Abstract
Clade A type 2 C protein phosphatases (PP2CAs) are core regulatory factors in the plant abscisic acid (ABA) signaling pathway. They play crucial roles in response to abiotic stress. However, a comprehensive understanding of the functions of individual members of rice PP2CA gene families remains limited. This study investigates the role of OsPP2C68 in response to abiotic stress. Our findings indicated that the OsPP2C68 is highly expressed in the embryo and endosperm, and is subcellular localized in the nucleus. OsPP2C68 knockout mutants reduced seed germination and root and stem lengths under ABA treatment. The mutants also exhibited higher stomatal closure rates, indicating increased sensitivity to ABA. In addition, the OsPP2C68 knockout mutants exhibited altered synthesis of osmolytes and antioxidant enzymes under drought and high salinity stress, along with the differential expression of genes associated with drought and salt stress responses, enhancing rice tolerance to drought and salt. These results collectively identify OsPP2C68 as a negative regulator in the rice ABA signaling pathway. It is responsive to drought and salt stress, and involved in regulating stomatal movement.
Similar content being viewed by others
Introduction
In natural environments, plants are frequently subjected to various abiotic stresses and other adverse factors, resulting in growth inhibition, developmental delays, reduced yields and loss of ecological value1. The manner in which plants respond to stress factors is intricately complex and initiated by the activation of an effective signaling system2. This condition triggers a series of efficient and intricate signal transduction mechanisms at the physiological, biochemical, and molecular levels to defend against various adversities3.
Abscisic acid (ABA) is a stress hormone in plants that mediates responses to environmental stresses4. When adversity occurs, plants initiate ABA-mediated signaling responses that affect plant growth and survival5. The ABA-dependent signaling pathway includes the following components: ABA receptors PYL/RCARs6, clade A type 2 C protein phosphatases7,8, SnRK29, bZIP transcription factors10 and ABA response cis-element(ABRE). In the absence of ABA, PP2CAs interact with SnRK2s to inhibit their activities, blocking ABA signal transduction11. In the presence of ABA, PP2CAs bind to the ABA receptor PYR1/PYL/RCAR to form a complex. This interaction leads to the activation of SnRK2s, initiating ABA-induced physiological responses12. The PP2CA gene families consists of serine/threonine protein phosphatases that play a crucial role in regulating ABA signal transduction through various mechanisms across different plant organs and tissues. They are integral components of the ABA signaling pathway, primarily exerting negative regulation. PP2CA proteins participate extensively in ABA-mediated processes, such as seed germination, dormancy, stomatal closure, and responses to environmental stresses13,14.
Rice is one of the crucial staple crops worldwide. However, adverse environmental conditions, such as drought, high salinity, and high temperatures, contribute significantly to reduced crop yields, posing a serious threat to food security. As climate change intensifies, these challenges are becoming increasingly detrimental to rice production15,16. At present, numerous PP2C genes have been identified in plants, such as Arabidopsis, rice, wheat, and tomato, which respond to abiotic stresses, such as ABA, drought, salt, and low temperatures17,18. The expression of AtPP2CG1 in Arabidopsis is induced by salt, drought, and ABA treatments. AtPP2CG1 positively regulates salt stress in an ABA-dependent manner19. Rice PP2CA gene families have 10 members20, and the functions of some gene family members have been initially clarified. Some genes have been found to respond to abiotic stresses, but the biological function of OsPP2C68 in rice remains unclear, and a comprehensive understanding is lacking at present.
Our current study focuses on the japonica rice variety, Nipponbare, as the research material. It utilizes biological techniques, such as Cas9 editing, to study the functions of the OsPP2C68. The results indicate that OsPP2C68 functions as a negative regulator in the rice ABA signaling pathway, responding to stresses, such as drought and salinity. Additionally, OsPP2C68 is involved in regulating ABA-mediated stomatal movement. This study enriches our understanding of the biological functions of PP2CA gene families.
Result
Isolation and generation of knockout variants of OsPP2C68
OsPP2C68 (LOC_Os09g15670) is located on rice chromosome 9, with a total length of 1407 bp, a CDS length of 1077 bp, and no intronFig. . 1a). It encodes 358 amino acids. OsPP2C68 has a PP2C structural domain, which is located in amino acid interval 92–352Fig. . 1b). To analyze the function of OsPP2C68, we constructed loss-of-function mutants of the OsPP2C68 transporter protein by using CRISPR-assisted genetic analysis in the background of Nipponbare. We obtained two knockout mutants, pp2c68-1 and pp2c68-2, where single base insertions occurred at the target sites, resulting in an altered amino acid sequence and a premature termination of translationFig. . 1c). To comprehensively study the function of OsPP2C68, we constructed overexpression lines and obtained seven positive plants. However, the overexpression lines exhibited slow growth, yellowing leaves during growth, and then gradual death, making them unsuitable for further research.
The gene structure of OsPP2C68 and CRISPR/Cas9 induced mutation of the OsPP2C68. (a) The gene structure of OsPP2C68. (b) Prediction of protein conserved domain. (c) The OsPP2C68 mutation site is shown on the gene structure. Comparison of base and amino acid sequences of pp2c68-1 and pp2c68-2 with WT. Premature stop codons in the deduced amino acid sequences of the mutant OsPP2C68 proteins are indicated with asterisks.
