Abstract
Carmine radish (Raphanus sativus L.) is cultivated in Fuling, Chongqing, for its red color. Dof-TFs are critical in regulating plant growth, development, stress responses, and signal transduction.This work comprehensively examined the structure, evolution, and expression of the carmine radish Dof gene and its behavior under cadmium (Cd) stress. The radish genome has 59 RsDofs, which are divided into nine clusters (A: 8, B1: 10, B2: 10, C1: 3, C2.1: 5, C2.2: 4, C3: 11, D1: 4, and D2: 4). Phylogenetic tree analysis revealed significant Dof gene family resemblance between Arabidopsis thaliana and Brassica napus. Perhaps segment duplication resulted in RsDof gene family expansion. Cd stress-induced RsDof expression patterns were studied using an RNA-seq atlas and qRT-PCR. The majority of RsDofs were tissue-specific and Cd-sensitive. The involvement of RsDof genes in Cd stress response and anthocyanin synthesis was verified using qRT-PCR. RsDof33 is involved in Cd stress response and anthocyanin synthesis. A. thaliana overexpressed the recombinant fusion protein RsDof33-GFP, which was localized to the nucleus, resulting in fewer rosette leaves, delayed flowering, and higher anthocyanin concentration. RsDof33-expressing plants had significantly higher transcript levels of the auxin biosynthetic genes YUCCA (AtYUC2), auxin efflux carrier (AtPIN4), and AtKNAT2, which are involved in leaf shape development, as well as AtPAL, AtCHS, AtCHI, AtDFR, AtLDOX, and AtUF3GT. These findings indicate that RsDofs are critical to plant development and stress responses.
Similar content being viewed by others
Introduction
Plants encounter several environmental concerns, with heavy metal toxicity, particularly from cadmium (Cd), being one of the most significant issues 1,2. Cd is detrimental to plants, disrupting their physiological systems, inducing stress, and impairing growth and productivity 3. Plants use specialized proteins known as transcription factors (TFs) to manage challenging conditions by regulating the expression of crucial genes that assist in stress response 4,5. A specific category of these facilitators, Dof TFs, is unique to plants and performs several functions in promoting growth, responding to stress, and synthesizing beneficial compounds 6.
Dof TFs possess a distinct component known as the zinc-finger DNA-binding domain (C2-C2 type) 7. Dof proteins consist of two primary components: a stable domain at the N-terminus that interacts with DNA and a variable domain at the C-terminus that regulates gene expression and interacts with other proteins 8. This adaptable design incorporates Dof elements into several critical functions, including seed germination, photoperiod regulation, plant development, hormonal activity, and the management of issues such as drought, salinity, and toxic metals 9.
Dof TFs have been extensively researched across several plant species. Dof factors in Arabidopsis thaliana regulate vascular tissue development (AtDof2.1), blooming time (AtDof5.8), and stress response (AtDof4.7) 10. Dof variables in crops such as rice and maize are associated with efficient nitrogen use, carbon management, and drought response 11,12. In plants such as soybean, they influence lignin synthesis and nodule formation, while in tomato, they facilitate ethylene production and fruit ripening. These studies demonstrate the many functions of Dof factors in plants 13. However, our understanding of their functions in less-researched plants and their involvement in assisting plants in managing heavy metals remains limited.
The Carmine radish (Raphanus sativus L.), cultivated extensively in Fuling, Chongqing, has vibrant scarlet roots 14. This hue derives from anthocyanins, which are unique compounds that assist in safeguarding the plant from harm inflicted by elements such as heavy metals 15. Although it is established that anthocyanins assist plants in managing stress, the regulatory mechanisms governing their synthesis under heavy metal stress remain incompletely understood. A recent study indicates that Dof transcription factors may play a crucial role in regulating anthocyanin synthesis by modulating the expression of specific genes 16. The genes implicated in anthocyanin biosynthesis include CHS, DFR, and UF3GT.
This study included a comprehensive investigation of Dof TFs across the carmine radish genome to elucidate their role in stress adaptation and anthocyanin biosynthesis. We identified 59 RsDof genes and categorized them into nine categories according to their evolutionary relationships. We examined the expression of these genes under cadmium stress via RNA sequencing and qRT-PCR, demonstrating that their activity varies according to the plant component and the stress condition. RsDof33 is a crucial gene for managing cadmium stress and synthesizing anthocyanins. Increasing the expression of this gene in Arabidopsis thaliana enhanced anthocyanin production and altered auxin synthesis in the plant.
These findings provide new insights into the diverse architectures and significant functions of Dof TFs in plants. They demonstrate how Dof TFs assist plants in managing stress and producing secondary metabolites such as anthocyanins. This study enhances our understanding of how plants manage stress induced by cadmium exposure. It proposes methods to modify plants to improve their stress resilience and increase anthocyanin production, perhaps benefiting agricultural yields.
Materials and methods
Identification of Dof genes in the carmine radish
The radish genomic sequence (http://www.radish-genome.org/) was retrieved from the radish genomic database. The Hidden Markov Model (HMM) for the Dof domain (PF02701) was obtained from Pfam56, and HMMER 3.0 was employed to identify potential Dof TFs in carmine radish using the annotated protein database. An HMM specific to carmine radish Dof proteins was constructed using hmmbuild, then employed to search for Dof candidates with an e-value cut-off of 1e-10. These candidates were further validated using the PlnTFDB and PlantTFDB databases and confirmed to contain the DOf domain using SMART, Prosca, and CDD. Several bioinformatics tools were utilized to analyze the physical and chemical properties, subcellular localization, and transmembrane characteristics of the confirmed RsDof proteins, including ProtParam (http://web.expasy.org/protparam) 17, WoLF PSORT (https://wolfpsort.hgc.jp/) 18, and TMHMM Server v2.0 (http://www.cbs.dtu.dk/services/TMHMM/) 19, respectively.
Analyzed of the conserved domain, the structure of RsDofs, and their phylogenetic evolution
Multiple protein sequence alignments of Dof domains were generated using ClustalW, and the results were optimized using GeneDoc software. The gene structure of RsDofs was visualized using Tbtools software, and their full-length protein sequences were analyzed. MEME suite 4.11.4 was used to identify conserved motifs with reference to Bailey, et al. 20, and sequence logos were created using the weblogo platform. Full-length protein sequences of Dof from eleven multiple species were downloaded from the Phytozome v13 database for phylogenetic tree analysis, including Raphanus sativus(Rs), Arabidopsis thaliana (At), Brassica napus(Bn), Solanum lycopersicum(Sl), Glycine max(Gm), Zea mays(Zm), Oryza sativa(Os), Amborella trichopoda(Atr), Populus trichocarpa(Pt), Sorghum biocolor(Sb) and Lotus corniculatus(Lc) 21. Then these sequences were aligned using MUSCLE with default settings, and an unrooted phylogenetic tree was constructed using the Maximum Likelihood method in MEGA-X (https://www.megasoftware.net/) with 1000 replicates bootstrap test 21,22.
Gene duplication and syntenic analysis of RsDofs
The RsDof TFs were allocated to the nine radish chromosomes through Circos visualization, utilizing physical location data extracted from the GFF file of the radish genome. The duplication events of RsDofs were assessed employing the Multiple Collinear Scanning Toolkit (MCScanX) 23. Furthermore, a collinearity examination was performed on the protein sequences of Arabidopsis, Brassica napus, and Raphanus sativus. To visualize the gene duplication and collinearity, Tbtool and Circos were used with default parameters 24. The results were depicted to illustrate the gene duplication events and collinearity relationships between the different species.
Cis-acting element analysis and protein interaction network prediction of RsDofs
The promoter sequences (upstream of 1.5 Kb) of RsDof TFs were analyzed using the Plant-CARE software to identify putative cis-acting elements 25. These elements provide insights into the potential regulatory mechanisms of RsDof TFs. Additionally, a protein interaction network was predicted for all putative RsDof protein sequences using the STRING web server 26, and the network was visualized using Cytoscape 27.
Expression patterns analysis of RsDofs by RNA-Seq data
The expression profiles of RsDof TFs in various tissues of carmine radish were explored using transcriptome data provided by the research lab (Jian Lab), encompassing inflorescence, fruit, flower, leaf, pericarp, and pulp 28. Furthermore, the expression patterns of RsDof TFs were investigated in response to heavy metal stress in radish(NCBI projects:). Transcriptome data from heavy metal-tolerant radish ‘XCB’ and heavy metal-susceptible radish ‘HX’ subjected to different treatments of Cd stress (0 h, 12 h, 24 h, 48 h, 72 h) were acquired. All raw data from the transcriptome libraries were collected from NCBI with accession number PRJNA832540. The triplicate data were averaged, log2-transformed (FPKM), and visualized by generating heatmaps using a plugin integrated within the Tbtools software.
Correlation network construction
The expression patterns of RsDof TFs and anthocyanin biosynthesis genes (ABGs) were investigated using RNA sequencing data from seven distinct radish cultivars. Heatmaps visualizing the expression patterns were generated using a Tbtools software plugin. Furthermore, significant variations in anthocyanin metabolites among the cultivars’ 'WW,' 'HX-'1', and 'HX-'3' were identified. A complex network was constructed to explore the relationships between ABGs, RsDof TFs, and anthocyanin metabolites. This network was established based on Pearson’s correlation coefficients, and correlations above 0.8 were visualized using Cytoscape software.
