted to classify the entire slide as a class. This instance-level MIL approach contributes to increasing the accuracy of the overall diagnostic result by identifying the diagnostic significance of each instance. This method is especially used to highlight and learn important areas in medical images (Fig. 5)27.
Flowchart of model learning and inference method through Multiple Instance Learning.
The training parameters were set with a batch size of 4, using Stochastic Gradient Descent (SGD) as the optimizer, a maximum of 10 epochs, an initial learning rate of 0.001, and a learning rate scheduler, StepLR, with a step size of 5 and gamma of 0.1. We trained on a set of 9,727 training WSIs, validated the model using a set of 1,250 validation WSIs.
The model was trained using the training set, with the validation set employed to monitor generalization during training by reducing loss. To ensure an unbiased evaluation, all final performance metrics were calculated on a separate test set that was not involved in the training process.
Statistical analysis methods
In this study, precision, sensitivity, F1-score, and AUC were utilized as performance evaluation metrics for the kidney tumor classification model. The F1-score represents the harmonic mean of precision and sensitivity, used to assess the balance between these two metrics. AUC indicates the model’s ability to distinguish between positive and negative classes. All indicators were through bootstrap sampling. Additionally, to evaluate the model, we calculated the weighted average value of AUC and F1 score for each class according to the amount of data samples. additionally, we used the accuracy index to determine the performance for the major subclasses of benign and malignant. This represents the proportion of true positive predictions among all predictions.
Finally, we confirmed the intermediate process of model inference through gradient-weighted class activation mapping (Grad-CAM) operation. Grad-CAM image is an image that visualize how much and where a convolution neural network (CNN) focuses its attention, and which parts of the image lead it to make specific decisions.
Results
We evaluated the model using a test set of 1,250 WSIs, constructed separately from the train set and validation set that used in train session, for normal, benign, and malignant classes. The class-specific metrics provided in Table 2; Fig. 6 illustrate the performance for each class, calculated with 95% confidence intervals (CI) using the bootstrap method. The reported metrics were achieved on a separate test set, which was not used during the training or validation stages. The validation set was used solely to monitor and adjust the model during training, ensuring it could generalize well. The test set provided an independent evaluation of the model’s final performance.
Receiver operating characteristic curve and AUC value based on the classification probability of each class. AUC, area under the curve.
The metrics below are all values output for the test set described above. For the normal class, the precision was 0.959 (CI: 0.958–0.960), sensitivity was 0.910 (CI: 0.908–0.911), and the F1-score was 0.934 (CI: 0.933–0.934). These metrics indicate a high level of accuracy and reliability in identifying normal tissues, demonstrating the model’s robustness in this category. In the benign class, precision was 0.620 (CI: 0.617–0.623), sensitivity was 0.764 (CI: 0.760–0.767), and the F1-score was 0.684 (CI: 0.682–0.687). Although the performance is lower compared to the normal class, the model still shows a reasonable ability to detect benign tumors, reflecting the challenges posed by the smaller dataset. The malignant class exhibited a precision of 0.887 (CI: 0.886–0.888), sensitivity of 0.869 (CI: 0.868–0.871), and an F1-score of 0.878 (CI: 0.877–0.879).
The weighted average F1-score, considering the number of tiles per class, was 0.879 (CI: 0.879–0.880), demonstrating strong overall performance. AUC values were 0.987 (CI: 0.986–0.987) for the normal class, 0.949 (CI: 0.949–0.950) for the benign class, and 0.957 (CI: 0.957–0.958) for the malignant class, with a overall average AUC of 0.964(CI: 0.948–0.98) and weighted average AUC of 0.969 (CI: 0.969–0.969). Based on the above performance indicators, our model showed excellent overall classification performance, particularly in distinguishing malignant and normal classes. These performance metrics underscore the potential utility of our models in the quantitative evaluation of renal tumor in clinical settings, highlighting their reliability and effectiveness in supporting pathological diagnostics.