Expression pattern and subcellular localization of OsPP2C68
Through RT-qPCR, we examined the expression level of OsPP2C68 and found that OsPP2C68 was expressed in roots, stems, flowers, leaves, nodes, shells, embryos, and endosperm. Its expression was highest in embryos and endosperm, while lowest in roots and stem tissues (Fig. 2a). Experiments in which the obtained homozygous OsPP2C68 knockout mutants that contained the proOsPP2C68:GUS vector were stained revealed that OsPP2C68 was expressed in the roots and leaves of seedlings. GUS staining was also observed in the nodes and leaves of mature plants, and in the embryos and endosperm of seeds, with deeper staining observed in embryos and endosperm (Fig. 2b). To determine the subcellular localization of OsPP2C68, the OsLC2-RFP nuclear marker plasmid was co-transferred with OsPP2C68-GFP into rice protoplasts. Observation via laser confocal microscopy indicated that OsPP2C68 was localized in the cell nucleus (Fig. 2c).
OsPP2C68 expression profile in different rice tissues and OsPP2C68 subcellular localization. (a) OsPP2C68 expression levels in mature rice tissues (root, stem, leaf, flower, node, hull, embryo and endosperm) by RT − qPCR. Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p < 0.01). (b) Histochemical GUS staining analysis of proOsPP2C68:GUS transgenic rice seedlings and plants. ((a), seed (b), leaf (c), hull (d), flower (e), node (f), root.) (c) Subcellular localization of OsPP2C68 − GFP in a rice protoplast. bars = 10 μm.
Analysis of the expression levels of OsPP2C68 under different abiotic stress conditions
The promoter analysis results showed that the promoter region of OsPP2C68 contained common cis-regulatory elements, such as CAAT-box and TATA-box, and specific regulatory elements that were responsive to ABA and MeJA (Supplementary Table S1). This result suggested that OsPP2C68 might be involved in responding to various abiotic stresses. To investigate the expression pattern of OsPP2C68 under different abiotic stresses, three-leaf stage seedlings of Nipponbare were treated with 100 µM ABA, 20% polyethylene glycol (PEG), 100 mM NaCl, and low temperature(4℃) at varying durations to analyze the expression level of OsPP2C68. The results indicated that the expression of OsPP2C68 was induced by ABA, PEG, NaCl, and low temperature (Fig. 3a-d).
Expression analysis of OsPP2C68 in different abiotic stresses in OsPP2C68 Knockout Mutants. (a) Expression under 100 µM ABA treatment. (b) Expression under 20% PEG treatment. (c) Expression under 100 mM NaCl treatment. (d) Expression under low temperature treatment. Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p < 0.01).
OsPP2C68 negatively regulates ABA response
To investigate the sensitivity of the OsPP2C68 knockout mutant to ABA, the germination rates of the OsPP2C68 knockout mutant and wild-type seeds under exogenous ABA treatment were statistically analyzed. The germination rates of the mutants pp2c68-1 and pp2c68-2 were the same as those of the wild type on 1/2 MS medium without ABA. On the 1/2 MS medium that contained 1, 2, 5, and 10 µM ABA, the germination rates of mutant seeds were significantly lower than that of the wild-type control (Fig. 4). The difference between the germination rates of the mutant and wild-type seeds gradually increased with an increase in exogenous ABA concentration.
Analysis of germination rate of OsPP2C68 knockout mutants and wild type under ABA treatment. (a) Germination of OsPP2C68 knockout mutants and WT in 0, 1, 2, 5 and 10 µM ABA medium for 6 days. (b-f) Germination rates of OsPP2C68 knockout mutants and WT in media containing 0 µM ABA (b), 1 µM ABA (c), 2 µM ABA (d), 5 µM ABA (e) and 10 µM ABA (f). (g) Seed germination rate of OsPP2C68 knockout mutants and wild type after treatment with 0, 1, 2, 5 and 10 µ m ABA for 108 h. Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p < 0.01).
Analysis and comparison of growth differences between the OsPP2C68 knockout mutants and wild-type seedlings after different exogenous ABA treatments during the seedling stage were conducted. The results showed that in the hydroponic solutions without ABA, no significant difference in growth was found between the wild-type and mutant pp2c68-1 and pp2c68-2 seedlings. However, in the hydroponic solutions with varying concentrations of ABA, the root length, stem length, and fresh weight of the mutants pp2c68-1 and pp2c68-2 were smaller than those of the wild type, and the differences became more evident with increasing ABA concentration. This finding suggests that the knockout of the OsPP2C68 gene enhances rice sensitivity to ABA, indicating that OsPP2C68 negatively regulates response to ABA (Fig. 5a-d).
We further analyzed stomatal status in the leaves of the wild-type and OsPP2C68 knockout mutant plants. No differences were noted in stomatal density and length between the knockout mutant and wild-type plants (Fig. 5f–g). Comparing stomatal aperture between the seedling-stage mutants and wild types under normal conditions, the percentage of closed stomata was slightly higher in the mutants, while the percentage of fully open stomata was lower compared with that in the wild type. In addition, the differences in the percentages of completely closed and fully open stomata between the knockout mutants and wild type became more significant after ABA treatment (Fig. 5h–i). These results indicate that OsPP2C68 can inhibit ABA signal transduction and further regulate stomatal movement mediated by ABA.
