Abstract
We determined the relationship between ferroptosis and immune cells in ankylosing spondylitis and the role of the Chinese herbal medicine Cassia twigs in treating ankylosing spondylitis. We analyzed clinical data on ankylosing spondylitis, transcriptome data, single-cell sequencing data, and genes related to ferroptosis and Cassia twigs. Clinical variables related to AS were selected through logistic regression analysis of the clinical data combined with machine learning. GSEA and enrichment analysis were performed on genes related to ferroptosis, combined with transcriptome data and drug-related genes, to identify the key genes and drug targets related to AS, as well as, the key immune cells. Then, the single-cell data and cell subtypes were analyzed. Finally, the interconnections between immune cells were analyzed through intercellular communication. Five variables, including neutrophils, were screened for clinical data analysis. The AUC of the experimental group was 0.859 and that of the validation group was 0.807. Ferroptosis gene NFE2L2 was identified as the final drug target of AS; it was upregulated in AS and downregulated in the control group by immunohistochemical verification, both of which were statistically significant (P < 0.001). Neutrophils were divided into two subgroups: high expression of NFE2L2 and low expression of NFE2L2. Through molecular docking, Cassia twigs were found to effectively act on the ferroptosis gene NFE2L2. Neutrophils act as important immune cells in AS. The twigs of the Chinese herb Cassia can treat AS by acting on the protein structure of the ferroptosis gene NFE2L2.
Similar content being viewed by others
Introduction
Ferroptosis1 is a type of non-apoptotic cell death characterized by iron-dependent lipid peroxidation. It is morphologically, biochemically, and genetically distinct from other forms of cell death, such as apoptosis, autophagy, and necrosis. Ferroptosis plays an important role in various diseases, including liver disease, neurological disorders, inflammatory diseases, and cancer2,3,4. Ferroptosis is often associated with immune cells. Its relationship with macrophages is well-established5,6. Macrophages are a group of immune cells that participate in host defense and help maintain tissue homeostasis through their unique functions, such as phagocytosis and endocytosis, cytokine secretion, and ROS production, mediating inflammation and regulating iron, lipid, and amino acid metabolism. Additionally, ferroptosis is involved in the pathological processes of various inflammatory diseases7,8,9. Ankylosing spondylitis (AS) is a chronic inflammatory disorder10,11 and is characterized by inflammation in the axial skeleton and sacroiliac joints.
As ferroptosis strongly influences the pathogenesis of inflammatory arthritis, including rheumatoid arthritis, osteoarthritis, gouty arthritis, ankylosing spondylitis, and many other inflammatory arthritis, the development of strategies for treating ferroptosis in patients with inflammatory arthritis may be effective12. Rong et al. divided AS into two subtypes based on the expression of key genes for ferroptosis and found that ferroptosis might play a regulatory role in AS13. The mechanism underlying ferroptosis in either osteoarthritis or ankylosing spondylitis is not fully understood. In another study, we found that neutrophils play an important role in AS6,14,15. Neutrophils can promote ferroptosis resistance of B cells by providing IL-6 through SLC7A11, and inhibition of SLC7A11 can significantly increase the ferroptosis of B cells and reduce B cell proliferation16. Some studies have shown that ferroptosis of neutrophils can lead to immunosuppression17.
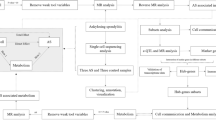
In this study, we assessed and clarified the associations among ferroptosis, AS, and neutrophils by integrating the ferroptosis gene with clinical data, transcriptome data, and single-cell data of AS. The immune role of neutrophils in AS was determined by cell communication analysis18,19,20. In our previous study, we also found that cassia twigs can be used to effectively treat AS; the most important component of cassia twigs is cinnamaldehyde21. Based on these findings, we examined whether the active components in cassia twigs treat AS by acting on the key protein of ferroptosis. The research process is shown in Fig. 1.
Study flowchart.
Materials and methods
Data
This retrospective study adhered to the guidelines of the Declaration of Helsinki, and all participants provided informed consent. Ethical approval was obtained from the FIRST AFFILIATED HOSPITAL of GUANGXI MEDICAL UNIVERSITY ETHICAL REVIEW COMMITTEE (Approval No.2024-E043-01).
Clinical data of patients
From 2012 to 2020, we screened patients based on whole blood routine following the modified New York criteria22 (AS diagnostic criteria) in our Department of Spinal Surgery.
Single-cell sequencing data
The samples used for single-cell sequencing were obtained from bone marrow blood obtained from the spine during surgery. The patient cohort included three patients diagnosed with AS and three patients diagnosed with thoracic/lumbar fractures. Detailed information on clinical patient inclusion criteria, sample processing, single-cell library preparation, and sequencing data processing is provided in the supplementary document.
Datasets
We performed an exhaustive search in the GEO database using the following search query: [“Ankylosing Spondylitis” (MeSH Terms) and mRNA (All Fields) AND “Homo sapiens” (Organism) AND “Expression profiling by array” (Filter)]. We included data from the inception of the database up to August 2021. We selected the GSE 73,754 dataset, based on the GPL10558 Illumina HumanHT-12 V4.0 Expression BeadChip, for our analysis. The genes associated with ferroptosis were obtained from the GSEA database and related references. We collected information on cinnamaldehyde from PubChem (https://pubchem.ncbi.nlm.nih.gov/), Comparative Toxicogenomics Database (CTD, https://ctdbase.org/) gathering and cinnamic aldehyde and SwissTargetPrediction website (http://www.swisstargetprediction.ch/) related genes.
Clinical data analysis
All patients provided clinical and cellular data for this study with informed consent. The study included 106 patients diagnosed with AS and 1333 non-AS patients from our hospital. The data were collected for the period January 2012 to August 2021. A comprehensive set of clinical data, including variables such as age, gender, and blood collection data, was collected for rigorous statistical analysis. The relationships among these variables that met the inclusion criteria are illustrated in Supplementary Fig. 2. To increase the robustness of the analysis, the data were randomly divided into a test group and a verification group (Table 1). The data was analyzed through logistic regression analysis and machine learning analysis.
