Abstract
Previous studies showed that the Eurasian sparrowhawk Accipiter nisus (Linnaeus, 1758) acts as the definitive host of the Sarcocystis halieti (Apicomplexa: Sarcocystidae) Gjerde, Vikøren et Hamnes, 2017. Herein, we report macrocysts of S. halieti in the pectoral muscle of a Sparrowhawk in the west of Iran. Between September 2019 and December 2023, a total of 55 wild birds belonging to 8 orders, 9 families, 18 genera, and 21 species were examined in Hamedan province, western Iran. In one Eurasian sparrowhawk, macroscopic cysts resembling rice grains were observed in the breast muscle of the bird. None of the other examined birds were infested with macrosarcocysts. DNA extracted from sarcocysts was molecularly characterized at four nuclear and mitochondrial DNA markers (the 18 S rRNA, 28 S rRNA, cox1 genes, and the ITS1 region). Sequence analysis confirmed the identity of the macrosarcocysts as Sarcocystis halieti. This is the first report of S. halieti in Asia. Sparrowhawk proved to act as both the definitive and intermediate host of Sarcocystis halieti. The phenomenon of the same species of Sarcocystis serving as both hosts is extremely rare among birds and needs to be studied further.
Similar content being viewed by others

Introduction
The genus Sarcocystis is one of the most species-rich genera in the phylum Apicomplexa, widely distributed worldwide. Members of the genus Sarcocystiscan infect poikilotherm animals (reptiles) as well as birds and mammals, including man1. These parasites have a two-host life cycle, which is based on animal prey-predator relationships. Sexual multiplication of the parasite takes place in definitive hosts, while asexual development occurs in intermediate hosts. The definitive host becomes infected by consuming animal tissues infected by mature sarcocsyts, whereas the intermediate host acquires the parasite by consuming food or water contaminated with sporocysts. Sarcocysts are mainly found in muscle tissues and CNS of the intermediate host, while sporocysts develop in the lamina propriaof the small intestine of the definitive host2.
To date, birds serve as intermediate hosts of 32 valid Sarcocystis species3. Sarcocystis rileyiforms macroscopic cysts resembling rice grains in the breast and other muscles of ducks4. Furthermore, two species, S. horvathi and S. wenzelican parasite chickens; the latter may induce encephalitis2,5. In wild birds, fatal diseases can be caused by S. falcatula and S. calchasi, which have low host specificity and can parasitize birds belonging to several distinct orders2.
Birds of prey are usually the definitive hosts of Sarcocystis species3. However, in the last decade, there has been an increasing number of reports of sarcocysts in tissues of birds of prey, indicating that these birds can serve as true intermediate hosts of Sarcocystis spp. Three species, S. falcatula, S. halieti, and S. wobeserihave been identified in the birds order Accipitriformes2,3,6,7, while S. falcatula and S. halietihave been detected in the birds order Strigiformes8,9.
Species of Sarcocystisare described in intermediate hosts based on morphological features of sarcocysts, including their walls, and genetic characteristics of the parasite investigated mainly by Sanger sequencing of informative loci2,3,10. Morphological examinations rely on light microscopy and/or complex and expensive electron microscopy. However, morphological studies alone are not sufficient to distinguish between evolutionarily close species with very similar morphological characteristics found in closely related hosts. For instance, even though four Sarcocystis species (S. columbae, S. halieti, S. lari, and S. wobeseri) are indistinguishable based on the usual morphological features, they have been discovered in the same species of bird, the herring gull (Larus argentatus)11. Therefore, DNA analysis is widely used in the identification and revision of Sarcocystis species. Notably, more than half of the Sarcocystis species parasitizing birds have been described in the last 15 years, largely due to the progress of molecular approaches and the increasing use of these techniques in parasitology and veterinary science10,11,12,13. Different loci, such as nuclear 18 S rRNA, 28 S rRNA, internal transcribed spacer 1 (ITS1), and mitochondrial cytochrome c oxidase subunit 1 (cox1), are the most widely applied for genetic characterization of Sarcocystis species10,14.
Iran is the 17th largest country in the world, with 550 avian species, almost equal to the richness of birds in Europe15,16. However, there is limited and scanty information about their parasite fauna, especially sarcocystosis, i.e., only four documents reporting Sarcocystis species in chickens, ducks, and pigeons17,18,19,20.
The Eurasian sparrowhawk Accipiter nisus (Linnaeus, 1758) is a relatively small bird of prey belonging to the family Accipitridae21,22. It is a widespread species in the temperate and subtropical parts of the Old World, with an estimated population of 1.5 million birds23. This species mainly preys on small forest birds22. In Iran, it is a fairly common breeding bird in Caspian forests and forests of northern Azarbaijan and a common winter visitor occurring throughout Iran, but chiefly in the north, central, and southwest16. The objective of the present study was to identify macroscopic cysts observed in the pectoral muscle of one Eurasian sparrowhawk necropsied in the west of Iran utilizing molecular methods.
Results
A total of 55 wild birds belonging to 8 orders, 9 families, 18 genera, and 21 species were examined (Table 1). In one Eurasian sparrowhawk bird examined on 13 June 2020, elongated spindle-shaped macroscopic cysts resembling rice grains were observed in the breast muscle of the bird (Fig. 1). None of the other examined birds were infested with macrocysts.
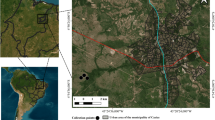
Samples were collected from Hamedan provinces in Iran. The map was drawn by using ArcGIS software version 10.3 (https://enterprise.arcgis.com/en/portal/).
Since the sparrowhawk was presented by the Provincial Department of Environment to the lab in the frozen state, histopathology could not be performed.
Figure 1 a) The Eurasian sparrowhawk (Accipiter nisus) was examined on June 13, 2020. b) Several elongated spindle-shaped macroscopic cysts resembling rice grains (arrows) were observed in the bird’s pectoral muscle.