To assess the model’s performance when data from lower categories were classified into higher categories, we calculated the accuracy for chromophobe RCC (86.80%), clear cell RCC (87.16%), and papillary RCC (86.77%)—subtypes that account for the majority of the dataset. Similarly, for benign tumors, angiomyolipoma achieved an accuracy of 76.99%, and oncocytoma 76.37%, with an overall average accuracy of 86.91% for the malignant category and 76.68% for the benign category (Fig. 7). We displayed representative tiles that were correctly classified by the model: normal (non-neoplastic) renal tissue as normal class (Fig. 8a), benign tumors as benign class (Fig. 8b: oncocytoma, Fig. 8c: angiomyolipoma), and malignant tumors as malignant class (Fig. 8d: papillary renal cell carcinoma, Fig. 8e: clear cell renal cell carcinoma, Fig. 8f: chromophobe renal cell carcinoma).
Accuracy of major subcategories in classify to higher class(malignant and benign).
Representative tiles showing accurate predictions and discrepancies. Panels (a-f) display tiles correctly classified as (a) normal tissue, (b-c) benign tumors (b): oncocytoma, (c): angiomyolipoma), and (d-f) malignant tumors (d): papillary renal cell carcinoma, (e): clear cell renal cell carcinoma, (f): chromophobe renal cell carcinoma). Panels (g-h) highlight misclassifications, with (e) a malignant tumor (clear cell renal cell carcinoma) incorrectly labeled as benign and (f) a benign tumor (oncocytoma) mistakenly predicted as malignant.
We have also displayed representative tiles where the model made misclassifications. Figure 8g shows a tile with a malignant tumor (clear cell renal cell carcinoma) that was incorrectly predicted as benign. The tumor cells in that tile show closely packed nests with granular, eosinophilic cytoplasm, which may have led our model to mistakenly classify it as oncocytoma, a benign tumor, due to their histological similarities that can also cause confusion for pathologists. Figure 8h shows a tile with a benign tumor (oncocytoma) that was incorrectly predicted as malignant. The tumor cells in this tile show a solid nest and cystic arrangement with eosinophilic cytoplasm, a feature occasionally observed in the eosinophilic variant of chromophobe renal cell carcinoma, potentially leading to the misclassification. Such eosinophilic tumors are often difficult to differentiate in pathologic diagnosis, making immunohistochemistry essential.
For the purpose of understanding the model’s decision-making process and identifying areas of focus, we applied Grad-CAM. This allowed us to clearly visualize the intensity of the model’s attention and determine which part of the image it focused on to predict a particular class. Figure 9 shows Grad-CAM images among correctly classified patches. For the normal class, it appears that our model identified normal renal tissue based on the histology of regularly arranged and loosely packed renal tubule structures (Fig. 9a). For the benign class, it seems that the model classified the oncocytoma as a benign tumor by recognizing the histologic feature of monomorphic tumor cell nests with granular eosinophilic cytoplasm (Fig. 9b). For the malignant class, our model appears to have classified the clear cell renal cell carcinoma as malignant based on the histologic features of closely packed monomorphic cells with nest and tubule structures, clear cytoplasm, and hemorrhagic tumor stroma (Fig. 9c).
Grad-Cam image and original image showing the attention intensity when the model infers, from left (a) normal tissue, (b) benign tumor (oncocytoma), (c) malignant tumor (clear cell renal cell carcinoma).
Discussion
We introduced a classification model designed to support the diagnostic evaluation of renal tissues, employing a dataset of digital pathology WSIs from nephrectomy specimens of over 2,535 patients from five institutions, providing a robust approach for distinguishing renal tissue types. Built upon the ResNet-18 architecture and incorporating the MIL method, the model is engineered to differentiate among renal tissues that are normal (non-neoplastic) renal tissue, benign tumors, and malignant tumors. It delivers superior performance metrics that demonstrate its capability in precisely classifying these distinct tissue types.