Phenotype and stomatal analysis of wild-type and OsPP2C68 knockout mutants under ABA treatment. (a) Growth of WT and OsPP2C68 knockout mutants in 0, 1, 2, 5, and 10 µM ABA hydroponic solution for 5 days. (b–d) Shoot length (b), root length (c), and fresh weight (d) of WT and OsPP2C68 knockout mutants in hydroponics containing 0, 1, 2, 5, and 10 µM ABA. (e) three stomatal types (f) Comparison of leaf stomatal density between WT and OsPP2C68 knockout mutants under normal conditions. (g) Comparison of leaf stomatal length between WT and OsPP2C68 knockout mutants under normal conditions. (h) Percentage of three stomatal types of WT and OsPP2C68 knockout mutants under normal conditions. (i) Percentage of three stomatal types of WT and OsPP2C68 knockout mutants under ABA treatment. Bars = 1 cm Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p < 0.01).
Analysis of drought tolerance in OsPP2C68 knockout mutant plants
To study the response of OsPP2C68 to drought stress, OsPP2C68 knockout mutant materials were subjected to mannitol osmotic stress during the seedling stage, and the root length, shoot length, and fresh weight were measured. The results showed that on 1/2 MS medium that contained 0 mM and 100 mM mannitol, no significant differences occurred in the shoot length, root length, and fresh weight between the mutant lines pp2c68-1 and pp2c68-2, and the wild-type seedlings. However, on 1/2 MS medium that contained 200 mM and 300 mM mannitol, the shoot length, root length, and fresh weight of the mutant lines pp2c68-1 and pp2c68-2 were significantly higher than those of the wild-type seedlings (Fig. 6). This result indicates that OsPP2C68 is involved in rice response to drought stress.
The phenotype identification of wild type and OsPP2C68 knockout mutants under different concentration mannitol treatment for 5 days. (a) The growth of OsPP2C68 knockout mutant and wild type under different mannitol treatment for 5 days. (b-d) Shoot length, root length and fresh weight statistics of OsPP2C68 knockout mutant and wild type under different mannitol treatment for 5 days. Bars = 1 cm Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p < 0.01).
Further experiments were conducted during the seedling stage by using the OsPP2C68 knockout mutant materials under PEG-simulated drought conditions. Wild-type and OsPP2C68 knockout mutant materials were grown in normal hydroponic solution until the three-leaf stage. Then, they were subjected to stress treatment with a hydroponic solution that contained 25% PEG6000 to induce severe leaf dehydration, followed by a recovery period of 10 days in normal hydroponic solution. The results showed that after recovery growth, the OsPP2C68 knockout mutant seedlings retained more green leaves and exhibited increased plant height (Fig. 7a). The survival rate of the pp2c68-1 and pp2c68-2 mutant rice seedlings was significantly higher than that of the wild type (Fig. 7b), indicating that mutation of the OsPP2C68 gene enhanced drought tolerance in rice.
When the wild-type and OsPP2C68 knockout mutant seedlings were grown under soil conditions until the reproductive stage, drought stress was initiated by withholding water until the leaves curled, wilted, and turned yellow, followed by rewatering. As shown in Fig. 7c, no significant difference was found between the OsPP2C68 knockout mutant and wild-type plants before drought stress. After rehydration, significant differences in growth were observed between the OsPP2C68 knockout mutants and wild-type plants. The mutant plants pp2c68-1 and pp2c68-2 had more green leaves, smooth leaves, and sturdy stems, while the wild-type plants exhibited severe leaf curling and even death. These results indicate that the OsPP2C68 knockout mutant plants exhibit drought tolerance, suggesting that OsPP2C68 plays a negative regulatory role in the drought tolerance of rice.
Phenotype of wild type and OsPP2C68 knockout mutants under drought stress. (a) Growth of wild type and OsPP2C68 knockout mutants seedlings under 25% PEG6000 stress. (b) Survival rate of WT and OsPP2C68 mutants testing in (a). Values are means ± SD (n = 3). (c) Phenotype of wild type and OsPP2C68 knockout mutants under soil drought stress. *: Significant level, 0.01 < p < 0.05.
Changes in physiological indicators under drought stress
The contents of free proline (Pro), malondialdehyde (MDA) and the activities of superoxide dismutase (SOD) and catalase (CAT), in the wild-type and OsPP2C68 knockout mutant seedlings were measured before and after drought stress. The results showed that after drought treatment, the Pro content and the activities of SOD and CAT significantly increased in the OsPP2C68 knockout mutants and wild-type seedlings. However, the knockout mutants exhibited significantly higher levels than the wild type. Conversely, the MDA content presented an opposite trend (Fig. 8a–d).
The plant tissues and cell membranes were damaged after stress treatment, leading to electrolyte leakage and reduced chlorophyll content. We separately measured changes in conductivity and chlorophyll content in the OsPP2C68 knockout mutant and wild-type seedlings before and after drought stress. The results showed that after drought treatment, conductivity levels significantly increased while chlorophyll content significantly decreased in the mutant and wild-type seedlings. However, conductivity in the mutant seedlings was significantly lower than that in the wild type, whereas chlorophyll content was significantly higher than that in the wild type (Fig. 8e–f).
