Abstract
Tobacco is one of the most important model plants. Plant height is one of the most important agronomic traits in tobacco. To better understand the population genetic structure and the genetic basis of plant height in tobacco, 437 tobacco germplasms were whole genome re-sequencing in this study. A total of 2,263,775 high-quality single nucleotide polymorphisms were identified. The analysis of the population genetic structure showed that tobacco germplasm could be divided into 8 clusters. In addition, gene flow was found between flue-cured tobacco and ustic tobacco, as well as between oriental tobacco and air-cured tobacco. Three genes associated with plant height were identified as candidate genes by genome-wide association study. This study provides valuable genetic resources for population structure analysis and elucidation of the genetic basis of various traits. It helps to improve the efficiency of molecular breeding.
Similar content being viewed by others
Introduction
Tobacco (Nicotiana tabacum L.) is an allotetraploid (2n = 4x = 48) by crossing Nicotiana sylvestris and Nicotiana tomentosiformis1. Tobacco is an important cash crop with leaves as the main product. It is widely cultivated between 55° N and 40° S latitudes2. Based on the processing after harvesting, tobacco is categorized into various types, including air-cured tobacco, sun-cured tobacco, and so on. Tobacco breeding has catered to commercial demands. Natural and artificial selection have led to the narrow genetic background of tobacco3, and an important model plant. In tobacco, the world’s first virus was discovered, called tobacco mosaic virus (TMV)4. Tobacco is sensitive to most of the known identified plant viruses and can be used to study plant-pathogenic microbial interactions. The method of Agrobacterium-mediated plant gene transformation was first developed in tobacco5. With the rapid development of sequencing technology and the completion of the assembly of tobacco genomes, the research on tobacco has been greatly promoted6. As the climate continues to change, the demand for new and improved tobacco varieties is increasing. Genetic diversity is essential for the continued development of breeding and the adaptation of plants to changing environments. Plant height (PH) is one of the most basic indicators in plant morphological investigations. PH affects yield to a certain extent. Linkage and association mapping methods have been used to locate quantitative trait loci in tobacco plant height7. Zhang et al. identified three sequence-related amplified polymorphism(SRAP) markers that may be associated with plant height8. Cheng et al. identified four SSR markers that may be associated with plant height7. Ikram et ·al. identified some candidate genes related to plant height using genotyping(GBS)9.
Whole genome resequencing is the sequencing of the genomes of different individuals of species with known genome sequences, and on this basis, the differences of individuals or populations are analyzed. Genome-wide association studies (GWAS) are a method to reveal the genetic basis of traits at the population level. These two methods are used in combination for plant analysis. A natural allele of a C2H2-type transcription factor was identified in rice by genome-wide association study, which conferred non-race-specific resistance to rice blast10. A ZmNAC78 gene was identified in maize through genome-wide association study, which regulates iron concentrations in maize kernels11. There have been studies to analyze tobacco by whole genome resequencing, but these studies are mainly based on SNP studies, with a small number of sequencing samples and relatively low sequencing depth12,13,14,15. Compared with the main food crops, the application of GWAS in tobacco is less studied. The analysis of the genetic structure of plant height through GWAS is conducive to the cultivation of excellent varieties suitable for plant height, which is helpful to cope with future weather changes, and lays an important foundation for the genetic study of complex quantitative traits.
In this study, a large amount of information on tobacco germplasm resources was provided by whole-genome resequencing of different tobacco germplasm, and the genetic diversity and relatedness among tobacco germplasm were further revealed by population genetic structure analysis of different tobacco species. The genome-wide association analysis was used to predict and analyze the genes related to tobacco plant height. This study will help to better understand the genetic diversity of cultivated tobacco germplasm and the genetic mechanisms of important agronomic traits, which will help breeders to develop more efficient and better quality variety development strategies.