Previous research has predominantly focused on utilizing pathology WSIs for classifying specific subcategories and nuclear grading within malignant renal tumors, achieving notable performance. However, there has not been effort aimed at classifying WSIs into broader categories, such as distinguishing between normal renal tissue, benign tumors, and malignant tumors. Contrastingly, our study analyzed a comprehensive dataset of 12,223 WSIs, encompassing a wide array of renal tissue types, significantly enhancing the model’s power, robustness, and broad applicability. This dataset includes 1,300 normal, 700 benign, 10,223 malignant WSIs, all sourced from five tertiary care hospitals28.
Our model’s strong performance metrics underscore its utility as a diagnostic support tool. Our model achieved high accuracy with F1-scores of 0.934 for normal tissue, 0.684 for benign tumors, and 0.878 for malignant tumors, along with a weighted average F1-score of 0.879 and an area under the receiver operating characteristic curve of 0.969. These results demonstrate its capability in accurately classifying renal tissue types. making it a valuable resource in various clinical settings. By incorporating this broader understanding, we emphasize that our AI model primarily focuses on aiding the classification of non-neoplastic, benign, and malignant categories. While it cannot replace the full diagnostic process, it can be a valuable tool in certain contexts, particularly in environments with limited pathology expertise. For instance, in smaller or remote hospitals where general pathologists manage nephrectomy specimens, the model can assist with initial classification, saving time and improving diagnostic confidence. Additionally, in high-volume centers, the AI can support quality control by flagging cases that may require further review, particularly in borderline diagnoses or during periods of heavy workload. This can serve as a safety net, helping to reduce diagnostic errors or oversight.
At the WSI level, classification uses one large piece of data rather than splitting it into smaller instances. One label is assigned to the entire image, and learning is also judged based on the entire image. This method has simple data preprocessing and can learn global features of the entire image, but if a specific region of the image is important, such as cancer cells, the features of that region may be diluted. So, in classification at the WSI level, providing precise location information of cancer cells using Grad-CAM images is challenging because the model considers not only cancer cells but also the histologically meaningless finding such as an artifact of the WSI during inference29.
However, we adopted the tile-level MIL approach instead of WSI level classification to focus on capturing and analyzing microscopic lesions and specific pathological features within small tile segments. This method significantly enhanced feature extraction in more detailed areas, thereby increasing the sensitivity to early and subtle pathological changes30. Recovering the tile results back to the full WSI would allow us to pinpoint the sets of key tiles, providing a more precise geographical mapping of cancerous regions within the WSIs. By classifying at the tile level, we ensured that the model focused on cancer tiles during training, minimizing the influence of the histologically meaningless findings in WSI and surrounding tissues. Using Grad-CAM that provide insights into the model’s attention during inference, we ensured that the model focused on key histologic features that define each class, such as arrangement and cytoplasmic characteristics of tumor cells as well as tumor stroma. This focus is attributed to the use of the tile-level MIL approach. By interpreting the Grad-CAM images, we could histologically understand the model’s inferences, making it a form of explainable artificial intelligence.
The disparity in performance across different tissue classifications could largely be attributed to the inherent difficulties in differentiating tumors with eosinophilic features, as evidenced by our review of Grad-CAM images. For instance, clear cell renal cell carcinoma with prominent eosinophilic features, could have led our model to mistakenly classify it as oncocytoma, a benign tumor, due to their histological similarities. Similarly, oncocytoma with solid nest and cystic arrangement, could have caused the model to make errors because these features are occasionally observed as a variant of renal cell carcinoma. Such confusions are challenges that pathologists often face in pathologic diagnosis.
Despite these errors in subtyping classes that show histological similarities, the performance disparity also stems from the limited dataset for benign tumors and the uneven distribution of benign class subcategories. Normal tissues were labeled across all WSIs, including those for benign and malignant tumors, due to the presence of peritumoral normal tissue, ensuring a dataset size comparable to that of malignant tumors. In contrast, labeling for benign tumors was confined to their specific WSIs, leading to a considerably smaller dataset. Enhancing the training dataset by acquiring additional benign tumor data or applying data augmentation techniques could potentially ameliorate this performance gap.