The above results indicate that the mutation of OsPP2C68 enhanced the synthesis of osmotic substances and antioxidant enzymes in rice plant and reduced damage to plant tissues and cell membranes, enhancing the drought tolerance of rice.
Analysis of changes in physiological indexes of wild type and OsPP2C68 knockout mutants under drought tolerance. (a) Pro content. (b) MDA content. (c) SOD activity. (d) CAT activity. (e) Relative electrolyte leakage. (f) Chlorophyll content. *Significant level, 0.01< p<0.05. **Extremely significant level, p<0.01.
Analysis of drought-related gene expression levels in the OsPP2C68 knockout mutant
To better explain the enhanced drought tolerance of rice at the molecular level due to OsPP2C68 gene knockout, we selected seven drought-related genes (OsAP37, OsP5CS1, OsNCED1, OsNCED2, OsNCED3, OsNCED4, OsNCED5 and OsABI5). Through RT-qPCR, we measured the expression changes of these genes before and after drought stress in the wild-type and OsPP2C68 knockout mutant plants. The quantitative results showed that before drought stress, no significant difference was found in the expression levels of the seven drought-related genes between the OsPP2C68 knockout mutants and the wild type. After drought stress treatment, the expression levels of these genes were significantly upregulated in the pp2c68-1 and pp2c68-2 mutants and wild type, however, the expression levels of OsABI5 were significantly downregulated (Fig. 9).
Analysis of drought relative gene in wild type and OsPP2C68 knockout mutants. (a) Relative expression of OsAP37, (b) Relative expression of OsP5CS1, (c) Relative expression of OsNCED1, (d) Relative expression of OsNCED2, (e) Relative expression of OsNCED3, (f) Relative expression of OsNCED4, (g) Relative expression of OsNCED5. (h) Relative expression of OsABI5. Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; *p < 0.05, **p < 0.01).
Analysis of salt tolerance in OsPP2C68 knockout mutant plants
Soil salinity is one of the critical factors that threatens rice food security. To investigate the response of OsPP2C68 to salt stress, OsPP2C68 knockout mutant materials were subjected to salt stress during the seedling stage. The results showed that on normal 1/2 MS medium without salt, the wild type and mutant lines pp2c68-1 and pp2c68-2 exhibited similar growth. However, under treatments of different salt concentrations, the shoot length, root length, and fresh weight of rice seedlings from the mutant lines pp2c68-1 and pp2c68-2 were significantly longer and heavier compared with the wild type (Fig. 10a–d). These experimental results indicate that the OsPP2C68 knockout mutants are more sensitive to salt stress during the germination stage.
After approximately 2 weeks of cultivation in hydroponic solution until the seedlings reached the three-leaf stage, the wild-type and OsPP2C68 mutant seedlings were subjected to salt stress by replacing the hydroponic solution with 100 mM NaCl for 4 days, followed by a 7-day recovery period. Upon recovery, the OsPP2C68 knockout mutant seedlings had more green leaves than the wild type (Fig. 10e). In addition, the survival rate of the pp2c68-1 and pp2c68-2 rice seedlings was significantly higher than that of the wild type (Fig. 10f). These results indicate that knocking out the OsPP2C68 gene enhances salt tolerance in rice.
Phenotype of wild type and OsPP2C68 knockout mutant under NaCl treatment. (a) Growth of wild type and OsPP2C68 knockout mutants under different concentration of NaCl treatment for 5 days. (b-d) Shoot length, root length and fresh weight statistics of OsPP2C68 wild type and mutant under different concentration of NaCl treatment for 5 days. (e) Phenotype of wild type and OsPP2C68 mutants before and after 100 mM NaCl treatment. (f) Survival rate of wild type and OsPP2C68 mutants after recovery. *: Significant level, 0.01 < p < 0.05.
Physiological indicators of OsPP2C68 knockout mutant seedlings under salt stress
In salt-stressed seedlings during the trefoil stage, physiological indicators (Pro, MDA, SOD, and CAT) were measured before and after stress in the OsPP2C68 knockout mutants and wild-type japonica rice seedlings. After salt stress treatment, the OsPP2C68 knockout mutants pp2c68-1 and pp2c68-2 exhibited significantly higher Pro content, and SOD and CAT activities compared with the wild type. In addition, MDA content was significantly lower in the knockout mutants compared with the wild type (Fig. 11a–d). Further determination of relative conductivity and chlorophyll content revealed that under 100 mM NaCl stress conditions, the OsPP2C68 knockout mutant seedlings and wild-type japonica rice seedlings demonstrated increased relative conductivity and decreased chlorophyll content. However, the knockout mutant seedlings exhibited lower relative conductivity and higher chlorophyll content compared with the wild type (Fig. 11e–f).
Analysis of changes in physiological indexes of wild type and OsPP2C68 mutant under NaCl tolerance. (a) Pro content. (b) MDA content. (c) SOD activity. (d) CAT activity. (e) Relative electrolyte leakage. (f) Chlorophyll content. *Significant level, 0.01 < p < 0.05.