Genetic identification and characterization of S. halieti from the Eurasian sparrowhawk in Iran
In the current study, partial sequences of four genetic loci of the Sarcocystis parasite from Iran were generated. Specifically, 255 bp 18 S rRNA, 842 bp 28 S rRNA, 828 bp cox1, and 505 bp ITS1 region (427 bp ITS1 and 78 bp 5.8 S rRNA) sequences generated were deposited in NCBI GenBank database under accession numbers (PQ270244 for 18 S rRNA, PQ270246 for 28 S rRNA, PQ276104 for ITS1, PQ274874 for cox1).
Based on a short 18 S rRNA sequence, the sarcocyst isolated from the Eurasian sparrowhawk in Iran showed 100% identity with numerous Sarcocystis spp. using predatory mammals (S. arctica, S. caninum, and S. svanai) or birds (S. calchasi, S. cornixi, S. corvusi, S. fulicae, S. halieti, S. turdusi, and S. wobeseri) as their intermediate hosts and birds as their definitive hosts. The variability of cox1 also appeared to be insufficient to identify the species of the isolated parasite, as 100% identity was found by comparing the sequence of the parasite studied in this work with those of S. calchasi, S. columbae, S. corvusi, and S. halieti (Table 2). Based on 28 S rRNA, the sequences of the studied parasite showed 99.9–100% similarity with S. halieti and less than 99.4% similarity with other Sarcocystis spp. Specifically, 99.3% similarity was established when comparing the analyzed parasite with Sarcocystis sp. ex Corvus corax, S. columbae, and S. corvusi, and 98.2–98.6% similarity was obtained when compared to S. calchasi, S. turdusi, and S. wobeseri. The 427 bp ITS1 sequence generated in the present work showed 96.3% similarity to that of Sarcocystis sp. isolate Skua-2016-CH isolated from the Chilean skua (Stercorarius chilensis), 94.6% similarity to that of Sarcocystis sp. ex Corvus corax, 92.5% similarity to that of Sarcocystis sp. ex Accipiter cooperii, 91.8% similarity to those of S. columbae and 90.5% similarity to that of S. corvusi. In summary, the genetic analysis showed that the sarcocysts found in the muscles of the Eurasian sparrowhawk were identified as S. halieti.
Phylogeny of S. halieti from Iran
In the 28 S rRNA phylogenetic tree, all the analyzed isolates of S. halieti, including the one obtained in the current study, were grouped together with a high bootstrap support (Fig. 2). However, the variability of this locus was not sufficient to determine the most closely related species to S. halieti. Phylogenetic relationships between S. halieti and three other taxa showing high sequence similarity (Sarcocystis sp. ex Corvus corax, S. columbae, and S. corvusi) were not resolved. Despite that, it was demonstrated that the above-mentioned taxa were sister species to S. wobeseri, S. calchasi, S. kutkienae, S. cornixi, S. turdusi, and S. fulicae cluster.
(a) The Eurasian sparrowhawk (Accipiter nisus) examined on June 13, 2020. (b) Several macroscopic cysts resembling rice grains (arrows) were observed in the bird’s pectoral muscle.
Figure 2 The phylogenetic relationships of S. halieti based on 28 S rRNA sequences. The ML tree was scaled according to the branch length and rooted on S. rileyi. GenBank accession numbers are given following the species name. Bootstrap support values ≥ 70 are displayed above branches.
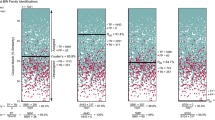
For the ITS1 phylogenetic investigation, only sequences of S. halieti that differed from each other were retrieved from the GenBank. These sequences, in combination with ones belonging to Sarcocystis spp. characterized by bird-bird life cycle and ones of undescribed species showing high similarity to S. halieti were compared with S. halieti from Iran. Altogether, 12 sequences of S. halieti were placed into one cluster with 88 bootstrap support values (Fig. 3). Also, some significant grouping of different S. halieti ITS1 sequences was noticed and analyzed in detail in the following section. Based on the ITS1 partial fragment, Sarcocystis sp. ex Stercorarius chilensis formed a sister branch to S. halieti. These two taxa were sister species to S. corvusi, S. columbae, and Sarcocystis sp. ex Accipiter cooperii. Finally, Sarcocystis sp. ex Corvus corax formed a sister branch to the above-mentioned species. Other examined species using birds as their intermediate and definitive hosts (S. wobeseri, S. calchasi, S. fulicae, S. turdusi, S. cornixi, and S. kutkienae) were phylogenetically distant from S. halieti group and were placed in a separate cluster.
The phylogenetic relationships of S. halieti based on 28 S rRNA sequences. The ML tree was scaled according to the branch length and rooted on S. rileyi. GenBank accession numbers are given following the species name. Bootstrap support values ≥ 70 are displayed above branches.
Figure 3 The phylogenetic placement of S. halieti based on ITS1 sequences. The ML tree was scaled according to the branch length and rooted on S. rileyi. The dashed line indicates that its length does not represent evolution distance. GenBank accession numbers are given after the species name. Bootstrap support values ≥ 70 are shown above branches.
Intraspecific genetic variability of S. halieti
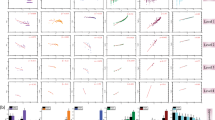
We have used eighty-two 427–428 bp and one hundred forty-five 350–351 bp long ITS1 sequences of S. halieti for the analysis of the intraspecific genetic variability of the species under investigation. In total, 11 (A1-A11) and 12 (B1-B12) different sequence variants of 427–428 bp and 350–351 bp fragments were ascertained. These sequence variants differed by up to eight and nine single nucleotide polymorphisms (SNPs), respectively. The median-joining network analysis showed that the most common A1 and A2 genotypes were found in 16 of 20 examined bird species which served as intermediate and definitive hosts of S. halieti (Fig. 4a). Based on both analyses ITS1 fragments relatively remote from most common genotypes A3 and A6 as well as B3, B4, and B12 were identified only in intermediate hosts of the parasite. Furthermore, all sequences from Brazil made A3, A6 or B3, B12 genotypes which differed from the other genotypes by at least four SNPs and apparently formed a distinct evolutionary lineage (Fig. 4). Based on the larger fragment analyzed (427–428 bp), S. halieti sequence from the Eurasian sparrowhawk in Iran demonstrated 100% identity with S. halieti isolates obtained from sarcocysts of the common starling (Sturnus vulgaris) (GenBank accession no.: MZ333538) and sporocysts of the red kite (Milvus milvus) (GenBank accession no.: MZ333537), both from Check Republic (Fig. 4b). The genotype established in Iran were closely related to genotypes detected in birds of the families Accipirtidae, Corvidae, Laridae, and Phalacrocoracidae from Lithuania and Spain and serving as intermediate hosts of S. halieti.