Ferroptosis analysis
Based on the GSE73754 dataset, we performed GSEA by integrating ferroptosis-related genes and used the limma, tidyr, and ggplot2 packages to calculate cell proportions and perform differential analysis to determine the expression differences of various immune cells between AS and the control group. Subsequently, the association between ferroptosis and immune cells was assessed. By constructing the protein interaction networ, the top 15 genes with degree values were selected as the importance genes of ferroptosis (Supplementary Table 3).
Cell clustering analysis, annotation, and visualization
Batch removal and dimensionality reduction were conducted using the “harmony” package, and the AS group was integrated with the normal control group for common analysis. KNN (Mutual Nearest Neighbor) correction was applied to mitigate potential batch effects again. Unsupervised cell clustering was performed using a graph-based approach with the resolution parameter set to 0.3 to identify different cell clusters. The “Find All Markers” function was used to identify the markers of each cluster. The identification criteria were as follows: (1) log fold change (log FC) > 1, and (2) p-value < 0.05. Additionally, t-SNE analysis, graph-based clustering, and labeling of clusters with the same cell type were performed to optimize cell types.
Drug target analysis and molecular docking
We conducted difference analysis(log FC > 1 and adj P Value < 0.05), Weighted Gene Co-Expression Network Analysis (WGCNA), and GSEA analysis on the downloaded transcriptome data. We identified the genes that were co-expressed by the three and were called AS genes. The selected cinnamaldehyde-related genes and disease genes were first combined for further analysis, and then, the most important target genes were obtained via intersection. Pharmacogenetic analysis and visualization were performed using the DC algorithm and Mcode tools in the STRING online database and cytoscape, these genes were called Drug genes. Clinical performance and expression differences of the genes were evaluated by the ROC curve and violin plots. In addition, docking analysis was performed using Discovery Studio software. The docking results were evaluated based on binding affinity (ΔG), the binding pose of the ligand within the active site of the target protein, hydrogen bonds, hydrophobic interactions, and other molecular contact interactions.
Immunohistochemistry analysis
The study subjects included interspinous ligaments from individuals diagnosed with AS as the experimental group, and interspinous ligaments from non-AS individuals undergoing spinal surgery as the control group. Immunohistochemistry was used to study the differential expression of central genes between the AS group and the control group. The interspinous ligament tissue was carefully processed, and immunohistochemical sections were successfully obtained. This complex process involved steps such as wax embedding, sectioning, antigen retrieval, antibody hybridization, color development, and tissue sealing. The antibodies used were sourced from anti-NFE2L2 (1:500, 80593; Proteintech). The specimens were then examined under a microscope, and images of both the AS and normal control groups were captured. ImageJ software was used to evaluate the positivity of all immunohistochemical images.
Cell communication analysis
Cell communication analysis revealed cell interactions and communication within the single-cell dataset. Based on the ‘cell Chat’ R package, the ‘Cell Chat DB. Human’ ligand-receptor interaction database was used to comprehensively analyze the differences in cell signaling between AS patients and the control group, including the number of cell subpopulations, intercellular communication strength, and functional differences.
Statistical analysis
In this study, statistical analyses were performed using R software (version 4.2.3). Logistic regression analysis, Random forest, SVM and LASSO regression were employed to identify clinical variables associated with AS, and ROC curve analysis was applied to evaluate the predictive ability of these variables. For differential expression analysis, transcriptome data were analyzed using the `limma` package in R, with criteria set as log FC > 1 and adj P Value < 0.05. GSEA and KEGG pathway analyses were conducted to identify key biological pathways related to ferroptosis and immune responses in AS. A t-test was performed to analyze the results of immunohistochemical validation, comparing NFE2L2 expression levels between the AS group and the control group. In single-cell analysis, Wilcoxon rank-sum tests were used to analyze cell subtype proportions to identify significant differences. Cell Chat was utilized to analyze intercellular signaling, and the software’s built-in statistical tests were used to compare interaction intensities.
Results
Neutrophils are a risk factor for predicting AS clinically
In total, 2,108 clinical samples were included, and data from 1,439 patients that met the inclusion criteria were collected. Following randomization principles, the clinical data that met the inclusion criteria were divided into a training group (Normal = 809, AS = 54) and a validation group (Normal = 524, AS = 52). Detailed information on the variables and groups is provided in Table 1. Univariate logistic regression analysis was performed to determine statistically significant differences with p-values < 0.05, and multivariate logistic regression analysis was performed to identify statistically significant differences with p < 0.05. The results of the logistic regression analysis are presented in Supplementary Table 1. To determine the most important variables, we used the random forest algorithm “%IncMSE.” The best regression effect was achieved by retaining the top 10 variables with the highest importance after tenfold cross-validation (Fig. 2A). The LASSO regression analysis of dependent variables (Fig. 2B) showed that 14 variables were significantly correlated with AS. We also used the SVM algorithm to analyze the data through ten-fold cross-validation. We obtained the output vector feature index and sorted the variables in descending order of usefulness, finally obtaining 13 key variables (Fig. 2C).
Neutrophils can serve as predictive risk factors for AS in clinical settings. (A-C) Results obtained from the RF, LASSO and SVM algorithm respectively. (D) The intersection of the four screening methods yielded five risk factors. (E) A dynamic prediction model was established using the selected parameters: PCTE, Pdw, NEUT, GRAN and Age. The red line represents the probability of AS occurrence at different values. (F-H) Calibration curve, ROC curve and clinical impact curve in the test group respectively. (I-K) Calibration curve, ROC curve and clinical impact curve in the verification group respectively.
