Abstract
Soluble N-ethylmaleimide-sensitive factor attachment protein receptors (SNAREs) facilitate intracellular vesicle trafficking and membrane fusion in eukaryotic organisms, significantly influencing fungal growth, development, and pathogenicity. Despite their importance, the specific functions of individual SNARE subunits remain largely unexplored. In Cytospora chrysosperma, a pathogen causing substantial damage to woody plants and leading to considerable ecological and economic impacts, 22 SNAREs have been identified. Yet, their functional roles have not been previously characterized. In this study, we focused on two SNARE-encoding genes, CcSec22 and CcSso1 that are supposed to mediate endoplasmic reticulum (ER)-Golgi and trans-Golgi Network (TGN)-secretory vesicle transport respectively. Notably, both genes demonstrate significantly elevated expression during the infection process. Targeted deletion of either of them resulted in retarded mycelial growth and remarkably decreased tolerance towards different cell-wall stressors. Interestingly, CcSec22 deletion mutants formed more hyphal branches while CcSso1 was required for morphological maintenance of conidiospores. More importantly, lack of either of them significantly reduced hyphal endocytosis and fungal virulence on host poplar. Collectively, our data highlighted that the SNARE genes CcSec22 and CcSso1 play pleiotropic roles in mycelial growth and development, stress responses, conidiation, endocytosis and in particular, they are required for the full virulence of C. chrysosperma. This study provides an insight into the role of SNARE proteins in C. chrysosperma pathogenesis.
Similar content being viewed by others
Introduction
Eukaryotic cells employ a highly conserved yet complex system of intracellular trafficking to sort and deliver cargos to their intended intracellular or extracellular destinations. This vesicular trafficking pathway is composed of four crucial steps: vesicle budding, transport, tethering, and fusion. To date, key components identified in vesicle fusion include soluble N-ethylmaleimide-sensitive factor attachment protein receptors (SNAREs), tethering complexes, Sec1/Munc18 (SM) family proteins, and Rab GTPases1. These elements play a pivotal role in the precise and efficient delivery of various cargos within the cell, a process fundamental to numerous cellular functions2,3,4. Among these components, SNARE proteins have been recognized for their specific role in mediating the transport and secretion of extracellular proteins crucial for pathogenesis in various fungal species. Structurally, they are distinguished by a specialized SNARE domain that typically contains 60–70 amino acids, organized in a series of heptad repeats5. SNARE proteins are primarily divided into two groups: v-SNAREs, which are integral to the membranes of transport vesicles, and t-SNAREs, located on the membranes of target cells6. Further classification of SNAREs hinges on specific amino acids in their SNARE domain, leading to the distinction between R-SNAREs, which contain arginine, and Q-SNAREs, characterized by the presence of glutamine7. The SNARE family encompasses a wide range of proteins. In the typical scenario, one R-SNARE forms a complex with three Q-SNAREs in the process of transporting cargo8,9,10,11.
Sec22 is an R-SNARE protein with diverse roles in eukaryotes, primarily localizing in the ER and Golgi to facilitate anterograde and retrograde transport. Recently, Sec22 homologs have been identified in several plant pathogenic fungi, where they play critical roles in pathogenicity. For instance, in Magnaporthe oryzae, Sec22 is essential for conidiogenesis, cell wall integrity, and host plant infection12. In Fusarium graminearum, the R-SNARE protein FgSec22 is crucial for hyphal growth, conidiation, deoxynivalenol (DON) production, and pathogenicity, which shows similarity as the Q-SNARE protein FgSyn813. Deleting the Sec22 homolog VdSec22 in Verticillium dahliae reduces virulence and disrupts the secretion of extracellular proteins involved in carbohydrate hydrolysis14. In Colletotrichum orbiculare, Sec22 mediates the secretion of virulence effectors to a biotrophic interface at the primary hyphal neck via exocytosis15. In the Ascomycete Sordaria macrospora, deleting Sec22 resulted in fewer ascospores, defects in ascospore pigmentation, and impaired germination, while vegetative growth remained normal. A Sec22-EGFP fusion construct, controlled by the native Sec22 promoter and terminator regions, was expressed during various stages of sexual development. In the Sec22 mutant, several development-related genes were deregulated, including three involved in melanin biosynthesis16. In Arthrobotrys oligospora, the SNARE protein AoSec22 coordinates mycelial growth, vacuole assembly, trap formation, stress response, and secondary metabolism17. All of these studies demonstrated that Sec22 was a key regulator of the secretory pathway.
Suppressors of Sec One (Sso1/2p) on the plasma membrane forms a t-SNARE complex with the cellular membrane protein Sec9p. The Sso1 of Saccharomyces cerevisiae plays a key role between the Golgi apparatus and the plasma membrane, promoting the plasma membrane and secretory vesicles fusion18. By overexpressing yeast Sso1 or Sso2 proteins, the Golgi-derived secretory vesicles can be targeted and fused to the plasma membrane to increase the production of saccharobacterial secreted proteins19. In addition, these two SNAREs proteins in S. cerevisiae are essential for the growth and development of cells. For example, Sso1 plays an important role in sporulation20,21. The absence of MoSso1 in M. oryzae led to reduced pathogenicity and deficient development of biotrophic interface complex (BIC). Compared with wild-type strains, MoSso1 mutant showed abnormal secretion of cytoplasmic effectors which rely on an unconventional secretory pathway22, indicating that MoSso1 is the key to cytoplasmic effector secretion and pathogenesis in M. oryzae22. In F. graminearum, vacuole-sorting protein Vps39 and SNARE proteins FgVam7 and FgSso1 act as vesicle complexes, which is required for vesicle transport, endocytosis and autophagy23. Thus, the growth, development, infection and the production of deoxynivalenol (DON) toxin were affected23. The pathogenicity of VdSso1 mutant in V. dahliae to cotton was significantly reduced. Comparative analysis of secreted proteins showed that the availability of pectin, cellulose, and xylan was reduced in VdSso1 mutants, meanwhile the cytotoxicity of the extracellular proteome of cotton leaves was also reduced24. In F. oxysporum, FolSso1 could regulate the processes of vegetative growth, reproduction and response to environmental stresses, however there were no significant regulatory effect on the pathogenesis25. These results indicate that SNAREs protein Sso1 plays important roles in fungal growth and development, conidial production, pathogenicity and secretion. Despite the acknowledged importance of SNAREs in the secretory pathway, their roles in fungal pathogenesis are still largely unknown.