Figure 4A median-joining network of S. halieti based on 351 bp (a) and 428 bp (b) ITS1 fragments. Hypothetical not-determined genotypes are represented by black rectangles. The circle area is proportional to the genotype frequency (A1 was identified for 97 sequences, A6-A11, B7-12 were detected for a single sequence each). Dashes display mutational steps. The different colors represent different hosts and geographic origins. BR Brazil, ES, Spain, LT Lithuania, CZ Czech Republic, IR Iran, and NO Norway.
The comparison of S. halieti ITS1 sequences originating from Brazil with those originating from Eurasia resulted in lower interspecific variability values (Table 3). However, such results should be considered with caution due to the small sample size from Brazil. Slightly higher genetic variability values were obtained when comparing the sample consisting of sequences obtained from definitive to those obtained from intermediate hosts (K = 2.29474 vs. 2.07192, Hd ± SD = 0.585 ± 0.034 vs. 0.457 ± 0.068, π ± SD = 0.00656 ± 0.00045 vs. 0.00592 ± 0.00095). However, strong differences in genetic variability were observed when comparing the variation of S. halieti in different bird families. Higher variation was observed in birds of the family Accipitridae when compared to those of the family Laridae serving as intermediate hosts of S. halieti, as well as in members of the family Accipitridae when compared to those of the family Corvidae serving as definitive hosts of the parasite. However, the Accipitridae as intermediate host samples were made up of only 4 sequences.
The genetic differentiation analysis based on FST pairwise calculations showed substantial genetic differences between some of the samples compared. Very high and significant genetic differentiation was obtained between Eurasia and Brazil samples (FST = 0.569, P < 0.001), Accipitridae and Laridae intermediate hosts samples (FST = 0.523, P = 0.003), and Accipitridae and Corvidae definitive hosts samples (FST = 0.351, P < 0.001). Moderate and significant genetic differentiation was assessed between intermediate hosts and definitive hosts samples (FST = 0.073, P = 0.001).
Discussion
Herein, we report the first data about the presence and, henceforth, possible circulation of S. halieti in Asia as confirmed by 18 S rRNA, 28 S rRNA, cox1, and ITS1 sequences obtained from macroscopic cysts in the pectoral muscle of the Eurasian sparrowhawk. We also prove that sparrowhawk acts as both the definitive and intermediate host of the parasite. In Iran, although numerous studies reported Sarcocystis species infecting food animals24,25, our study is the first research work on sarcocystosis in wild birds. In the only four previous studies from the country, researchers examined domestic birds, i.e. pigeons Columbia livia domestica (3 studies), chickens Gallus gallus domesticus, and ducks Anas platyrhynchos domesticus (1 study). For the first time, Khordadmehr and colleagues reported that two of 94 pigeons in Yazd province located in the central desert were infected with microscopic Sarcocystiscysts in the tunica muscularis mucosa of the gizzard19. Shortly after that, Shahbazi and her team performed the first PCR-based study on pigeons’ microcysts in a northwestern region and reported a 99% genetic similarity with S. calchasi17. In the same region, Khordadmehr and colleagues reported SarcocystisDNA in the brain and the gizzard of 1 / 100 examined pigeons18. In that study, sequencing of the 28 S rRNA gene product was not successful, but consensus sequences of cox1 displayed 99.1% nucleotide similarity and 96% query cover with two sequences available in the GenBank i.e., S. columbae ex. common wood pigeon Columba palumbus from Lithuania (accession number: MH138312) and S. corvusi ex. Eurasian jackdaw Corvus monedula from Lithuania (accession number: MH138314). In another study, microscopy examination following the digestion method revealed Sarcocystisbradyzoites in the breast muscle of 100% of 18 backyard ducks and 94.78% of 37 backyard chickens in Mazandaran, the northern part of Iran20. Considering the shortage of information about the occurrence, prevalence, and distribution of Sarcocystis species infecting avian hosts in Iran, further studies, especially PCR-sequencing studies on wild birds, are recommended.