The variables with the greatest significance, as selected via logistic regression analysis, LASSO, SVM, and random forest, are summarized in Supplementary Table 2. The intersection of the results obtained from these four methods was visualized using a Venn diagram (Fig. 2D), which revealed five predictors: PCTE, Pdw, NEUT, GRAN, and Age. These predictors were used to formulate a predictive model (Fig. 2E). To evaluate the model’s effectiveness, calibration curves (Fig. 2F), ROC curves (Fig. 2G), and clinical impact curves (Fig. 2H) were generated using the test group. The area under the ROC curve (AUC) was calculated to be 0.859, which indicated a favorable predictive performance. The calibration curve analysis showed that the values predicted by the nomogram closely matched the actual measurements. The C-index of the model was determined to be 0.878 (95% CI: 0.826–0.929). For internal validation, the validation group was used. The calibration curves, ROC curves, and clinical impact curves are illustrated in Fig. 2I-K. Through internal validation, the AUC was calculated as 0.807, which indicated a robust predictive performance. The results of the calibration curve analysis confirmed the alignment of the nomogram-predicted values with actual measurements, and the C-index was found to be 0.822 (95% CI: 0.764–0.879). The NEUT and GRAN showed the proportion of neutrophils in the bloodstream. The prediction model emphasized the significant role of neutrophils in AS.
Neutrophils are the key immune cells in AS
After the raw data were screened and normalized, 31,242 high-quality single-cell transcriptomes were obtained from three AS and three control samples. We conducted preliminary dimensionality reduction on six clinical samples, and the results of dimensionality reduction are shown on the left side of Fig. 3A. After integrating three samples, each from the AS group and the control group, dimensionality reduction was performed again. The specific results are shown in the middle and right of Fig. 3A. To eliminate the batch effect during single-cell preparation, we selected the first 2,000 highly variable genes and performed harmonic batch removal and dimensionality reduction on the data again, the results are shown in Fig. 3B and C. Through cell clustering, 16 clustering results were obtained, and the results were visualized using TSNE (Fig. 3D). By annotating the clustering results, six cell types were obtained(Fig. 3E), including monocyte, NK cell, neutrophils, B cell, GMP (granulocyte-monocyte progenitor), and CMP (common myeloid progenitor). To identify the specific immune cells associated with AS, we performed immunoinfiltration analysis on the downloaded dataset GSE73754. The distribution of various types of cells in GSE73754 and the control group is shown in Fig. 3F. The T cell, NK cell, and neutrophil in AS were statistically significant compared to the control group (P < 0.05) (Fig. 3G), and the number of neutrophils was significantly higher in patients with AS. The results of clinical data analysis, single-cell analysis, and transcriptome analysis showed that neutrophils play a key role in the pathogenesis of AS.
Single-cell atlas of AS and annotation, clustering, visualization. (A) Comparison of cell distribution between AS and control group in single cell samples. Each dot represents a cell. (B) Using harmony to eliminate batch effects. (C) Keep the feature variance map of the first 2000 genes. (D) Formation of 16 clusters, with each number corresponding to a unique cluster color. (E) Annotation of the 16 clusters resulting in the identification of 6 distinct cell types. (F) Cell distribution of AS and control group in GSE73754. (G) Immunoinfiltration analysis of GSE73754 showed that the expression of neutrophils in AS was significantly different from that in the control group (P < 0.001).
Ferroptosis gene is involved in the pathogenesis of AS
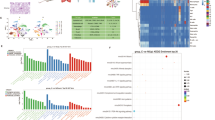
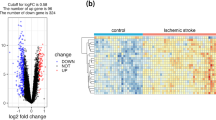
We obtained differential genes (DIFF) by conducting differential analysis and GSEA enrichment analysis on the GSE73754 dataset. The results of GSEA showed the first five cases of intermediate upper and lower regulation of GSE73754, respectively (Supplementary Fig. 1A). The first five upregulated outcomes included acute myeloid leukemia, growth hormone synthesis, secretion, and action, leukocyte transendothelial migration, mucin type O-glycan biosynthesis, and platelet activation. By analyzing the ferroptosis gene set, we found statistically significant differences in the expression of the ferroptosis singnal in AS (P < 0.01) (Supplementary Fig. 1B). We also found that in AS, the expression of neutrophils was negatively correlated with the expression of ferroptosis (Supplementary Fig. 1C), which indicated that ferroptosis was involved in the immune regulation of AS. Thus, we continued our follow-up analysis to test this hypothesis. In the GSEA, a ridge plot was used to show the correlation pathways that were highly correlated with AS in GSE73754, including the FoxO signaling pathway and the JAK − STAT signaling pathway, both of which had P-adj < 0.05 (Supplementary Fig. 1D).
In WGCNA, a clustering tree was obtained (based on the best soft threshold and expression profile data; the top half was the hierarchical clustering tree of genes, and the bottom half was the gene module, which divided the genes near the cluster into one module). The height represented the distance between the observed values and the cluster, and the dynamic tree was the dynamic module corresponding to the height (Supplementary Fig. 1E). Cluster analysis was conducted for six modules. The correlation and confidence of all genes in each module were calculated, and all statistically significant modules (P < 0.05) were selected to conduct the WGCNA. Finally, four modules were obtained (see Supplementary Fig. 1F). By selecting the differential genes and conducting the WGCNA, we obtained the DIFF and WGCNA genes, respectively. Combined with the genes related to ferroptosis, the intersection of the 18 genes was obtained based on the Wayne diagram (Supplementary Fig. 1G). These genes were called the core genes of the disease (see Supplementary Table 4 for the names of the specific genes). Through the GO and KEGG analyses of the disease genes, we found that those genes mainly participated in the biological process of the disease (Supplementary Fig. 3A and B) and were closely related to the ferroptosis signaling pathway (Supplementary Fig. 3C-F).