Plant material and experiment design
To verify the expression patterns of selected genes in response to cadmium stress, quantitative real-time polymerase chain reaction (qRT-PCR) analysis was conducted. The expression of RsDof35-a, RsDof15-a, RsDof1-a, RsDof1-b, RsDof10, RsDof11, RsDof21-a, RsDof21-b, RsDof33, RsDof14-a, RsDof9-a, and RsDof32-a genes was examined across various developmental stages of carmine radish, including the seedling stage (SS), initial expansion (IE), full expansion (FE), bolting stage (BS), initial flowering stage (IFS), full-bloom stage (FBS), and podding stage (PS). The RNAseq data were collected from NCBI with accession number (PRJNA565866) 15. This approach aimed to explore the potential role of these genes in radish development and anthocyanin biosynthesis.
RNA isolation and qRT-PCR expression analysis
In 2020, the experimental plants were cultivated in a glasshouse at the research facilities of Yibin University in Yibin, China. Two varieties of radishes, namely the heavy metal-tolerant ‘XCB’ and the heavy metal-susceptible 'HX,' were selected for the study. These radish plants were exposed to various durations of heavy metal stresses. Root samples from seedlings under control conditions (CK) and under cadmium stress were collected at different time intervals (0 h, 12 h, 24 h, 48 h, and 72 h) with three replicates, following a previous experimental procedure (Gao et al., 2020). Additionally, pulp samples from distinct developmental stages ["SS_root," "IE_root," "FE_root," “BS_root”, “IFS_root”, “FBS_root”, and “PS_root”] of carmine radish were harvested in triplicates. These RNA samples were then utilized for qRT-PCR analysis. Twelve candidate TFs (RsDofs) showing significant expression changes were chosen based on their tissue expression profiles and were confirmed through qRT-PCR. Primer design was conducted using Primer 5.0 software, and the radish actin gene was employed as an internal reference gene (Table S1). The qRT-PCR assays were conducted following the standard protocol of the ABI7500 system, as outlined by 29, with triplicate technical replicates. Expression levels were quantified as fold-changes using the relative quantitative approach (2^-ΔΔCT), following the methodology described by Bustin, et al. 30. Furthermore, RNA was isolated from Arabidopsis thaliana leaves using the RNAprep Pure Plant Kit (Tiangen). First-strand cDNA synthesis was performed using the PrimeScriptTM RT Master Mix (TaKaRa) according to the manufacturer’s instructions. The expression levels of RsDof33 and other genes in wild-type (WT) and transgenic lines were assessed using specific primers (Table S1). Arabidopsis Actin2 was used as the internal control for quantification. Additionally, qRT-PCR was employed to examine auxin biosynthetic genes (AtYUC2, AtYUC4, and AtYUC6), auxin transport genes (AtAUX1, AtPIN1, AtPIN3, and AtPIN4), as well as genes associated with leaf morphogenesis (AtKNAT2, AtKNAT6, AtDPA4, and AtCUC2) in OE1 transgenic lines, followed by OE7 and OE8, along with the WT. Moreover, relative expression analysis of anthocyanin biosynthesis genes (ABGs) by qRT-PCR in the leaves of wild-type and transgenic lines (OE7 and 8) involved in anthocyanin biosynthesis was also conducted for identification purposes.
Subcellular localization of RsDof33 and construction of over-expression (OE) of RsDof33 in A. thaliana
For the subcellular localization analysis of RsDof33, Arabidopsis thaliana protoplasts were utilized as the experimental system. The coding sequence of RsDof33 was fused with the green fluorescent protein (GFP), under the control of a CMV35S promoter, within an expression plasmid. Subsequently, this construct was introduced into Arabidopsis protoplasts for subcellular localization assessment, following the methodology outlined by Yoo, et al. 31. An empty GFP gene was employed as a negative control. Confocal fluorescent imaging was conducted using a confocal scanning laser inverted microscope (LSM510 Meta; Carl Zeiss, Jena, Germany). To construct the over-expression plasmid, the full-length open reading frame (ORF) of RsDof33 was cloned into the linearized vector p1300-GFP. The RsDof33 coding sequence was integrated at the XbaI site of the p1300-GFP vector, replacing the CaMV35S promoter, thereby generating the p1300-RsDof33-GFP construct. The Agrobacterium-mediated floral dip method, as detailed by Clough and Bent 32, was employed for the genetic transformation of Arabidopsis. Transgenic Arabidopsis seeds were screened on MS medium supplemented with antibiotics, and T1-positive transgenic plants were identified by PCR amplification using specific primers with leaf genomic DNA as the template. T3 transgenic plants were obtained through successive screening across three generations. Total protein extraction from p1300-RsDof33-GFP T3 transgenic plants was carried out using a designated extraction buffer (Tris–HCl, pH 7.5, 50 mM; NaCl, 100 mM; EDTA, pH 8.0, 1 mM; glycerol, 10%; SDS, 0.5%). Western blot analyses were conducted utilizing GFP antibody (ABMART), with total protein from Arabidopsis serving as the control.
Data statistical analysis
A variance analysis was performed to evaluate differences in the relative expression levels of the genes across the various samples, followed by multiple comparisons using Duncan’s least significant range (LSR) tests in the SPSS statistical software. Different letters denoted significant differences at the p = 0.05 level. Specifically, the presence of a letter (a, b, c, d, e, f, or g) indicated a significant difference from a column lacking a common superscript letter (P < 0.05). Conversely, columns labeled with the same letter were not significantly different at the 5% level, based on Duncan’s multiple range test.
Results
Identification and physiochemical characteristics of RsDofs in radish
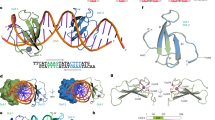
Through comprehensive searching of the radish genome using HMM with the Dof domain (PF02701), 59 RsDof family members were identified, and further validation via SMART, Prosca, and CDD. Following the distribution of Dof genes of their closet orthologs in A. thaliana, RsDofs were named RsDof1 to RsDof36-b. Their responding physical and chemical properties were identified as varied in those RsDofs and shown in Table S2. Analysis of RsDof proteins showed that lengths of amino acid, molecular weights, and isoelectric points ranged from 167 to 437 aa, 18.9 to 431.5 kDa, 4.81 (RsDof2) to 9.9 (RsDof7-a) respectively. Subcellular localization prediction indicated that 46 RsDof proteins were localized in the nucleus, while the remaining proteins were found in the chloroplast (9), mitoplast (2), and cytoplasm (1). Additionally, to investigate the sequence characteristics of RsDof proteins, a multiple sequence alignment analysis was performed to construct the Dof structural domain using Clustal W. It is indicated that the typical CX2CX21CX2C motif was highly conserved contained in RsDof ‘proteins’ structural domain sequences, where four Cys residues could be covalently combined by Zn2+ (Fig. 1).
Multiple sequence alignment of Dof domain for RsDof proteins in radish. (A) Extraction of conserved domains of Dof proteins (B) the conserved domains shown by WebLogo.
Phylogenetic analysis and classification of RsDofs
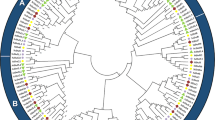
A comprehensive analysis of the evolutionary relationships among RsDof proteins involved the alignment of 572 Dof protein sequences from Arabidopsis (36), Brassica napus (112), and Radish (59 identified RsDofs) to construct a phylogenetic tree (Fig. 2). The resulting phylogenetic tree delineated the RsDof homologs into nine groups (A to I), further subdivided into 14 subgroups: A1, A2, B, C, D1, D2, D3, D4, E1, E2, F, G, H, and I (Fig. 2). Group A emerged as the largest clade encompassing 56 members, while Group B represented the smallest clade with only seven members. Notably, a significant proportion of Dofs from O. sativa and S. bicolor clustered within Group F, indicating their close evolutionary relationship as grass species.
Phylogenetic tree of Dof proteins in Amborella trichopoda, Sorghum bicolor, Zea mays, Solanum lycopersicum, Oryza sativa, Lotus corniculatus, Glycine max, Populus trichocarpa, Arabidopsis thaliana, Brassica napus and Raphanus sativus. This phylogenetic tree was generated using the neighbor-joining method with 1000 bootstraps in MEGA. Eleven Dof subfamilies were classified with different colors, and their names are marked in the corresponding positions. The circle, five-pointed star, and square text indicate A. thaliana, B. napus, and R. sativus proteins, respectively.
Chromosome distribution, gene duplication, and Synteny analysis of RsDofs
To elucidate the evolutionary mechanisms underlying the expansion of RsDofs genes in radish, their chromosomal localization and gene duplication events were investigated. The analysis revealed an uneven distribution of RsDofs on the chromosomes, with 59 RsDofs identified across nine chromosomes. Notably, Chr 5 harbored the highest number of RsDofs (10), while Chr 3 contained the fewest (2) RsDof genes (Fig. 3). Furthermore, the study identified 30 segmental duplications and two sets of tandem duplication (RsDof24-f-RsDof24-d and RsDof24-e-RsDof24-b) among RsDofs gene pairs. These duplication events are likely key contributors to the expansion of the gene family in radish (Fig. 3, Table S3).
Chromosomal locations of RsDof genes and synteny analysis of interchromosomal relationships of RsDof genes in radish. The gray blocks indicate parts of the radish chromosomes. The red lines indicate segment duplicated RsDof gene pairs. Blue text indicates tandem duplicated RsDof gene pairs.
Gene structure and conserved motifs of RsDofs
To assess the diversity and conservation of the 59 RsDofs genes in radish, their conserved motifs were analyzed using MEME, resulting in the prediction of ten distinct motifs (ranging from 19 to 50) (Figure S1). Notably, Motif 1 was identified as the conserved structural domain of Dof, characterized by the CX2CX21CX2C single zinc finger structure. Additionally, comparable motif structures of RsDofs proteins were observed within evolutionary branches of the phylogenetic tree, indicating conservation among certain motif structures.