Analysis of salt-related gene expression levels in OsPP2C68 knockout mutants
To validate the improved salt stress tolerance of OsPP2C68 at the molecular level, we selected nine salt stress-related genes (OsDREB6, OsHKT1, OsNAC9, OsbZIP23, OsbZIP71, OsNCED2, OsNCED3, OsNCED4, and OsNCED5) and detected expression changes in the wild-type and OsPP2C68 knockout mutant seedlings before and after salt stress. Under normal conditions, no significant difference was found in the expression levels of the nine salt-related genes between the mutant and wild-type seedlings. Meanwhile, the expression levels of the nine genes were upregulated in the mutants pp2c68-1 and pp2c68-2, and the wild type; they were significantly higher in the mutant than in the wild type after salt stress treatment (Fig. 12). This result suggests that the nine genes may play roles in the salt stress response regulated by OsPP2C68, or OsPP2C68 may regulate their expression to enhance rice resistance to salt stress.
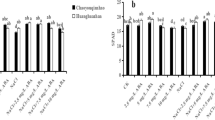
Analysis of relative expression of NaCl relative gene in wild type and OsPP2C68 mutants. (a) Relative expression of OsDREB6, (b) Relative expression of OsHKT1, (c) Relative expression of OsNAC9, (d) Relative expression of OsbZIP23, (e) Relative expression of OsbZIP71, (f) Relative expression of OsNCED2, (g) Relative expression of OsNCED3, (h) Relative expression of OsNCED4, (i) Relative expression of OsNCED5. Data represent mean ± SD (n = 3). Asterisks indicate statistically significant differences (Student’s t-test; * p < 0.05, ** p< 0.01).
Discussion
PP2CAs are important negative regulators in the ABA signaling pathway, and different PP2CA proteins participate in the transduction of ABA signals through distinct mechanisms21,22. The rice PP2CA gene families consists of 10 members. These rice PP2CAs have been reported to be sensitive to ABA. Some members of this family regulate various traits, such as seed germination, plant height, root length, fertility, and thousand-grain weight23,24,25. Research has also found functional redundancy among certain PP2CAs. For example, after knocking out OsPP2C49, sensitivity to ABA did not change compared with the wild type26. This study demonstrates that the knockout mutant of OsPP2C68 exhibits phenotypes related to seed germination, seedling growth, stomatal movement, and responses to abiotic stress, indicating the important role of this gene in regulating ABA signal transduction. In earlier phases of this research, overexpression plants of OsPP2C68 were also generated. However, these materials exhibited slow growth during development and eventually died, preventing further investigation. The reasons behind these observations require further analysis.
ABA is an important regulator of stomatal movement. Research has identified OsPP2C09, OsPP2C50, and OsPP2C53 in rice as major negative regulators of ABA signaling during stomatal closure. In addition, the OsPP2C09 mutant exhibits significantly reduced stomatal length and increased stomatal density25,27. This study found that OsPP2C68 also negatively regulates stomatal movement, but it does not affect stomatal density or length. This result indicates that OsPP2C68 and OsPP2C09 have distinct regulatory mechanisms for stomata.
This study discovered that the expression of OsPP2C68 is induced by drought and salt stress. Following drought and salt treatments, the survival rates of the OsPP2C68 knockout mutants were significantly higher than that of the wild type. The mutants exhibited increased Pro content, enhanced activities of SOD and CAT, and reduced levels of MDA and electrolyte leakage. These findings indicate that the knockout mutants promote the synthesis of osmolytes and antioxidant enzymes under drought and salt stress conditions, enhancing the plant’s ability to scavenge reactive oxygen species and reduce cellular membrane damage, mitigating the detrimental effects of drought and salt stress. Furthermore, under drought and salt stress, the knockout mutants of OsPP2C68 exhibited upregulation in the expression of drought- and salt-related genes. These results indicate that the knockout of OsPP2C68 affects the synthesis of osmolytes and antioxidant enzymes within the plant, and the expression of genes related to drought and salt stress response. Consequently, this condition enhances the drought and salt tolerance of rice plants.
At present, many experimental studies have utilized members of the PP2C family as bait to screen for proteins that interact with PP2Cs in the ABA signaling pathway. This approach aims to identify ABA receptors or downstream transcription factors involved in the ABA signaling pathway. Such effort has significantly advanced the understanding of the functions of PP2CAs and laid the foundation for deciphering the intricate regulatory mechanisms of the ABA signaling pathway. However, further research is necessary to investigate the interacting proteins of OsPP2C68 in the ABA signaling pathway.
Materials and methods
Plant growth conditions and stress treatments
To examine the transcript levels of the OsPP2C68 gene under various abiotic stresses and hormone treatments, the transcript levels of the OsPP2C68 gene under different stress conditions were studied using the japonica rice variety, Nipponbare, as the research material. The seeds were sterilized, allowed to germinate for 2 days at 28 °C, and subsequently grown hydroponically in a growth chamber The hydroponic solution followed the method of the International Rice Research Institute28. During the third-leaf stage of seedling development, treatments with 100 µM ABA, 20% polyethylene glycol (PEG), 100 mM NaCl, and cold stress(4℃) were applied at varying durations. Leaf samples were collected at specified time points, and total RNA was extracted from the leaves for further analysis.
RNA isolation and qRT-PCR analysis
Total RNA was extracted from Nipponbare by using TRIzol reagent, and complementary DNA (cDNA) was synthesized using a Thermo Fisher reverse transcription kit. RT-qPCR was performed using an ABI PRISM 7500 qPCR instrument and ChamQ Universal SYBR qPCR Master Mix. Triple quantification was performed for each cDNA sample. The experiment employed OsActin as an internal reference control for RT-qPCR, and relative gene expression was analyzed using the 2−ΔΔCt method29.
