Abstract
Prediction of isocitrate dehydrogenase (IDH) mutation status and epilepsy occurrence are important to glioma patients. Although machine learning models have been constructed for both issues, the correlation between them has not been explored. Our study aimed to exploit this correlation to improve the performance of both of the IDH mutation status identification and epilepsy diagnosis models in patients with glioma II-IV. 399 patients were retrospectively enrolled and divided into a training (n = 279) and an independent test (n = 120) cohort. Multi-center dataset (n = 228) from The Cancer Imaging Archive (TCIA) was used for external test for identification of IDH mutation status. Region of interests comprising the entire tumor and peritumoral edema were automatically segmented using a pre-trained deep learning model. Radiomic features were extracted from T1-weighted, T2-weighted, post-Gadolinium T1 weighted, and T2 fluid-attenuated inversion recovery images. We proposed an iterative approach derived from LASSO to select features shared by two tasks and features specific to each task, before using them to construct the final models. Receiver operating characteristic (ROC) analysis was employed to evaluate the model. The IDH mutation identification model achieved area under the ROC curve (AUC) values of 0.948, 0.946 and 0.860 on the training, internal test, and external test cohorts, respectively. The epilepsy diagnosis model achieved AUCs of 0.924 and 0.880 on the training and internal test cohorts, respectively. The proposed models can identify IDH status and epilepsy with fewer features, thus having better interpretability and lower risk of overfitting. This not only improves its chance of application in clinical settings, but also provides a new scheme to study multiple correlated clinical tasks.
Similar content being viewed by others
Introduction
Gliomas are the most common and lethal malignant primary brain tumors with significant mortality and morbidity1. Glioma-associated epilepsy (GAE) is frequently observed in patients with gliomas2,3 and significantly affects their quality of life. The etiology of GAE is likely multifactorial and complex and remains poorly comprehended4. Previous study5 suggested that response to successive antiepileptic drugs (AED) regimens is poorer than in the non–tumor-associated epilepsy population. If glioma patients have preoperative seizures, lack of gross total resection and postoperative disabling seizures, timely AED up-titration is recommended. If seizures persist, then either re-resection or a comprehensive epilepsy program assessment to identify and remove the ictal onset zone should be performed5. A better understanding of the mechanisms and early identification of GAE is crucial to protect neurocognition, restrict disease progression, and improve the quality of life of patients3,6.
The isocitrate dehydrogenase (IDH) gene is mutated in more than 70% of grade II and III gliomas7. IDH mutation status was suggested by the World Health Organization (WHO)8 as an essential biomarker for glioma subtyping and is associated with survival rate. IDH1 and IDH2 mutations are genetic alterations that occur frequently in gliomas patients and IDH1 mutations are more commonly associated with glioma than IDH2 mutations9. The classification of IDH into IDH-wild type (IDH-W) and IDH-mutant type (IDH-M) has now become an integral part of the molecular diagnosis of gliomas10. The current procedure for the identification of IDH mutation status with tissue samples is invasive and expensive. Therefore, a non-invasive alternative is highly desirable. Both classical machine learning (ML) and deep learning (DL) have been used in the non-invasive prediction of IDH mutation in glioma patients. While classical ML combining radiomics and clinical features has become a powerful tool in identifying IDH mutations11, DL models have also achieved remarkable success due to its ability to automatically learn relevant features from magnetic resonance imaging (MRI) data12.
Many studies have shown that IDH mutations are frequently associated with preoperative seizures in patients with grade II13,14,15 and grade III–IV gliomas13,16,17. D-2-hydroxyglutarate (D2HG) product of mutant IDH is similar in structure to glutamate, a widespread excitatory neurotransmitter in the central nervous system. Therefore, D2HG may disturb excitatory and inhibitory regions in the brain, subsequently triggering epilepsy18. IDH-M gliomas are more prone to develop seizures than IDH-W gliomas, and tumor growth may stimulate seizures, which may promote tumor growth in return19. This has important implications for the personalized management of GAE, as inhibitors targeting IDH-M may also improve antiepileptic therapy in patients with IDH-M gliomas. Utilizing correlation between IDH mutation and epilepsy identification tasks could improve the clinical diagnosis and treatment of glioma patients.