Based on the primer pairs used for the genetic characterization of Sarcocystis species at the four most popular loci, i.e., 18 S rRNA, 28 S rRNA, ITS1, and cox1, we obtained partial length sequences of the amplified fragments. This was due to relatively faint PCR fragments, which could be the result of improperly preserved biological material26. Indeed, the infected sparrowhawk was found dead in nature (unknown time-lapse between the death to collection by the Provincial Department of Environment) and was presented to the lab frozen. Nevertheless, the 842 bp and 505 bp long fragments of the 28 S rRNA and ITS1 region, respectively, were sufficiently variable to enable accurate identification of the Sarcocystis species as S. halieti (Table 2; Figs. 2 and 3). In contrast, using 18 S rRNA and cox1 sequences it turned out that the parasite belonged to the genus Sarcocystis and was most closely related to species using birds and predatory mammals as their intermediate hosts. Previously, it was also shown that mitochondrial cox1, as well as the RNA polymerase B gene of the apicoplast genome, are not variable enough for the separation of Sarcocystis species employing birds as their intermediate and definitive hosts10,14,27. Nonetheless, the cox1 marker is suitable for the distinguishment of Sarcocystis species S. anasi, S. albifronsi, S. rileyi, and S. wenzeli, which spread between birds as intermediate hosts and placental mammals as definitive hosts4,5,28,29. The cox1, suggested as a barcoding marker for animals, is used as a golden standard for the identification of Sarcocystis spp. forming sarcocysts in ungulates30,31,32. However, this locus is not variable enough for the separation of Sarcocystisfrom small mammals and predatory mammals33,34. Thus, it is clear that Sarcocystis from birds using different definitive hosts (birds or mammals) have evolved independently of each other at different rates. Studies by other researchers have also confirmed that ITS1 is most suitable for the identification of Sarcocystis spp. characterized by a bird-bird lifecycle12,13,14,35. Also, 28 SrRNA can be used for the diagnosis of these groups of parasites10,36. In the current work, it was demonstrated that ITS1 is more suitable than 28 S rRNA for understanding the phylogenetic relationships of analyzed species. However, it should be noted that due to large ITS1 length variability in different Sarcocystis species, it is hard to generate ITS1 alignment. Therefore, the ITS1 is more suitable for uncovering phylogenetic relationships between closely related taxa than for making phylogenetic inferences about more remote groups10. Based on ITS1 and/or 28 S rRNA, sequences of S. halieti had the highest similarity with several undescribed Sarcocystis species. The phylogenetic analysis showed that S. halieti clearly represents different species compared to Sarcocystis sp. ex Accipiter cooperii and Sarcocystis sp. ex Corvus corax (Fig. 3). Despite the very high 96.3% ITS1 sequence similarity of Sarcocystis sp. ex Stercorarius chilensis with S. halieti, Sarcocystis sp. formed a separate branch in the phylogenetic tree and further investigations are needed to resolve the taxonomic status of this parasite.
The historical confusion on the S. halieti parasite should also be clarified here. Recently, Sarcocystis species have been characterized generally on the basis of the morphology of sarcocysts found in intermediate hosts and the comparison of DNA sequences of several genetic loci13,37. However, this was not the case with S. halieti. Initially, the 18 S rRNA, 28 S rRNA, and ITS1 sequences of Sarcocystis sp., named Sarcocystis sp. ex Phalacrocorax carbo, appeared in the GenBank database in 2012 with accession numbers JQ733511–JQ733513. Sarcocysts of this parasite were isolated from the muscles of the great cormorant collected in Lithuania. However, the name of the parasite was not given, as a morphological characterization of sarcocysts was not performed10. Subsequently, Norwegian researchers identified several Sarcocystis species using DNA sequencing in the intestinal mucosa of one white-tailed sea eagle (Haliaeetus albicilla)14. The comparison of cox1, 18 S rRNA, 28 S rRNA, and ITS1 sequences showed the identity of one of the detected parasites with Sarcocystis sp. ex Phalacrocorax carbo, and this organism was named S. halieti. Only later, the morphology of sarcocysts of S. halietiwas characterized in detail with the help of light and transmission electron microscopy10.
Previously, it was reported that the ITS1 marker shows a high level of intraspecific variability for Sarcocystis spp. using ungulates as intermediate hosts38. Furthermore, for some Sarcocystisspecies of this group, intraspecific and interspecific differences with ITS1 overlap39. Thus, ITS1 is not an appropriate locus for the genetic characterization of Sarcocystis spp. from ungulates30,31,39. In contrast, the intraspecific variability of avian Sarcocystis spp. within ITS1 is relatively low and does not hinder species identification. In general, no or minor (up to 0.1%) intraspecific variation of avian Sarcocystis spp. was observed within rRNA and cox1 loci. Whereas intraspecific differences of these Sarcocystis species in ITS1 do not exceed 3%. Furthermore, based on ITS1, the minimum interspecific difference between two valid Sarcocystis species that are characterized by a bird-bird life cycle is not less than 5%41. So, ITS1 is a suitable locus for the discrimination of these Sarcocystis species. Notably, the highest variability in ITS1 among avian Sarcocystis spp. was observed particularly for S. halieti10,11,40,41,42. These findings enabled us to perform a detailed population genetic analysis of S. halieti samples based on the aligned ITS1 sequences retrieved from GenBank. The median-joining network analysis indicated that several sequences of S. halieti from Brazil represent separate evolutionary lineages (Fig. 4). Furthermore, some evolutionary lineages were observed only in intermediate hosts of S. halieti. Analyses of genetic variability and genetic differentiation also showed that some samples of S. halieti genetically differ depending on geographical and host factors. In general, investigations on the population genetics of Sarcocystis species are limited. The intraspecific genetic variability of these parasites has been examined using amplified fragment length polymorphism (AFLP), microsatellites, and DNA sequence data43,44,45. The dependence of genetic differences on the geography of Sarcocystisparasites has been reported previously1,2,4. Furthermore, it was shown that Sarcocystis species parasitizing the same host may differ in the level of genetic variability11. However, the genetic variability pattern of the same Sarcocystis species in different hosts is inadequately understood and should be further examined. More parasite isolates from the same geographical area but from different hosts would be desirable for such studies.
Recording the Eurasian sparrowhawk as the intermediate host of S. halieti is of significance since this bird species was confirmed as the definitive host of S. halietivery recently46. In Lithuania, 5/16 (31.3%) of the intestinal scraping samples of Eurasian sparrowhawks were positive by molecular analysis for sporocysts/oocysts of S. halieti. In addition to the Eurasian sparrowhawk, sarcocysts of S. halietiwere also found in the muscles of five more species of birds of prey belonging to the families Accipitridae and Strigidae6,9,42,47. To the best of our knowledge, including this study, S. halieti has been detected in 16 species of birds belonging to seven families and six orders (Table 4)6,9,10,11,35,40,41,42,47,48. Our study presents the first identification of S. halieti in Asia while it was previously found in the intestines of 10 birds from the families Accipitridae and Corvidae in Europe and North America (Table 4)14,27,46,48,49,50. However, in the case of corvids, oocysts but not sporocysts were observed in the intestines of birds49. Since the genus Sarcocystis differs from other members of the family Sarcocystidae, such as Toxoplasma, Besnoitia, Hammondia, Neospora, etc., by endogenous sporulation of oocysts in definitive hosts2,51, therefore, transmission experiments are needed to clarify whether corvids are true definitive hosts of S. halieti and several other Sarcocystis spp. forming sarcocysts in the muscles of birds. Intriguingly, three bird species, Eurasian sparrowhawk, common raven (Corvus corax), and hooded crow (Corvus cornix), might potentially serve as both intermediate and definitive hosts of S. halieti41,46,49.