As the model was designed to classify broad categories such as benign, malignant, and normal, there were limitations in distinguishing between aggressive and indolent variants within the malignant class. To enhance the model’s practical utility, follow-up research should focus on expanding the dataset to include more diverse morphologies, such as sarcomatoid and rhabdoid features, and incorporating metadata to classify normal tissue, benign tumors, and malignant tumors along with their subcategories or degrees of differentiation. Additionally, optimizing the algorithm to improve overall classification performance is essential. These advancements will lead to more reliable models and provide flexible metrics that can be applied across a wider range of diagnostic and research settings.
In summary, our study classified renal tissue into three categories: normal, benign tumor, and malignant tumor, utilizing instance-level MIL along with large-scale datasets collected from multiple hospitals. This approach provides a broader categorization of renal tissue types rather than solely subclassifying malignant tumors, with a average AUC of 0.964. The use of Grad-CAM for explainable AI ensured that our model focused on key histologic features, such as the arrangement and cytoplasmic characteristics of tumor cells as well as tumor stroma. This not only paves the way for developing practical diagnostic support models that can be seamlessly integrated into actual pathological diagnostic processes but also highlights the importance of constructing and utilizing large-scale datasets, like those used in this study, for application in various diagnostic environments.
Data availability
The data used to support the findings of this study are available upon request from the corresponding authors.
References
Padala, S. A. et al. Epidemiology of renal cell carcinoma. World J. Oncol. 11, 79 (2020).
Siegel, R. L., Kimberly, D. & Miller and Ahmedin Jemal. Cancer statistics, CA: a cancer journal for clinicians 69.1 (2019): 7–34. (2019).
Lu, M. Y. et al. Federated learning for computational pathology on gigapixel whole slide images. Medical Image Analysis 20.1 : 71-90 (2022). (2022).
Lu, M. Y. et al. Federated learning for computational pathology on gigapixel whole slide images. Medical image analysis 76, 102298 (2022). (2022).
Wu, J. et al. A precision diagnostic framework of renal cell carcinoma on whole-slide images using deep learning. IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, 2021. (2021).
Abu Haeyeh et al. Development and evaluation of a novel deep-learning-based framework for the classification of renal histopathology images. Bioengineering 9.9 : 423. (2022).
Tabibu, Sairam, P. K., Vinod & Jawahar, C. V. Pan-renal cell carcinoma classification and survival prediction from histopathology images using deep learning. Sci. Rep. 9 (1), 10509 (2019).
Chanchal, A. et al. A novel dataset and efficient deep learning framework for automated grading of renal cell carcinoma from kidney histopathology images. Sci. Rep. 13 (1), 1–16 (2023).
Murugan, P. et al. Protocol for the examination of resection specimens from patients with renal cell carcinoma. version 4.2.0.0. College of American Pathologists (CAP); Preprint at (2024). https://documents.cap.org/protocols/Kidney_4.2.0.0.REL_CAPCP.pdf
WHO. Urinary and male genital tumours WHO classification of tumours, 5th Edition, Volume 8 (WHO Classification of Tumours Editorial Board). WHO, (2022).
Amin, M. B. et al. The eighth edition AJCC cancer staging manual: continuing to build a bridge from a population-based to a more personalized approach to cancer staging. Cancer J. Clin. 67 (2), 93–99 (2017).
Jiang, Y. et al. Emerging role of deep learning-based artificial intelligence in tumor pathology. Cancer Commun. 40 (4), 154–166 (2020).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25 (1), 44–56 (2019).
Delahunt, B. et al. The International Society of Urological Pathology (ISUP) grading system for renal cell carcinoma and other prognostic parameters. Am. J. Surg. Pathol. 37 (10), 1490–1504 (2013).
Park, J. et al. Dr. answer Ai for prostate cancer: predicting biochemical recurrence following radical prostatectomy. Technol. Cancer Res. Treat. 20, 15330338211024660 (2021).