MRI is an indispensable preoperative examination for patients with glioma due to its high-resolution multiplanar structural information. Currently, multiparametric MRI (mpMRI) is routinely employed for the diagnosis and delineation of tumor compartments. Radiomics extracts large number of quantitative features from medical images and uses them to build predictive models that relate image features to the target information in an objective, repeatable, and non-invasive manner20. Previous studies have used radiomics for both IDH mutation status identification21 and GAE occurrence identification22,23 based on MRI. However, these studies examined these two issues separately, failing to take advantage of their latent correlation. Since IDH mutation status and GAE are closely related, we hypothesized that models for these two targets may share some common radiomics features and building models with shared features may lead to improved performance and better understanding of the models.
Materials and methods
Data
This study was approved by the institutional review board and the requirement for written informed consent was waived for its retrospective nature. Our research was carried out in accordance with the Declaration of Helsinki. We enrolled 399 consecutive patients with glioma who underwent MRI scanning before surgery at the hospital between August 2016 and September 2019 as the main cohort. The inclusion criteria and exclusion criteria are shown in Fig. 1. Preoperative diagnosis of GAE was based on clinical manifestations, electroencephalography (EEG), and imaging findings. IDH mutation, including the IDH1 or IDH2 mutation in the resected tumor was identified using immunohistochemistry. Patients from the hospital were scanned using 3.0 T MRI scanners (Magnetom Trio TIM/Prisma, Verio or Skyra, Siemens Healthcare; Discovery 750, GE Medical Systems). The T1-weighted image (T1WI), T2-weighted image (T2WI), and post-Gadolinium T1 weighted image (T1Gd) were scanned with a turbo spin-echo sequence (TR/TE 2000–3800/90–120 ms), spin-echo sequence (TR/TE 200–220/2–3 ms), and spin-echo sequence (TR/TE 250/2.5 ms), respectively. T2 fluid-attenuated inversion recovery image (T2-FLAIR) images were scanned with TI = 2400–2500 ms, TE = 81–135 ms, and TR = 8000–8500 ms. For all scans, image matrix = 256 × 256, field of view (FOV) = 240 × 240 mm2, section thickness = 5 mm, and intersection gap = 1 mm.
Flowchart to illustrate the selection of the main cohort of the study.
The internal dataset was randomly split into a training cohort (n = 279) and an internal test cohort (n = 120). Repeated random splitting was used to ensure primary clinical characteristics, such as age (p = 0.334), gender (p = 0.603), and tumor grade (p = 0.934) exhibited no significant difference (p < 0.05 was regarded as statistically significant) between their distributions in the training and internal test cohorts.
A multi-center dataset from The Cancer Imaging Archive (TCIA)24 was used as external test cohort for IDH mutation status identification. The clinical information is provided by The Cancer Genome Atlas (TCGA)25. The data is screened for the availability of gender, age, IDH mutation status, preoperative T1WI, T2WI, T1Gd and T2-FLAIR images. The final external test cohort includes 228 patients.
Tumor segmentation and image preprocessing
To assess the performance of automatic segmentation, region of interest (ROI) containing the entire tumor and peritumoral edema was manually drawn on the T2-FLAIR images in the main cohort using the ITK-SNAP (version 3.6.0) software by two experienced neuroradiologists. For each patient, T1WI, T2WI, and T1Gd images were aligned to T2-FLAIR images using Elastix26,27. Before feature extraction, all images were normalized to the range of [0,1] and processed with the Contrast Limited Adaptive Histogram Equalization (CLAHE) algorithm28 to standardize the intensity variation between different MRI systems.
For automatic tumor segmentation, brain extraction was performed using HD-BET29, then a pre-trained nnU-Net model30 was used to automatically segment tumor ROIs comprising the entire tumor and peritumoral edema.
Radiomics feature extraction
Feature extraction was performed using the PyRadiomics (version 3.0)31 package in Python (version 3.7.6). For each case, features were extracted from auto-segmented ROI, including 14 shape features extracted from ROI mask, 18 first-order statistical features, and 75 texture features extracted from each of the original T1WI, T2WI, T1Gd, and T2-FLAIR images and corresponding Laplacian of Gaussian (LoG) filtered images, by three kernel sizes (1.5, 3.0 and 5.0 mm). The extracted texture features included those based on the gray level co-occurrence matrix (GLCM), gray level run length matrix (GLRLM), gray level size zone matrix (GLSZM), neighboring gray tone difference matrix (NGTDM), and gray level dependence matrix (GLDM). Age and gender were also used in model construction. All the features were normalized using z-score. For feature understandability and stability, we deliberately avoided using wavelet derived features.