Materials and methods
Plant materials, planting and phenotypic data
This study classifies tobacco populations based on the processing methods after leaf harvesting. Six types of tobacco germplasm resources were collected, including representative cultivated varieties. A total of 437 tobacco accessions were collected for this study, including 278 flue-cured tobacco (FT), 72 sun-cured tobacco (ST), 7 air-cured tobacco (AT), 38 burley tobacco (BT), 41 oriental tobacco (OT) and 1 rustic tobacco (RT). All accessions were planted at the experimental field of the Yunnan Academy of Tobacco Agricultural Sciences during the summer seasons in 2017 and 2018. Normal seeds were selected and sowed in deeply ploughed fields with proper moisture content (15–20%). The seed was planted in three-row plots in a randomized complete block design with three replications for each environment. Only one accession was planted in each plot, and the plots were 5 m in length with a row spacing of 0.4 m. The space between two plots was 0.5 m. All field management, including watering, weed management, and fertilization, was performed according to the usual local management in each test location for all accessions during the growing period. We cultivated a large number of plants and selected healthy ones for phenotypic measurements in the final stage. The same phenotyping procedure and scoring standards were adopted in all the environments. Phenotypic measurements were conducted using uniform tools and equipment to ensure consistent measurement conditions for all plants. The Plant Heighttraits of 372 accessions was characterized (276 flue-cured tobacco, 72 sun-cured tobacco, 7 air-cured tobacco, 11 burley tobacco, 5 oriental tobacco and 1 yellow flower tobacco), which related to yield, architecture, and quality with miss rate < 10%. Phenotypic data for each accession were defined as the average of the three replicates in the same environment. These plants were all evaluated at the same locations for two years (2017 and 2018).
DNA isolation and genome sequencing
Genomic DNA was extracted from fresh leaves of a single cultivated tobacco using the CTAB method. A total of 1.5 µg DNA from each accession was used as input for a DNA sequencing library, which was generated using the Truseq Nano® DNA HT sample preparation Kit (Illumina USA) following the manufacturer’s recommendations. Index codes were added to attribute sequences to each sample. The libraries were sequenced on the Illumina Novoseq platform to obtain 150 bp paired-end reads.
SNP calling
To obtain reliable reads, the raw reads (fastq format) were subjected to a series of quality control procedures to remove the low-quality reads (mainly resulting from base-calling duplicates and adapter contamination). About 19.46 Tb (~ 49.76 Gb per sample) of high-quality reads (Q20 > = 94.12% and Q30 > = 86.73%) were obtained.
The clean reads were mapped to the reference genome of N. tabacum with Burrows–Wheeler Aligner (version 0.7.8) software with the command ‘mem -t 4 -k 32 -M’16,17. The alignment results were converted to BAM files using SAMtools software18. The potential PCR duplicates were removed using the SAMtools command ‘rmdup’. Only the pair with the highest mapping quality will be retained if multiple read pairs have identical external coordinates. After alignment, SNP calling on a population scale was performed using a Baysesian approach, as implemented in the package SAMtools. A raw population genotype file with SNPs and InDel was created in this step. The SNPs supported by at least four mapped reads, mapping quality > = 20 and Phred-scaled genotype quality > = 5 with less than 0.1 missing data were considered as high-quality SNPs used for subsequent analyses. The method for InDel calling was similar to that for SNP calling, and only InDels of < = 5 bp were taken into account. Consequently, the identified SNPs and InDel were further annotated according to the reference genome using ANNOVAR (version 2013-05-20) software19. Based on the genome annotation, SNPs were categorized as being located in exonic regions (overlapping with a coding exon), splicing sites (within 2 bp of a splicing junction), intronic regions (overlapping with an intron), upstream and downstream regions (within a 1 kb region upstream or downstream from the transcription start site), and intergenic regions. The SNPs in coding exons were further grouped into synonymous or nonsynonymous mutations. The SNPs causing gain of a stop codon, loss of a stop codon or splicing were designated as large-effect SNPs.
To exclude false variants, SNPs were filtered by GATK (version 4.2.0.0) software20 based on the following parameters: QUAL < 30.0||MQ < 40.0||QD < 2.0||FS > 60.0||MQRankSum < − 12.5||ReadPosRankSum < − 8.0; Then SNPs were filtered by vcftools (version 0.1.13) based on the following parameters: max-missing > 0.9||mac < 3||minQ < 30||minDP < 3||maf < 0.05.