Cytospora chrysosperma, the causative agent of stem canker, poses a substantial threat to a variety of perennial trees worldwide, severely impacting forest ecosystems. Recent advances in understanding the pathogenesis of C. chrysosperma on poplar trees have underscored the critical role of the many virulence factors in its pathogenic processes26,27,28,29,30,31,32. Notably, several effector candidates from C. chrysosperma, including members of the Cysteine-rich secretory proteins, Antigen 5, and Pathogenesis-related 1 proteins (CAP) and Glycoside Hydrolase 12 (GH12) families, have been implicated in fungal virulence and manipulation of the plant immune system33,34,35. However, the mechanisms governing their secretion remain unexplored.
In previous studies, 22 SNARE proteins have been identified and classified in C. chrysosperma36, yet a comprehensive understanding of their functions is lacking. Consequently, this study aims to clarify the functions of two SNARE proteins, CcSec22 and CcSso1 in C. chrysosperma. Our approach involved multiple analytical methods applied to targeted gene disruption mutants. We discovered that the deletion of either the CcSec22 or CcSso1 gene leads to a marked reduction in both vegetative growth and resistance to a variety of cell-wall stressors. Furthermore, we found that CcSec22 plays a significant role in shaping the morphology of hyphal branching, whereas CcSso1 is crucial for spore morphology. Most notably, our study reveals that both CcSec22 and CcSso1 are essential for efficient endocytosis and the overall virulence of the fungus.
Materials and methods
Bioinformatic analysis
The gene information of CcSec22 (GME3972_g) and CcSso1 (GME1900_g) were taken from previously published data, and their functional domains were confirmed with the default parameters of InterProScan (https://www.ebi.ac.uk/interpro/search/sequence/) and HMMER (https://www.ebi.ac.uk/Tools/hmmer/). A BLAST algorithm was used to search for published orthologs of Sec22 or Sso1 in PHI-base (http://phi-blast.phi-base.org), combined with manual mining of literature. The resulting sequences were downloaded from the GenBank database, followed by sequence alignment and phylogenetic analysis with the MEGA 6.0 software and the alignment was further processed with the Genedoc software.
Strains and culture conditions
The wild-type (WT) fungus C. chrysosperma (CFCC 89981), isolated from poplar, was used in this study. The wild-type and the mutant strains constructed in this study were cultured on potato dextrose agar (PDA, 200 g potato, 20 g dextrose, 15 g agar) media at 25–30 °C in the dark. Liquid PDB medium was used to cultivate the fungal mycelia for DNA extraction, RNA isolation and protoplast preparation. Fresh hyphal blocks were incubated in yeast extract peptone dextrose (YEPD, 20 g peptone, 10 g yeast extract, and 20 g glucose) medium for downstream cell wall digestion to produce protoplast. In addition, TB3 (3 g yeast extract, 3 g casamino acid, 200 g sucrose, 7 g agar) medium was used to culture transformants. The Escherichia coli Dh5α strains were used for vector construction and grown on Luria–Bertani agar plate at 37 °C in the dark.
RNA extraction and gene expression analysis
To analyze the expression of CcSec22 and CcSso1 gene during infection processes, RNA samples were extracted from wild-type hyphal mass cultured in PDB with poplar twig tissues for 0 (immediately after inoculation), 1, 2, 3, 4, 5 and 6 days. All RNA samples were isolated with TRIzol reagent (Invitrogen, USA) and purified with a the PureLink RNA Mini Kit (Invitrogen, USA) following the instructions of the manufacture. RNA integrity was confirmed by agarose gel electrophoresis. First-strand cDNA was synthesized from 1 μg of RNA with SuperScript IV reverse transcriptase (Invitrogen) according to the manufacturer’s instructions, followed by RT-qPCR with ABScript II One Step SYBR Green RT-qPCR Kit (Abclonal) using a bio-rad CFX Real-Time PCR Detection Systems. The CcActin gene served as the endogenous control. Relative gene expression levels were calculated by the 2-ΔΔCt method. Each gene assay was performed in biological triplicate with four independent technical replicates each.
Targeted gene deletion of CcSec22 and CcSso1
CcSec22 and CcSso1 were disrupted by homologous recombination using split-marker strategy (Fig. S2A)37. Briefly, the upstream and downstream fragments of either the CcSec22 or CcSso1 gene were amplified from C. chrysosperma DNA with paired primers, CcSec22-5Ffor/CcSec22-5Frev and CcSec22-3Ffor/Cc22-3Frev or CcSso1-5Ffor/CcSso1-5Frev and CcSso1-3Ffor/CcSso1-3Frev, respectively. The hygromycin-resistant gene HPH was used as a selection marker, which included approximately 20 bp of overlap with the 5′ and 3′ flanking sequences of CcSec22 or CcSso1, respectively. The target gene fragment for CcSec22 or CcSso1 disruption was obtained by fusion of the upstream and downstream fragments and two-thirds of the hygromycin cassette by overlap PCR using the primers, CcSec22-5Ffor/HY-R and YG-F/CcSec22-3Frev or CcSso1-5Ffor/HY-R and YG-F/CcSso1-3Frev, respectively.