Feature selection and model construction
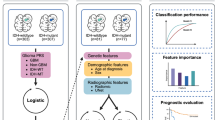
Study workflow is shown in Fig. 2. First, to reduce feature dimensions, Pearson correlation coefficient (PCC) values between all features were calculated, and if the PCC value between two features was higher than 0.99, we randomly removed one of them. For feature selection, Least absolute shrinkage and selection operator (LASSO) was applied to each task to select task-specific features, and multi-task LASSO32 was used to select features shared by two tasks. For both LASSO and multi-task LASSO, we used bootstrapping to improve the robustness of feature selection. That is, each experiment was replicated 100 times using random resampling with replacement over the training cohort. In each repetition, we randomly select 80% cases in the training cohort for model construction, and 20% cases for cross-validation. Models with the validation area under the receiver operating characteristic curve (AUC) ≥ 0.7 were retained and features that were selected more than 80 times in the retained models were kept for subsequent analysis. The penalty term coefficient \(\:\lambda\:\) used in LASSO was empirically set to 0.001.
Workflow of the study. Radiomics features were extracted from ROIs segmented with a pretrained model. Single-task LASSO was utilized to select task specific features for each task. Multi-task LASSO was utilized to select features shared by two tasks. Finally, task-specific and shared features were combined to develop a radiomics model for identifying IDH mutations status and glioma-associated epilepsy (GAE).
Finally, the shared and task-specific features were combined to construct the final model with LASSO for each task. Hyper-parameter \(\:\lambda\:\) was iterated between 0.1 and 0.0001, and determined by 1-SE rule to reduce number of features used in the model. Since the final models for two tasks retained their specific feature while sharing some of the features, we call this approach collaborative multi-task (CMT) learning.
For comparison, two single-task LASSO models and a multi-task LASSO model were also built with the same feature set and hyperparameters to identify the IDH mutation status and epilepsy.
In the entire process of all model construction, only the training cohort was used. Our code is available at: https://github.com/wangyidada/CMTLasso.git.
Evaluation of models
Receiver operating characteristic (ROC) curve analysis, confusion matrix and waterfall plot were used to evaluate the performance of the models, with both the internal and external test cohorts. Accuracy, sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were also calculated at a cutoff value that maximized the value of the Youden index in the training cohort. DeLong test was used to pairwise compare the ROCs between different models. To investigate the model characteristics, Pearson correlation coefficient values between all features selected by either IDH mutation status model or GAE identification model were calculated and visualized.
Statistical analysis
Age is reported as the mean and range, and the difference in its distribution between the GAE and non-GAE groups and between the IDH-M and IDH-W groups was assessed using the Mann-Whitney U test. Gender and tumor grade were reported as frequencies and proportions, and differences between groups were assessed using the Pearson chi-square test. Statistical analysis was performed using SciPy (version 1.5.2) and statistical significance was defined as p-value < 0.05. The Radiomics Quality Score (RQS)20 was used for evaluating the quality of our study.
Results
Demographics
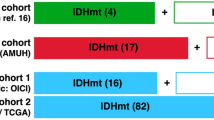
The main clinical and radiological characteristics of patients in the three cohorts are summarized in Tables 1 and 2. The number of patients in the training, internal, and external test cohorts were 279 (IDH-M/IDH-W: 146/133; GAE/non-GAE: 173/106), 120 (IDH-M/IDH-W: 60/60; GAE/non-GAE: 75/45), and 228 (IDH-M/IDH-W: 82/146), respectively. The tables indicate that there were significant differences (p-value < 0.001) in age and tumor grade between the IDH-M with IDH-W groups, as well as in the GAE and non-GAE groups. Statistically, young patients and patients with low-grade glioma (LGG) had a higher chance of having IDH-M type and were more likely to develop epilepsy.
Performance of tumor segmentation
The pre-trained nnU-Net achieved a mean Dice of 0.847 ± 0.153 in the internal dataset. Figure 3 shows the segmentation results and ground truth for three patients with WHO grades II-IV.
Visual comparison of segmentation results and the ground truth of 3 random patients with grade II (patient1), III (patient2) and IV (patient3), respectively, on T2-FLAIR images.
Performance of single-task LASSO models
For IDH mutation status identification, the single-task LASSO model utilized 23 features and achieved AUC values of 0.932 [95% confidence interval (CI), 0.901–0.958], 0.940 [95% CI,0.901–0.976] and 0.882 [95% CI, 0.831–0.926] in the training, internal test and external test cohorts, respectively.