Plasmid construction and plant transformation
To construct the gene knockout strain, we utilized the CRISPR/Cas9 system for targeted genome modification in rice30,31, A 20 bp single guide RNA sequence (GGTGTGCTGTGATTCTGCGG) was cloned into the pOs-sgRNA vector and then subcloned into the Cas9 vector pYLCRISPR/Cas9Pubi-H. The CRISPR/Cas9 construct was transformed into Nipponbare.
A 2000 bp fragment upstream of the OsPP2C68 start codon was amplified from rice genomic DNA. The amplified promoter fragment was digested by EcoRI and HindIII enzymes and cloned into the pCAMBIA1301-GUS vector. The OsPP2C68 promoter-GUS constructs were transformed into Nipponbare, and OsPP2C68 knockout mutants of rice were obtained. All transgenic rice plants were obtained using agrobacterium-mediated transformation32.
GUS staining and subcellular localization
During the rice maturation stage, various tissues from OsPP2C68 promoter-GUS transgenic plants were collected for GUS staining analysis by following the previously described experimental procedure33,34. The OsPP2C68-GFP vector plasmid was constructed by ligating the coding sequence (CDS) of the OsPP2C68 gene to the green fluorescent protein (GFP) expression vector pCAMBIA1390 with a ubiquitin promoter driven. Transfection of the nuclear red fluorescent reference vector plasmid OsLC2-RFP and OsPP2C68-GFP into rice protoplasts was conducted, followed by incubation in darkness at 28 °C for 16–24 h. Observations were made using a laser scanning confocal microscope LSM 880 to identify the site where fluorescence was expressed35,36.
ABA sensitivity assay
The sensitivity of seed germination to ABA was determined by inoculating seeds onto 1/2 MS medium that contained ABA. Fully filled and uniform seeds of the wild type and OsPP2C68 knockout mutants were selected for seed germination assay. The hulled mutant and wild-type seeds were sterilized, dried, and inoculated into 1/2 MS medium without ABA and with 1, 2, 5, and 10 µM ABA for germination, with 20 seeds per dish in three replicates. Seed germination was counted once at 12 h. The seed germination rate was determined by the number of seeds in each dish.
Response of Transgenic plants to abiotic stress
To analyze the gene for stress tolerance, seeds of the wild-type and OsPP2C68 knockout mutants were inoculated and grown on 1/2 MS medium for 3 days at 28 °C in an incubator under 14 h light/10 h dark cycles. Seeds with uniform growth were transferred to 1/2 MS medium that contained different concentrations of ABA, mannitol, and NaCl for 5 days. Then, all the seedlings were counted for germination percentage, shoot length, root length and fresh weight.
In this study, the wild-type and OsPP2C68 knockout mutants were grown in normal hydroponic solution until the three-leaf stage at 28 °C in an incubator under 14 h light/10 h dark cycles. Thereafter, they were transferred to hydroponic solutions that contained 100 µM ABA, 25% PEG, and 100 mM NaCl for stress treatments37,38,39,40. After the appearance of phenotypic differences, the plants were returned to normal hydroponic solution for a recovery period of 7–10 days. Subsequently, the survival rates of rice seedlings were calculated. Survival rate was determined as the ratio of the number of surviving plants in each hydroponic container to the total number of plants subjected to treatment. Each experiment was repeated three times.
Conducting reproductive stage soil drought stress analysis on the wild-type and OsPP2C68 knockout mutants involved growing seedlings under soil conditions until the reproductive stage. The seedlings were subjected to water drought stress until leaf wilting, curling, and yellowing occurred. Subsequently, they were rewatered for recovery growth. Finally, phenotypic data were measured and statistically analyzed to assess the effects of drought stress on both genotypes.
Observation of rice leaf stomata
We selected rice flag leaves of roughly the same size, washed the middle part of the back of each leaf with distilled water, dried the leaves, and then evenly applied a 2–3 cm long layer of transparent nail polish. After the nail polish had dried to form a film, we slowly peeled it off by using tweezers and placed it on a microscope slide for observation. The types of rice stomata could be classified into three categories: fully open, partially open, and fully closed stomata. We observed the types of rice stomata under a microscope, evaluating stomatal density (number/mm²) by using scanning electron microscopy images of the wild-type and OsPP2C68 knockout mutant leaf samples. Each measurement was repeated using three random fields of view (each with an area of 0.24 mm²). The length and width of the stomata were measured using the same stomatal apparatus, conducting three independent repetitions.
Enzyme activity assay
The activities of superoxide dismutase (SOD), peroxidase (POD), and catalase (CAT) were determined in accordance with their respective assay kit instructions (Jiancheng Bioengineering Institute, Nanjing, China). Plant tissue was ground in ice-cold phosphate buffer (pH 7.4) at nine times the volume (1 g tissue: 9 mL buffer), followed by centrifugation of the sample at 3500 rpm for 10 min. Absorbance of the supernatant was measured at 420, 550, and 595 nm.
Determination of free proline, malondialdehyde and chlorophyll content
Free Pro content was measured in accordance with the method used by Song et al.41. MDA content was measured following the method described by Duan et al.42. The chlorophyll content of rice plants was measured by grinding 0.1 g of fresh leaves, and the absorbance of each sample was tested via colorimetry at 652 nm by using a UV 2400 ultraviolet/visible spectrophotometer43。.