Protoplast generation and transformation were performed as described previously38. Briefly, high-quality protoplasts of the wild-type strain were generated using Driselase and Lysing enzyme, and the recombinant fragments were then transformed into protoplasts, followed by selection on TB3 agar medium supplemented with 25 μg/ml hygromycin. The successful disruptions were verified by PCR assays with the primer pairs External-CcSec22for/External-CcSec22rev and Internal-CcSec22for/Internal-CcSec22rev, or External-CcSso1for/External-CcSso1rev and Internal-CcSso1for/Internal-CcSso1rev, respectively.
For the complementation of the mutant strains, a fragment containing the entire length of the CcSec22 or CcSso1 coding region along with its native promoter and terminator regions was PCR amplified using the primer pairs CcSec22-Compfor/ CcSec22-Comprev, or CcSso1-Compfor/ CcSso1-Comprev, respectively. The resulting PCR products were co-transformed into protoplasts of the ΔCcSec22-8 or ΔCcSso1-7 strain along with a geneticin-resistant cassette. Transformants were cultured in TB3 medium supplemented with 25 μg/ml hygromycin and 50 μg/ml geneticin. successful complementation was confirmed by PCR with the primer pair Internal-CcSec22for/Internal-CcSec22rev, or Internal-CcSso1for/Internal-CcSso1rev, respectively. The complementation strain was named ΔCcSec22/C or ΔCcSso1/C in this study.All primers used in the present study are listed in Table S1.
Analysis of vegetative growth, conidiation and stress tolerance
To investigate differentiation among the wild-type and mutant strains, 5-mm-diameter mycelial plugs, derived from the leading edge of the 3-days-old colonies, were inoculated onto PDA plates and then subjected to morphology and size analysis. To determine the levels of stress resistance, cultures were grown on PDA plates amended with 300 μg/ml calcofluor white (CFW) and 300 μg/ml Congo red (CR) for 60 h. The relative growth inhibition rate of the fungal strains was calculated using the following formula: inhibition rate = (CFW or CR − CK)/CK, where CFW represents the mean diameter of the fungi grown on PDA plates supplemented with 300 μg/ml CFW, CR represents the mean diameter of the fungi grown on PDA plates supplemented with 300 μg/ml CR, and CK represents the mean diameter of the fungi grown on PDA plates without any additives. The pycnidia yield of each strain were checked from the 25-day-old cultures grown on PDA plate. Three biological replicates were analyzed for each treatment.
Microscopy image processing
To examine hyphal morphology, strains were grown on a thin layer of PDA medium on the microscope slides for 2 days, followed by light microscopical observation. To examine endocytosis, FM4-64 (Thermo fisher, Waltham, MA, USA) was firstly dissolved in sterile water at the final concentration of 5 μM. Hyphae grown on slides for 2 dpi were stained with 10 μL 5 μM FM4-64 at room temperature and then observed without glass cover under a fluorescence microscope immediately with excitation spectra at 535 ± 20 nm and emission spectra at 610 ± 30 nm. Images were captured at 0 min, 5 min, 10 min, 15 min, 20 min, and up to 30 min39. To analyze endocytosis process, fluorescence intensities (gray values) in the cell at 0 or 20 min were quantified using the Plot Profile function in Fiji. These values were then exported and processed in Excel. The signal differences between the membrane (two peaks) and the cytosol (region between the two peaks) were compared among different strains at 0 and 20 min, respectively.
To examine the conidial morphology, pycnidia were picked and smashed in sterile water to release individual spores. Later, the conidiospores were observed under a light microscope. These experiments were performed with three biological replicates and five technical replicates for each treatment.
Infection assays
Cytospora chrysosperma, an opportunistic pathogen, initiates colonization through wounds when host trees grow weak. For the pathogenicity test, 15-cm-long healthy annual branches of the susceptible species Populus × canadensis were selected and scalded with a 5-mm-diameter hot iron bar, then they were inoculated with 5-mm-diameter mycelial plugs of C. chrysosperma wild-type strains, deletion mutants and complemented mutants from the leading edge of the colonies at 3 dpi. After inoculation, the twigs sealed with sealing film were placed in trays with distilled water to maintain humidity at 25 °C in the dark. In the following days, the twigs were sprayed with water for promotion of pathogen infection. Lesions of inoculated twigs were photographed and measured at 4 dpi.
Statistical analysis
All experimental data were presented as the mean ± standard deviation (SD) of at least three biological measurements. The differences between treatments were statistically evaluated by one-way ANOVA analysis of variance using Prism 10 (GraphPad, San Diego, CA, USA). Differences were considered statistically significant if p value < 0.05.