Phylogenetic tree, population structure and gene flow
A phylogenetic tree was constructed to elucidate the genetic relationships among different taxa. The analysis was performed using FastTree (version 2.1.10) software21, ML trees were constructed, and visualization was performed using R software. PCA analysis using Plink (version 1.90) software22 and visualization using R software. Population structure analysis was performed using ADMIXTURE (version 1.3) software23 and visualized using R software.
Gene flow between the six tobacco taxa was evaluated on the basis of SNPs with gene flow counts (m) from 0 to 6 using TreeMix (version 1.12) software24.
Genome-wide association study
Kinship analysis was performed with GEMMA (version 0.98.4) software25. GWAS analysis uses GEMMA’s mixed linear mode (MLM) and used both the PCA matrix and the kinship matrix as the covariate. Manhattan plot and QQ plot were visualized using R software.
The initial screening of plant height-related SNPs was carried out with a threshold of 5*10−8, and 353 SNPs associated with plant height were screened. Then further annotation was performed using SnpEff (version 5.0-1) software26 to obtain detailed information and prediction of the gene in which the SNP is located, and the CDS sequence of the gene in which the SNP is located was placed on NCBI for comparison to further screen for genes highly associated with the plant. The LD blocks were used to identify candidate gene regions using the LDBlockShow (version 1.4) software27.
Identification and annotation of candidate genes
GO enrichment analysis was used to identify all GO terms that were significantly enriched in genes associated with strain height identified in the GWAS analysis. All 117 genes associated with strain height were used for GO term classification and KEGG enrichment analysis using the eggNOG-mapper28,29. To further understand the biological functions of strain height-related genes, TBtools software was used to visualize the KEGG pathway30. Three genes with a high degree of presumed influence were further analyzed.
Result
Whole-genome resequencing and mapping
A total of 437 tobacco germplasm from different geographical locations were selected for genome sequencing, of which 372 germplasm were subjected to plant height determination (Supplementary Tables S1 and S4). These samples were sequenced with an average depth of 13× using the Illumina platform, filtered the raw reads to remove low-quality sequences and redundant reads, and obtained 21.90 Tb of raw bases (average 50.30 Gb per sample) and 21.84 Tb of clean bases (average 50.17 Gb per sample). The Q20 reads ranged from 94.12 to 98.43% with an average of 97.46%, and Q30 reads ranged from 86.73 to 97.22% with an average of 95.62%, indicating that the sequencing data were of good quality (Supplementary Table S1). The GC content ranged from 38.68 to 43.86%, averageing 39.68%. Comparisons were made with the G306 reference genome with an average mapping rate of 99.59% (78.53 to 99.86%) (Supplementary Table S2).
Variations calling and distribution
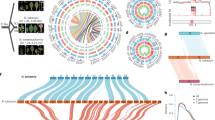
After completing sequencing, SNP locus variants were identified in the tested germplasm using SAMtools software, and 17,862,966 SNPs were identified; the identified SNPs were filtered and 2,263,775 high-quality SNPs were obtained for subsequent analysis. Analysis of the predicted mutation types of these SNPs showed that A/G and C/T transitions were the most frequent among the twelve possible single-base mutations, accounting for 19.77% and 19.71% of the total, respectively. Among the 2,263,775 SNPs, 1,546,844 were transitions and 716,931 SNPs were transversions. The ratio of transitions to transversions (Ts/Tv) was 2.16. further analysis of the distribution of SNPs in the genome revealed that 81.38% were located in intergenic regions, 3.62% in introns, 1.08% in exons, and 0.15% and 0.21% in 5′ and 3′ UTRs (Supplementary Table S3). The number of SNPs on LG1 was 1,443,328, the most among all chromosomes, while the number of SNPs on LG19 was 395,872, the least among all chromosomes, with an average of 744,290.25 SNPs per chromosome (Fig. 1).
Distribution of SNP. (A) Chromosomes of reference genome. (B) Genetic density distribution. (C) GC density distribution. (D) Distribution of SNP density. (E) Distribution of INDEL density. (F) Distribution of Tajima’s D. (G) Gene collinearity of reference genome. LG, chromosome.