Indeed, the phenomenon of the same species of Sarcocystis serving as both hosts is extremely rare among birds and was previously described only for corvid-associated S. cornixi and S. kutkienaethat were found in both the muscles37,41,52and intestines49 of these omnivorous birds. As discussed above, the latter findings should be checked using transmission experiments. Nevertheless, the phenomenon that some Sarcocystis species might use the same bird species to complete their life cycle, encompassing both sexual and asexual phases, should be addressed in further studies.
Since the description of S. halietiin 201714, new host records have proliferated over several years in different countries and avian families (Table 3). Since this species can form cysts not only in muscles but also in the brain of intermediate hosts and encephalitis, most likely caused by S. halieti has been reported in the little owl (Athene noctua) in Germany9, further research on S. halieti, covering host diversity, intraspecific variability, and histopathology, are needed.
There were some limitations in the present study, such as molecular detection and characterization of Sarcocystis only for the macrocysts and not in different muscle and brain tissues and feces because of resources, and also the impossibility of histopathological and electron microscopy because of the bird’s condition. However, the strength is that we could establish a multi-loci PCR system for the detection of Sarcocystis in domestic and wild animals and birds for the first time in Iran that can facilitate future molecular-based studies in the country and region.
The phylogenetic placement of S. halieti based on ITS1 sequences. The ML tree was scaled according to the branch length and rooted on S. rileyi. The dashed line indicates that its length does not represent evolution distance. GenBank accession numbers are given after the species name. Bootstrap support values ≥ 70 are shown above branches.
Conclusions
Herein, we report the first data about the presence and possible circulation of S. halieti in Asia as confirmed by 18 S rRNA, 28 S rRNA, cox1, and ITS1 sequences obtained from macroscopic cysts in the pectoral muscle of the Eurasian sparrowhawk in Iran. We also prove that sparrowhawk acts as both the definitive and intermediate host of the parasite. The phenomenon of the same species of Sarcocystis serving as both hosts is extremely rare among birds and needs more future investigations. Considering the shortage of information about the occurrence, prevalence, and distribution of Sarcocystis species infecting avian hosts in Iran, further studies, especially PCR and DNA sequencing studies on wild birds, are recommended.
Methods
Study area and sampling
Birds included in this study were examined between September 2019 and December 2023 in Hamedan, Iran (34.7983° N, 48.5148° E) (Fig. 5). Hamedan has a hot-summer Mediterranean climate (Köppen-Geiger classification Csa) with a yearly rainfall of ca. 443 mm. The birds were either euthanized by the Provincial Department of Environment because of general health failure or were found dead in the environment. The time lapse from death to collection of birds could not be estimated; however, only fresh carcasses were examined. Individual birds were sent to the Laboratory of Parasitology, Faculty of Veterinary Medicine, Bu-Ali Sina University, in sealed plastic bags for examination. The bird identifications were made using the reference book Atlas of Birds of Iran15. The birds underwent a standard examination for searching chewing lice (Published elsewhere)53 and then were skinned for the observation of macroscopic sarcocysts. Detected macrosarcocysts were cut with a 1–2 mm margin with sterile scalpel blades and preserved in sterile Eppendorf tubes at −20 C until further examination.
A median-joining network of S. halieti based on 351 bp (a) and 428 bp (b) ITS1 fragments. Hypothetical not-determined genotypes are represented by black rectangles. The circle area is proportional to the genotype frequency (A1 was identified for 97 sequences, A6-A11, B7-12 were detected for a single sequence each). Dashes display mutational steps. The different colors represent different hosts and geographic origin. BR Brazil, ES, Spain, LT Lithuania, CZ Check Republic, IR Iran, NO Norway.
DNA extraction, PCR protocols, and sequencing
Tissue specimens (ca. 200 mg) were minced using sterile surgical scalpel blades in disposable plastic Petri dishes, then transferred into 1.5 Eppendorf microtubes and homogenized using disposable plastic applicators, followed by three freeze/thaw cycles. Finally, the homogenates were vortexed with sterile glass beads. Genomic DNA was extracted using FavorPrep™Tissue Genomic DNA Extraction Mini Kit (Favorgen, Pingtung, Taiwan) according to the manufacturer’s instructions. DNAs were tested for the presence of Sarcocystis spp. DNA by conventional PCRs targeting fragments of the 18 S rRNA of apicomplexan parasites, e.g., Toxoplasma gondii, Neospora caninum, eimeriid coccidia, isosporid coccidia, Sarcocystis, Cryptosporidium and Hammondia51, cox132, 28 SrRNA29,36, and ITS129,50. The primers and conditions utilized in this study are described in Table 5. PCRs were performed using 2 µL from the prepared DNA as the template with the Taq DNA Polymerase Master Mix RED (Ampliqon, Odense, Denmark) in a SimpliAmp thermal cycler (Applied Biosystems, Waltham, MA, USA). For all reactions, the DNA of Sarcocystis bovifelis isolated from one naturally-infected cattle slaughtered in the Hamedan industrial abattoir served as a positive control, and distilled water as the negative control.
The PCR amplification products were documented using a UV Imager (Transilluminator, Vilber Lourmat, France) after electrophoresis in a 1% agarose gel (SinaClon, Tehran, Iran) gels stained with DNA Safe Stain (SinaClon, Tehran, Iran) at 100 V for 60 min. The cPCR amplicons were sequenced by an Applied Biosystems 3500 Genetic Analyzer (Thermo Fisher Scientific, MA, USA) in Codon Genetics Laboratory (Tehran, Iran).