Results
Bioinformatic analysis of CcSec22 and CcSso1 in C. chrysosperma
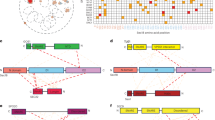
The genome of C. chrysosperma encodes 22 SNAREs as identified previously36. The number of genes encoding vesicle-fusion components is highly conserved among C. chrysosperma, V. dahliae and yeast14,40. A putative model of the associations of SNARE complexes with vesicle trafficking in C. chrysosperma was constructed by identifying vesicle-fusion component orthologs between C. chrysosperma and V. dahliae or Saccharomyces cerevisiae (Fig. 1). Vesicle-fusion component play a central role in extracellular trafficking, and, therefore, may be necessary for the transport of effectors and other proteins important for pathogenesis. Certain vesicle-fusion components (Qa, CcSed5; Qb, CcBos1; Qc, CcBet1; R, CcSec22/CcYkt6) are predicted to participate in trafficking from the endoplasmic reticulum (ER) to the cis-Golgi, while others mediate the movement of secretory vesicles to the plasma membrane (PM) (Qa, CcSso1/2; Qb/Qc, CcSec9; R, CcSnc1) (Fig. 1).
Illustration of the SNARE association in vesical fusion steps in C. chrysosperma. ER, endoplasmic reticulum; TGN, Trans-Golgi network.
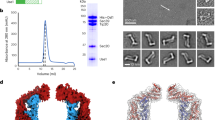
CcSec22 encodes a polypeptide of 215 amino acids18, containing a SNARE-like longin domain (light blue, 133-211 aa, PF13774), a profilin-like globular domain consisting of a five-stranded antiparallel β-sheet that is sandwiched by an α-helix on one side, and two α-helices on the other. CcSec22 has an additional synaptobrevin domain (blue, 36-117 aa, PF00957), formed by 5 β-sheets and 7 α-helixes (Fig. 2A). The full length of CcSso1 is 335 aa, which encodes a 50 aa SNARE domain (light pink, 70-260 aa, PF05739) and a 190 aa syntaxin domain (pink, 70-260 aa, PF00804), tertiarily formed by 7 α-helixes only (Fig. 2B). And both of them have a C-terminal transmembrane (TM) domain as indicated in grey (196-214 aa, 221-314 aa) respectively in Fig. 2.
Tertiary structure and phylogenetic tree of Sec22 or Sso1 from reported fungi. (A) Phylogenetic tree of Sec22 or Sso1 proteins was constructed based on alignment of the full sequences of Sec22 from fungi: Cytospora chrysosperma (CcSec22, GME3972_g and CcSso1, GME1900_g) Magnaporthe oryzae (MoSec22, MGG_04050.6 and MoSso1, MGG_04090.6), Sordaria macrospora (SmSec22, SMAC_06625), Fusarium graminearum (FgSec22, XP_011323737.1 and FgSso1, FGSG_00950)), Colletotrichum orbiculare (CoSec22, AB778552), Verticillium dahlia (VdSec22, VDAG_08386 and VdSso1, VDAG_00278), and Arthrobotrys oligospora (AoSec22, AOL_s00676g350). (B) Tertiary structure of CcSec22 and CcSso1. Both CcSec22 and CcSso1 contain 3 different domains, as indicated in different colors: SNARE-like longin domain in light blue, synaptobrevin domain in blue, syntaxin domain in light pink, SNARE domain in pink and TM domain in grey.
A highest of 73.36% amino acid sequence identity was found between CcSec22 and VdSec22 (VDAG_08386) with PHI-base blast program, and the best match of CcSso1 to an annotated Sso1 protein was that of V. dahliae (VDAG_00278; 53.59% identity in amino acid sequence), followed by FgSso1 (GnSyn1, FGSG_00950, 31.88%). Besides, we selected functionally reported Sec22 and Sso1 homologs from phytopathogenic fungi for sequence alignment. All selected Sec22 homologs shared the conserved Arginine (R) residue (Fig. 2A, Fig. S1A) and Sso1 the Glutamine (Q) residue (Fig. 2A, Fig. S1B). In addition, Sec22 homologs share high sequence similarities and protein size, with deviation of only one or two amino acids (Fig. S1A). Further processing of these two alignments for phylogenetic analysis showed that CcSec22 and CcSso1 are highly homologous to MoSec22 and VdSso1 respectively and are well grouped into 2 categories (Fig. 2A).
Role of CcSec22 or CcSso1 in mycelial growth and conidiation
To evaluate the role of CcSec22 or CcSso1 in the growth and development of C. chrysosperma, their mutant strains were generated by replacing the whole CcSec22 or CcSso1 coding region with the hygromycin phosphotransferase resistance (HPH) marker gene using the split-marker strategy (Fig. S2A). Mutant strains were selected and confirmed by PCR analysis. Two deletion mutants of each gene, ΔCcSec22-8 and ΔCcSec22-27, and ΔCcSso1-7 and ΔCcSso1-17 were selected for downstream analysis. For complementation, ΔCcSec22-8 and ΔCcSso1-7 were used as receptor strains respectively (Fig. S2B,C).
The CcSec22 or CcSso1 mutant showed delayed vegetative growth in comparison to the wild-type strain on PDA medium at 60 h post inoculation (hpi) (Fig. 3A,B). In particular, the colony morphology of ΔCcSec22 mutant was abnormally dense as compared to the wild-type and complemented strains (Fig. 3A). To understand the mechanism behind, microscopic analysis was employed to observe the hyphal morphology. The ΔCcSec22 mutant showed hyper-branching phenotype with more lateral branches (Fig. 3C). In contrast CcSso1 deletion mutants exhibited neither colonial nor branching morphological disturbance (Fig. 3C). The quantification of individual hyphal branches in each of the strain were shown in Fig. S3.
Comparison of mycelial growth and conidial morphology of wild-type and mutant strains in C. chrysosperma. (A) Representative colony of the wild-type and mutant strains on PDA plates. (B) Quantification of colony diameter in different strains. The error bars represent the standard deviations based on three independent biological replicates with three technical replicates each. The different letters indicate significant differences at P < 0.05. (C) Hyphal branching in the tested strains cultured on slides coated with PDA media (scale bar = 50 μm). (D) Microscopic images (scale: 10 μm) of conidia.