The distribution of motifs revealed distinct patterns among subgroups, with motifs 10, 7, 5, and 7 exclusives to the B1 subgroups, motifs 2 and 10 to the B2 subgroups, and motifs 8 and 4 to the C3 and A subgroups, respectively (Fig. 4A,B). Furthermore, analysis of intron–exon structures indicated minimal variation in the number of introns (ranging from 0 to 1) among RsDof genes. Specifically, 26 and 33 RsDofs members were found to possess either one or no introns, predominantly located upstream of the Dof structural domain (zf-dof) (Fig. 4C). This was further supported by the phylogenetic tree, which demonstrated similar exon–intron patterns within the same gene groups.
Structural analysis of Dof genes in radish. (A) a phylogenetic tree of RsDof protein was generated using the neighbor-joining method with 1000 bootstraps in MEGA. (B) MEME (suite 4.11.4) was used to identify the conserved motifs based on the RsDof protein sequences, and a colored box numbered (1 to 20) is indicated in each motif at the bottom. (C) The Dof domain was identified using NCBI CD-hit, the Dof domain was constructed using TBtools software, and different groups of the Dof family are divided with different colors. (D) Exon–intron structure of RsDOF genes. Green boxes indicate untranslated 5′- and 3′-regions; yellow boxes indicate exons; black lines indicate introns.
Promoter analysis of RsDof genes
In order to elucidate the transcriptional regulation by RsDof gene family members, the cis-regulatory elements within their promoters were investigated. Notably, essential elements such as the TATA-box and CAAT-box were identified in all RsDof genes, while elements associated with plant growth and development, including Zein metabolism regulation (O2-site) and meristem expression (CAT-box), were also prevalent. Moreover, a diverse array of phytohormone-responsive elements, such as ABREs for abscisic acid, TGACG-motif and CGTCA-motif for MeJA, TCA-element for salicylic acid, and P-box for gibberellin, were observed in the promoters of RsDof genes.
This suggests that RsDof genes are subject to regulation by various phytohormones and potentially participate in plant hormone signaling pathways. Furthermore, the presence of elements responsive to anaerobic induction (ARE), light responsiveness (G-box, GT1-motif, Box-4, and AE-box), as well as defense and stress response elements (TC-rich) in the promoters of RsDof genes further indicates their involvement in plant growth, development, stress responses, and defense mechanisms. In summary, the analysis of cis-regulatory elements in the promoters of RsDof genes underscores their involvement in diverse biological processes, including plant growth and development, stress responses, and phytohormone signaling pathways (Fig. 5, Table S4).
Prediction of cis-regulatory elements in the promoter regions of the RsDof gene family. Different cis-elements are represented by different colors. Cis-element analysis of the promoter sequences (1500 bp) upstream of the RsDof genes was performed using PLACE (http://www.dna.affrc.go.jp/PLACE/.)
Expression patterns of RsDofs in different tissues of radish
The RNA-seq data revealed three distinct groups of Dof genes based on their expression patterns in different tissues of the carmine radish. Group 1 Dof genes were primarily expressed in the pulps, while Group 2 genes were specifically expressed in the flowers and inflorescences. Meanwhile, Group 3 encompassed genes with tissue-specific expression in various parts of the plant, such as the pericarp, flower, leaf, and inflorescence. For group 1, most of the RsDofs were specially expressed in pulps of carmine radish, including RsDof1-a, RsDof1-b, RsDof33, RsDof35-a, RsDof13-a, RsDof27-c, RsDof31, RsDof15-d, RsDof7-b, RsDof24-c, RsDof13-b, RsDof15-a, RsDof29-a, and RsDof27-b.
For group 2, most of RsDofs were specially expressed in flower (RsDof4-a, RsDof14-c, RsDof34-b, RsDof17, RsDof11, RsDof21-b, RsDof5-c, RsDof13-c, RsDof21-a, RsDof5-b, RsDof9-b, RsDof14-a) and inflorescence (RsDof34-a, RsDof19-b, RsDof5-a, RsDof4-a, RsDof17) of carmine radish. For group 3, different RsDofs were specially expressed in different tissues, including RsDof2 and RsDof32-b in pericarp, RsDof32-a, RsDof4-c, RsDof30-a and RsDof9-c in flower, RsDof10, RsDof4-c, RsDof30-a and RsDof9-c in leaf, and RsDof32-a, RsDof24-d, RsDof24-b, RsDof36-a, RsDof29-b and RsDof35-b in flower respectively. Notably, flowers and pulps were found to be the primary sites of Dof gene expression, suggesting their significant roles in flower development and the biosynthesis of anthocyanins. (Fig. 6, Table S5).
Heatmap and hierarchical clustering of RsDof genes in florescence, fruit, flower, leaf, pericarp and pulp. The expression values were calculated by RPKM measurement, and log2 transformed before generating heatmaps.
Responses of RsDOfs genes to heavy metal cadmium (Cd) treatments
The study investigated the modulation of RsDof gene expression in response to heavy metal stress in radish, utilizing transcriptome data from the heavy metal-tolerant cultivar ‘XCB’ and the heavy metal-susceptible cultivar 'HX.' Hierarchical clustering analysis was employed to analyze the expression patterns of RsDof genes. Notably, several RsDof transcripts exhibited significant alterations in response to cadmium stress, including RsDof5-b, RsDof35-a, RsDof15-a, RsDof1-a, RsDof13-b, RsDof14-a, RsDof14-b, RsDof30-a, RsDof32-b, RsDof17, RsDof9-a, and RsDof11, RsDof21-a, RsDof21-b, RsDof33, RsDof32-a, RsDof14-a, and RsDof9-b et al.,) (Fig. 7A, Table S6).
Responses of RsDOfs genes to Cd treatments in two contrast cultivars. (A) Heatmap and hierarchical clustering of RsDof genes responsive to Cd stress in two cultivars under a series of treatments. (B) The expression patterns of the candidate RsDof genes (RsDof1-a, RsDof1-b, RsDof33, RsDof35-a, RsDof13-a, RsDof27-c, RsDof31, RsDof15-d, RsDof7-b, RsDof24-c, RsDof13-b, RsDof15-a, RsDof29-a and RsDof27-b) in two cultivars responsive to Cd stress were validated using qRT-PCR technology. Three independent biological repeats were performed, and all data points are the means of three biological replicates ± standard error (SE). (C). Correlation between RNA-seq and qRT-PCR data was conducted. Each RNA-seq expression data was plotted against that from quantitative real-time PCR and fit into a linear regression. Both x- and y-axes were shown in log2 scale.
Of those, the key roles of these genes (RsDof35-a, RsDof15-a, RsDof1-a, RsDof1-b, RsDof10, RsDof11, RsDof21-a, RsDof21-b, RsDof33, RsDof14-a, RsDof9-a, and RsDof32-a) responsive to Cd heavy metal stress were validated by qRT-PCR. The qRT-PCR results revealed significant up-regulation of RsDof35-a and RsDof15-a in the ‘HX’ cultivar under cadmium stress while exhibiting complex expression dynamics in the ‘XCB’ cultivar over different treatment durations. Conversely, RsDof1-a and RsDof14-a showed significant up-regulation in the ‘HX’ cultivar but down-regulation in the ‘XCB’ cultivar under cadmium stress. Additionally, RsDof1-b, RsDof11, RsDof21-b, RsDof10, and RsDof21-a were down-regulated in the ‘HX’ cultivar under cadmium stress, with varying expression patterns observed in the ‘XCB’ cultivar. Notably, RsDof33, RsDof9-a, and RsDof32-a were consistently down-regulated in both radish cultivars, indicating their putative negative regulatory roles in response to heavy metal stress (Fig. 7B). Correlation between the RNA-seq and qPCR data indicated that both results were basically consistent for gene expression profiles (Fig. 7C, R = 0.76), which confirmed the high reliability of the RNA-seq data obtained in our study.
Expression profiles of the candidate RsDof genes related to anthocyanin biosynthesis
To investigate the key roles of these genes in anthocyanin biosynthesis, young fleshy roots of the carmine radish “Hongxin 1” were used for RNA-Seq validation at different stages of fleshy root development ((seedling stage (SS), initial expansion (IE), full-expansion (FE), bolting stage (BS), initial. flowering stage (IFS); full-bloom stage (FBS) and podding stage (PS))). The expression patterns of RsDof genes were compared to the dynamics of anthocyanin profiles in the fleshy roots. The results showed that the transcripts of RsDof35-a, and RsDof33 were upregulated in various growth stages of fleshy roots, from initial expansion to full bloom, but decreased in the podding stage. This pattern was consistent with the dynamics of anthocyanin profiles in the fleshy roots of carmine radish, particularly for RsDof33 (Fig. 8A,B). RsDof1-b and RsDOf1-a were found to have a significant decrease from the seedling stage (SS) to the bolting stage (BS), followed by an increase. On the other hand, the expression of RsDof32-a was significantly increased from the seedling stage (SS) to full expansion (FE) and then increased. For the transcript of RsDof14-a and RsDof9-a, which showed similar trends (increased from SS to IE, and then decreased to FE, increased FE to FBS, and last decreased to PS stage), and RsDof15-a and RsDof21-a were significantly enriched only in the seedling stage, while RsDof21-a was more enriched in both the seedling stage and initial expansion stage. RsDof10 and RsDof11 were more enriched in the seedling stage, initial expansion stage, and full bloom stage, and the seedling stage, initial expansion stage, and initial flowering stage, respectively (Fig. 8A). Overall, the expression patterns of RsDof33 showed consistency with the dynamics of anthocyanin profiles in the fleshy roots of carmine radish, suggesting that RsDof33 may play a role in the regulatory mechanism of anthocyanin synthesis in the fleshy roots.
