Abstract
Cells continuously sense their surroundings to detect modifications and generate responses. Very often changes in extracellular concentrations initiate signaling cascades that eventually result in changes in gene expression. Increasing stimulus strengths can be encoded in increasing concentration amplitudes or increasing activation frequencies of intermediaries of the pathway. In this paper we show that the different way in which amplitude and frequency encoding map environmental changes endow cells with qualitatively different information transmission capabilities. While amplitude encoding is optimal for a limited range of stimuli strengths, frequency encoding can transmit information with equal reliability over much broader ranges. The qualitative difference between the two strategies stems from the scale invariant discriminating power of the first transducing step in frequency codification. The apparently redundant combination of both strategies in some cell types may then serve the purpose of expanding the span over which stimulus strengths can be reliably discriminated. In this paper we discuss a possible example of this mechanism in yeast.
Similar content being viewed by others
Introduction
Cells continuously sense their surroundings to detect modifications and generate appropriate responses. Environmental changes are often due to changes in concentrations which induce further changes in intracellular components producing a signaling cascade. There are different strategies that cells use to “interpret” and react to environmental changes. On occasions, the intensity of the external stimulus is encoded in the amplitude of the concentration of active molecules in the following steps of the signaling pathway1,2,3. In others, it is encoded in the frequency with which the molecules of one or more steps switch between being active and inactive4,5,6,7. When the end response involves modifications of gene expression these different strategies may result in different dynamics of transcription factor (TF) nuclear translocation. Namely, even upon stepwise changes in the concentration of extracellular effectors, the nuclear TF fraction can remain elevated for a certain time (mimicking the dynamics of the environment) or display a non-trivial pulsatile behavior6,7,8,9. The question then arises of how the two types of encoding differ and under what circumstances one of them could be better suited than the other6,10,11,12.
Previous studies7,13,14 showed that cells may use the same TF to modulate the expression of different sets of genes depending on the TF’s nuclear translocation dynamics (amplitude or frequency). Dynamics may then serve the purpose of multiplexing information transmission15. What matters in this case is whether different external stimuli can be reliably encoded in different dynamics of TF’s nuclear fractions allowing their identification14. In a previous work16 we focused on the gene regulatory part of the process and approached the problem applying information theory17 to the steps that go from the TF’s nuclear fraction to mRNA production. More specifically, we computed the Mutual Information (MI) between the amplitude, duration or frequency of the TF’s nuclear fraction and the mRNA produced. Given two random variables, X and Y, MI quantifies the amount of information that is gained, on average, about one of them from observing the other. Namely, it can be expressed as \(MI=H(X)-H(X\vert Y)=H(Y)-H(Y\vert X)\) where H(X) and H(Y) are the marginal entropies of the two variables and \(H(X\vert Y)\) and \(H(Y\vert X)\) are the conditional ones (see Methods for a mathematical definition in terms of the probabilities of the variables). MI has been widely applied across various fields due to its ability to detect dependencies without assuming any specific model for the data. MI has proven to be very versatile in uncovering intricate relationships in complex biological systems. It has been used to infer gene regulatory interactions18,19 and to quantify the information transmitted between neurons or between stimuli and neural responses, providing insights into neural coding and brain connectivity20,21. In our previous work16, using a two state promoter model, we found that the parameters that maximized MI lied in the same region of parameter space for both amplitude and frequency encodings and that the two strategies mainly differed in their sensitivity to changes in promoter parameters16, making frequency modulation better suited for signal identification without the need to incorporate extra regulatory motifs. In the present paper, we broaden our perspective and analyze the differences and similarities between both encodings when the signaling network between the stimulus and the TF is included. The main result derived from this study is that amplitude and frequency encodings have qualititavely different information transmission capabilities due to the different way in which they map external stimuli. Namely, this different mapping equips frequency encoding with a scale invariant discriminating power of stimuli strenghts as opposed to an intensity-dependent power for amplitude codification. In this way, when considering the processing from external stimulus to gene expression, amplitude encoding is optimal for a limited range of stimuli strengths, while the information transmission capability of frequency encoding can remain relatively invariant with this strength, depending on the various timescales involved. The combination of both mechanisms to encode the same type of stimulus, which has been observed in certain systems, can then serve to enlarge the range over which stimulus strengths can be reliably discriminated. In this paper we discuss one possible such example in S. cerevisiae cells.