Phylogenetic analysis
The generated DNA sequences were checked for their quality with the help of Chromas 2.6.5 (https://technelysium.com.au/wp/chromas/). Afterward, sequences obtained with forward and reverse primers were assembled into consensus sequences using freely available CLC Sequence Viewer v. 8.0 (QIAGEN, Aarhus, Denmark). The resulting sequences were compared with those of various Sarcocystis spp. available in NCBI GenBank using the nucleotide BLAST, Mega BLAST, and discontiguous Mega BLAST options (http://blast.ncbi.nlm.nih.gov/, accessed on 16 August 2024)54.
The phylogenetic analysis of the 28SrRNA and ITS1 region was performed using MEGA11 v. 11.0.13 software55. Based on BLAST comparison, it was demonstrated that our sequences showed the highest similarity with those of Sarcocystis spp. using birds as their intermediate and definitive hosts. Therefore, these sequences have been taken as ingroup. Meanwhile, sequences of S. rileyi forming macrocysts in the muscles of ducks and transmitted via predatory mammals were chosen as the outgroup for both phylogenetic analyses. The ClustalW algorithm incorporated into MEGA11 was used to get multiple alignments. The final alignment of the partial 28SrRNA fragment contained 19 sequences and 846 nucleotide positions, including gaps. The alignment of the partial ITS1 region lacking the 5’ part consisted of 31 sequences and 528 nucleotide positions, including gaps. The phylogenetic trees were reconstructed using the Maximum Likelihood (ML) method. The Tamura 3 parameter + G (gamma distribution) evolutionary model56 was suggested by MEGA11 based on the calculated lowest Bayesian Information Criterion values of checked models. The robustness of the resulting phylogenies was tested using the bootstrap test (1000 bootstrap replicates).
The analysis of intraspecific genetic variability
To assess geographic, intermediate, and definitive host factors for the intraspecific evolution of the identified Sarcocystis species, we performed a phylogenetic network analysis using NETWORK v. 10.2.0.0 software (https://www.fluxus-engineering.com/sharenet.htm, accessed on 16 August 2024). DnaSP v. 6 software57 was used for the estimation of intraspecific genetic variability of the identified Sarcocystis species. The number of haplotypes (h), the number of polymorphic sites (S), the average number of nucleotide differences (K), the haplotype diversity (Hd), the nucleotide diversity (π), and the standard deviation (SD) for Hd and π were calculated for several subsets divided by the hosts and geographic origin of the parasite. The genetic differentiation between selected sample pairs was evaluated with Arlequin v. 3.5.2.258 using fixation index (FST). The significance of the resulting FST values was tested by 10,000 permutations at the 95% confidence level.
Data availability
The datasets generated and analyzed during the current study are available in the NCBI—GenBank—Nucleotide platform (https://www.ncbi.nlm.nih.gov/genbank/) and can be accessed through accession numbers: PQ270244 for 18 S rRNA, PQ270246 for 28 S rRNA, PQ276104 for ITS1, and PQ274874 for cox1. Any additional data are available from the corresponding author, [AS] on request.
Abbreviations
- PCR:
-
polymerase chain reaction
- cox1:
-
cytochrome c oxidase subunit I
- 18S rRNA:
-
18 S ribosomal RNA
- 28S rRNA:
-
28 S ribosomal RNA
- ITS1:
-
internal transcribed spacer 1
References
Prakas, P. et al. Molecular identification of four Sarcocystis species in cattle from Lithuania, including S. hominis, and development of a rapid molecular detection method. Parasit Vectors 13, 610 (2020).
Dubey, J. P., Calero-Bernal, R., Rosenthal, B. M., Speer, C. A. & Fayer, R. Sarcocystosis of Animals and Humans (CRC, 2016). https://doi.org/10.1201/b19184
Prakas, P., Calero-Bernal, R. & Dubey, J. P. Sarcocystis infection in domestic and wild avian hosts: inseparable flight partners. Vet. Parasitol. 110413 https://doi.org/10.1016/j.vetpar.2025.110413 (2025).
Prakas, P. et al. Sarcocystis spp. macrocysts infection in wildfowl species in Eastern Baltic Region: Trends in prevalence in 2011–2022. Animals 13, 2875 (2023).
Pan, J. et al. Morphological and molecular characterization of Sarcocystis wenzeli in chickens (Gallus gallus) in China. Parasit. Vectors. 13, 512 (2020).
Prakas, P. et al. Molecular identification of Sarcocystis halieti in the muscles of two species of birds of prey from Spain. Parasit. Vectors. 14, 414 (2021).
Shadbolt, T., Pocknell, A., Sainsbury, A. W., Egerton-Read, S. & Blake, D. P. Molecular identification of Sarcocystis wobeseri-like parasites in a new intermediate host species, the white-tailed sea eagle (Haliaeetus albicilla). Parasitol. Res. 120, 1845–1850 (2021).
Wünschmann, A., Rejmanek, D., Cruz-Martinez, L. & Barr, B. C. Sarcocystis falcatula-associated encephalitis in a free-ranging Hornedhorned owl (Bubo virginianus). J. Vet. Diagn. Invest. 21, 283–287 (2009).
Maier-Sam, K. et al. Encephalitis associated with Sarcocystis halieti infection in a free-ranging little owl (Athene noctua). J. Wildl. Dis. 57, 712–714 (2021).
Prakas, P., Butkauskas, D., Švažas, S. & Stanevičius, V. Morphological and genetic characterisation of Sarcocystis halieti from the great cormorant (Phalacrocorax carbo). Parasitol. Res. 117, 3663–3667 (2018).
Prakas, P., Butkauskas, D. & Juozaitytė-Ngugu, E. Molecular identification of four Sarcocystis species in the herring gull, Larus argentatus, from Lithuania. Parasit. Vectors. 13, 2 (2020).
El-Morsey, A. et al. Sarcocystis chloropusae (protozoa: Sarcocystidae) n. sp. from the common moorhen (Gallinula chloropus) from Egypt. Parasitology 142, 1063–1065 (2015).