Population structure
The 2,263,775 high-quality SNPs obtained after filtering were used for population structure analysis. PCA showed that the percentage of overall variation explained by PC1 and PC2 was 44.95% and 18.73%, respectively (Fig. 2A). Population genetic evolution analysis found that different types of tobacco germplasm were clustered, which was consistent with the results of PCA analysis (Fig. 2B). The kinship of 437 tobacco germplasm points was also analyzed, and the results showed that the direct kinship of different tobacco germplasm is very complex, which is consistent with the results of population structure analysis (Fig. 2C, Supplementary Fig. 1). Further, ADMIXTURE was used for population structure analysis, in which the number of clusters is usually determined based on the cross-validation error rate, and the number of clusters with the lowest cross-validation error rate is the optimal number of clusters. The cross-validation error rate was lowest when K = 8, which indicated that the 437 tobacco germplasm could be divided into 8 clusters (Fig. 2D).
Analysis of group structure. (A) Principal component analysis. (B) Phylogenetic tree. BT Burley tobacco, green; RT rustic tobacco, orange; FT flue-cured tobacco, pink; AT air-cured tobacco, red; ST sun-cured tobacco, light blue; OT oriental tobacco, dark blue. (C) Population genetic structure analysis. (D) Cross-validation error rate of population genetic structure analysis.
Gene flow analysis
Gene flow between these six tobacco taxa was analyzed. According to the variance explained analysis, when allowing for two mixing events (m = 2) (Supplementary Fig. 2) the results were optimal, gene flow was extensive between FT and RT, and OT and AT, which most likely reflects the many common genomic components they share due to hybridization during domestication and breeding (Fig. 3).
Detection of gene flow among six tobacco taxa. Lines represent gene flow; arrows indicate the direction of gene flow. Scale bar shows the mean, standard error of entries in the sample covariance matrix as tenfold. Color bars show migration weights: red indicates strong gene flow, and yellow indicates weak gene flow.
GWAS of plant height
On the basis of the 2,263,773 SNPs, GWAS analysis of plant height was performed using two types of data. The first category was phenotype data from two years (Supplementary Table S4). The other category was the overall performances of the 379 accessions were predicted as the best linear unbiased prediction (BLUP) using a mixed linear model (MLM). The present study used two years of phenotypic records to perform GWAS on plant height. A strong GWAS peak on chromosome 1 was identified in the Manhattan chart (Fig. 4A). The Q–Q plot reflects the good results of the correlation analysis (Fig. 4B).
Genome-wide association study (GWAS). (A) GWAS of plant height. (B) Q–Q plot of plant height.
Prediction of candidate genes for plant height
A total of 353 SNPs highly significantly associated with the plants were screened using P < 5*10−8 as the threshold. These 353 SNPs were further annotated and were found to have 3 putative impacts of LOW and above (Supplementary Tables S5 and S6). These three SNPs were LG1:155162419, LG1:152799784 and LG1:153204197, and the variant annotation types were: missense variant, missense variant and splice region variant & intron variant, respectively, in the CDS sequences of the SNPs were also BLASTed at NCBI, and the results were Nicotiana tabacum alpha-farnesene synthase-like (LOC107765628), Nicotiana tabacum uncharacterized LOC107776276 (LOC107776276) and Nicotiana tabacum tetrahydrocannabinolic acid synthase-like (LOC107807513). To further confirm that these three SNPs were associated with plant height, SNP typing was subsequently investigated and haplotype results for all three SNPs showed significant differences in plant height between mutated and unmutated plants (Fig. 5).
The haplotypes of the three SNPs were studied separately. LG1:152799784 had haplotype A, and the plant height was higher and significant compared to the unmutated one (Fig. 5A), and LD analysis showed linkage (Fig. 5D). LG1:153204197 also had haplotype A. The plant height was higher and significant compared to the unmutated one (Fig. 5B), and LD analysis showed strong linkage (Fig. 5E). The haplotype of LG1:155162419 was T, and the plant height was also taller and significant compared to the unmutated one (Fig. 5C), and LD analysis showed the same strong linkage (Fig. 5F).