For epilepsy identification, the single-task LASSO model utilized 17 features and achieved AUC values of 0.902 [95% CI, 0.865–0.936] and 0.871 [95% CI, 0.798–0.933] in the training and internal test cohorts, respectively. Detailed performance metrics of the two single-task models are listed in Tables 3 and 4, respectively. The ROC curves are shown in Fig. 4. The features retained in the two models and their corresponding weights are shown in Supplementary Figure S1. No shared features were selected by both the IDH mutation status and epilepsy models, which means single-task models use a total of 40 features for two tasks.
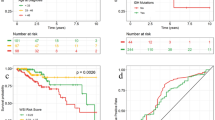
IDH mutation status and epilepsy identification results by single task (ST), multi-task (MT) and collaborative multi-task (CMT) models in different cohorts. (a–c) ROC curves for IDH mutation status identification by CMT, MT and ST models; (d–f) ROC curves for epilepsy identification by CMT, MT and ST models; (g–i) waterfall plots for the distribution of epilepsy identification in the internal test cohort, IDH mutation status identification in the internal test cohort, and IDH mutation status identification in the external test cohort by CMT models, respectively. (j–l) confusion matrix of epilepsy identification in the internal test cohort, IDH mutation status identification in the internal test cohort, and IDH mutation status identification in the external test cohort by CMT models, respectively. ROC = receiver operating characteristic; AUC = area under curve; GAE = glioma-associated epilepsy.
Performance of multi-task LASSO model
The multi-task LASSO model used a shared set of 30 features and achieved training AUCs of 0.927 [95% CI, 0.897–0.953] and 0.902 [95% CI, 0.864–0.934], internal test AUCs of 0.938 [95% CI,0.892–0.974] and 0.870 [95% CI, 0.809–0.972] for IDH mutation status identification and epilepsy identification, respectively. The multi-task model achieved AUC of 0.867 [95% CI, 0.817–0.915] in the external test cohort for identification of IDH mutation status. Detailed performance metrics of the models are also listed in Tables 3 and 4. The ROC curves are shown in Fig. 4. The weights of all the features for both tasks in the multi-task model are shown in Supplementary Figure S2.
Performance of CMT LASSO models
For IDH mutation status identification, the CMT LASSO model contained 20 features and achieved AUC values of 0.948 [95% CI, 0.923–0.970], 0.946 [95% CI, 0.902–0.982] and 0.860 [95% CI, 0.806–0.910] in the training, internal test and external test cohorts, respectively. For epilepsy identification, the CMT LASSO model contained 16 features and achieved AUC values of 0.924 [95% CI, 0.893–0.952] and 0.880 [95% CI, 0.811–0.937] in the training and internal test cohorts, respectively. Detailed performance metrics of the two models are listed in Tables 3 and 4 for comparison. The ROC curves, confusion matrix, and waterfall plots are shown in Fig. 4. The features included in the two models and their corresponding weights are shown in Fig. 5. Eight features were shared by both the IDH mutation status and epilepsy models, which means a total of 28 features were used for two tasks.
Features and their corresponding weights used in final CMT models for identifying epilepsy occurrence (a) and IDH mutation status (b). Green and purple bars stand for weights for shared and task-specific features, respectively.
In the internal test cohort, there was no significant difference between CMT and ST LASSO models for IDH mutation status (p = 0.672) and epilepsy (p = 0.507) identification, and there was significant differences between CMT and MT LASSO models (p < 1e-5) in both two tasks. Overall, the proposed model outperformed the both single task and multi-task LASSO models. PCC values between all features selected by two models were shown Fig. 6, for ST, MT, and CMT schemes separately. Our study derived an RQS score of 19 (36 in total) (Supplementary Table S1).
Pearson correlation coefficient (PCC) values between features selected by IDH and GAE models constructed with different approaches: single task (ST) (a), multi-task (MT) (b), and collaborative multi-task (CMT) (c) models for IDH mutation status and glioma-associated epilepsy (GAE) identification. Feature names are represented as sequence name for brevity. The detailed feature names and PCC values are shown in enlarged version in Supplementary Figures S3, S4, and S5.