Expanding signaling capabilities with dynamics22 is characteristic of the universal second messenger calcium, Ca2+, which encodes different inputs in different spatio-temporal distributions of its free cytosolic concentration23,24,25 and differentially regulates gene expression depending on its dynamics26,27. Interestingly, Ca2+ is involved in various signaling pathways that result in TF’s nuclear fractions pulsatile behaviors6,8,28. Although TF oscillations might not be a mere reflection of those of intracellular Ca2+, they share some common properties. Sequences of intracellular Ca2+ pulses elicited by constant concentrations of external effectors have been observed to be very stochastic in different cell types, particularly in those in which Ca2+ release from the endoplasmic reticulum through Inositol 1,4,5-trisphosphate (IP3) receptors (IP3Rs) is involved29,30,31. It was argued32 that this stochasticity occurs because pulses arise via random Ca2+-channel openings which yield localized Ca2+ elevations that eventually nucleate to produce a global increase in Ca2+. This behavior is characteristic of spatially extended excitable systems in which “extreme events” (Ca2+ spikes or pulses) are triggered by noise and subsequently amplified through space33,34. We briefly remind here that excitable systems are characterized by a stable stationary state and a threshold which, if surpassed due to a perturbation, a long excursion in phase space (a spike) is elicited before the system relaxes to its stable fixed point35. Excitability has been associated with intracellular Ca2+ patterns36,37,38,39 and with the dynamics of pulsatile TFs40,41. In the case of Ca2+ pulses, the interspike time intervals have been observed to be the sum of a fixed component (due to spike duration and refractoriness) and a stochastic one of average, \(T_{av}\), that decreases exponentially with the effector’s concentration31. We have recently derived this dependence using a noise-driven excitable model of intracellular Ca2+ pulses42, both numerically and analytically applying Kramer’s law43. A similar dependence can be obtained for other noisy excitable systems44. These studies imply that, when pulse sequences occur in noise-driven excitable systems, an exponential dependence between the mean interpulse frequency and stimulus strength can be expected. Interestingly, the TF Crz1 in yeast exhibits bursts of nuclear localization6 whose mean frequency can be shown to increase exponentially with extracellular Ca2+, the external effector in this case. Other TFs that exhibit pulsatile nuclear localization have frequencies that are convex increasing functions of the external stimulus strength7,9 and might, in principle, depend exponentially on such strength. We have not seen a thourough analysis of this dependence outside the realm of Ca2+ signaling, but based on this discussion, here we assume that frequency encoding entails an exponential dependence of mean frequency with external input strength42,45.
In this paper we use the simplest possible model to compare the information transmission capabilities of amplitude and frequency encoding upon a constant external stimulus. It is a model with a first step in which the TF’s nuclear fraction is related to the external stimulus with a different mapping and nuclear TF dynamics depending on the codification. The analytic results that we present only depend on this first step. For the simulations with which we analyze the information transmission from the external stimulus to gene expression this first step is supplemented with the simple transcription model considered in our previous work16 which goes from the nuclear TF concentration to the mRNA produced over a fixed time frame. The only difference between amplitude and frequency encoding in this second part is due to the different nuclear TF dynamics. For the transcription step we use a two-state promoter model with a TF dependence of the transition rates originally obtained from data fitting13 which we derived from a mechanistic model16 and parameter values that are based on our previous studies16. We then compute the mutual information, MI, between the mRNA produced and the stimulus strength, \(I_{ext}\), using various \(I_{ext}\) distributions that are defined over the same compact support but differ in their mode, mean and median while keeping approximately the same variance. The first indication of the qualitative difference between amplitude and frequency encoding is reflected in the different dependence of MI with the median of the \(I_{ext}\) distribution obtained numerically. We then derive analytic results which show that the reason for this difference can be traced back to the different properties of the mapping from stimulus strength to nuclear TF’s fraction of both encodings which endows frequency codification with a scale invariant discriminating power. We then study how the subsequent steps in the processing of the response can limit this scale invariance identifying two key limiting timescales. We finally discuss how the combination of both encodings can expand the range over which external stimulus strengths can be reliably discriminated focusing on a possible example in the pheromone respose pathway in cells of S. cerevisiae.