Except morphological observation, we further investigated the roles of CcSec22 or CcSso1 in conidiation and conidial morphology. The wild-type strain and ΔCcSec22 mutant produced a bar-like spore; in contrast, the conidia of the ΔCcSso1 were morphologically abnormal with a rounder shape (Fig. 3D).
CcSec22 is required for cell wall integrity
To investigate the cause of defects resulting from loss of CcSec22 or CcSso1, the structural integrity of the cell wall and membrane was examined. Our data indicate that either ΔCcSec22 or ΔCcSso1 mutants showed significantly delayed growth with irregular colony morphology on CFW or CR plates as compared to the wild-type and complementation strains (Fig. 4). These findings suggest that CcSec22 and CcSso1 are both important for cell wall stress resistance in the fungus.
Defects of CcSec22 or CcSso1 deletion mutants towards stress responses. (A) Colonial morphology of fungal strains under cell wall-perturbing stress. CK serves as the control, with no cell wall stressors added to the media. (B) Relative growth inhibition rate of fungal colonies on PDA plates supplemented with indicated concentrations of Congo red (CR) and calcofluor white (CFW). The different letters indicate a significant difference at p < 0.05.
Deletion of CcSec22 or CcSso1 affects endocytosis
As both CcSec22 and CcSso1 are SNARE proteins involved in vacuole assembly and membrane fusion, we performed staining with FM4-64, an endocytic tracer. At 0 min, the dye accumulated both at plasma membrane and punctate and patchy structures in wild-type strain, but specifically at the plasma membrane in either CcSec22 or CcSso1 deletion mutant, indicating that the internalization of the dye had not begun in mutants, but in wild-type strain (Fig. 5A). Quantification of fluorescence intensity at 0 min revealed a significantly greater difference between membrane and cytosolic signals in the deletion mutants (ΔCcSec22: 76.9; ΔCcSso1: 72.5) compared to the wild type (50.1) (Fig. 5B). At 20 min, punctate and patchy structures slightly started to appear in cytoplasm in both deletion mutants. While the dye was dispersed in the whole hyphae of wild-type at 20 min, indicating the uptake of the dye and functional endocytosis process in hyphae (Fig. 5A). This observation was further supported by the reduced signal differences between the membrane and cytosol across all strains at 20 min (WT: 39.9; ΔCcSec22: 50.6; ΔCcSso1: 32.3) (Fig. 5B).
Involvement CcSec22 or CcSso1 in endocytosis and transport of effector proteins. (A) Representative hyphal endocytosis. Hyphal grown on slides were stained using 5 μM FM4-64, and images were photographed at different time points (0–20 min) using fluorescence microscope. (B) Lines showing the fluorescence intensity in membrane and cytoplasm of each strain at different time points.
The two membrane-fusion components CcSec22 and CcSso1 are required for full virulence of C. chrysosperma
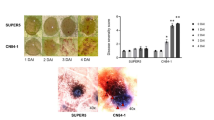
Individual characterization using RT-qPCR showed that both CcSec22 and CcSso1 were upregulated after 3 dpi. Specifically, CcSec22 reached its highest expression at 4 dpi with a two-fold increase, and CcSso1 peaked at 6 dpi with a three-fold increase (Fig. 6A). These results suggest their potential involvement in pathogenicity, albeit at a relative later stage.
Pathogenicity test of deletion mutants. (A) The expression pattern of CcSec22 or CcSso1 gene during infection process. RT-qPCR was performed to quantify the expression profile of CcSec22 or CcSso1 gene after C. chrysosperma infection. Lines represent the mean values of three replicates ± standard deviation (SD). Different small letters indicate significant differences at P < 0.05. (B) Representative infection symptoms on detached poplar twigs inoculated with the wild-type, mutant, and complemented strains at 4 dpi. (C) Diameter of lesions produced by the different strains on twigs tested in (B). The different letters on the bars indicate a significant difference at P < 0.05.
To assess the role of membrane-fusion components in C. chrysosperma pathogenesis, the susceptible poplar variety was inoculated with two independent CcSec22 and CcSso1 deletion mutants as mentioned before along with the wild-type strain. As expected, the CcSec22 deletion strains were less virulent on poplar, whereas the wild-type strain caused severe symptoms (Fig. 6A,B). Similarly, targeted replacement of CcSso1 also significantly reduced virulence on poplar twigs (Fig. 6A,B). Furthermore, two independent complementation transformant strains revealed that reintroduction of CcSec22 and CcSso1 with native promoter to ΔCcSec22-8 and ΔCcSso1-7 mutants, respectively, restored their virulence, comparable to that of the wild-type strain when inoculated on poplar. These results therefore suggested that CcSec22 or CcSso1 is required for full virulence of C. chrysosperma on susceptible poplar, indicating that the vesicle-fusion components involved in trafficking play important roles in the pathogenicity and virulence of C. chrysosperma.
Discussion
In this study, we characterized two SNARE homolog proteins, Sec22 and Sso1, in C. chrysosperma, focusing on their roles in pathogenesis-related phenotypes. Our gene-targeted replacement approach revealed that deletion of CcSec22 or CcSso1 leads to a range of developmental defects, highlighting their involvement in cell wall integrity maintenance. While the precise functions of CcSec22 and CcSso1 in membrane fusion and trafficking remain to be fully determined, our findings underscore their essential roles in fungal endocytosis and pathogenicity in this species.