Expression profiles of the candidate RsDof genes related to anthocyanin biosynthesis and Sub-localisation analysis of RsDof33 genes. (A)The expression patterns of the candidate RsDof genes (RsDof1-a, RsDof1-b, RsDof33, RsDof35-a, RsDof13-a, RsDof27-c, RsDof31, RsDof15-d, RsDof7-b, RsDof24-c, RsDof13-b, RsDof15-a, RsDof29-a, and RsDof27-b) in the pulp of fleshy roots obtained from the dynamics development stage of fleshy roots (seedling stage (SS), initial expansion (IE), full-expansion (FE), bolting stage (BS), initial. flowering stage (IFS); full-bloom stage (FBS) and podding stage (PS)) in carmine radish """Hongxin 1″"" using qRT-PCR. Relative gene expression levels were normalized against actin transcript levels, and log2 scale for fold change of gene expression in the development stage of fleshy roots comprising of ""“SS”"", "“IE”", “FE”,” “BS”,” “IFS”,” “FBS” and “PS” was shown. The standard error calculated from three biological replicates and significant (P < 0.05) difference identified by uncorrected Fisher’s LSD test in multiple comparisons after two-way ANOVA are indicated by error bars and stars, respectively. (B) Digram of Vector construction of p1300-RsDof 33-GFP. C Sub-localisation analysis of RsDof 33 TF.
Additionally, it was found that RsDof33 was responsive to Cd stress. Therefore, RsDof33 was selected for further gene function analysis in carmine radish. An expression plasmid containing RsDof33 with a GFP reporter and a CMV35S promoter was constructed (Fig. 8B). and then transformed into Arabidopsis protoplasts. The recombinant fusion protein RsDof33-GFP was observed to be localized in the nucleus and dispersed into the cytoplasm as small granules using laser confocal microscopy (Fig. 8C).
Correlation network analysis for anthocyanin biosynthesis genes, RsDOF transcription factors, and anthocyanin metabolisms
In the study of radish RsDOF gene regulation, metabolomics data identified four metabolites (Pelargonidin-3-O-glucoside, Cyanidin-3-O-glucoside, Pelargonidin-3,5-O-diglucoside, and Peonidin-3-O-rutinoside), alongside 12 anthocyanin biosynthesis genes (Rs4CL3, RsC4H2, RsCHI3, RsCHS1, RsFLS5, RsMYB113.2, RsPAL1.1, RsPAL1.4, RsPAL1.5, RsPAL2, RsTT8.1, and RsTT8.2) and four RsDOF transcription factors (RsDof33, RsDof30a, RsDof20a, RsDof9a), across diverse radish cultivars ( WW_NM (white radish white skin and white flesh root, WW), HX-1 (Hongxin red skin and white flesh root, RW), HX-2 (Hongxin red skin and pinky flesh root, RP), HX-3 (Hongxin red skin and red flesh root, RR), WG-1 (Waguan red skin and white flesh root, RW), WG-2(Waguan red skin and pinkly flesh root, RP), and WG-3 (Waguan red skin and red flesh root, RR) (Fig. 9A,B). A network integrating these elements and relevant miRNAs exhibited limited significant associations, except for specific interactions such as miR8005e-RsDOf33 (Fig. 9C).
Correlation network analysis was performed involving structural genes, RsDOF transcription factors, and anthocyanin content. (A) The expression patterns of 17 candidate RsDof genes in diverse radish cultivars. (B)The expression patterns of anthocyanin biosynthesis genes in diverse radish cultivars. (C) A network integrating the candidate RsDof genes and anthocyanin biosynthesis genes with relevant miRNAs. (D) A correlation network incorporating the four metabolites, 12 anthocyanin biosynthesis genes, and four RsDOF TFs was established. Anthocyanin metabolism was represented by red ellipse boxes, RsDOF transcription factors by blue ellipse boxes, and anthocyanin biosynthetic genes by green diamond boxes. Red solid linesRed solid lines denoted positive regulation, while blue lines indicated negative regulation. Edges were depicted when the linear correlation coefficient exceeded 0.8 with a p-value below 0.05.
Subsequently, a correlation network was established, incorporating the four metabolites, 12 anthocyanin biosynthesis genes, and four RsDOF TFs, which revealed Pearson correlation coefficients exceeding 0.8 (Fig. 9D). Notably, positive correlations with anthocyanin contents were observed for 11 anthocyanin biosynthesis genes, with RsPAL1.1, RsCHS1, and RsTT8.1 exhibiting the highest positive correlations. At the same time, RsFLS5 displayed negative correlations with anthocyanin contents (Table S7). Among the RsDOF TFs, alterations in transcript levels were noted for RsDof33, RsDof30a, RsDof20a, and RsDof9a, with RsDof33 showing the strongest positive correlation. Significant correlations were observed between RsDof33 and the 11 anthocyanin biosynthesis genes, followed by RsDof30a, RsDof20a, and RsDof9a (Table S8). Furthermore, robust positive associations were identified between levels of RsDof33, RsDof30a, RsDof20a, and RsDof9a with anthocyanin biosynthesis genes (Table S9), implying the potential involvement of RsDof33 in regulating anthocyanin biosynthesis.
Overexpression of RsDof33 decreases rosette leaf number and shortens flowing time, as well as anthocyanin biosynthesis in A. thaliana
To functionally understand the involvement of RsDOf33 in plant growth and development, we generated RsDOf33-overexpressing (OE) plants. Compared to wild-type (WT) plants, the overexpression lines exhibited different growth phenotypes, as shown in Fig. 9A. Among these lines, OE1 (Leaf number: 8), OE7 (6), and OE-1 (4) had fewer rosette leaves but promoted early flowering, with OE8 showing the most significant effect (Fig. 10A). We found anthocyanin enriched in OE1, OE7, and OE-8, respectively, especially for OE-8. On the other hand, OE3 (14), OE4 (12), and OE6 (12) had a similar number of rosette leaves and moderately promoted early flowering. Lastly, OE2 (20) and OE5 (18) had more rosette leaves but only slightly promoted early flowering. To further analyze the relative expression level of the RsDOf33 gene, qRT-PCR was performed on the transgenic pure lines (OE1, OE2, OE3, OE4, OE5, OE6, and OE7).
RsDof33 regulates rosette leaf number by upregulating the expression of some genes related to auxin and leaf shape development and anthocyanin biosynthesis. (A) Different growth phenotypes in overexpression line plants. (B) QRT-PCR further analyzed the relative expression level of RsDof33 for those transgenic pure lines. (C) Western bolting (WB) detection of RsDOf33 protein in the OE1 transgenic lines, followed by OE7 and OE8. (D). The expression levels of auxin biosynthetic genes (AtYUC2, AtYUC4 and AtYUC6), auxin transport genes (AtAUX1, AtPIN1, AtPIN3, and AtPIN4), as well as genes related to leaf shape development (AtKNAT2, AtKNAT6, AtDPA4, and AtCUC2) were examined in OE plants. (E). The expression levels of anthocyanin biosynthetic genes were examined in OE plants.
The results indicated a significant increase in RsDOf33 gene expression in the OE8 transgenic lines, followed by OE7 and OE1 (Fig. 10B,C , Figure S2). This suggests that higher expression of the RsDOf33 TF makes the plants more susceptible to promoting early flowering and increasing anthocyanin accumulation but reduces the number of rosette leaves (Fig. 10A,C, Figure S3). Previous studies have underscored the pivotal role of auxin in leaf development. Hence, we delved into whether RsDOf33 governs leaf development by modulating auxin signaling. We scrutinized the expression levels of auxin biosynthetic genes (AtYUC2, AtYUC4, and AtYUC6), auxin transport genes (AtAUX1, AtPIN1, AtPIN3, and AtPIN4), and genes associated with leaf shape development (AtKNAT2, AtKNAT6, AtDPA4, and AtCUC2). Our analysis unveiled a notable rise in the transcript levels of the auxin biosynthetic gene YUCCA (AtYUC2) and an auxin efflux carrier (AtPIN4) in plants expressing RsDOf33 compared to wild-type Arabidopsis thaliana plants.
Conversely, the expression levels of auxin transport genes (AtAUX1) and auxin efflux carriers (AtPIN3) were diminished (Fig. 10D). Moreover, we noted a significant elevation in the transcript levels of the leaf shape development gene AtKNAT2 in 35S::RsDOf33 plants (OE8) in comparison to wild-type A. thaliana plants (Fig. 10D).
To assess whether RsDof33 regulates anthocyanin biosynthesis, we validated the relative expression levels of anthocyanin biosynthesis-related genes (DFR, PAP1, PAL, C4H, 4CL3, CHS, CHI, F3H, F3’H, FLS, LDOX, and UF3GT) in A. thaliana. Our findings indicated significantly augmented transcript levels of the anthocyanin biosynthesis-related genes AtPAL, AtCHS, AtCHI, AtDFR, AtLDOX, and AtUF3GT in RsDOf33-expressing plants compared to wild-type A. thaliana plants, particularly in OE8. At the same time, AtFLS exhibited decreased expression (Fig. 10E).
These outcomes suggest that the alterations induced by RsDOf33 in auxin biosynthesis and polar transport could impact leaf development and anthocyanin biosynthesis. The up-regulation of specific genes linked to leaf shape development and anthocyanin biosynthesis potentially influences rosette leaf number and anthocyanin content in A. thaliana.
Discussion
Molecular characterization and evolution of radish Dof TFs
Researchers have elucidated the pivotal functions of Dof TFs in diverse plant developmental processes and their responsiveness to varied abiotic and biotic stresses 33,34. Nonetheless, a comprehensive exploration of Dof TFs in carmine radish is lacking, and their precise functionalities remain ambiguous. Hence, a genome-wide investigation was undertaken, culminating in the identification of 59 potential RsDofs. This count exceeds those found in rice (30 OsDofs) 35, Arabidopsis (36 AtDofs) 35, tomato (34 SiDofs) 33, cucumber (36 CsDofs) 36, and potato (35 StDofs) 37, yet falls short of the numbers reported in banana (74 MaDofs) 38 and Chinese cabbage (76 BraDofs) 39. The disparity in the quantity of Dof transcription factors across plant species aligns with data indicating that family sizes might significantly range across various groupings 40. Certain species possess a more significant number of genes, perhaps correlating with their adaptability to many settings and ecological functions 41.