Methods
The model
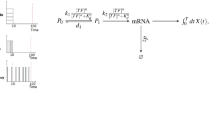
We show in Fig. 1a scheme of the model considered in the paper. A constant external stimulus can either elicit a single pulse of TF nuclear traslocation (amplitude encoding) or a sequence of pulses (frequency encoding). Namely, during the simulation time, \(0\le t\le T=100\, {\text{min}}\), the TF nuclear concentration, [TF], is given by:
where H is the step function (\(H(x)=1\) for \(x\ge 0\) and 0 otherwise) and \(\zeta\) and the inter-pulse time interval, \(\tau \equiv t_{i+1}-t_i\), are random variables described later in more detail. The external stimulus strength, \(I_{ext}\), determines \(A_{TF}\) and \(\langle \tau \rangle\) for each encoding, respectively, according to:
where \(EC_{50}\), h, \(T_{{\text{min}}}\), \(\kappa\) and b are fixed parameters. In the model all the concentrations, \(I_{ext}\), and the parameters in Eqs. (3) and (4), with the exception of \(T_{{\text{min}}}\), are dimensionless. The rationale for choosing Eqs. (1)–(4) as well as the meaning of the various parameters are explained later in this Section. We have not included a detailed dynamical system to derive TF from \(I_{ext}\) since it would only add to the transient of the TF dynamics without affecting the (100 min) mRNA time integral that we take as the output of the process ( see e.g., the example of Msn2 in yeast where the transients last for \(\sim 5-8\,{\text{min}}\)7).
[TF] modulates the transcription rate according to the model depicted in the second row of Fig. 113,16, where \(P_0\) and \(P_1\) represent the two (effective) states of the promoter and transcription proceeds only when the promoter is in the \(P_1\) state at rate, \(k_2 [TF]^nP_1/(K_d^n+ [TF]^n)\) (as explained later, we also use \(P_0\) and \(P_1\) to denote the probabilities that the system is in one or the other state), while mRNA is degraded at rate, \(d_2\). This part of the model as well as the indicator of gene expression (the accumulated mRNA) are the same as those used in our previous study16 where we performed an extensive search of optimal parameter values for information transmission through the transcription step. The parameters of the transcription step used in the present paper were chosen based on this previous study. We describe the model in more detail in what follows.
The model. An external stimulus elicits a single pulse or a sequence of pulses of nuclear TF translocation in amplitude and frequency encoding, respectively. The stimulus strength, \(I_{ext}\), (the input) determines the mean concentration, \(A_{TF}\), of the single TF pulse and the mean time, \(\langle \tau \rangle\), between successive pulses in amplitude and frequency encoding, respectively, according to the expressions inside the blue box (Eqs. 3 and 4). The rationale for these choices as well as the meaning of the various parameters are explained later in Methods. The resulting nuclear TF concentration, [TF], depicted for each encoding in the blue box, modulates the transcription rate according to the two-state promoter model presented in the second row which, in turn, determines the mRNA time integral that is taken as the indicator of gene expression, i.e., the output (third row). The transcription model is the same as the one considered in our previous work16. We recall here that \(P_0\) and \(P_1\) denote the two promoter states as well as the probability that the promoter is in each of these states, that the two arrows connecting \(P_0\) and \(P_1\) in the scheme represent the transition between the two states while the arrow that goes from \(P_1\) to mRNA is used to mean that the mRNA production proceeds at a rate given by \(k_2 [TF]^nP_1/(K_d^n+ [TF]^n)\).
Transcription
Transcription is modeled equally for frequency and amplitude encoding using the effective two-state promoter model13,16 depicted in the second row of Fig. 1. Using \(P_0\) and \(P_1\) to denote the probabilities that the promoter is in one or the other of its two states, noticing that \(P_0=1-P_1\), and recalling that the arrow connecting \(P_1\) with mRNA in the scheme means that the mRNA production proceeds at rate, \(k_2 [TF]^nP_1/(K_d^n+ [TF]^n)\), the dynamical equations that are used to simulate the model read:
where [TF] is the nuclear TF concentration (in dimensionless units), \(P_1\) represents the probability that the promoter is in its active TF-bound state and X(t) is the random number of mRNA molecules at time, t, that can take on the values \(N=\{0,1,2\ldots \}\). Equation (5) is a deterministic equation for the probabilities and Eq. (6) is a stochastic equation for the random variable, X. The extensive search of “optimal” parameter values performed before for the transcription model16 showed that MI achieves its maximum in the same bulk region of parameter space for amplitude and frequency encoding. The search did not yield sharp maxima either16. Thus, in this paper we use a fixed set of parameter values for the transcription step within this previously determined region, limiting the exploration of parameters to those determining the mapping from the external stimulus or the \(I_{ext}\) distribution. The simulations are then performed with time step \(dt=0.1\,{\text{min}}\), integration time \(T=100\,{\text{min}}\), \(n=10\), \(K_d=40\) (in the same dimensionless units as [TF]), \(k_1=1/{\text{min}}\), \(d_1=0.01/{\text{min}}\), \(k_2=10/{\text{min}}\) and \(d_2=0.12/{\text{min}}\) and using Euler’s method to integrate Eq. (5). In all cases, the output is:
From the external stimulus to the nuclear TF
For amplitude modulation, [TF] is modeled as a single pulse of 10 min duration and (dimensionless) amplitude (see Eq. (1)):
with \(\zeta\) a Gaussian distributed random variable of standard deviation, \(\sigma _\zeta =10\), and \(A_{TF}\) related to the external input strength, \(I_{ext}\), (typically, the dimensionless concentration of an external ligand), by the cooperative Hill function (3). Equation (3) commonly describes the input-output patterns observed in signaling cascades46 and, in the context of the present model, aggregates in one step the various processes that go from the external stimulus to the TF’s nuclear fraction. \(EC_{50}\) and \(I_{ext}\) are measured in the same dimensionless units, which not necessarily coincide with those of [TF].