Discussion
Typical radiomics studies treat each problem independently, ignoring the correlations between associated problems. On the other hand, multi-task learning (MTL) can jointly learn multiple related tasks so that the knowledge contained in a task can be leveraged by other tasks33. However, MTL uses a common set of features for all tasks, ignoring the unique characteristics of each task, which may lead to a sub-optimal model performance. Thus, we proposed an approach to combine classic LASSO and multi-task LASSO to select both shared and task-specific features to build collaborative multitask models without performance compromise. To the best of our knowledge, this is the first attempt to construct two radiomics models for correlated targets using both task-specific and shared radiomics features.
Epilepsy is a common and disabling symptom of many types of brain tumors, associated with tumor location, patient age, and histopathological subtype14. D2HG product of IDH mutation may increase neuronal activity by mimicking the activity of glutamate on the NMDA receptor, and IDH mutation gliomas are more likely to cause epilepsy in patients13. IDH1/2 mutations were reported to be an independent predictor for epilepsy, as a presenting feature in low-grade gliomas patients15. Patients affected by IDH mutant gliomas present more frequently with epileptic activity than patients with IDH wild-type gliomas9. Therefore, identifying IDH-mutant status is helpful for identifying treatments for epilepsy.
Manual delineation of ROIs is inevitably influenced by radiologists’ experience and subjective judgment, leading to the variability in extracted radiomics features. Thus, auto-segmentation was used to alleviate the workload and subjectivity of manual delineation. The pre-trained nnU-Net achieved good performance in segmenting the whole tumor ROIs, as demonstrated not only by the quantitative metrics in the internal dataset, but also by the performance of downstream diagnosis models.
Many studies have reported that the radiomics features extracted from MRI in patients with gliomas identified IDH mutation status with an AUC between 0.80 and 0.9611,34,35, and epilepsy with an AUC between 0.82 and 0.87 22,23,36,37. The details of these studies are summarized in Supplementary Tables S2 and S3. While the performance achieved by the proposed CMT learning ranked among the top, the major difference between this study and previous ones is that the correlation between IDH mutations and epilepsy was exploited to find the shared feature set. This minimized the number of features used, simplifying radiomics models and lowering the risk of overfitting. Different from previous multi-task studies38,39, we also made use of task-specific features to improve the performance of model. In the internal dataset, the CMT model achieved the highest AUC values for both tasks. Over the external dataset, the CMT model achieved a performance comparable to ST model for IDH mutations identification with smaller number of features (20 vs. 23), which reduced the model complexity. Interestingly, incorporation of task-specific features not only increased the performance of the model, but also reduced the total number of features used from 30 of multi-task model to 28 of CMT models, implying the importance of combining shared features with task-specific features. The reduction makes the model easier to understand and increases the chance of finding useful, generalizable biomarkers. More importantly, the features shared by the two tasks can be used to perform correlations association analysis between different tasks. The weights of most features have the same sign in both tasks, suggesting that inherent connection might exist between IDH-M and GAE.
Our results suggested that features from preoperative routine mpMRI were important biomarkers for gliomas since they are associated with both IDH mutations and epilepsy. Figure 5 outlined that the feature extracted from T1Gd and T2-FLAIR sequences exhibited high importance scores. Previous study40 also found that T1Gd was the optimal MRI sequence for IDH mutation status identification, T2-FLAIR sequence was moderately suitable, and T1WI and T2WI sequences were identified as the least suitable options. Age was shared by two tasks and negatively correlated with IDH-M/epilepsy, which is consistent with the observation that LGG is more common in younger patients and is correlated with a higher chance of having IDH-M/epilepsy than HGG patients41,42. Although tumor grade is an important biomarker for both tasks, its confirmation requires invasive surgery or biopsy, which limits the clinical use of the model. Therefore, we did not include tumor grade as a clinical characteristic in the model building.
Figure 6 visualizes the correlation between features retained in the models, reflecting feature redundancy in the model. It can be seen from Fig. 6a, in models built with single task LASSO, features retained in IDH model and GAE model are heavily correlated though there are no feature overlapping between these two models. This demonstrated that a failure to reveal the connections between two tasks. Multi-task model in Fig. 6b makes full use of the associations of the two tasks, however, there is still heavy redundancy in the model, implying an over-complex model and sub-optimal performance. In comparison, in CMT models in Fig. 6c, feature redundancy is reduced greatly. This is in accordance with the fact that CMT models used fewer number of features in total and in each model.