Consistent with previous research, the alignment of RsDof protein sequences unveiled a markedly conserved Dof domain (Fig. 1), signifying the evolutionary conservation of Dof TFs across plant species. The preservation of the Dof domain is crucial since it regulates the activation and deactivation of genes, as shown by several research investigations 40,42.
The gene architecture, protein composition, exon–intron arrangement, and phylogenetic support the categorization of these 59 RsDof proteins into nine primary clusters (A-I), akin to categorizations in other plant taxa. Through phylogenetic scrutiny of dicots, monocots, and non-vascular plants, the carmine radish Dof gene family was partitioned into nine subfamilies (Groups A–I), with Group D comprising only four RsDof members (RsDof34a, RsDof34b, RsDof10, and RsDof2), aligning with AtDof34, AtDof10, and AtDof2, respectively. This classification aligns with the evolutionary trends seen in other species, whereby Dof subfamilies often exhibit similar sequences and functions 43. The exon–intron configuration serves as supportive evidence for deducing evolutionary ties among genes or organisms 44. RsDof genes within the same subfamily typically show analogous exon–intron patterns, while discrepancies are evident across distinct subfamilies. The variations in exon–intron architecture may provide significant insights into the functional and regulatory disparities across related gene families 45. The motif distribution among Dof proteins mirrors their evolutionary relationships as deduced from the phylogenetic tree 46.
The findings indicate that the sequence structure of motif 1, as detected by MEME, is consistent with the Dof domain, potentially implicated in binding to specific promoter sequences (Fig. 4). Motif 1 has a consistent sequence that remains unchanged across several plant species 47. This region facilitates protein binding to DNA and regulates critical processes in plant growth. This has been shown in research using Arabidopsis and rice 48. Moreover, within the same subfamily of RsDofs, there is a presence of at least one or two conserved motif variants with consistent spatial arrangements. At the same time, distinct disparities are evident among different subfamilies, implying functional resemblances among Dof members within a shared subfamily. This indicates that Dof TFs may regulate the same genes or processes in smaller groups. Studies on other plants indicate that members of these smaller groups often collaborate similarly 49.
Additionally, the structural conservation of RsDof genes within subfamilies is congruent with analogous patterns observed in other plant species such as Arabidopsis and rice 50, barley 51, sorghum 52, and purple false brome 53. Investigators have discovered the same outcomes in cotton plants. Researchers identified that certain Dof genes belonging to specific categories assist plants in managing stress 54. Furthermore, segmental duplication appears to have exerted a more substantial influence than tandem duplication in propelling the evolution of the Dof gene family. Our results suggest that 30 duplicate pairs of RsDof genes arose from segmental duplications, with an additional two pairs originating from tandem duplications in radish, which is similar to previous observations in Arabidopsis, rice, and apple 55,56,57. Segmental duplication plays a more significant role than tandem duplication in the evolutionary development of Dof genes, a tendency seen across several plant species 58. This indicates that extensive duplications in plant genomes significantly contribute to the proliferation of gene families.
Potential roles of RsDof TFs in tissue differentiation and organ development
Numerous studies have demonstrated that changes in spatiotemporal expression patterns are key indicators of functional divergence in duplicated genes 59. In our study, we found that most RsDofs were predominantly expressed in flowers and pulp, suggesting their crucial role in developing these tissues. Other species have shown the same outcomes, with Dof transcription factors significantly influencing the development of floral structures and tissue formation 60. Interestingly, duplicated genes (RsDof13-a, RsDof13-b, RsDof13-c) exhibited exceptionally different expression patterns arising from segmental duplication events, indicating that the functional divergence may have provided genetic sources with novel biological functions during the evolution of the RsDof gene family. The phenomenon of duplicated Dof genes in many plants acquiring novel functions has been seen. This is thought to facilitate the expansion of gene families and the emergence of novel functionalities 61. These findings suggest that expanding RsDof genes may contribute to increased biological complexity.
Furthermore, phylogenetic analysis revealed that ten RsDof genes (RsDof5a/RsDof5b/RsDof5c/RsDof9a/RsDof9b/RsDof12/RsDof17/RsDof30a/ RsDof30b/RsDof33) showed a high similarity to the five A. thaliana Cycling DOF Factors (CDF1(AtDof33), CDF2(AtDof30), CDF3(AtDof17), CDF4(AtDof12), CDF5(AtDof9) and COG1(AtDof5)) in the Group I subfamily. These findings align with previous studies indicating that analogous CDF genes regulate several aspects of plant development, including flowering time, circadian rhythms, and environmental responses 62. Among these, the transcripts of RsDof5a, RsDof5b, RsDof5c, RsDof9a, RsDof9b, RsDof17, and RsDof30-a were identified enriched in flowers. At the same time, RsDof33 was specifically accumulated in the pulp of carmine radish (Fig. 3). RsDof33's functionality in plant tissue suggests its potential to enhance the development of fleshy roots and the synthesis of anthocyanins. This parallels the influence of CDF genes in other plant species on these processes. Moreover, RsDof33 has been identified as participating in the developmental stages of fleshy roots associated with anthocyanin biosynthesis. Previous research has demonstrated that CDF1 (AtDof33) suppresses the transcription of Constans (CO), consequently impeding flowering in A. thaliana 63. In accordance with this concept, the RsDof33 genes may also play a role in the growth and development of flowers. They may regulate crucial genes involved in transitioning from a plant that produces leaves to one that generates flowers and seeds. Hence, it is likely that these RsDof33 genes also play roles in flower development.
Moreover, RsDof genes homolog to the transcript of A. thaliana Dof factor OBPs were distributed in different subgroups, were separately distributed in subgroup B1, G, B2 and A1 respectively. OBP1-like factors regulate cellular division and development across several plant species 64. Of those, RsDof1b and RsDof1a were found enriched in the pulp of carmine radish, while RsDof32a and RsDof32b were found enriched in pericarp, but RsDof18, RsDof20a and RsDof20b did not detected in all the tissues. This expression pattern in specific tissues corresponds with previous research indicating that Dof transcription factors had distinct functions at various developmental stages, such as tissue expansion and angiogenesis.
Furthermore, RsDof1a and RsDof1b have been implicated in the growth stages of fleshy roots associated with anthocyanin biosynthesis. In Arabidopsis, OBP1 is recognized for regulating cell division and facilitating cell transition via interactions with genes related to the cell cycle 65. This indicates that RsDof1a and RsDof1b in carmine radish may possess similar functions. It has been documented that the A. thaliana Dof factor OBP1 is involved in the overall regulation of cell division and plays a pivotal role in cell cycle re-entry by serving as a transcriptional regulator of key cell cycle genes 66. In addition, OBP-like factors regulate the transition from cell division to cell differentiation in plant roots and stems 67. Analysis of promoter activity indicates that AtDof5.8 may be involved in the initial stages of leaf vasculature formation, given that its promoter sequences harbor cis-elements for stage-specific expression during vascular development 68. This comprehension of the role of Dof TFs with findings from previous studies indicates that Dof factors contribute to regulating blood vessel tissue development 40. In our investigation, many meristem-specific activation-related elements known as CCGTCC-box were identified in the promoter regions of RsDof genes within Group E, with each gene containing more than 2.3 copies (Table S5). These promoter characteristics are often associated with genes facilitating root and shoot development. They have also been associated with growth and regeneration in other flora 69. This observation suggests that these RsDof genes may also play a role in various processes associated with fleshy root development in carmine radishes. Given that CCGTCC-boxes are crucial for regulating growth in meristematic regions, the RsDof genes in Group E may play a significant role in root development and tissue formation in the carmine radish.
Expression and functions of RsDof genes in response to heavy metal stress
Previous studies have shown that Dof transcription factors play important roles in various abiotic stress responses, particularly in response to plant hormones such as auxins and gibberellins 70. In addition to auxins and gibberellins, hormones, including cytokinins and ethylene, also regulate Dof gene activity during periods of stress. This indicates that these hormones contribute to plant adaptation to stress. Auxin, for example, can be induced by different abiotic stresses and can promote the expression of stress-defense genes 71,72 through ABA and other pathways. Auxins, a category of plant hormones, regulate Dof genes to assist plants in managing water and salt stress 73. Dof genes have been observed in several plant species, indicating their significant and adaptable functions in helping plants manage stress 49. In potatoes, the StDof genes exhibited either ABA-dependent or ABA-independent expression patterns 37. A comparable regulation of Dof genes, contingent upon ABA, has been identified in rice and barley 74. This indicates that Dof TFs may be significant in plants’ ABA signaling pathways under environmental stress.
We found that the promoter regions of the 59 RsDof family members contained several phytohormone regulation-related elements, including the TCA-element, TGA-element, GARE-motif, ABRE, and CGTCA-motif (Table S4). These components are often seen in genes that respond to stress 75. These genes are activated when plants encounter specific environmental stressors. This indicates that RsDofs likely influence plant responses to stress via hormonal mechanisms. Also, suggests that RsDofs are likely involved in phytohormone responses and may serve as important regulators in plant hormone pathways. Dof genes responsive to ABA are crucial for regulating antioxidant systems and stress-activated signaling pathways 76.