For frequency modulation, [TF] is modeled as a sequence of 1 min-duration square pulses of (dimensionless) amplitude 100 plus a random variable, \(\zeta\), as in the case of amplitude encoding (see Eq. (2)), and stochastic interpulse time intervals,
with \(T_{{\text{min}}}\) fixed31 and \(\eta\) exponentially distributed with rate parameter exponentially dependent on \(I_{ext}\)42,45 so that:
with \(\kappa , b >0\). In this case b is measured in the inverse of the (dimensionless) units of \(I_{ext}\).
Input, output and mutual information
In this paper we compute MI between the external stimulus strength, \(I_{ext}\), and the mRNA produced, O, over the time course of the simulation (Eq. (7) with \(T=100\,{\text{min}}\)). MI can be written in terms of the marginal and joint probability densities of these two variables, \(p_I\), \(p_O\) and \(p_{I,O}\), respectively, as:
MI is computed numerically using an \(I_{ext}\) distribution of compact support, \([I_m,I_M]\), and the Jackknife method47,48. This method is commonly used to derive entropy estimates since it reduces the bias of the Maximum Likelihood (ML) estimator of \(x\log {x}\)47,48. It consists of taking a linear regression of the ML estimator for various sample sizes, with the slope being the bias of the ML estimator and the intercept being the Jackknife estimator. For the computation we used \(T_{{\text{min}}}=5, 10\, {\text{min}}\), various values of \(\kappa\) and the \(I_{ext}\) distribution, \(p_I\), derived from either one of these expressions:
with \(0\le x\le 1\) a stochastic variable distributed according to the Beta-distribution:
for different choices of \(\alpha , \beta >1\) s.t. \(\alpha + \beta \ge 3\), so as to obtain different values for the median of \(I_{ext}\). The Beta-distribution (Eq. 14) is often used to model fractional quantities. It is the posterior distribution that is obtained when applying a Bayesian approach to estimate the probability of a Bernoulli process from \(\alpha +\beta -2\) observations49. Assuming that the external stimulus corresponds to a ligand that, upon binding to a membrane receptor, triggers a signaling cascade, the probability of the Bernoulli process can be interpreted as that of a ligand’s molecule being within a certain (interaction) distance from the membrane (when Eq. (12) is used). This probability will be proportional to the ligand’s concentration. Fixing \(\alpha +\beta\) and choosing different values of \(\alpha\) will then correspond to having different (mean) ligand concentrations nearby the cell. This is illustrated in Fig. 2.
Cumulative distribution function (CDF) of the stimulus strength, \(I_{ext}\), obtained with Eq. (12) (a), (c) and Eq. (13) (b) combined with Eq. (14) for 10 equally spaced \(\alpha\) values increasing from left to right and \(\alpha + \beta\) fixed ((a), (b): \(1.2\le \alpha \le 3\) and \(\alpha +\beta =4\); (c): \(1.4\le \alpha \le 5\) and \(\alpha +\beta =6\)). Varying \(\alpha\) and \(\beta\) for increasing values of \(\alpha +\beta \ge 3\) yields broader ranges of the \(I_{ext}\) median and standard deviation (with smaller values for the latter: \(\sim 0.19-0.22\) in (a) and \(\sim 0.14-0.19\) in (b)). Values such that \(\alpha +\beta = 4,5\) present a good balance of medians that cover relatively well the [0,1] interval with relatively invariant and not too small variances.