Haplotype analysis of three SNPs. (A) Haplotype analysis of LG1:152799784. (B) Haplotype analysis of LG1:153204197. (C) Haplotype analysis of LG1:155162419. (D) 50 kb LD Block of LG1:152799784. (E) 50 kb LD Block of LG1:153204197. (F) 50 kb LD Block of LG1:155162419.
Functional prediction of candidate genes by GWAS
GO functional enrichment analysis was performed on the genes identified in the GWAS analysis that were associated with strain height. The results showed that the enrichment was mainly in molecular functions and biological processes (Fig. 6). The molecular functions mainly included sesquiterpene synthase activity, terpene synthase activity, carbon-oxygen lyase activity, acting on phosphates, carbon-oxygen lyase activity, lyase activity; biological processes mainly include sesquiterpene metabolic process, sesquiterpene biosynthetic process, terpene biosynthetic process, terpene metabolic process, hydrocarbon biosynthetic process, etc. (Fig. 6A). Further, GO enrichment analysis was then performed for the three genes predicted to have a high level of impact, with GO enrichment results for evm.model.LG1.5430 includs magnesium ion binding, terpene synthase activity, lyase activity, and then GO enrichment results for evm.model.LG1.5499 included oxidoreductase activity, flavin adenine dinucleotide binding, obsolete oxidation-reduction process, FAD binding. However, no enrichment results for evm.model.LG1.5437 were available (Supplementary Table S7).
In order to understand the biological pathways of the predicted genes associated with plant height, KEGG pathway enrichment analysis on these genes were performed. The results showed that they are mainly involved in Sesquiterpenoid and triterpenoid biosynthesis, Metabolism of terpenoids and polyketides, Metabolism, Diterpenoid biosynthesis, and Aminoacyl-tRNA biosynthesis, etc. (Fig. 6B). Further, KEGG enrichment analysis of the three genes showed that evm.model.LG1.5430 was mainly associated with alpha-farnesene synthase, evm.model.LG1.5499 was mainly associated with cysteine desulfurase, while evm.model. LG1.5437 remained unenriched (Supplementary Table S8).
GO and KEGG enrichment analysis of candidate genes. (A) GO Term enrichment analysis of candidate genes. (B) KEGG Pathway enrichment analysis of candidate genes.
Discussion
In this study, 437 tobacco germplasms were whole-genome resequencing and 2,263,775 high-quality SNPs were found. Analysis of population structure revealed that the genetic relationship between tobacco germplasm was very complex (Fig. 2C), which may be caused by interspecific hybridization during the breeding process. The ADMIXTURE analysis found that tobacco could be divided into 8 clusters (Fig. 2D), which were significantly different from the 6 species groups we provided (Fig. 2B). This difference may be due to the classification method based on morphology and baking modulation methods.
Gene flow exists between different types of tobacco. Gene flow is a major factor affecting the purity of crop varieties31, the results of gene flow analysis of six tobacco species showed that gene flow exists between FT and RT, and between OT and AT (Fig. 3). It can be seen in the phylogenetic tree and PCA plot that FT and RT are partially clustered together, and OT and AT are partially clustered together. This suggests that there may be direct purposeful crosses between different species of tobacco during the breeding process (Fig. 2B). In addition, gene flow occurs through three routes: pollen, seed, and nutrient propagule-mediated32,33, and pollen-mediated gene flow is influenced by physical and environmental conditions; pollen-mediated gene flow does not occur between wild-type and cultivated plants when grown under greenhouse conditions31, but bees and other insects can spread tobacco pollen in between flowers34, so pollen-mediated gene flow may have occurred during breeding or growing in the field.