Relative conductivity measurement
The procedure involved weighing 0.1 g of clean rice leaf samples by using an electronic balance. The leaves were then cut into approximately uniform-sized fragments by using scissors and placed in 15 mL centrifuge tubes that contained 10 mL of double-distilled water. The tubes were left to soak overnight for 12 h. After soaking, the electrical conductivity (EC) of the solution was measured using a conductivity meter and recorded as R1. A boiling water bath was boiled for 0.5 h. After the water had cooled down, conductivity value was measured and recorded as R2 (relative conductivity = R2/R1 × 100%).
Data analyses
All data were expressed as mean standard deviation and represented by error bars. Data analysis was completed by comparing raw data from all individuals by using the Microsoft Excel program. Graphs were created using GraphPad Prism 8.0 and Adobe Photoshop.
Primers
The sequences of the primers used in the study are listed in Supplementary Table S2.
Data availability
All data were available with in this article and its supplementary files. All constructs and transgenic plants are available upon request. The OsPP2C68 gene sequences are available from the China Rice Data Center (http://www.ricedata.cn/index.htm) using the accession numbers LOC_Os09g15670. Source data are provided with this paper.
References
Zhu, J. K. Abiotic stress signaling and responses in plants. J. Cell. 167, 313–324 (2016).
Nykiel, M., Gietler, M., Fidler, J. & Prabucka, B. &M. Labudda, abiotic stress signaling and responses in plants. J. Plants 12, 3405 (2023).
Pérez-Clemente, R. M. et al. Biotechnological Approaches to Study Plant Responses to Stress. J. BioMed Research International. 1–10 (2013) (2013).
Leung, J. et al. Arabidopsis ABA response gene ABI1: features of a calcium-modulated protein phosphatase. J. Sci. 264, 1448–1452 (1994).
Chen, K. et al. Abscisic acid dynamics, signaling, and functions in plants. J. J. Integr. Plant. Biology. 62, 25–54 (2020).
Xiaojie et al. Characterization and functional analysis of pyrabactin Resistance-Like abscisic acid receptor family in rice. J. Rice. 8, 1–13 (2015).
Singh, A., Giri, J. & Kapoor, S. Tyagi &G.K. Pandey, protein phosphatase complement in rice: genome-wide identification and transcriptional analysis under abiotic stress conditions and reproductive development. J. BMC Genomics. 11, 1–18 (2010).
Kuhn, J. M., Boisson-Dernier, A. & Dizon, M. B. Maktabi &J.I. Schroeder, the protein phosphatase AtPP2CA negatively regulates abscisic acid signal transduction in arabidopsis, and effects of abh1 on AtPP2CA mRNA. J. Plant. Physiol. 140, 127–139 (2006).
A, K., I, W. & E, K. D. G, SnRK2 protein kinases-key regulators of plant response to abiotic stresses. J. Omics: J. Integr. Biology. 15 (12), 859–872 (2011).
Nijhawan, A., Jain, M., Tyagi, A. K. & Khurana, J. P. Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. J. Plant. Physiol. 146, 333 (2008).
Umezawa, T. et al. Type 2 C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. J. Proc. Natl. Acad. Sci. 106, 17588–17593 (2009).
Hauser, F., Waadt, R. & Schroeder, J. I. Evolution of abscisic acid synthesis and signaling mechanisms. J. Curr. Biology. 21, 346–355 (2011).
Nakabayashi, K., Okamoto, M., Koshiba, T., Kamiya, Y. & Nambara, E. Genome-wide profiling of stored mRNA in Arabidopsis thaliana seed germination:: epigenetic and genetic regulation of transcription in seed. J. Plant. J. 41, 697–709 (2005).
Wasilewska, A. et al. An update on abscisic acid signaling in plants and more. J. Mol. Plant. 1, 198–217 (2008).
Ortas, I., Rafique, M. & Çekiç, F. Ö. Do mycorrhizal Fungi enable plants to Cope with abiotic stresses by overcoming the detrimental effects of salinity and improving drought tolerance?? J. Symbiotic Soil. Microorganisms. 60, 391–428 (2021).
Zhou, H. et al. Insights into plant salt stress signaling and tolerance. J. J. Genet. Genomics. 51, 16–34 (2024).
Tör, M., Lotze, M. T. & Holton, N. Receptor-mediated signalling in plants: molecular patterns and programmes. J. J. Experimental Bot. 60, 3645–3654 (2009).
Li, A., Wang, X., Leseberg, C. H., Jia, J. & Mao, L. Biotic and abiotic stress responses through calcium-dependent protein kinase (CDPK) signaling in wheat (Triticum aestivum L). J. Plant. Signal. Behav. 3, 654–656 (2008).
Liu, X. et al. AtPP2CG1, a protein phosphatase 2 C, positively regulates salt tolerance of Arabidopsis in abscisic acid-dependent manner. J. Biochem. Biophys. Res. Commun. 422, 710–715 (2012).
Xue, T. et al. Genome-wide and expression analysis of protein phosphatase 2 C in rice and Arabidopsis. J. BMC Genomics 9, 1–21 (2008).