Here we would like to appeal to researchers developing radiomics models to pay more attention to the intrinsic quality of the models, including the complexity of the model, and the redundancy of the features retained in model, rather than just focusing on the classic performance metrics. It was reported in a systematic review that underpowered studies, which included more features than the sample size could support, lead inevitably to false-positive results43. For example, it has been suggested that the “rule of thumb” that there should be minimum 10 samples in the smaller group for each feature included in the model, should be used as an initial step to determine the sample size44. Reducing the feature number to reduce the risk of overfitting was also included in the widely-used radiomics quality score20. Among models with no significantly different performances, a simpler one using fewer features should be preferred. This idea is reflected in the popular 1-SE rule for feature selection and explains why the CMT models are deemed better than single-task models. Since the redundancy in the features used in the model is positively associated with the number of features used, visualization of the redundancy as in Fig. 6 may help us to determine whether the model can be further simplified.
Our study also has some limitations. Firstly, external validation was used only for IDH status identification. So, further validation with a larger, multi-institutional dataset covering both tasks is warranted. Furthermore, the clinical relevance of simultaneous prediction of IDH status and epilepsy is not yet clear. While this problem can be explored further in the future, we also hope the proposed CMT scheme could be validated in more studies where the simultaneous predictions of multiple tasks are more clinically relevant. Secondly, due to the limited dataset size, we did not conduct stratified studies on grade II, III, and IV glioma patients, which we believe can further clarify the implications of this study to patients with different grade gliomas. Thirdly, we built all models with features extracted from the whole tumor. Therefore, further studies using features from sub-regions of the tumor, such as the enhanced components, necrosis, and edema, might improve the model performance and provide more insight into the disease. Finally, the proposed CMT scheme can be validated in more studies on different problems.
In conclusion, we developed a novel radiomics approach for the identification of IDH mutation status and GAE using age and radiomic features extracted from mpMRI. The proposed approach exploited the underlying connections between these two problems, reduced the risk of overfitting, improved the model performance, and can potentially been used to find valuable generalized biomarkers for multiple clinical problems. The proposed model may be used as a convenient and non-invasive tool for the diagnosis and treatment of patients with gliomas.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Weller, M. et al. Glioma. Nat. Rev. Dis. Primers. 1, 15017. https://doi.org/10.1038/nrdp.2015.17 (2015).
van Breemen, M. S. M., Wilms, E. B. & Vecht, C. J. Epilepsy in patients with brain tumours: Epidemiology, mechanisms, and management. Lancet Neurol. 6, 421–430. https://doi.org/10.1016/S1474-4422(07)70103-5 (2007).
Englot, D. J., Chang, E. F. & Vecht, C. J. Epilepsy and brain tumors. Handb. Clin. Neurol. 134, 267–285. https://doi.org/10.1016/B978-0-12-802997-8.00016-5 (2016).
Rosati, A. et al. Epilepsy in cerebral glioma: timing of appearance and histological correlations. J. Neuro-Oncol. 93, 395–400. https://doi.org/10.1007/s11060-009-9796-5 (2009).
Neal, A., Morokoff, A., O’Brien, T. J. & Kwan, P. Postoperative seizure control in patients with tumor-associated epilepsy. Epilepsia 57, 1779–1788. https://doi.org/10.1111/epi.13562 (2016).
Kemerdere, R. et al. Low-grade temporal gliomas: Surgical strategy and long-term seizure outcome. Clin. Neurol. Neurosurg. 126, 196–200. https://doi.org/10.1016/j.clineuro.2014.09.007 (2014).
Ohgaki, H. & Kleihues, P. Population-based studies on incidence, survival rates, and genetic alterations in astrocytic and oligodendroglial gliomas. J. Neuropath Exp. Neur. 64, 479–489. https://doi.org/10.1093/jnen/64.6.479 (2005).
Wesseling, P. & Capper, D. WHO 2016 classification of gliomas. Neuropath Appl. Neuro. 44, 139–150. https://doi.org/10.1111/nan.12432 (2018).
Solomou, G., Finch, A., Asghar, A. & Bardella, C. Mutant IDH in Gliomas: Role in Cancer and Treatment options. Cancers 15 https://doi.org/10.3390/cancers15112883 (2023).
Louis, D. N. et al. The 2021 WHO classification of tumors of the Central Nervous System: a summary. Neuro-Oncology 23, 1231–1251. https://doi.org/10.1093/neuonc/noab106 (2021).
Zhao, J. et al. Diagnostic accuracy and potential covariates for machine learning to identify IDH mutations in glioma patients: Evidence from a meta-analysis. Eur. Radiol. 30, 4664–4674. https://doi.org/10.1007/s00330-020-06717-9 (2020).