SNARE proteins are the essential components of the vesicles transport system including budding, transportation and membrane fusion which involve in cargos internal transport, endocytosis, and external secretion and so on41. By regulating the fusion of vesicles that containing important cell wall materials at specific sites within the growing tip of the hypha, SNARE proteins are generally involving in the fungal growth including hyphal development, sporulation, and fungal morphogenesis42,43. Additionally, SNARE proteins also play a pivotal role in polarized growth in fungi, enabling the extension and branching of hyphae44. Therefore, numerous studies have reported that deletion of SNARE proteins would reduce vegetative growth and sporulation, affect fungal morphogenesis, result in abnormal hyphal branching14,45. Here, we also found that deletion of CcSec22 or CcSso1 would significantly retard fungal growth, especially CcSec22. And the branching of hyphae of CcSec22 deletion mutants was inordinate compared to the wild type and complemented strains, which may be crucial for fungal colonization and nutrient acquisition (Fig. 3C). On the other hand, the defects of SNARE deletion mutants in endocytosis would also affect fungal growth through acting on nutrient uptake process46. For example, the MoVam7 from Magnaporthe oryzae and FgVam7 from F. graminearum are both required for endocytosis, deletion of Vam7 would compromise the ability of endocytosis44,47. In addition to the fungal systems, SNARE proteins were also critical. They are required for cellular growth, division, movement, and cell–cell communication. Dysfunction of SNARE proteins will results in a various of human diseases, such as neurodegenerative disorders, cancer, diabetes and infections48. In this study, deletion of CcSec22 or CcSso1 would also compromise fungal endocytosis, the extracellular fluorochrome FM4-64 was normally uptaken into the fungal cytoplasm, while the fluorochrome was trapped in the fungal membrane in CcSec22 or CcSso1 deletion mutants (Fig. 5).
Cell wall integrity is the prerequisite for fungal adaptation and survival. By regulating the transcription of cell wall related materials such as chitin to the right regions, SNARE proteins contribute to the establishment and maintenance of the fungal cell wall, which is essential for providing structural support and protection against environmental stressors49. Missing of SNARE proteins would damage cell wall integrity and resulted in more sensitive to cell wall stressors50,51,52. For example, the Q-SNARE protein FgSyn8 from F. graminearum is required for cell wall integrity, the growth of ΔFgSyn8 mutant was conspicuously reduced in response to the cell wall damaging agents53. Similarly, CcSec22 or CcSso1 deletion mutants were more sensitive to the CR and CFW compared to the wild type.
The roles of SNARE proteins in fungal pathogenicity are clear, which contribute to the formation of specific infection structure such as appressorium and the secretion of numerous virulence factors through exocytosis and so on54. The MoVam7 deletion mutants could not form appressorium, resulting in the loss of fungal pathogenicity in the unwound rice leaves44. Furthermore, the SNARE proteins are involved in DON accumulation in phytopathogenic Fusarium species43,51. Several of SNARE proteins also participate in the secretion of secondary metabolite in fungi17. Notably, the MoSso1 is involved in a novel form of secretion system, which contributes to the secretion of cytoplasmic effectors in M. oryzae55. Additionally, the SNARE proteins also participate into the fungal pathogenic processes through affecting the expansion of hypha among the plant cells. Herein, the observed upregulation of CcSec22 and CcSso1 during infection implicates their specific functions in pathogenicity, likely associated with increased vesicular trafficking demands during host invasion. In pathogenic fungi, this process is crucial for the secretion of effector proteins and enzymes necessary for host penetration, a dependency underscored by the induction of these SNAREs during infection. In this study, deletion of CcSec22 or CcSso1 significantly reduced fungal pathogenicity (Fig. 6). Collectivelly, we found that the SNARE proteins in various pathogenic fungi share many conserved roles such as fungal growth, stress responses, endocytosis, exocytosis and pathogenicity. Although the sequences’ identity of Sec22 or Sso1 was low among different fungal species, the domain structures of them are highly conserved, which may contribute to their similar functions among fungi (Fig. 2).
Overall, this study illuminates the complex roles of CcSec22 and CcSso1 in C. chrysosperma, from cellular processes to host interactions. Future research should delve into the mechanistic aspects of these SNARE proteins, their interactomes, and specific cargo, to fully elucidate their roles in fungal biology and pathogenesis. Insights into how these SNAREs modulate C. chrysosperma's pathogenicity could pave the way for novel antifungal strategies.
Data availability
The sequences of CcSec22 and CcSso1 could be found in NCBI database with the accession number: ROW02800.1 and ROV93767.1 respectively.
References
Stanton, A. E. & Hughson, F. M. The machinery of vesicle fusion. Curr. Opin. Cell Biol. 83, 102191 (2023).
Cui, L. et al. Vesicle trafficking and vesicle fusion: Mechanisms, biological functions, and their implications for potential disease therapy. Mol. Biomed. 3(1), 29 (2022).
Koumandou, V. L., Dacks, J. B., Coulson, R. M. R. & Field, M. C. Control systems for membrane fusion in the ancestral eukaryote; evolution of tethering complexes and SM proteins. BMC Evol. Biol. 7(1), 29 (2007).
Ostrowicz, C. W., Meiringer, C. T. & Ungermann, C. Yeast vacuole fusion: a model system for eukaryotic endomembrane dynamics. Autophagy 4(1), 5–19 (2008).
Weimbs, T., Mostov, K., Low, S. H. & Hofmann, K. A model for structural similarity between different SNARE complexes based on sequence relationships. Trends Cell Biol. 8(7), 260–262 (1998).