Hirayama, T. & Shinozaki, K. Perception and transduction of abscisic acid signals: keys to the function of the versatile plant hormone ABA. J. Trends Plant. Sci. 12, 343–351 (2007).
Saez, A. et al. Gain-of-function and loss-of-function phenotypes of the protein phosphatase 2 C HAB1 reveal its role as a negative regulator of abscisic acid signalling. J. Plant. J. 37, 354–369 (2004).
Bhatnagar, N. et al. The protein phosphatase 2 C clade A protein OsPP2C51 positively regulates seed germination by directly inactivating OsbZIP10. J. Plant. Mol. Biology. 93, 389–401 (2016).
Song, J. et al. PROTEIN PHOSPHATASE 2C08, a negative regulator of abscisic acid signaling, promotes internode elongation in rice. J. Int. J. Mol. Sci. 24, 10821 (2023).
Miao, J. et al. OsPP2C09, a negative regulatory factor in abscisic acid signalling, plays an essential role in balancing plant growth and drought tolerance in rice. J. New. Phytologist. 227, 1417–1433 (2020).
Liu, K. et al. Histone deacetylase OsHDA706 increases salt tolerance via H4K5/K8 deacetylation of OsPP2C49 in rice. J. J. Integr. Plant. Biology. 65, 1394–1407 (2023).
Min, M. K. et al. Two clade a phosphatase 2Cs expressed in guard cells physically interact with abscisic acid signaling components to induce stomatal closure in rice. J. Rice. 12, 1–13 (2019).
Li, J. et al. The OsAKT1 channel is critical for K + uptake in rice roots and is modulated by the rice CBL1-CIPK23 complex. J. Plant. Cell. 26, 3387–3402 (2014).
Dooms, M., Chango, A. & Abdel-Nour, A. Quantitative PCR (qPCR) and the guide to good practices MIQE: adapting and relevance in the clinical biology context. J. Ann. De Biol. Clinique. 72, 265–269 (2014).
Xie, K., Zhang, J. & Yang, Y. Genome-wide prediction of highly specific guide RNA spacers for CRISPR–Cas9-mediated genome editing in model plants and major crops. J. Mol. Plant. 7, 926 (2014).
Usman, B., Nawaz, G., Zhao, N., Liu, Y. & Li, R. Generation of high yielding and fragrant rice (Oryza sativa L.) lines by CRISPR/Cas9 targeted mutagenesis of three homoeologs of cytochrome P450 gene family and OsBADH2 and transcriptome and proteome profiling of revealed changes triggered by mutations. J. Plants. 9, 788 (2020).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. J. Plant. J. 6, 271–282 (1994).
Rovira, A., Sentandreu, M., Nagatani, A., Leivar, P. & Monte, E. The sequential action of MIDA9/PP2C.D1, PP2C.D2, and PP2C.D5 is necessary to form and maintain the Hook after germination in the dark. J. Front. Plant. Sci. 12, 636098 (2021).
Ai, P. et al. Two rice phosphate transporters, OsPht1; 2 and OsPht1; 6, have different functions and kinetic properties in uptake and translocation. J. Plant. J. 57, 798–809 (2009).
Li, S. et al. Short panicle1 encodes a putative PTR family transporter and determines rice panicle size. J. Plant. J. 58, 592–605 (2009).
Wang, X. et al. Rice potassium transporter OsHAK8 mediates K + Uptake and translocation in response to low K + Stress. J. Front. Plant. Sci. 12, 730002 (2021).
Susilawati, P. N. et al. Application of consecutive polyethylene glycol treatments for modeling the seminal root growth of rice under water stress. J. Sci. Rep. 12, 2096 (2022).
Samtani, H., Sharma, A. & Khurana, P. Overexpression of HVA1 enhances drought and heat stress tolerance in Triticum aestivum doubled haploid plants. J. Cells 11, 912 (2022).
Yang, S. et al. A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice. J. BMC Plant. Biology 19, 1–15 (2019).
Huang, Y. et al. 9-cis-Epoxycarotenoid dioxygenase 3 regulates plant growth and enhances Multi-Abiotic stress tolerance in rice. J. Front. Plant. Sci. 9, 162 (2018).
Song, S. Y., Chen, Y. & Chen, J. Dai &W.-H. Zhang, physiological mechanisms underlying OsNAC5-dependent tolerance of rice plants to abiotic stress. J.Planta 234, 345 (2011).
Duan, J. et al. OsMIOX, a myo-inositol Oxygenase gene, improves drought tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L). J. Plant. Sci. 196, 143–151 (2012).
Heath, R. L. & Packer, L. Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. J. Archives Biochem. Biophys. 125, 185–198 (1968).
Acknowledgements
This work is supported by the Natural Science Foundation of Hunan Province in China (2022JJ30377).
Author information
Authors and Affiliations
Contributions
L.C. participated in the selection of articles. B.W. and Y.L. conceived and conducted the experiments; B.Z. and H.X. participated in analyzing the results; J.C., F.W., W.L., M.L. provided assistance during the experiments. B.W. and Y.L. wrote the paper. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, B., Luo, Y., Zhong, B. et al. The abscisic acid signaling negative regulator OsPP2C68 confers drought and salinity tolerance to rice. Sci Rep 15, 6730 (2025). https://doi.org/10.1038/s41598-025-91226-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-91226-2