Choi, Y. S. et al. Fully automated hybrid approach to predict the mutation status of gliomas via deep learning and radiomics. Neuro-Oncol. 23, 304–313. https://doi.org/10.1093/neuonc/noaa177 (2021).
Chen, H. et al. Mutant IDH1 and seizures in patients with glioma. Neurology 88, 1805–1813. https://doi.org/10.1212/Wnl.0000000000003911 (2017).
Liubinas, S. V. et al. IDH1 mutation is associated with seizures and protoplasmic subtype in patients with low-grade gliomas. Epilepsia 55, 1438–1443. https://doi.org/10.1111/epi.12662 (2014).
Stockhammer, F. et al. IDH1/2 mutations in WHO grade II astrocytomas associated with localization and seizure as the initial symptom. Seizure-Eur J. Epilep. 21, 194–197. https://doi.org/10.1016/j.seizure.2011.12.007 (2012).
Toledo, M. et al. Epileptic features and survival in glioblastomas presenting with seizures. Epilepsy Res. 130, 1–6. https://doi.org/10.1016/j.eplepsyres.2016.12.013 (2017).
Yang, Y. et al. An analysis of 170 glioma patients and systematic review to investigate the association between IDH-1 mutations and preoperative glioma-related epilepsy. J. Clin. Neurosci. 31, 56–62. https://doi.org/10.1016/j.jocn.2015.11.030 (2016).
Moussawi, K., Riegel, A., Nair, S. & Kalivas, P. W. Extracellular glutamate: functional compartments operate in different concentration ranges. Front. Syst. Neurosci. 5, 94. https://doi.org/10.3389/fnsys.2011.00094 (2011).
Huberfeld, G. & Vecht, C. J. Seizures and gliomas - towards a single therapeutic approach. Nat. Rev. Neurol. 12, 204–216. https://doi.org/10.1038/nrneurol.2016.26 (2016).
Lambin, P. et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 14, 749–762. https://doi.org/10.1038/nrclinonc.2017.141 (2017).
Bhandari, A. P., Liong, R., Koppen, J., Murthy, S. V. & Lasocki, A. Noninvasive determination of IDH and 1p19q status of Lower-grade gliomas using MRI radiomics: A systematic review. Am. J. Neuroradiol. 42, 94–101. https://doi.org/10.3174/ajnr.A6875 (2021).
Gao, A. K. et al. Radiomics for the prediction of Epilepsy in patients with frontal glioma. Front. Oncol. 11 https://doi.org/10.3389/fonc.2021.725926 (2021).
Bai, J. et al. Radiomics Nomogram improves the prediction of Epilepsy in patients with gliomas. Front. Oncol. 12 https://doi.org/10.3389/fonc.2022.856359 (2022).
Clark, K. et al. The cancer imaging archive (TCIA): Maintaining and operating a public information repository. J. Digit. Imaging. 26, 1045–1057. https://doi.org/10.1007/s10278-013-9622-7 (2013).
Ceccarelli, M. et al. Molecular profiling reveals biologically discrete subsets and pathways of progression in diffuse glioma. Cell 164, 550–563. https://doi.org/10.1016/j.cell.2015.12.028 (2016).
Klein, S., Staring, M., Murphy, K., Viergever, M. A. & Pluim, J. P. W. Elastix: A toolbox for intensity-based Medical Image Registration. IEEE T Med. Imaging 29, 196–205. https://doi.org/10.1109/Tmi.2009.2035616 (2010).
Shamonin, D. P. et al. Fast parallel image registration on CPU and GPU for diagnostic classification of Alzheimer’s disease. Front. Neuroinform. 7 https://doi.org/10.3389/fninf.2013.00050 (2014).
Reza, A. M. Realization of the contrast limited adaptive histogram equalization (CLAHE) for real-time image enhancement. J. VLSI Signal. Process.-Syst. Signal. Image Video Technol. 38, 35–44. https://doi.org/10.1023/B:VLSI.0000028532.53893.82 (2004).
Isensee, F. et al. Automated brain extraction of multisequence MRI using artificial neural networks. Hum. Brain Mapp. 40, 4952–4964. https://doi.org/10.1002/hbm.24750 (2019).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211. https://doi.org/10.1038/s41592-020-01008-z (2021).
van Griethuysen, J. J. M. et al. Computational Radiomics system to decode the radiographic phenotype. Cancer Res. 77, E104–E107. https://doi.org/10.1158/0008-5472.Can-17-0339 (2017).