Máca, O. & González-Solís, D. Sarcocystis cristata sp. nov. (Apicomplexa, Sarcocystidae) in the imported great blue turaco Corythaeola cristata (Aves, Musophagidae). Parasit. Vectors. 14, 56 (2021).
Gjerde, B., Vikøren, T. & Hamnes, I. S. Molecular identification of Sarcocystis halieti n. sp., Sarcocystis lari and Sarcocystis truncata in the intestine of a white-tailed sea eagle (Haliaeetus albicilla) in Norway. Int. J. Parasitol. Parasites Wildl. 7, 1–11 (2018).
Kaboli, M. et al. The Atlas of Birds of Iran (Department of Environment of Iran, Tehran, 2016).
Khaleghizadeh, A. et al. Birds of Iran: Annotated Checklist of the Species and Subspecies (Iranian Research Institute of Plant Protection, Tehran, 2017).
Shahbazi, P., Alizadeh Sefat, M. & Katiraii, F. Molecular detection of Sarcocystis spp. in pigeon. Iran. J. Vet. Med. 12, 93 (2018).
Khordadmehr, M. et al. Natural infection with Toxoplasma gondii, Neospora Caninum and Sarcocystis species in domestic pigeons (Columba livia domestica) in Iran. Comp. Immunol. Microbiol. Infect. Dis. 93, 101946 (2023).
Khordadmehr, M., Ranjbar, V. R., Shahbazi, P. & Zeinoddin, M. Co-infection of Sarcocystis sp. and hadjelia truncata in fantail pigeons (Columba livia Domestica). Bulg. J. Vet. Med. 21, 115–121 (2018).
Vahedi Noori, N., Salehi, A., Razavi, M. & Masoumi, M. Investigation of parasitic Sarcocystis infection in native poultry carcasses in north part of Iran, Mazandaran (Amol). J. Vet. Sci. Med. 7, 3 (2019).
Hagemeijer, E. & Blair, M. The EBCC Atlas of European Breeding Birds: Their Distribution and Abundance (T&AD Poyser, 1997).
Newton, I. Highlights from a long-term study of Sparrowhawks. Br. Birds. 101, 607–623 (2008).
Zawadzka, D. & Zawadzki, J. Breeding populations and diets of the sparrowhawk Accipiter nisus and the hobby Falco subbuteo in the Wigry National Park (NE Poland). Acta Ornithol. 36, 25–31 (2001).
Hamidinejat, H. et al. Prevalence and distribution patterns of Sarcocystis in camels (Camelus dromedarius) in Yazd province, Iran. J. Parasit. Dis. 37, 163–165 (2013).
Hajimohammadi, B. et al. Occurrence and distribution of Sarcocystis parasite isolated from sheep in Yazd Province, Iran. J. Commun. Health Res. 3, 205–210 (2014).
Rojas, A., Germitsch, N., Oren, S., Sazmand, A. & Deak, G. Wildlife parasitology: sample collection and processing, diagnostic constraints, and methodological challenges in terrestrial carnivores. Parasit. Vectors. 17, 127 (2024).
Mayr, S. L. et al. Accipiter hawks (Accipitridae) confirmed as definitive hosts of SarcocTurdusiurdusi, Sarcocystis cornixi and Sarcocystis sp. ex Phalacrocorax carbo. Parasitol. Res. 115, 3041–3047 (2016).
Kutkienė, L., Prakas, P., Sruoga, A. & Butkauskas, D. Description of Sarcocystis anasi sp. nov. and sarcocystis albifronsi sp. nov. in birds of the order Anseriformes. Parasitol. Res. 110, 1043–1046 (2012).
Gjerde, B. Molecular characterisation of Sarcocystis Rileyi from a common eider (Somateria mollissima) in Norway. Parasitol. Res. 113, 3501–3509 (2014).
Rudaitytė-Lukošienė, E. et al. Sarcocystis spp. diversity in the roe deer (Capreolus capreolus) from Lithuania and Spain. Parasitol. Res. 119, 1363–1370 (2020).
Rudaitytė-Lukošienė, E., Prakas, P. & Butkauskas, D. Molecular identification of seven Sarcocystis species in red deer (Cervus elaphus) from Lithuania. Parasitol. Int. 85, 102419 (2021).
Gjerde, B. Phylogenetic relationships among Sarcocystis species in cervids, cattle and sheep inferred from the mitochondrial cytochrome c oxidase subunit I gene. Int. J. Parasitol. 43, 579–591 (2013).
Juozaitytė-Ngugu, E. et al. Identification of Sarcocystis and Trichinella species in muscles of gray wolf (Canis lupus) from Lithuania. Vet. Sci. 11, 85 (2024).
Kirillova, V. et al. Identification and genetic characterization of Sarcocystis arctica and Sarcocystis lutrae in red foxes (Vulpes vulpes) from Baltic States and Spain. Parasit. Vectors. 11, 173 (2018).
Sato, A. P. et al. Molecular characterization of Sarcocystis spp. in seabirds from southern Brazil. Parasitol. Int. 90, 102595 (2022).
Mugridge, N. et al. Phylogenetic relationships of the genus Frenkelia: a review of its history and new knowledge gained from comparison of large subunit ribosomal ribonucleic acid gene sequences. Int. J. Parasitol. 29, 957–972 (1999).
Prakas, P., Butkauskas, D. & Juozaitytė-Ngugu, E. Molecular and morphological description of Sarcocystis kutkienae sp. nov. from the common raven (Corvus corax). Parasitol. Res. 119, 4205–4210 (2020).
Cerqueira-Cézar, C. K. et al. Histopathological, morphological, and molecular characterization of Sarcocystis species in elk (Cervus elaphus) from Pennsylvania, USA. Parasitol. Res. 117, 3245–3255 (2018).