Shimizu, H. & Keiichi, I. Nakayama. Artificial intelligence in oncology. Cancer Sci. 111 (5), 1452–1460 (2020).
Park, T. et al. Artificial intelligence in urologic oncology: the actual clinical practice results of IBM Watson for Oncology in South Korea. Prostate Int. 11 (4), 218–221 (2023).
Karimi, D. et al. Deep learning with noisy labels: exploring techniques and remedies in medical image analysis. Med. Image. Anal. 65, 101759 (2020).
Otsu, N. A threshold selection method from gray-level histograms. Automatica 11.285-296 : 23–27. (1975).
Bejnordi, B. et al. Stain specific standardization of whole-slide histopathological images. IEEE Trans. Med. Imaging. 35 (2), 404–415 (2015).
Goodfellow, I. et al. Generative adversarial nets. Adv. Neural. Inf. Process. Syst. 27 (2014).
Breen, J. et al. Generative adversarial networks for stain normalisation in histopathology. Applications of generative AI. Cham: Springer International Publishing, 227–247. (2024).
He, K. et al. Deep residual learning for image recognition. Proceedings of the IEEE conference on computer vision and pattern recognition. (2016).
Agarap, A. F. Deep learning using rectified linear units (relu). arXiv preprint arXiv:1803.08375 (2018).
Srivastava, N. et al. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15 (1), 1929–1958 (2014).
Chlap, P. et al. A review of medical image data augmentation techniques for deep learning applications. J. Med. Imaging Radiat. Oncol. 65, 545–563 (2021).
Ilse, M., Tomczak, J. & Welling, M. Attention-based deep multiple instance learning. International conference on machine learning. PMLR, (2018).
Althnian, A. et al. Impact of dataset size on classification performance: an empirical evaluation in the medical domain. Appl. Sci. 11 (2), 796 (2021).
Selvaraju, R. R. et al. Grad-cam: Visual explanations from deep networks via gradient-based localization. Proceedings of the IEEE international conference on computer vision. (2017).
Li, J. et al. A multi-resolution model for histopathology image classification and localization with multiple instance learning. Comput. Biol. Med. 131, 104253 (2021).
Acknowledgements
We thank the reviewers for their opinions and give credit to our co-authors for their hard work. This work was supported by the Technology Innovation Program (K_G012001187801, “Development of Diagnostic Medical Devices with Artificial Intelligence-Based Image Analysis Technology”) funded by the Ministry of Trade, Industry and Energy (MOTIE, Korea). Additional support was provided by the GRRC program of Gyeonggi Province [GRRC-Gachon2023(B01), Development of AI-based Medical Imaging Technology], the National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT (MSIT, Korea) (Grant No. RS-2022-00166555), and a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health and Welfare, Republic of Korea (Grant No. RS-2021-KH113146). This work was also supported by the Export-Oriented “2023 Small and Medium Business Technology Development (R&D) Support Project” (Grant No: RS-2023-00280710) funded by the Ministry of SMEs and Startups (MSS, Korea).
Author information
Authors and Affiliations
Contributions
Chan Kwon Jung and Kwang Gi Kim contributed to the study conception and design. Material preparation, data collection and analysis were performed by Seung Wan Moon, Jisup Kim, Young Jae Kim, Sung Hyun Kim, Chi Sung An. The first draft of the manuscript was written by Seung Wan Moon and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Moon, S.W., Kim, J., Kim, Y.J. et al. Leveraging explainable AI and large-scale datasets for comprehensive classification of renal histologic types. Sci Rep 15, 1745 (2025). https://doi.org/10.1038/s41598-025-85857-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-85857-8
Keywords
This article is cited by
-
Multicenter study of CT-based deep learning for predicting preoperative T staging and TNM staging in clear cell renal cell carcinoma
BMC Cancer (2025)
-
Evidential deep learning-based ALK-expression screening using H&E-stained histopathological images
npj Digital Medicine (2025)
-
Multi-institutional validation of AI models for classifying urothelial neoplasms in digital pathology
Scientific Reports (2025)