Ungar, D. & Hughson, F. M. SNARE protein structure and function. Annu. Rev. Cell Dev. Biol. 19, 493–517 (2003).
Sutton, R. B., Fasshauer, D., Jahn, R. & Brunger, A. T. Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 Å resolution. Nature 395(6700), 347–353 (1998).
Ossig, R. et al. Exocytosis requires asymmetry in the central layer of the SNARE complex. EMBO J. 19(22), 6000–6010 (2000).
Wang, T., Li, L. & Hong, W. SNARE proteins in membrane trafficking. Traffic 18(12), 767–775 (2017).
Burri, L. & Lithgow, T. A complete set of SNAREs in yeast. Traffic 5(1), 45–52 (2004).
Koike, S. & Jahn, R. SNARE proteins: Zip codes in vesicle targeting?. Biochem. J. 479(3), 273–288 (2022).
Song, W. et al. R-SNARE homolog MoSec22 is required for conidiogenesis, cell wall integrity, and pathogenesis of Magnaporthe oryzae. PLoS One 5(10), e13193 (2010).
Adnan, M. et al. R-SNARE FgSec22 is essential for growth, pathogenicity and DON production of Fusarium graminearum. Curr. Genet. 66(2), 421–435 (2020).
Wang, J. et al. Snare-encoding genes Vdsec22 and Vdsso1 mediate protein secretion required for full virulence in Verticillium dahliae. Mol. Plant Microbe Interact. MPMI 31(6), 651–664 (2018).
Irieda, H. et al. Colletotrichum orbiculare secretes virulence effectors to a biotrophic interface at the primary hyphal neck via exocytosis coupled with Sec22-mediated traffic. Plant Cell 26(5), 2265–2281 (2014).
Traeger, S. & Nowrousian, M. Functional analysis of developmentally regulated genes chs7 and sec22 in the Ascomycete Sordaria macrospora. G3 Bethesda 5(6), 1233–1245 (2015).
Zhu, Y. et al. SNARE protein AoSec22 orchestrates mycelial growth, vacuole assembly, trap formation, stress response, and secondary metabolism in Arthrobotrys oligospora. J. Fungi (Basel) 9(1), 75 (2023).
Aalto, M. K., Ronne, H. & Keränen, S. Yeast syntaxins Sso1p and Sso2p belong to a family of related membrane proteins that function in vesicular transport. EMBO J. 12(11), 4095–4104 (1993).
Ruohonen, L., Toikkanen, J., Tieaho, V., Outola, M. & Soderlund, H. S. K. Enhancement of protein secretion in Saccharomyces cerevisiae by overproduction of Sso protein, a late-acting component of the secretory machinery. Yeast 13(4), 337–351 (1997).
Yuan, Q. & Jäntti, J. Functional analysis of phosphorylation on Saccharomyces cerevisiae syntaxin 1 homologues Sso1p and Sso2p. PLoS One 5(10), e13323 (2010).
Oyen, M., Jäntti, J. & Keränen, S. H. R. Mapping of sporulation-specific functions in the yeast syntaxin gene SSO1. Curr. Genet. 45(2), 76–82 (2004).
Giraldo, M. C., Dagdas, Y. F. & Gupta, Y. K. Two distinct secretion systems facilitate tissue invasion by the rice blast fungus Magnaporthe oryzae. Nat. Commun. 2013, 4 (1996).
Li, B. et al. The FgVps39-FgVam7-FgSso1 complex mediates vesicle trafficking and is important for the development and virulence of Fusarium graminearum. Mol. Plant Microbe Interact. MPMI 30(5), 410–422 (2017).
Wang, J. & Tian, L. DD Z: SNARE-encoding genes VdSec22 and VdSso1 mediate protein secretion required for full virulence in Verticillium dahliae. Mol. Plant Microbe Interact. MPMI 31(6), 651–664 (2018).
Yang, S., Hai-Xia, Y., Sheng-Xuan, L. & Zong-Hua, W. Functional analysis of the SNARE protein FolSso1 in Fusarium oxysporum f. sp. lycopersici. Acta Phytopathol. Sin. 49, 530–538 (2019).
Yu, L., Yang, Y., Qiu, X., Xiong, D. & Tian, C. The mitogen-activated protein kinase module CcSte11-CcSte7-CcPmk1 regulates pathogenicity via the transcription factor CcSte12 in Cytospora chrysosperma. Stress Biol. 4(1), 1–19 (2024).
Yu, L., Yang, Y., Xiong, D. & Tian, C. Phosphoproteomic and metabolomic profiling uncovers the roles of CcPmk1 in the pathogenicity of Cytospora chrysosperma. Microbiol. Spectr. 10(4), e0017622 (2022).
Yu, L. et al. Comparative transcriptomic analysis of MAPK-mediated regulation of pathogenicity, stress responses, and development in Cytospora chrysosperma. Phytopathology 113(2), 239–251 (2023).
Xiong, D., Yu, L., Shan, H. & Tian, C. CcPmk1 is a regulator of pathogenicity in Cytospora chrysosperma and can be used as a potential target for disease control. Mol. Plant Pathol. 22(6), 710–726 (2021).
Wang, Y. & Wang, Y. Oxalic acid metabolism contributes to full virulence and pycnidial development in the poplar canker fungus Cytospora chrysosperma. Phytopathology 110(7), 1319–1325 (2020).
Han, Z., Yu, R., Xiong, D. & Tian, C. A Sge1 homolog in Cytospora chrysosperma governs conidiation, virulence and the expression of putative effectors. Gene 778, 145474 (2021).