Cassia twigs can treat AS by acting on NFE2L2, a key gene related to ferroptosis
We found 18 drug target intersections between the two websites based on the Wayne diagram. Due to delays in reliability and information on each drug target web page, we combined the drug targets screened from the two sites under the name drug(Supplementary Table 5). Subsequently, the drug target genes were intersected with the key genes of the disease, and three key genes were obtained, including NFE2L2, GCLM, and TXNRD1. The intersection results are shown in Fig. 4A-B. Through the STRING online analysis tool and the Cytoscape tool, the Cytohub plug-in was used to build a PPI network of interactions between drug targets and disease genes (Fig. 4C). Using the MCODE plug-in, the indirect links between drug targets and other genes were calculated, and five genes (Fig. 4D), including SRXN1, SLC7A11, GLRX, GSTM1, and GSTA4, were obtained, which might affect the efficacy of Cinnamomum cassia in AS through indirect interaction with drug targets. The expression of three drug target genes and five indirectly acting genes in GSE73754 was calculated, and the expression of the drug target in the AS group was found to be statistically significant compared to that in the control group (P < 0.01) (Fig. 4E). The efficacy of drug targets was evaluated by plotting the ROC curve and calculating the area under the curve (AUC). We found that the AUC of GCLM, NFE2L2, and TXNRD1 were 0.746, 0.766, and 0.731, respectively (Fig. 4F-H), these were used as key reference values.
(A) Cinnamaldehyde related genes were screened through two drug component databases, among which 18 overlapping genes and 362 common genes were found. (B) The intersection of cinnamaldehyde related genes and AS differential genes obtained 3 co-expressed drug target genes. (C) The interaction network of cinnamaldehyde genes and disease genes was mapped using cytoscape software. Green for drug genes, red for disease genes. (D) GCLM and TXNRD1 can promote the therapeutic effect of cinnamaldehyde on AS through indirect interaction with other five drug genes. (E) The expression of 3 direct drug targets and 5 indirect drug targets in GSE73754 showed that NFE2L2, GCLM and TXNRD1 were highly expressed in AS, with statistical differences compared with the control group (P < 0.001). (F-H) ROC curves representing GCLM, NFE2L2, and TXNRD1 respectively. NFE2L2 has the highest AUC value of 0.766.
We used the STRING tool and the DC algorithm in Cytoscape to obtain the top 15 genes. By combining single-cell annotation results, we analyzed the interactions of key ferroptosis-related genes within each cell cluster in the AS single-cell sequencing data and found statistically significant results (P < 0.05) (Fig. 5A). We also intersected drug target genes with neutrophil-specific expression genes and identified NFE2L2 as the most important core gene. Similarly, we found that NFE2L2 was statistically significant in most cells in the AS single-cell sequencing data (P < 0.05) (Fig. 5B), and visualized its expression in each cell using t-SNE (Fig. 5C). The expression of the first 15 core genes of ferroptosis in each cell is shown in Supplementary Fig. 4A, we found that NFE2L2 was expressed at higher levels in neutrophils. Therefore, to verify the expression of NFE2L2 in AS, we conducted an immunohistochemistry analysis and found that the expression of NFE2L2 in the AS group was significantly higher than that in the control group (P < 0.001) (Fig. 5D). Based on the results of the previous analysis, we classified the expression of NFE2L2 in neutrophils into two subtypes: the high-expression group and the low-expression group (Fig. 5E). The distribution of the two subtypes is shown in Fig. 5F. To comprehensively understand the differences between the two subpopulations, we conducted GO and KEGG analyses and found no significant difference between them (Supplementary Fig. 4B and C). Finally, to elucidate the interaction between cinnamaldehyde and NFE2L2, we performed molecular docking analysis and found that their binding energy is less than − 3.2 kcal/mol, with significant hydrogen bonds present, indicating that the receptor-ligand complex is relatively stable (Fig. 5G).
(A) The expression of ferroptosis related genes in annotation results was statistically significant in pairwise comparison between cells of the AS single-cell sequencing data (P < 0.05). (B) The expression of NFE2L2 in annotation results was statistically significant in neutrophils compared with other cells of the AS single-cell sequencing data (P < 0.05). (C) NFE2L2 scatterplot expressed in the annotation results, the darker the color, the greater the expression. (D) Immunohistochemistry showed that the expression of NFE2L2 in AS was higher than that in control group, with a statistically significant difference (P < 0.001). (E) According to the expression of NFE2L2, neutrophils were divided into two subgroups. (F) Distribution of neutrophils in two subgroups. (G) The left and right images show the 3D and 2D docking results of cinnamaldehyde with the NFE2L2 molecule. In the 3D diagram, hydrogen bond interactions between the two are shown, with the green intensity indicating stronger interaction strength. In the 2D diagram, clear hydrogen bonds are visible.
Neutrophils are closely related to other cells in AS
To further assess the intercellular interactions in AS, we compared the number and intensity of annotated intercellular interactions in the AS and control groups. We found that the number of cell interactions was 703 in the control group and 587 in the AS group (Fig. 6A). The intensity of cell interaction was 13.631 in the control group and 11.632 in the AS group (Fig. 6B). Additionally, the number and intensity of interactions between neutrophils and other cells were significant in the control and AS groups. By constructing a heat map of cell interactions, we more intuitively found that neutrophils interacted significantly with other cells when they acted as signal transmitters (Fig. 6C-D). We identified and compared the signal paths of the AS group and the control group by comparing the information flow of each signal path. Information flow was defined by the sum of the communication probabilities between all pairs of cells in the network (i.e., the total weight in the network). Important signaling pathways were ranked based on inferred differences in the overall information flow between the AS group and the control group network. We found that the four most prominent signaling pathways in AS included JAM, CSF, SEMA3, and TENASCIN (Fig. 6E). Finally, we analyzed the probability of ligand-receptor pair-mediated communication between neutrophils and other cell populations by comparing them with the control groups. We found that in the AS group, the neutrophils communicated with other cell populations through various signaling pathways and ligand-receptor pairs (P < 0.05) (Fig. 6F).
Comparison of cellular interactions between AS and control group. (A) The left side represents a shell diagram of the number of cell interactions in the AS and control groups, with thicker lines representing more interactions. The interaction bar graph is represented on the right. (B) The left side represents a shell diagram of the intensity of cell interactions in AS and control groups, with thicker lines representing greater interaction intensity. The interaction bar graph is represented on the right. (C) The interaction between cells in AS and control group was shown by heat map. The darker the color, the greater the effect. (D) AS cell interactions with control group. (E) Results of information flow visualization of cell-cell interactions between AS and control group. (F) Comprehensive analysis shows receptor ligand pairs of neutrophils interacting with other cells. The size of the dots represents statistical differences, and the color of the dots represents the importance of the receptor ligand pairs.