Additionally, some RsDofs were found to be associated with the functional terms "response to oxidative stress" and "oxidation–reduction process," indicating their potential involvement in ROS-mediated signaling. Plants subjected to heavy metal stress often generate ROS as a component of their typical response 77,78. Dof genes regulate how plants eliminate ROS across various plant species 79. In wheat, eight TaDof genes were significantly upregulated under heavy metal stress and were suggested to play essential roles in ROS clearance pathways 80. Dof genes in rice have been identified as regulators of antioxidant enzymes in response to toxic metals 81. This indicates that these genes may enhance plant resilience to stress. However, in our study, RsDof33, RsDof9-a, and RsDof32-a were significantly down-regulated in both radish cultivars under heavy metal stress. This disparity indicates that some Dof genes are activated in other species by exposure to heavy metals. This indicates that the function of Dof proteins in stress responses may vary according to the species or the specific tissue type. This suggests that RsDof genes may have more prominent roles in plant organ development rather than in various abiotic stress responses. This indicates that the RsDof genes in red radishes may have a diminished response to stress. This may be due to compensatory actions of other genes or proteins or the involvement of different genes that react to stress. Therefore, further investigation is needed to elucidate the detailed molecular mechanisms of RsDof TFs. Future research should focus on elucidating the collaborative functions of Dof genes and their interactions with other stress-responsive components. This will assist in identifying methods to enhance the stress resistance of radish plants.
Functional of RsDof33 involved in leaf development and anthocyanin biosynthesis through affecting auxin signaling
To explore the potential role of RsDof33, its ortholog in Arabidopsis, known as CDF, was investigated. Among the identified orthologs, AtCDF1 and AtCDF5 in Arabidopsis have been demonstrated to facilitate hypocotyl elongation under short-day conditions 82. In analogous circumstances, CDF1 and CDF5 have been seen to influence growth responses regulated by the diurnal cycle in other plants, such as rice 83. They regulate the height of the rice plant in response to certain light conditions. Conversely, Arabidopsis CDF2 and PIF4 (PHYTOCHROME-INTERACTING FACTOR 4) form a complex that co-regulates the downstream target gene YUCCA8, thus promoting hypocotyl elongation 84. PIF4 has been shown to collaborate with other proteins that regulate gene expression, including HY5. This relationship influences plant growth in response to light. In our investigation, the expression patterns of RsDof33 transcripts exhibited concordance with the dynamics of anthocyanin profiles during fleshy root development in carmine radish. Previous studies have shown that CDF family members regulate the synthesis of the pigment anthocyanin, which occurs when plants are exposed to light or exhibit certain growth patterns 85. This is particularly applicable to plants with succulent roots and stems. We proposed that RsDof33 may play a role in orchestrating the development of fleshy roots in carmine radishes.
In our investigation, overexpression of the RsDof33 gene in Arabidopsis accelerated early flowering and anthocyanin content and reduced rosette leaf count. This characteristic aligns with findings from several studies. Overexpression of Dof TFs, such as AtDOF4.2, in Arabidopsis results in accelerated flowering and elevated flavonoid concentrations 86. Additionally, we established a correlation between the RsDof33 gene and auxin biosynthetic genes in transgenic Arabidopsis plants. Notably, the expression levels of the auxin biosynthetic genes YUCCA (AtYUC2) and the auxin efflux carrier AtPIN4 were markedly augmented in RsDof33-expressing plants, in contrast to wild-type Arabidopsis plants. Additional research has shown that increasing the activity of Dof genes results in a corresponding rise in YUC genes 87. This indicates that Dof proteins consistently regulate auxin biosynthesis.
Concurrently, the expression of the auxin transport genes AtAUX1, AtPIN1, and AtPIN3 were diminished, while the expression of AtYUC4 and AtYUC6 remained largely unaltered in the transgenic plants (Fig. 10D). RsDof33 influences the activation of certain YUC genes, likely regulating auxin levels in various regions of the plant. The alterations in auxin are crucial for the development of leaves and roots 88.
Furthermore, the transcript levels of AtKNAT2 were significantly elevated in the 35S::RsDof33 plants compared to wild-type Arabidopsis plants (Fig. 10D). KNAT2 is a gene that is part of the KNOTTED1-like homeobox family 89. It regulates leaf growth and the arrangement of veins inside the leaves 90. When RsDof33 enhances the activity of KNAT2, it may contribute to the initiation of development and the morphology of leaves during the formation of rosette leaves. Based on these observations, we propose that the collaborative action of RsDof33 and AtPIN4 is necessary for robustly activating the transcription of AtYUC2 in the leaves during the development of rosette leaves. Previous studies have shown that PIN proteins collaborate with transcription factors, such as Dof, to regulate auxin distribution and affect growth processes, especially during the first phases of leaf and root development 91. Further studies in Arabidopsis have demonstrated that the coordinated action of CDF2 and PIF4 is essential for activating the transcription of YUC8 in cotyledons, leading to increased auxin biosynthesis and promoting hypocotyl growth 84,92. PIF4 and CDF2 regulate the activity of other genes crucial for using and transporting auxin in plants. This facilitates plant growth following their environment, such as adapting to variations in light and temperature. Therefore, a comparable mechanism is likely involved in regulating rosette leaf development by RsDof33. This indicates that RsDof33 may interact with light-responsive pathways to regulate auxin-related developmental processes.
Additionally, it is noteworthy that CYCLING DOF FACTORS (CDF) are temporally regulated by the circadian clock and are involved in repressing photoperiodic flowering and tuberization, as well as promoting hypocotyl elongation and regulating abiotic stress responses (Kloosterman 93,94,95,96,97. CDFs, particularly CDF1 and CDF2, are recognized for their influence on daily biological rhythms and their association with many light-regulated activities 98. This influences the timing of flowering in plants and their responses to stressors.
In addition, previous research revealed that the Arabidopsis Dof transcription factor AtDOF4.2 impacts flavonoid accumulation and the expression of genes associated with flavonoid biosynthesis, indicating a potential regulatory role of Dof TFs in flavonoid production. This finding suggests that Dof TFs may regulate the synthesis of secondary metabolites, such as flavonoids and anthocyanins, by modulating auxin activity in the plant. However, the precise molecular mechanisms underlying this regulation remain largely unexplored and require further investigation. In our study, we found transcript levels of the anthocyanin biosynthesis-related genes in RsDOf33-expressing plants compared to wild-type A. thaliana plants, particularly in OE1. At the same time, AtFLS exhibited decreased expression (Fig. 10E). Other investigations have shown that when RsDof33 is hyperactive, genes such as PAL, CHS, and DFR, which are implicated in flavonoid biosynthesis, exhibit increased activity. This association indicates that Dof proteins may contribute to flavonoid biosynthesis. These outcomes suggest that the alterations induced by RsDOf33 in auxin biosynthesis and polar transport could impact leaf development and anthocyanin biosynthesis. Finally, further investigation is necessary to elucidate the interactions of RsDof33 with other proteins that regulate anthocyanin synthesis and foliar development.
Conclusion
In this investigation, a total of 59 RsDof genes were identified and classified into nine subfamilies labeled A (8), B1(10), B2(10), C1(3), C2.1(5), C2.2(4), C3(11), D1(4), and D2(4). These subfamilies demonstrated similar exon–intron structures within their respective groups while exhibiting variations across different subfamilies. The analysis also unveiled segmental duplications and two pairs of tandem duplications of RsDof gene pairs, indicating diverse degrees of positive selection pressure on the Dof gene family during evolution in radish. Additionally, the type and quantity of cis-acting elements indicated that RsDof genes may be regulated by various hormones and environmental factors. RNA-seq data analysis and qRT-PCR showed that RsDof genes display distinct expression specificity and preferences in different organs and developmental stages, suggesting potential roles of RsDOfs in tissue differentiation and organ development.
Furthermore, overexpression of the RsDof33 gene in Arabidopsis led to early flowering time promotion and enhanced anthocyanin content while reducing the number of rosette leaves. Subsequent investigations revealed that the cooperation between RsDof33 and AtPIN4 is essential for activating AtYUC2 transcription in leaf development, indicating the involvement of RsDof33 in rosette leaf development. Overall, this study provides valuable insights into the molecular mechanisms and functional characteristics of the RsDof gene family, contributing to a comprehensive understanding of their roles in plant development, growth, and anthocyanin biosynthesis.
Data availability
All raw data from the transcriptome libraries were deposited at NCBI with accession number PRJNA832540.
References
El-Sappah, A. H. et al. Plants’ molecular behavior to heavy metals: From criticality to toxicity. Front. Plant Sci. https://doi.org/10.3389/fpls.2024.1423625 (2024).
Abbas, M. et al. Genome-wide analysis and expression profiling of SlHsp70 gene family in Solanum lycopersicum revealed higher expression of SlHsp70-11 in roots under Cd2+ stress. Front. Biosci.-Landmark 27, 186 (2022).
El-Sappah, A. H. & Rather, S. A. in Plant abiotic stress physiology 25–46 (Apple Academic Press, 2022).
El-Sappah, A. H. et al. Heat stress-mediated constraints in maize (Zea mays) production: challenges and solutions. Front. Plant Sci. 13, 879366 (2022).
El-Sappah, A. H. et al. Interplay of silymarin and clove fruit extract effectively enhances cadmium stress tolerance in wheat (Triticum aestivum). Front. Plant Sci. 14, 1144319 (2023).
Baldoni, E., Genga, A. & Cominelli, E. Plant MYB transcription factors: Their role in drought response mechanisms. Int. J. Mol. Sci. 16, 15811–15851 (2015).
Luo, T. et al. Genome-wide identification and functional analysis of Dof transcription factor family in Camelina sativa. BMC Genomics 23, 812 (2022).
Fang, Z. et al. Genome-wide identification, structure characterization, and expression profiling of Dof transcription factor gene family in wheat (Triticum aestivum L). Agronomy 10, 294 (2020).
Lal, M. K. et al. Mechanistic concept of physiological, biochemical, and molecular responses of the potato crop to heat and drought stress. Plants 11, 2857 (2022).