Compared to previous studies, our study used a larger number of tobacco species, expanded the sample size, and identified more SNPs through whole genome sequencing averaging over 10X. Plant height is a complex quantitative trait, and three SNPs related to plant height were identified by GWAS, namely LG1:155162419, LG1:152799784, and LG1:153204197. They are all located in chromosome 1 (Fig. 4). The haplotype results showed a significant difference in plant height between mutated and non-mutated plants (Fig. 5). GO term and KEGG pathway analysis found that they were involved in a variety of biosynthetic processes. The regulation mechanism of the three genes on plant height is worthy of further study. The study provided tremendous information on tobacco germplasm genetic resources and identified 17,862,966 SNPs, which will provide the basis for more accurate identification of disease-causing gene variants in the near future and facilitate new breeding strategies such as marker-assisted or genomic selection and low-cost genome editing for favourable traits for a cost-effective and environmentally friendly tobacco industry.
Data availability
The data supporting the findings of this study are available from the corresponding author (lyang@sdau.edu.cn).
References
Wang, J. et al. High-quality assembled and annotated genomes of Nicotiana tabacum and Nicotiana benthamiana reveal chromosome evolution and changes in defense arsenals. Mol. Plant https://doi.org/10.1016/j.molp.2024.01.008 (2024).
Jia, L. et al. Study on the change of global ecological distribution of Nicotiana tabacum L. based on MaxEnt model. Front. Plant Sci. 15, 1–11. https://doi.org/10.3389/fpls.2024.1371998 (2024).
Lewis, R. S. & Nicholson, J. S. Aspects of the evolution of Nicotiana tabacum L. and the status of the United States Nicotiana Germplasm Collection. Genet. Resour. Crop Evol. 54, 727–740. https://doi.org/10.1007/s10722-006-0024-2 (2006).
Creager, A. N. H., Scholthof, K.-B.G., Citovsky, V. & Scholthof, H. B. Tobacco mosaic virus: Pioneering research for a century. Plant Cell 11, 301–308. https://doi.org/10.1105/tpc.11.3.301 (1999).
Barton, K. A., Binns, A. N., Matzke, A. J. M. & Chilton, M.-D. Regeneration of intact tobacco plants containing full length copies of genetically engineered T-DNA, and transmission of T-DNA to R1 progeny. Cell 32, 1033–1043. https://doi.org/10.1016/0092-8674(83)90288-x (1983).
Sierro, N. et al. The tobacco genome sequence and its comparison with those of tomato and potato. Nat. Commun. 5, 1–9. https://doi.org/10.1038/ncomms4833 (2014).
Cheng, L. et al. Quantitative trait loci mapping for plant height in tobacco using linkage and association mapping methods. Crop Sci. 55, 641–647. https://doi.org/10.2135/cropsci2014.05.0404 (2015).
Zhang, J. et al. Genetic diversity of agronomic traits and association analysis with SRAP markers in flue-cured tobacco (Nicotiana tabacum) varieties from China and abroad. Acta Agron. Sin. 38, 1029–1041 (2012).
Ikram, M. et al. Genetic dissection of tobacco (Nicotiana tabacum L.) plant height using single-locus and multi-locus genome-wide association studies. Agronomy 12, 1047. https://doi.org/10.3390/agronomy12051047 (2022).
Li, W. et al. A natural allele of a transcription factor in rice confers broad-spectrum blast resistance. Cell 170, 114–126. https://doi.org/10.1016/j.cell.2017.06.008 (2017).
Yan, P. et al. Biofortification of iron content by regulating a NAC transcription factor in maize. Science 382, 1159–1165. https://doi.org/10.1126/science.adf3256 (2023).
Thimmegowda, G. C. et al. Whole genome resequencing of tobacco (Nicotiana tabacum L.) genotypes and high-throughput SNP discovery. Mol. Breed. 121, 1–10. https://doi.org/10.1007/s11032-018-0876-0 (2018).
Tong, Z. et al. Construction of a high-density genetic map with whole genome sequencing in Nicotiana tabacum L. Genomics 112, 2028–2033. https://doi.org/10.1016/j.ygeno.2019.11.015 (2020).
Yang, H., Geng, X., Zhao, S. & Shi, H. Genomic diversity analysis and identification of novel SSR markers in four tobacco varieties by high-throughput resequencing. Plant Physiol. Biochem. 150, 80–89. https://doi.org/10.1016/j.plaphy.2020.02.023 (2020).