Discussion
AS is an arthritic disease that affects the spine and sacroiliac joints23. Many researchers have investigated its immune environment. Most immune cells and cytokines are involved in the pathogenesis of AS. Specifically, the IL-23/IL-17 pathway plays a crucial role in the occurrence and development of diseases24. The immune cells involved in the initiation, progression, and regulation of ankylosing spondylitis include dendritic cells, macrophages, natural killer cells, and neutrophils25. In this study, we found that neutrophils are not only a predictor of the diagnosis of AS but also an important type of immune cell that participates in the immune process of AS. Using single-cell sequencing technology, we found that when compared to the normal group, in the AS group, neutrophils interacted more strongly with other cells and reached other cells through multiple ligand-receptor pairs. Neutrophils are involved in dynamic behavior and immunomodulatory properties of inflammatory processes and are important cells associated with innate and acquired immune responses against infectious agents26,27. The products released by neutrophils during inflammation can also recruit and/or activate other cell types28, such as monocytes, endothelial cells, T and B lymphocytes, NK cells, etc. Thus, the close relationship between neutrophils and human immunity is important for maintaining balance in the body and defending against external threats. Neutrophils are a core member of the immune system and play a vital role in the body’s immune defense29,30. They participate in and regulate the immune response in various ways to effectively respond to invasion and keep the body healthy31. Neutrophils release different types of chemicals at the site of inflammation, including chemokines and cytokines, which guide other immune cells, such as monocytes and lymphocytes, toward the affected area. This process assists the action of the immune system at the site of injury or infection, thus accelerating the recovery and removal of pathogens32. Concerning the regulation of the immune system, neutrophils are also involved in immune cell interactions and signaling33,34. They work together with other immune cells, such as lymphocytes and monocytes, to form a tightly coordinated immune network35. This cell-cell interaction and messaging help in precisely regulating the immune response; thus, ensuring an appropriate and efficient response against different types of threats36.
Ferroptosis is also known as iron-dependent cell death1. Ferroptosis is a specific form of cell death that is closely related to iron accumulation and metabolism. It is considered to be a form of cell death independent of traditional apoptosis, necrosis, and other cell death pathways. In ferroptosis, the accumulation and abnormal oxidation state of iron in cells leads to lipid peroxidation, membrane rupture, and damage to the intracellular organs, ultimately resulting in cell death4. This process is related to lipid peroxidation in the cell membrane and usually involves the breakdown of some antioxidant defense system. Neutrophils are the main performers of the immune system30, and their relationship with ferroptosis is important for understanding immune cell biology and the mechanism underlying the response to infection. Neutrophils release oxygen free radicals in response to infection. In this study, we found a negative correlation between ferroptosis and neutrophils in AS, and the differential genes in AS were significantly associated with the ferroptosis signaling pathway. This suggests that ferroptosis is involved in the immune process of AS and may serve as a prerequisite for neutrophils to exert their immune functions, such as phagocytosis and oxidative killing. Therefore, exploring the role of ferroptosis in the immune response of AS may provide new insights and directions for the treatment of AS.
Cassia twig is a commonly used herb in traditional Chinese medicine37,38. It has multiple functions, such as warming Yang and relieving surface, regulating qi and blood, regulating channels, and activating collaterals. It is closely related to the human immune system39,40, and its characteristics of warming Yang and relieving surface complement each other with the regulatory function of the immune system; these characteristics jointly maintain the overall physical health of individuals. Cinnamomum officinale may have a multilayer relationship with the functions of neutrophils through its multiple effects41,42. In our previous study, we found that cassia twigs are closely related to AS21, and cinnamaldehyde, the main component of cassia twigs, plays the most important role. Our study found that the ferroptosis-related gene NFE2L2 has molecular binding sites with cinnamaldehyde, providing a potential direction and approach for the treatment of AS with traditional Chinese medicine. Several studies have shown that NFE2L2 is frequently activated after infection43. NFE2L2 plays a critical role in ferroptosis through three key pathways: the synthesis and metabolism of glutathione/glutathione peroxidase 4, iron metabolism, and lipid processes44,45,46. It is involved in the pathological processes of various inflammatory diseases47,48,49,50, including systemic lupus erythematosus, atherosclerosis, rheumatoid arthritis, and ulcerative colitis. In addition, studies have shown that NFE2L2 is closely related to immune cells, especially the neutrophils. Abnormal expression of TIPE2 might target the NFE2L2/HO-1 pathway and inhibit M1 macrophage-associated neutrophil inflammation in asthma51. In studies on antineutrophil cytoplasmic antibody-associated vasculitis, it was found that neutrophils deficient in NFE2L2 exhibited enhanced neutrophil extracellular trap (NET) formation, exacerbating the development of vasculitis52. Furthermore, in pharmacological studies on inflammatory diseases53,54,55, numerous drugs have been found to influence neutrophil migration, differentiation, and activity by targeting NFE2L2-related genes or signaling pathways. These studies have shown that NFE2L2 is closely related to immune regulation, and neutrophils. Further studies on the specific role of NFE2L2 in the neutrophils of patients with AS might provide a theoretical basis and drug target for treating such patients.
Conclusion
In this study, we found that the neutrophil content is an important clinical information in AS. Through single-cell sequencing and transcriptome analysis, we also showed that neutrophils are important immune cells in patients with AS and are closely related to other immune cells. By conducting a network pharmacological analysis, we also found that NFE2L2, a key gene related to ferroptosis, is a potential therapeutic target for AS. Cassia twig, a traditional Chinese medicine, can effectively act on NFE2L2-related proteins and neutrophils associated with AS. These findings suggested that cassia twigs can provide a new strategy for treating ankylosing spondylitis.