Le Hir, R. & Bellini, C. The plant-specific Dof transcription factors family: New players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. 4, 164 (2013).
Ranjan, R. & Yadav, R. Targeting nitrogen use efficiency for sustained production of cereal crops. J. Plant Nutr. 42, 1086–1113 (2019).
Hirel, B., Tétu, T., Lea, P. J. & Dubois, F. Improving nitrogen use efficiency in crops for sustainable agriculture. Sustainability 3, 1452–1485 (2011).
Kumari, P., Ojha, R., Varshney, V., Gupta, V. & Salvi, P. Biotechnological Advances for Disease Tolerance in Plants 337–362 (Springer, Berlin, 2024).
Peng, H. & Gao, J. The whole genome assembly and evolution analyze of carmine radish (Raphanus sativus L.) Mitochondrion. Mitochondrial DNA. Part B, Resources 5, 2252–2253. https://doi.org/10.1080/23802359.2020.1772136 (2020).
Song, X., Gao, J. & Peng, H. Transcriptomic dynamics changes related to anthocyanin accumulation in the fleshy roots of carmine radish (Raphanus sativus L.) characterized using RNA-Seq. PeerJ 9, e10978. https://doi.org/10.7717/peerj.10978 (2021).
Gao, J., Peng, H., Chen, F., Luo, M. & Li, W. Genome-wide analysis of transcription factors related to anthocyanin biosynthesis in carmine radish (Raphanus sativus L.) fleshy roots. PeerJ 7, e8041 (2019).
Garg, V. K. et al. MFPPI - Multi FASTA ProtParam Interface. Bioinformation 12, 74–77. https://doi.org/10.6026/97320630012074 (2016).
Horton, P. et al. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 35, W585-587. https://doi.org/10.1093/nar/gkm259 (2007).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 305, 567–580. https://doi.org/10.1006/jmbi.2000.4315 (2001).
Bailey, T. L. et al. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 37, W202-208. https://doi.org/10.1093/nar/gkp335 (2009).
Goodstein, D. M. et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178-1186. https://doi.org/10.1093/nar/gkr944 (2012).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evolut. 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Wang, Y. et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49. https://doi.org/10.1093/nar/gkr1293 (2012).
Krzywinski, M. et al. Circos: An information aesthetic for comparative genomics. Genome Res. 19, 1639–1645. https://doi.org/10.1101/gr.092759.109 (2009).
Lescot, M. et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30, 325–327. https://doi.org/10.1093/nar/30.1.325 (2002).
von Mering, C. et al. STRING: A database of predicted functional associations between proteins. Nucleic Acids Res. 31, 258–261. https://doi.org/10.1093/nar/gkg034 (2003).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Gao, J. et al. Genome-wide characterization and expression analysis of GRAS genes in carmine radish (Raphanus sativus L.). Sci. Hortic. 308, 111532 (2023).
Gao, J. et al. A putative pathogen-resistant regulatory pathway between MicroRNAs and candidate target genes in maize. J. Plant Biol. 58, 211–219. https://doi.org/10.1007/s12374-014-0572-5 (2015).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Yoo, S. D., Cho, Y. H. & Sheen, J. Arabidopsis mesophyll protoplasts: A versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565–1572. https://doi.org/10.1038/nprot.2007.199 (2007).
Clough, S. J. & Bent, A. F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. The Plant J. Cell. Mol. Biol. 16, 735–743. https://doi.org/10.1046/j.1365-313x.1998.00343.x (1998).
Cai, X. et al. Genome-wide analysis of plant-specific Dof transcription factor family in tomato. J. Integrative Plant Biology 55, 552–566. https://doi.org/10.1111/jipb.12043 (2013).
Corrales, A. R. et al. Characterization of tomato cycling Dof factors reveals conserved and new functions in the control of flowering time and abiotic stress responses. J. Exp. Bot. 65, 995–1012. https://doi.org/10.1093/jxb/ert451 (2014).
Lijavetzky, D., Carbonero, P. & Vicente-Carbajosa, J. Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evolut. Biol. 3, 17. https://doi.org/10.1186/1471-2148-3-17 (2003).
Wu, Z. et al. Genome-wide identification and expression profile of Dof transcription factor gene family in pepper (Capsicum annuum L). Front. Plant Sci. 7, 574. https://doi.org/10.3389/fpls.2016.00574 (2016).
Venkatesh, J. & Park, S. W. Genome-wide analysis and expression profiling of DNA-binding with one zinc finger (Dof) transcription factor family in potato. Plant Physiol. Biochem. PPB 94, 73–85. https://doi.org/10.1016/j.plaphy.2015.05.010 (2015).
Feng, B. H. et al. The banana fruit Dof transcription factor MaDof23 acts as a repressor and interacts with MaERF9 in regulating ripening-related genes. J. Exp. Bot. 67, 2263–2275. https://doi.org/10.1093/jxb/erw032 (2016).
Ma, J., Li, M. Y., Wang, F., Tang, J. & Xiong, A. S. Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genom. 16, 33. https://doi.org/10.1186/s12864-015-1242-9 (2015).
Noguero, M., Atif, R. M., Ochatt, S. & Thompson, R. D. The role of the DNA-binding one zinc finger (DOF) transcription factor family in plants. Plant Sci. 209, 32–45 (2013).
Lehti-Shiu, M. D., Panchy, N., Wang, P., Uygun, S. & Shiu, S.-H. Diversity, expansion, and evolutionary novelty of plant DNA-binding transcription factor families. Biochimica et Biophysica Acta (BBA)-Gene Regulatory Mechanisms 1860, 3–20 (2017).
Gupta, S. et al. Insights into structural and functional diversity of Dof (DNA binding with one finger) transcription factor. Planta 241, 549–562 (2015).
Liao, B. et al. The role of liriodendron Dof gene family in abiotic stress response. Plants 13, 2009 (2024).
Malviya, N. et al. Genome wide in silico characterization of Dof gene families of pigeonpea (Cajanus cajan (L) Millsp.). Mol Biol Rep 42, 535–552. https://doi.org/10.1007/s11033-014-3797-y (2015).
Xu, G., Guo, C., Shan, H. & Kong, H. Divergence of duplicate genes in exon–intron structure. Proc. Natl. Acad. Sci. 109, 1187–1192 (2012).
Kushwaha, H., Gupta, S., Singh, V. K., Rastogi, S. & Yadav, D. Genome wide identification of Dof transcription factor gene family in sorghum and its comparative phylogenetic analysis with rice and Arabidopsis. Mol. Biol. Rep. 38, 5037–5053. https://doi.org/10.1007/s11033-010-0650-9 (2011).
Hayden, C. A. & Jorgensen, R. A. Identification of novel conserved peptide uORF homology groups in Arabidopsis and rice reveals ancient eukaryotic origin of select groups and preferential association with transcription factor-encoding genes. BMC Biol. 5, 1–30 (2007).
Ouyang, X. et al. Genome-wide binding site analysis of FAR-RED ELONGATED HYPOCOTYL3 reveals its novel function in Arabidopsis development. The Plant Cell 23, 2514–2535 (2011).
Liu, Y. et al. Genome-wide analysis of wheat DNA-binding with one finger (Dof) transcription factor genes: evolutionary characteristics and diverse abiotic stress responses. BMC Genom. 21, 1–18 (2020).
Wang, H. W. et al. The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J. For Cell Mol. Biol. 52, 716–729. https://doi.org/10.1111/j.1365-313X.2007.03268.x (2007).
Stepanova, A. N. et al. The Arabidopsis YUCCA1 flavin monooxygenase functions in the indole-3-pyruvic acid branch of auxin biosynthesis. The Plant cell 23, 3961–3973. https://doi.org/10.1105/tpc.111.088047 (2011).
Cook, S. D. et al. Auxin biosynthesis: Are the Indole-3-acetic acid and phenylacetic acid biosynthesis pathways mirror images?. Plant physiology 171, 1230–1241. https://doi.org/10.1104/pp.16.00454 (2016).
Trainotti, L., Tadiello, A. & Casadoro, G. The involvement of auxin in the ripening of climacteric fruits comes of age: The hormone plays a role of its own and has an intense interplay with ethylene in ripening peaches. J. Exp. Bot. 58, 3299–3308. https://doi.org/10.1093/jxb/erm178 (2007).
Wang, Z. et al. Unveiling the molecular mechanisms of γ-polyglutamic acid-mediated drought tolerance in cotton through transcriptomic and physiological analyses (2024).
Cannon, S. B., Mitra, A., Baumgarten, A., Young, N. D. & May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 4, 10. https://doi.org/10.1186/1471-2229-4-10 (2004).
Yu, L. et al. Ancient rapid functional differentiation and fixation of the duplicated members in rice Dof genes after whole genome duplication. The Plant J. Cell Mol. Biol. 108, 1365–1381. https://doi.org/10.1111/tpj.15516 (2021).
Zhang, Z., Yuan, L., Liu, X., Chen, X. & Wang, X. Evolution analysis of Dof transcription factor family and their expression in response to multiple abiotic stresses in Malus domestica. Gene 639, 137–148. https://doi.org/10.1016/j.gene.2017.09.039 (2018).
Huang, Y.-L. et al. The impact of tandem duplication on gene evolution in Solanaceae species. J. Integr. Agric. 21, 1004–1014 (2022).
Hellsten, U. et al. Accelerated gene evolution and subfunctionalization in the pseudotetraploid frog Xenopus laevis. BMC Biol. 5, 31. https://doi.org/10.1186/1741-7007-5-31 (2007).
Rueda-López, M., García-Gutiérrez, A., Cánovas, F. M. & Ávila, C. The family of Dof transcription factors in pine. Trees 27, 1547–1557 (2013).