Wen, D. S., Yu, L., Xiong, D. G. & Tian, C. M. Genome-wide identification of bZIP transcription factor genes and functional analyses of two members in Cytospora chrysosperma. J. Fungi 8(1), 34 (2022).
Han, Z., Xiong, D., Xu, Z., Liu, T. & Tian, C. The Cytospora chrysosperma virulence effector CcCAP1 mainly localizes to the plant nucleus to suppress plant immune responses. mSphere 6(1), e00883-00820 (2021).
Han, Z., Xiong, D., Schneiter, R., Tian, C. The function of plant PR1 and other members of the CAP protein superfamily in plant-pathogen interactions. Mol. Plant Pathol. (2023).
Xu, Z., Luo, Z., Xiong, D., Gao, M. & Tian, C. A glycoside hydrolase 12 protein from Cytospora chrysosperma triggers plant immunity but is not essential to virulence. Phytopathol. Res. 5(1), 32 (2023).
Li, X., Xiong, D. & Tian, C. Genome-wide identification, phylogeny and transcriptional profiling of SNARE genes in Cytospora chrysosperma. J. Phytopathol. 169(7–8), 471–485 (2021).
Catlett, N. L., Lee, B.-N., Yoder, O. C. & Turgeon, B. G. Split-marker recombination for efficient targeted deletion of fungal genes. Fungal Genet. Rep. 50(1), 9–11 (2003).
Liu, L. et al. Genetic transformation system of Cytospora chrysosperma, the causal agent of poplar canker. Microbiol. China 44, 2487–2497 (2017).
Wang, X., Lu, D. & Tian, C. CgEnd3 regulates endocytosis, appressorium formation, and virulence in the poplar anthracnose fungus Colletotrichum gloeosporioides. Int. J. Mol. Sci. 22(8), 4029 (2021).
Jahn, R., Lang, T. & Südhof, T. C. Membrane fusion. Cell 112(4), 519–533 (2003).
Gupta, G. D. & Health, I. B. Predicting the distribution, conservation, and functions of SNAREs and related proteins in fungi. Fungal Genet. Biol. 36(1), 1–21 (2002).
Kienle, N., Kloepper, T. H. & Fasshauer, D. Phylogeny of the SNARE vesicle fusion machinery yields insights into the conservation of the secretory pathway in fungi. BMC Evol. Biol. 9, 19 (2009).
O’Mara, S. P. et al. The Fusarium graminearum t-SNARE Sso2 is involved in growth, defense, and don accumulation and virulence. Mol. Plant Microbe Interact. MPMI 33(7), 888–901 (2020).
Dou, X. et al. MoVam7, a conserved SNARE involved in vacuole assembly, is required for growth, endocytosis, ROS accumulation, and pathogenesis of Magnaporthe oryzae. PLoS One 6(1), e16439 (2011).
Zhao, W. et al. Genome-wide identification of Phytophthora sojae SNARE genes and functional characterization of the conserved SNARE PsYKT6. Fungal Genet. Biol. 48(3), 241–251 (2011).
Adnan, M. et al. SNARE protein Snc1 is essential for vesicle trafficking, membrane fusion and protein secretion in fungi. Cells 12(11), 1547 (2023).
Zhang, H. et al. SNARE protein FgVam7 controls growth, asexual and sexual development, and plant infection in Fusarium graminearum. Mol. Plant Pathol. 17(1), 108–119 (2016).
Yoon, T. Y. & Munson, M. SNARE complex assembly and disassembly. Curr. Biol. CB 28(8), R397–R401 (2018).
Li, B. et al. The t-SNARE protein FgPep12, associated with FgVam7, is essential for ascospore discharge and plant infection by trafficking Ca2+ ATPase FgNeo1 between Golgi and endosome/vacuole in Fusarium graminearum. PLoS Pathog. 15(5), e1007754 (2019).
Adnan, M., Islam, W., Zhang, J., Zheng, W. & Lu, G. D. Diverse role of SNARE protein Sec22 in vesicle trafficking, membrane fusion, and autophagy. Cells 8(4), 337 (2019).
Li, B., Gao, Y., Mao, H. Y., Borkovich, K. A. & Ouyang, S. Q. The SNARE protein FolVam7 mediates intracellular trafficking to regulate conidiogenesis and pathogenicity in Fusarium oxysporum f. sp. lycopersici. Environ. Microbiol. 21(8), 2696–2706 (2019).
Pelham, H. R. SNAREs and the specificity of membrane fusion. Trends Cell Biol. 11(3), 99–101 (2001).
Adnan, M. et al. Q-SNARE protein FgSyn8 plays important role in growth, DON production and pathogenicity of Fusarium graminearum. Microb. Pathog. 140, 103948 (2020).
Qi, Z. et al. The syntaxin protein (MoSyn8) mediates intracellular trafficking to regulate conidiogenesis and pathogenicity of rice blast fungus. New Phytol. 209(4), 1655–1667 (2016).
Giraldo, M. C. et al. Two distinct secretion systems facilitate tissue invasion by the rice blast fungus Magnaporthe oryzae. Nat. Commun. 2013, 4 (1996).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32471869).
Author information
Authors and Affiliations
Contributions
D.X. designed the experiment. Z.H., Z.L. and X.L. conducted the experiments. Z.H., Z.L. and D.X. wrote the draft manuscript. C.T. and D.X. reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, Z., Luo, Z., Li, X. et al. SNARE genes CcSec22 and CcSso1 coordinate fungal growth, sporulation, cell wall stress tolerance, endocytosis and full virulence in Cytospora chrysosperma. Sci Rep 15, 5034 (2025). https://doi.org/10.1038/s41598-025-88584-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88584-2