Limitation
In this study, we integrated single-cell sequencing data, transcriptome data, ferroptosis-related genes, and network pharmacology to identify NFE2L2 as a key ferroptosis-related gene in Neutrophils in AS. We found that the essential component of Cassia twig, cinnamaldehyde, could potentially target NFE2L2 to exert therapeutic effects on AS. However, the molecular docking results revealed that the binding energy between Cassia twig and NFE2L2 was − 3.2 kcal/mol. While this indicates a potential interaction, it does not meet the widely accepted threshold of -5 kcal/mol, which typically signifies strong binding affinity. This limitation underscores the need for further validation to determine the biological significance of this interaction. Future research should build upon the current findings by conducting in vitro and in vivo experiments, such as functional activity evaluations and structural analyses, to further validate the interaction between Cassia twig and NFE2L2, thereby clarifying its therapeutic potential in targeting NFE2L2 for the treatment of AS.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- AS:
-
Ankylosing spondylitis
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- WBC:
-
White blood cell
- RBC:
-
Red blood cell
- Hb:
-
Hemoglobin
- HCT:
-
Hematocrit
- ESR:
-
Erythrocyte sedimentation rate
- Mcv:
-
Mean corpuscular volume
- PLT:
-
Platelet
- MCH:
-
Mean corpuscular hemoglobin
- MCHC:
-
Mean corpuscular hemoglobin concentration
- Mpv:
-
Multi-purpose vehicle
- Pdw:
-
Platelet distributionwidth
- GRAN/NEUT:
-
Neutrophil
- LYMF:
-
Lymphocyte
- LRR:
-
Lymphatic return rate
- MONO/MONO1:
-
Monocyte
- EON/EON1:
-
Eosinophil
- BP:
-
Biological Processes
- CC:
-
Cellular component
- MF:
-
Molecular function
- CI:
-
Confidence interval
- AUC:
-
Area under the curv
- LASSO:
-
Least absolute shrinkage and selection operator
- RF:
-
Random Forest
References
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149 (5), 1060–1072. https://doi.org/10.1016/j.cell.2012.03.042 (2012).
Haschka, D., Hoffmann, A. & Weiss, G. Iron in immune cell function and host defense. Semin. Cell Dev. Biol. 115, 27–36. https://doi.org/10.1016/j.semcdb.2020.12.005 (2021).
Tang, D., Chen, X., Kang, R. & Kroemer, G. Ferroptosis: molecular mechanisms and health implications. Cell Res. 31 (2), 107–125. https://doi.org/10.1038/s41422-020-00441-1 (2021).
Pu, F. et al. Ferroptosis as a novel form of regulated cell death: implications in the pathogenesis, oncometabolism and treatment of human cancer. Genes Dis. 9 (2), 347–. https://doi.org/10.1016/j.gendis.2020.11.019 (2022).
Yang, Y. et al. Interaction between macrophages and ferroptosis. Cell. Death Dis. 13 (4), 355. https://doi.org/10.1038/s41419-022-04775-z (2022).
Jiang, J. et al. Upregulated of ANXA3, SORL1, and neutrophils may be key factors in the Progressionof Ankylosing Spondylitis. Front. Immunol. 13, 861459. https://doi.org/10.3389/fimmu.2022.861459 (2022).
Song, J. et al. Molecular mechanism of interleukin-17A regulating airway epithelial cell ferroptosis based on allergic asthma airway inflammation. Redox Biol. 68, 102970. https://doi.org/10.1016/j.redox.2023.102970 (2023).
Li, M. et al. Ferroptosis triggers airway inflammation in asthma. Ther. Adv. Respir. Dis. 17, 17534666231208628. https://doi.org/10.1177/17534666231208628 (2023).
Zhao, H., Tang, C., Wang, M., Zhao, H. & Zhu, Y. Ferroptosis as an emerging target in rheumatoid arthritis. Front. Immunol. 14, 1260839. https://doi.org/10.3389/fimmu.2023.1260839 (2023).
Lockwood, M. M. & Gensler, L. S. Nonradiographic axial spondyloarthritis. Best Pract. Res. Clin. Rheumatol. 31(6), 816. https://doi.org/10.1016/j.berh.2018.08.008 (2017).
Simone, D., Al Mossawi, M. H. & Bowness, P. Progress in our understanding of the pathogenesis of ankylosing spondylitis. Rheumatology (Oxford, England). 57 (suppl_6), vi4–vi9. https://doi.org/10.1093/rheumatology/key001 (2018).
Chang, S., Tang, M., Zhang, B., Xiang, D. & Li, F. Ferroptosis in inflammatory arthritis: a promising future. Front. Immunol. 13, 955069. https://doi.org/10.3389/fimmu.2022.955069 (2022).
Rong, T., Jia, N., Wu, B., Sang, D. & Liu, B. New insights into the Regulatory Role of Ferroptosis in Ankylosing Spondylitis via Consensus clustering of ferroptosis-related genes and Weighted Gene Co-expression Network Analysis. Genes 13 (8). https://doi.org/10.3390/genes13081373 (2022).
Sun, X. et al. Identification of clinical heterogeneity and construction of a novel subtype predictive model in patients with ankylosing spondylitis: an unsupervised machine learning study. Int. Immunopharmacol. 117, 109879. https://doi.org/10.1016/j.intimp.2023.109879 (2023).
Zhou, C. et al. Immune cell infiltration-related clinical diagnostic model for Ankylosing Spondylitis. Front. Genet. 13, 949882. https://doi.org/10.3389/fgene.2022.949882 (2022).
Wang, Z. et al. Neutrophil-derived IL-6 potentially drives ferroptosis resistance in B cells in lupus kidney. Mediat. Inflamm. 2023, 9810733. https://doi.org/10.1155/2023/9810733 (2023).
Flemming, A. Neutrophil ferroptosis causes immunosuppression. Nat. Rev. Immunol. 23 (1), 6. https://doi.org/10.1038/s41577-022-00820-2 (2023).
An, C., Wang, X., Song, F., Hu, J. & Li, L. Insights into intercellular receptor-ligand binding kinetics in cell communication. Front. Bioeng. Biotechnol. 10, 953353. https://doi.org/10.3389/fbioe.2022.953353 (2022).