Wang, X. et al. Genome-wide identification and characterization of Dof gene family in Salvia miltiorrhiza. Ornam. Plant Res. 4(1), 0–0. https://doi.org/10.48130/opr-0024-0030 (2024).
Renau-Morata, B. et al. CDF transcription factors: Plant regulators to deal with extreme environmental conditions. J. Exp. Bot. 71, 3803–3815 (2020).
Lucas-Reina, E., Romero-Campero, F. J., Romero, J. M. & Valverde, F. An evolutionarily conserved DOF-CONSTANS module controls plant photoperiodic signaling. Plant Physiol. 168, 561–574. https://doi.org/10.1104/pp.15.00321 (2015).
Fu, J. et al. Research article Dissecting grain yield pathways and their interactions with grain dry matter content by a two-step correlation approach with maize seedling transcriptome. (2010).
Ramirez-Parra, E. et al. The transcription factor OBP 4 controls root growth and promotes callus formation. New Phytol. 213, 1787–1801 (2017).
Skirycz, A. et al. The DOF transcription factor OBP1 is involved in cell cycle regulation in Arabidopsis thaliana. Plant J. Cell Mol. Biol. 56, 779–792. https://doi.org/10.1111/j.1365-313X.2008.03641.x (2008).
Malhan, D., Bhatia, S. & Yadav, R. K. Genome wide gene expression analyses of Arabidopsis shoot stem cell niche cell populations. Plant Signaling Behav. 10, e1011937 (2015).
Konishi, M. & Yanagisawa, S. Sequential activation of two Dof transcription factor gene promoters during vascular development in Arabidopsis thaliana. Plant Physiol. Biochem. PPB 45, 623–629. https://doi.org/10.1016/j.plaphy.2007.05.001 (2007).
Singha, D. L. et al. Harnessing tissue-specific genome editing in plants through CRISPR/Cas system: Current state and future prospects. Planta 255, 1–17 (2022).
Baumann, K., De Paolis, A., Costantino, P. & Gualberti, G. The DNA binding site of the Dof protein NtBBF1 is essential for tissue-specific and auxin-regulated expression of the rolB oncogene in plants. The Plant cell 11, 323–334. https://doi.org/10.1105/tpc.11.3.323 (1999).
Rock, C. D. & Sun, X. Crosstalk between ABA and auxin signaling pathways in roots of Arabidopsis thaliana (L.) Heynh. Planta 222, 98–106. https://doi.org/10.1007/s00425-005-1521-9 (2005).
Park, J. E. et al. GH3-mediated auxin homeostasis links growth regulation with stress adaptation response in Arabidopsis. J. Biol. Chem. 282, 10036–10046. https://doi.org/10.1074/jbc.M610524200 (2007).
Cai, M. et al. Allele specific expression of Dof genes responding to hormones and abiotic stresses in sugarcane. PloS One 15, e0227716 (2020).
Mondal, R., Kumar, A. & Chattopadhyay, S. K. Structural property, molecular regulation, and functional diversity of glutamine synthetase in higher plants: a data-mining bioinformatics approach. The Plant J. 108, 1565–1584. https://doi.org/10.1111/tpj.15536 (2021).
Panahi, B. The molecular core responses to multiple abiotic stresses in barley (Hordeum vulgare L.) Insights from global transcriptome analysis. South Afr. J. Bot. 177, 237–250. https://doi.org/10.1016/j.sajb.2024.11.039 (2025).
Pastori, G. M. & Foyer, C. H. Common components, networks, and pathways of cross-tolerance to stress. The central role of “Redox” and abscisic acid-mediated controls. Plant Physiol. 129, 460–468. https://doi.org/10.1104/pp.011021 (2002).
Gou, C. et al. Integrative application of silicon and/or proline improves Sweet corn (Zea mays L. saccharata) production and antioxidant defense system under salt stress condition. Sci. Rep. 13, 18315. https://doi.org/10.1038/s41598-023-45003-8 (2023).
El-Sappah, A. H. et al. Genome-wide identification and expression analysis of metal tolerance protein (MTP) gene family in soybean (Glycine max) under heavy metal stress. Mol. Biol. Rep. 50, 2975–2990. https://doi.org/10.1007/s11033-022-08100-x (2023).
Ruta, V. et al. The DOF transcription factors in seed and seedling development. Plants 9, 218 (2020).
Washio, K. Functional dissections between GAMYB and Dof transcription factors suggest a role for protein-protein associations in the gibberellin-mediated expression of the RAmy1A gene in the rice aleurone. Plant Physiol. 133, 850–863. https://doi.org/10.1104/pp.103.027334 (2003).
Zulfiqar, F. & Ashraf, M. Antioxidants as modulators of arsenic-induced oxidative stress tolerance in plants: An overview. J. Hazard. Mater. 427, 127891. https://doi.org/10.1016/j.jhazmat.2021.127891 (2022).
Martín, G. et al. The photoperiodic response of hypocotyl elongation involves regulation of CDF1 and CDF5 activity. Physiol. Plant 169, 480–490. https://doi.org/10.1111/ppl.13119 (2020).
Henriques, R. et al. The antiphasic regulatory module comprising CDF5 and its antisense RNA FLORE links the circadian clock to photoperiodic flowering. New Phytol. 216, 854–867. https://doi.org/10.1111/nph.14703 (2017).
Gao, H. et al. PIF4 enhances DNA binding of CDF2 to co-regulate target gene expression and promote Arabidopsis hypocotyl cell elongation. Nature Plants 8, 1082–1093. https://doi.org/10.1038/s41477-022-01213-y (2022).
Yang, S. et al. Transcriptome-based analysis reveals a role for PpCDF5 in the promotion of anthocyanin accumulation at a low nighttime temperature. Hortic. Plant J. https://doi.org/10.1016/j.hpj.2024.08.003 (2024).
Gu, F. et al. Identification of Dof transcription factors in Dendrobium huoshanense and expression pattern under abiotic stresses. Front Genet 15, 1394790. https://doi.org/10.3389/fgene.2024.1394790 (2024).
Bernardi, J. et al. Impaired Auxin Biosynthesis in the defective endosperm18 mutant is due to mutational loss of expression in the ZmYuc1 Gene encoding endosperm-specific YUCCA1 protein in Maize. Plant Physiol. 160, 1318–1328. https://doi.org/10.1104/pp.112.204743 (2012).
Saini, S., Sharma, I., Kaur, N. & Pati, P. K. Auxin: A master regulator in plant root development. Plant Cell Rep. 32, 741–757. https://doi.org/10.1007/s00299-013-1430-5 (2013).
Pautot, V. et al. KNAT2: Evidence for a Link between knotted-like genes and carpel development. Plant Cell 13, 1719–1734. https://doi.org/10.1105/tpc.010184 (2001).
Semiarti, E. et al. in Progress in Biotechnology Vol. 18 (eds Noriyuki Morohoshi & Atsushi Komamine) 63–68 (Elsevier, 2001).
Mounet, F. et al. Down-regulation of a single auxin efflux transport protein in tomato induces precocious fruit development. J. Exp. Bot. 63, 4901–4917. https://doi.org/10.1093/jxb/ers167 (2012).
Kohnen, M. V. et al. Neighbor detection induces organ-specific transcriptomes, revealing patterns underlying hypocotyl-specific growth. Plant Cell 28, 2889–2904. https://doi.org/10.1105/tpc.16.00463 (2016).
Kloosterman, B. et al. Naturally occurring allele diversity allows potato cultivation in northern latitudes. Nature 495, 246–250. https://doi.org/10.1038/nature11912 (2013).
Imaizumi, T., Schultz, T. F., Harmon, F. G., Ho, L. A. & Kay, S. A. FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science (New York) 309, 293–297. https://doi.org/10.1126/science.1110586 (2005).
Fornara, F. et al. Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Dev. Cell 17, 75–86. https://doi.org/10.1016/j.devcel.2009.06.015 (2009).
Martín, G. et al. Circadian waves of transcriptional repression shape PIF-regulated photoperiod-responsive growth in Arabidopsis. Curr Biol 28, 311-318.e315. https://doi.org/10.1016/j.cub.2017.12.021 (2018).
Fornara, F. et al. The GI-CDF module of Arabidopsis affects freezing tolerance and growth as well as flowering. Plant. J. Cell. Mol. Biol. 81, 695–706. https://doi.org/10.1111/tpj.12759 (2015).
Imaizumi, T. Arabidopsis circadian clock and photoperiodism: time to think about location. Curr. Opin. Plant. Biol. 13, 83–89. https://doi.org/10.1016/j.pbi.2009.09.007 (2010).
Funding
This work was supported by the Science and Technology Plan Research Project of Yibin (2024NY002), and the Qihang Project of Yibin University (2023QH17)..
Author information
Authors and Affiliations
Contributions
Conceptualization, Caiming Gou, Jia Li and Ahmed H. El-Sappah; Methodology, Hua Peng, Ahmed H. El-Sappah, Jia Li and Bo Chen; Software, Caiming Gou, and Jia Li; Formal Analysis, Hua Peng, Caiming Gou, Hua Peng, Guoting Cheng and Bo Chen, ; Resources, Caiming Gou; Writing – Original Draft Preparation, Caiming Gou, Jia Li, and Ahmed H. El-Sappah; Writing – Review & Editing, Caiming Gou, Zhangfei Zheng, Jia Li and Ahmed H. El-Sappah; Funding Acquisition, Caiming Gou; All authors have read and approved the manuscript for publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gou, C., Li, J., Chen, B. et al. Genome wide identification of Dof transcription factors in Carmine radish reveals RsDof33 role in cadmium stress and anthocyanin biosynthesis. Sci Rep 15, 4766 (2025). https://doi.org/10.1038/s41598-025-88308-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88308-6