Argyriou, A., Evgeniou, T. & Pontil, M. Multi-task feature learning. Adv. Neural. Inf. Process. Syst. 19 https://doi.org/10.7551/mitpress/7503.003.0010 (2006).
Zhang, Y. & Yang, Q. A. Survey on Multi-task Learning. IEEE T Knowl. Data Eng. 34, 5586–5609. https://doi.org/10.1109/Tkde.2021.3070203 (2022).
Truong, N. C. et al. Two-stage training framework using multicontrast MRI radiomics for IDH mutation status prediction in glioma. Radiology: Artif. Intell. 6, e230218. https://doi.org/10.1148/ryai.230218 (2024).
Hosseini, S. et al. MRI-based radiomics combined with deep learning for distinguishing IDH-mutant WHO grade 4 astrocytomas from IDH-wild-type glioblastomas. Cancers 15, 951. https://doi.org/10.3390/cancers15030951 (2023).
Liu, Z. Y. et al. Radiomics analysis allows for precise prediction of epilepsy in patients with low-grade gliomas. Neuroimage-Clin. 19, 271–278. https://doi.org/10.1016/j.nicl.2018.04.024 (2018).
Wang, Y. Y. et al. Predicting the type of tumor-related epilepsy in patients with low-grade gliomas: a radiomics study. Front. Oncol. 10 https://doi.org/10.3389/fonc.2020.00235 (2020).
Rao, N., Cox, C., Nowak, R. & Rogers, T. T. Sparse overlapping sets lasso for multitask learning and its application to fMRI analysis. Adv. Neural. Inf. Process. Syst. 2202–2210. https://doi.org/10.48550/arXiv.1311.5422 (2013).
Obozinski, G., Taskar, B. & Jordan, M. I. Joint covariate selection and joint subspace selection for multiple classification problems. Stat. Comput. 20, 231–252. https://doi.org/10.1007/s11222-008-9111-x (2010).
Kasap, D. N. G. et al. Comparison of MRI sequences to predict mutation status in gliomas using radiomics-based machine learning. Biomedicines 12 (2024). https://doi.org/10.3390/biomedicines12040725
Ichimura, K. et al. IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro-Oncology 11, 341–347. https://doi.org/10.1215/15228517-2009-025 (2009).
Pallud, J. et al. Epileptic seizures in diffuse low-grade gliomas in adults. Brain 137, 449–462. https://doi.org/10.1093/brain/awt345 (2014).
Chalkidou, A., O’Doherty, M. J. & Marsden, P. K. False Discovery Rates in PET and CT studies with texture features: a systematic review. Plos One 10 https://doi.org/10.1371/journal.pone.0124165 (2015).
Halligan, S., Menu, Y. & Mallett, S. Why did reject my radiomic biomarker paper? How to correctly evaluate imaging biomarkers in a clinical setting. Eur. Radiol. 31, 9361–9368. https://doi.org/10.1007/s00330-021-07971-1 (2021).
Funding
This work was supported by the Henan Provincial Medical Science and Technology Tackling Program (LHGJ20230181) and Xing-Fu-Zhi-Hua Foundation of ECNU.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Y. W., J.C. and G.Y. were involved in the conceptualization. J.B., H. Z. and Y.S. contributed to the literature screening and selection. J.C., A.G., Y. Z. and G.Z. extracted the data and assessed the quality. Y.W., H.Y. and C.W. contributed to the statistical analysis. Y.W., A.G. and H.Y. assisted in the original draft of the manuscript. G.Y. checked and edited the manuscript. G.Y. and J.C. contributed to the supervision. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
Institutional Review Board approval was obtained.
Consent to participate
Written informed consent was waived by the Institutional Review Board because of the retrospective nature of our study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Gao, A., Yang, H. et al. Using partially shared radiomics features to simultaneously identify isocitrate dehydrogenase mutation status and epilepsy in glioma patients from MRI images. Sci Rep 15, 3591 (2025). https://doi.org/10.1038/s41598-025-87778-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-87778-y
Keywords
This article is cited by
-
Machine learning-based models for predicting glioma-associated epilepsy: a systematic review and meta-analysis
Discover Oncology (2025)
-
Diagnostic performance of deep learning for predicting glioma isocitrate dehydrogenase and 1p/19q co-deletion in MRI: a systematic review and meta-analysis
European Radiology (2025)