Cheng, C., Chen, W., Jin, H. & Chen, X. A review of single-cell RNA-Seq annotation, integration, and cell-cell communication. Cells 12 (15). https://doi.org/10.3390/cells12151970 (2023).
Shao, X., Lu, X., Liao, J., Chen, H. & Fan, X. New avenues for systematically inferring cell-cell communication: through single-cell transcriptomics data. Protein Cell. 11 (12), 866–880. https://doi.org/10.1007/s13238-020-00727-5 (2020).
Wei, W. et al. Network pharmacology combined with molecular docking and experimental validation to explore the potential mechanism of Cinnamomi ramulus against ankylosing spondylitis. Ann. Med. 55 (2), 2287193. https://doi.org/10.1080/07853890.2023.2287193 (2023).
Moll, J. M. & Wright, V. New York clinical criteria for ankylosing spondylitis. A statistical evaluation. Ann. Rheum. Dis. 32 (4), 354–363. https://doi.org/10.1136/ard.32.4.354 (1973).
Ritchlin, C. & Adamopoulos, I. E. Axial spondyloarthritis: new advances in diagnosis and management. BMJ 372, m4447. https://doi.org/10.1136/bmj.m4447 (2021).
Zhu, W. et al. Ankylosing spondylitis: etiology, pathogenesis, and treatments. Bone Res. 7, 22. https://doi.org/10.1038/s41413-019-0057-8 (2019).
Rezaiemanesh, A. et al. Immune cells involved in the pathogenesis of ankylosing spondylitis. Biomed. Pharmacotherapy = Biomedecine Pharmacotherapie. 100, 198–204. https://doi.org/10.1016/j.biopha.2018.01.108 (2018).
Navegantes, K. C. et al. Immune modulation of some autoimmune diseases: the critical role of macrophages and neutrophils in the innate and adaptive immunity. J. Transl Med. 15 (1), 36. https://doi.org/10.1186/s12967-017-1141-8 (2017).
Odobasic, D., Kitching, A. R. & Holdsworth, S. R. Neutrophil-mediated regulation of Innate and adaptive immunity: the role of Myeloperoxidase. J. Immunol. Res. 2016, 2349817. https://doi.org/10.1155/2016/2349817 (2016).
Thieblemont, N., Wright, H. L., Edwards, S. W. & Witko-Sarsat, V. Human neutrophils in auto-immunity. Semin. Immunol. 28 (2), 159–173. https://doi.org/10.1016/j.smim.2016.03.004 (2016).
Schoen, J. et al. Neutrophils’ Extracellular Trap Mechanisms: From Physiology to Pathology. Int. J. Mol. Sci. 23 (21). https://doi.org/10.3390/ijms232112855 (2022).
Rawat, K. & Shrivastava, A. Neutrophils as emerging protagonists and targets in chronic inflammatory diseases. Inflamm. Research: Official J. Eur. Histamine Res. Soc. [et al]. 71 (12), 1477–1488. https://doi.org/10.1007/s00011-022-01627-6 (2022).
Loh, W. & Vermeren, S. Anti-inflammatory neutrophil functions in the resolution of inflammation and tissue repair. Cells 11 (24). https://doi.org/10.3390/cells11244076 (2022).
Meng, Y., Sun, J., Zhang, G., Yu, T. & Piao, H. Approaches for neutrophil imaging: an important step in personalized medicine. Bioengineered 13 (6), 14844–14855. https://doi.org/10.1080/21655979.2022.2096303 (2022).
García-Ortiz, A. & Serrador, J. M. ERM Proteins at the crossroad of leukocyte polarization, Migration and Intercellular Adhesion. Int. J. Mol. Sci. 21 (4). https://doi.org/10.3390/ijms21041502 (2020).
Futosi, K., Fodor, S. & Mócsai, A. Neutrophil cell surface receptors and their intracellular signal transduction pathways. Int. Immunopharmacol. 17 (3), 638–650. https://doi.org/10.1016/j.intimp.2013.06.034 (2013).
Feng, X. et al. CD_99 G1 neutrophils modulate osteogenic differentiation of mesenchymal stem cells in the pathological process of ankylosing spondylitis. Ann. Rheum. Dis. https://doi.org/10.1136/ard-2023-224107 (2023).
Kazzaz, N. M., Sule, G. & Knight, J. S. Intercellular Interactions as regulators of NETosis. Front. Immunol. 7, 453. https://doi.org/10.3389/fimmu.2016.00453 (2016).
Han, R. et al. Conventional disease-modifying anti-rheumatic drugs combined with Chinese Herbal Medicines for rheumatoid arthritis: a systematic review and meta-analysis. J. Traditional Complement. Med. 12 (5), 437–446. https://doi.org/10.1016/j.jtcme.2022.01.005 (2022).
Lee, J. & Lim, S. Anti-inflammatory, and anti-arthritic effects by the twigs of Cinnamomum cassia on complete Freund’s adjuvant-induced arthritis in rats. J. Ethnopharmacol. 278, 114209. https://doi.org/10.1016/j.jep.2021.114209 (2021).
Jia, W. et al. Ramulus Cinnamomi essential oil exerts an anti-inflammatory effect on RAW264.7 cells through N-acylethanolamine acid amidase inhibition. J. Ethnopharmacol. 317, 116747. https://doi.org/10.1016/j.jep.2023.116747 (2023).
Pan, B. et al. Cinnamomi Ramulus inhibits the growth of colon cancer cells via Akt/ERK signaling pathways. Chin. Med. 17 (1), 36. https://doi.org/10.1186/s13020-022-00588-6 (2022).
Yang, C. L. et al. Anti-inflammatory and antibacterial activity constituents from the stem of Cinnamomum validinerve. Molecules (Basel Switzerland). 25 (15). https://doi.org/10.3390/molecules25153382 (2020).