Wang, Y. et al. Construction of a SNP fingerprinting database and population genetic analysis of cigar tobacco germplasm resources in China. Front. Plant Sci. 12, 618133. https://doi.org/10.3389/fpls.2021.618133 (2021).
Edwards, K. D. et al. A reference genome for Nicotiana tabacum enables map-based cloning of homeologous loci implicated in nitrogen utilization efficiency. BMC Genom. 18, 448. https://doi.org/10.1186/s12864-017-3791-6 (2017).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595. https://doi.org/10.1093/bioinformatics/btp698 (2010).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, 1–4. https://doi.org/10.1093/gigascience/giab008 (2021).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164–e164. https://doi.org/10.1093/nar/gkq603 (2010).
McKenna, A. et al. The genome analysis toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. https://doi.org/10.1101/gr.107524.110 (2010).
Poon, A. F. Y., Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490. https://doi.org/10.1371/journal.pone.0009490 (2010).
Chang, C. C. et al. Second-generation PLINK: Rising to the challenge of larger and richer datasets. GigaScience 4, 1–16. https://doi.org/10.1186/s13742-015-0047-8 (2015).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664. https://doi.org/10.1101/gr.094052.109 (2009).
Tang, H., Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, 1–48. https://doi.org/10.1371/journal.pgen.1002967 (2012).
Zhou, X. & Stephens, M. Genome-wide efficient mixed-model analysis for association studies. Nat. Genet. 44, 821–824. https://doi.org/10.1038/ng.2310 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92. https://doi.org/10.4161/fly.19695 (2014).
Dong, S.-S. et al. LDBlockShow: A fast and convenient tool for visualizing linkage disequilibrium and haplotype blocks based on variant call format files. Brief. Bioinform. 22, 1–6. https://doi.org/10.1093/bib/bbaa227 (2021).
Cantalapiedra, C. P. et al. eggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol. Biol. Evol. 38, 5825–5829. https://doi.org/10.1093/molbev/msab293 (2021).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592. https://doi.org/10.1093/nar/gkac963 (2023).
Chen, C. et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol. Plant 16, 1733–1742. https://doi.org/10.1016/j.molp.2023.09.010 (2023).
Kim, D.-S., Song, I. & Ko, K. Low risk of pollen-mediated gene flow in transgenic plants under greenhouse conditions. Hortic. Environ. Biotechnol. 59, 723–728. https://doi.org/10.1007/s13580-018-0074-3 (2018).
Org, Z. The Economics of Genetically Modified Crops (2009).
Kang, Y., Shin, Y. K., Park, S.-W. & Ko, K. The Economics of Genetically Modified Crops (2016).
Loureiro, I. et al. Pollen-mediated gene flow in the cultivation of transgenic cotton under experimental field conditions in Spain. Ind. Crops Prod. 85, 22–28 (2016).
Acknowledgements
This work was supported by the Science Project 110202101042 (JY-19)/2022530000241007, the Foundation of Innovation Team Project for Modern Agricultural Industrious Technology System of Shandong Province (SDAIT-25-01) , and the platform construction for tobacco genome breeding (2017YN05). We thank Supercomputing Center in Shandong Agricultural University for technical support.
Author information
Authors and Affiliations
Contributions
Conceptualization, Long Yang, Fangchan Jiao; methodology, Long Yang; software, Yuanheng Wang; validation, Yanru Song, Yuanheng Wang; formal analysis, Yongping Li; investigation, Jie Ding, Xingfu Wu; resources, Fangchan Jiao; data curation, Yanru Song, Ying Liu; writing—original draft preparation, Yuanheng Wang; writing—review and editing, Yanru Song, Ying Liu, Haozhen Li; visualization, Long Yang; supervision, Long Yang; funding acquisition, Fangchan Jiao, Long Yang. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, Y., Wang, Y., Liu, Y. et al. Whole genome re-sequencing in 437 tobacco germplasms identifies plant height candidate genes. Sci Rep 15, 4734 (2025). https://doi.org/10.1038/s41598-025-88064-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88064-7