Gjerde, B. Molecular characterisation of Sarcocystis Bovifelis, Sarcocystis bovini n. sp., Sarcocystis hirsuta and Sarcocystis cruzi from cattle (Bos taurus) and Sarcocystis sinensis from water buffaloes (Bubalus bubalis). Parasitol. Res. 115, 1473–1492 (2016).
Juozaitytė-Ngugu, E. & Prakas, P. The richness of Sarcocystis species in the common gull (Larus canus) and black-headed gull (Larus ridibundus) from Lithuania. Parasitologia 3, 172–180 (2023).
Juozaitytė-Ngugu, E., Butkauskas, D., Švažas, S. & Prakas, P. Investigations on Sarcocystis species in the leg muscles of the bird family Corvidae in Lithuania. Parasitol. Res. 121, 703–711 (2022).
Prakas, P. et al. First report of Sarcocystis halieti (Apicomplexa) in bearded vulture (Gypaetus barbatus). Vet. Res. Commun. 48, 541–546 (2024).
Asmundsson, I. M., Dubey, J. P. & Rosenthal, B. M. A genetically diverse but distinct north American population of Sarcocystis neurona includes an overrepresented clone described by 12 microsatellite alleles. Infect. Genet. Evol. 6, 352–360 (2006).
Elsheikha, H. M., Schott, H. C. & Mansfield, L. S. Genetic variation among isolates of Sarcocystis neurona, the agent of protozoal myeloencephalitis, as revealed by amplified fragment length polymorphism markers. Infect. Immun. 74, 3448–3454 (2006).
Stojecki, K., Karamon, J., Sroka, J. & Cencek, T. Molecular diagnostics of Sarcocystis spp. infections. Pol. J. Vet. Sci. 15, 589–596 (2012).
Šukytė, T., Butkauskas, D., Juozaitytė-Ngugu, E., Švažas, S. & Prakas, P. Molecular confirmation of Accipiter birds of prey as definitive hosts of numerous Sarcocystis species, including Sarcocystis sp., closely related to pathogenic S. calchasi. Pathogens 12, 752 (2023).
Llano, H. A. B. et al. Molecular screening for Sarcocystidae in muscles of wild birds from Brazil suggests a plethora of intermediate hosts for Sarcocystis falcatula. Int. J. Parasitol. Parasites Wildl. 17, 230–238 (2022).
Máca, O. & González-Solís, D. Role of three bird species in the life cycle of two Sarcocystis spp. (Apicomplexa, Sarcocystidae) in the Czech Republic. Int. J. Parasitol. Parasites Wildl. 17, 133–137 (2022).
Juozaitytė-Ngugu, E. et al. The role of birds of the family corvidae in transmitting Sarcocystis protozoan parasites. Animals 11, 3258 (2021).
Rogers, K. H., Arranz-Solís, D., Saeij, J. P. J., Lewis, S. & Mete, A. SarcocCalchasilchasi and other Sarcocystidae detected in predatory birds in California, USA. Int. J. Parasitol. Parasites Wildl. 17, 91–99 (2021).
Ho, M. S. et al. Identification of bovine Neospora parasites by PCR amplification and specific small-subunit rRNA sequence probe hybridization. J. Clin. Microbiol. 34, 1203–1208 (1996).
Kutkiene, L., Prakas, P., Sruoga, A. & Butkauskas, D. Sarcocystis in the birds family Corvidae with description of Sarcocystis cornixi sp. nov. from the hooded crow (Corvus cornix). Parasitol. Res. 104, 329–336 (2009).
Bahiraei, Z. et al. Chewing lice of wild birds in Iran: new data and a checklist of avian louse species reported in Iran. Front. Vet. Sci. 10, 1324619 (2024).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 38, 3022–3027 (2021).
Tamura, K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G + C-content biases. Mol. Biol. Evol. 9, 678–687 (1992).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Excoffier, L., Laval, G. & Schneider, S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol. Bioinform Online. 1, 47–50 (2007).
Acknowledgements
The authors wish to thank Mehdi Safikhani, Erfan Moeinifar, Majid Shabanloo, Reza Daneshpajooh, Alireza Mohammadi, Javad Noori Azhar (Provincial Department of Agriculture, Hamedan, Iran), and Mr. Shirdarreh for their kind cooperation in sample collection; Atabak Roohi-Aminjan and Mohammad Babaei (Bu-Ali Sina University, Hamedan, Iran) and Abolghasem Khaleghizadeh (Iranian Research Institute of Plant Protection, Agricultural Research, Education and Extension Organization, Tehran, Iran) for the confirmation of bird species; Zahra Bahiraei, Marzieh Rezaee-Hamid, Fatemeh Nikbin and the staff of Laboratory of Parasitology, Faculty of Veterinary Medicine, Bu-Ali Sina University for their assistance.
Funding
This work was supported by the Bu-Ali Sina University, Hamedan, Iran [Grant Number 99–187, 1400–10209, 2022 − 1490, 2023- 402188 to AS].
Author information
Authors and Affiliations
Contributions
Alireza Sazmand: Conceptualization, Methodology, Investigation, Formal Analysis, Writing - Original Draft, Writing – Review & Editing, Supervision, Project Administration, Funding Acquisition. Leili Moradi: Investigation. Pardis Almasi: Investigation. Reza Nabavi: Investigation. Petras Prakas: Investigation, Formal Analysis, Writing - Original Draft, Writing – Review & Editing.
Corresponding author
Ethics declarations
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Ethics approval
Ethical approval was not required for the study involving animals in accordance with the local legislation and institutional requirements because examined birds were euthanized by a certified veterinarian of the Provincial Department of Environment because of general health failure prior to transfer to the Faculty of Veterinary Medicine, Bu-Ali Sina University for research purposes.
Consent to participate
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sazmand, A., Moradi, L., Almasi, P. et al. First report of Sarcocystis halieti in Asia: the genetic confirmation in muscles of the Eurasian sparrowhawk (Accipiter nisus) from Iran. Sci Rep 15, 6989 (2025). https://doi.org/10.1038/s41598-025-90666-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-90666-0