Hagenlocher, Y., Hösel, A., Bischoff, S. C. & Lorentz, A. Cinnamon extract reduces symptoms, inflammatory mediators and mast cell markers in murine IL-10(-/-) colitis. J. Nutr. Biochem. 30, 85–92. https://doi.org/10.1016/j.jnutbio.2015.11.015 (2016).
Sun, Y. et al. Xinfeng capsule inhibits inflammation and oxidative stress in rheumatoid arthritis by up-regulating LINC00638 and activating Nrf2/HO-1 pathway. J. Ethnopharmacol. 301, 115839. https://doi.org/10.1016/j.jep.2022.115839 (2023).
Han, H., Zhang, G., Zhang, X. & Zhao, Q. Nrf2-mediated ferroptosis inhibition: a novel approach for managing inflammatory diseases. Inflammopharmacology 32 (5), 2961–2986. https://doi.org/10.1007/s10787-024-01519-7 (2024).
Dodson, M., Castro-Portuguez, R. & Zhang, D. D. NRF2 plays a critical role in mitigating lipid peroxidation and ferroptosis. Redox Biol. 23, 101107. https://doi.org/10.1016/j.redox.2019.101107 (2019).
Babaei-Abraki, S., Karamali, F., Nasr-Esfahani, M. H. & Ferroptosis The functions of Nrf2 in human embryonic stem cells. Cell. Signal. 106, 110654. https://doi.org/10.1016/j.cellsig.2023.110654 (2023).
Wruck, C. J. et al. Role of oxidative stress in rheumatoid arthritis: insights from the Nrf2-knockout mice. Ann. Rheum. Dis. 70 (5), 844–850. https://doi.org/10.1136/ard.2010.132720 (2011).
Chen, Z. et al. Inhibition of Nrf2/HO-1 signaling leads to increased activation of the NLRP3 inflammasome in osteoarthritis. Arthritis Res. Therapy. 21 (1), 300. https://doi.org/10.1186/s13075-019-2085-6 (2019).
Lin, Z. et al. Artemisinin analogue SM934 protects against lupus-associated antiphospholipid syndrome via activation of Nrf2 and its targets. Sci. China Life Sci. 64 (10), 1702–1719. https://doi.org/10.1007/s11427-020-1840-1 (2021).
Bian, W., Wei, L. & Wang, K. Carthamin yellow alleviates dextran sodium sulfate-induced ulcerative colitis by repairing the intestinal barrier and activating the Nrf2/GPX4 axis. Int. Immunopharmacol. 141, 113020. https://doi.org/10.1016/j.intimp.2024.113020 (2024).
Chen, J. K. et al. Artemisia argyi extract ameliorates IL-17A-induced inflammatory response by regulation of NF-κB and Nrf2 expression in HIG-82 synoviocytes. Environ. Toxicol. 37 (11), 2793–2803. https://doi.org/10.1002/tox.23637 (2022).
Ueda, Y. et al. Transcription factor Nrf2 activation regulates NETosis, endothelial injury, and kidney disease in myeloperoxidase-positive antineutrophil cytoplasmic antibody-associated vasculitis. Kidney Int. 105 (6), 1291–1305. https://doi.org/10.1016/j.kint.2024.03.010 (2024).
Seidel, U., Huebbe, P., Rimbach, G. & Taurine A Regulator of Cellular Redox Homeostasis and skeletal muscle function. Mol. Nutr. Food Res. 63 (16), e1800569. https://doi.org/10.1002/mnfr.201800569 (2019).
Lu, J., Xiong, G., Li, H., Zhang, D. & Zhang, X. Nuclear factor erythroid 2-Related factor 2 intervenes the release of Neutrophil Extracellular traps during Lipopolysaccharide-Induced Acute Lung Injury in mice. Mediat. Inflamm. https://doi.org/10.1155/2024/8847492 (2024).
Li, F. et al. Curculigoside mitigates dextran sulfate sodium–induced colitis by activation of KEAP1–NRF2 interaction to inhibit oxidative damage and autophagy of intestinal epithelium barrier. Int. J. Mol. Med. 52 (5). https://doi.org/10.3892/ijmm.2023.5310 (2023).
Funding
This study was supported by the National Natural Science Foundation of China(Grant/Award number: 8236090175), Joint Project on Regional High-Incidence Diseases Research of Guangxi Natural Science Foundation (Grant/Award number: 2023JJA140227), the “Medical Excellence Award” Funded by the Creative Research Development Grant from the First Affiliated Hospital of Guangxi Medical University, and Guangxi Young and Middle aged Teacher’s Basic Ability Promoting Project(Grant/Award Number: 2023KY0115).
Author information
Authors and Affiliations
Contributions
Conceptualization: Tianyou Chen, Chong Liu; Methodology: Tianyou Chen, Jichong Zhu; Formal analysis and investigation: Chenxing Zhou, Bin Zhang; Writing - review and editing: Tianyou Chen, Chong Liu; Funding acquisition: Chong Liu; Data curation: Zhenwei Yang, Jiang Xue; Resources: Shaofeng Wu, Chengqian Huang; Software: Sitan Feng; Supervision: Xinli Zhan, Chong Liu; Validation: Xinli Zhan, Jiarui Chen; Writing - original draft preparation and Visualization: Jichong Zhu.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
We confirm that all subjects and /or their legal guardians provided written informed consent for participation in this study. Prior approval of the study was obtained from the FIRST AFFILIATED HOSPITAL of GUANGXI MEDICAL UNIVERSITY ETHICAL REVIEW COMMITTEE(Approval No.2024-E043-01). The study complies with the Declaration of Helsinki. Written informed consent was obtained from all individual patients included in the study, including those from whom the single-cell sequencing samples were derived.
Consent for publication
All authors have consented to publication and agree to be accountable for the research presented.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, T., Zhu, J., Zhan, X. et al. Investigating ferroptosis-related genes NFE2L2 in neutrophils for ankylosing spondylitis: therapeutic potential of cassia twigs. Sci Rep 15, 8233 (2025). https://doi.org/10.1038/s41598-025-88775-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-88775-x