As shown in Fig. 2, the range of median \(I_{ext}\) values that can be explored by changing \(\alpha\) when Eq. (12) is used varies approximately linearly between 0 and 1. In order to explore a wider range of values, we decided to probe Eq. (13) for which the range varies logarithmically (Fig. 2b). Increasing values of \(\alpha +\beta \ge 3\) yield more widely spread medians and smaller values of the standard deviation as illustrated in Fig. 2. In the paper we show the results obtained with the distributions of Fig. 2a because their medians cover reasonably well the whole interval, [0, 1], with a relatively invariant and not too small standard deviation (between 0.19 and 0.22).
Computation of the distribution function of the intermediaries of the response
In the approach followed in this paper the intermediaries of the response are the amplitude, A, or the interpulse time, \(\tau\), (or, equivalently, the frequency \(\nu \equiv 1/\tau\)) of the TF’s nuclear concentration in the case of amplitude and frequency encoding, respectively. The amplitude, A, is given by Eqs. (3), (8) where \(\zeta\) is normally distributed with standard deviation, \(\sigma _\zeta\). This implies that A is Gaussian distributed with mean \(A_{TF}\) and standard deviation, \(\sigma _\zeta\). The interpulse time, \(\tau\), is given by Eq. (9) with \(\eta\) exponentially distributed with mean, \(\langle \eta \rangle = T_{IP}-T_{{\text{min}}}\), where \(T_{IP}\) is given by Eq. (10). A and \(\tau\) (or \(\nu\)) can be thought of as the “input” of the transcription model. Given that their distributions are known for a given value of the external stimulus, \(I_{ext}\), and that the \(I_{ext}\) distribution is known as well, it is possible to derive the A and \(\tau\) or \(\nu\) distributions that are fed into the transcription model in our approach. In particular, in the paper we show the results obtained for the choice of \(I_{ext}\) distribution given by Eqs. (12) and (14). In such a case, the cumulative distribution functions (CDF) of A and \(\nu\) can be computed as:
with f given by Eq. (14), \(A_{TF}\) by Eq. (3), \(T_{IP}\) by Eq. (10) and where we are assuming in Eq. (15) that the integral in a from \(-\infty\) to 0 is negligible for most values of A. We use these expressions to compute numerically these two CDFs (which are shown in Fig. 4c,d in the "Results" section).
Results
Amplitude and frequency encodings yield qualitatively different dependences between MI and stimulus strength
We show in Fig. 3 the values, MI (Eq. 11), obtained for amplitude encoding using the 10 \(I_{ext}\) distributions of Fig. 2a, plotting MI as a function of the corresponding medians, Med\((I_{ext})\), for different choices of h and \(EC_{50}\) in Eq. (3). Qualitatively similar results were obtained for the distributions of Fig. 2b and for other values of h and \(EC_{50}\) (see Supplementary note). We observe in Fig. 3 that MI is maximum at Med\((I_{ext})\le EC_{50}\) which approaches \(EC_{50}\) from below as h increases (Fig. 3b). It is also apparent that MI remains within a small percent of this maximum for a limited range of Med\((I_{ext})\) (around \(EC_{50}\) for h large enough, as illustrated in Fig. 3a).
Mutual Information between O (Eq. 7) and \(I_{ext}\) for the 10 \(I_{ext}\) distributions of Fig. 2a as a function of Med(\(I_{ext}\)). (a) and (b) correspond to amplitude encoding for which Eq. (3) was used with \(h=8\) and \(EC_{50}=0.25, 0.5, 0.75, 1\) in (a) and with \(EC_{50}=0.5\) and \(h= 1, 2, 8\) in (b). (c) and (d) correspond to frequency encoding for which Eq. (10) was used with \(T_{{\text{min}}}=5\,{\text{min}}\), \(b=4\), \(\kappa = 9\) (squares), \(T_{{\text{min}}}=5\,{\text{min}}\), \(b=6\), \(\kappa = e^4\) (circles) and \(T_{{\text{min}}}=10\,{\text{min}}\), \(b=4\) and \(\kappa = 9\) (asterisks) in (c) and with \(T_{{\text{min}}}=5\,{\text{min}}\), \(b = 4\) (squares), \(b= 6\) (triangles) and \(\kappa = \exp (b)\) in (d). In all cases, the results are depicted with symbols and joined by curves for the sake of clarity .
The situation observed in Fig. 3a,b for amplitude encoding is qualitatively different from the one derived for frequency encoding, provided that \(\kappa\) and b in Eq. (10) are such that the bulk of the \(T_{IP}\) distribution allows the discrimination of nearby mean frequencies and includes values that are not too large, so that the probability of eliciting one pulse during the finite time of the simulation (100 min) is non-negligible. These two conditions can be satisfied simultaneously, as illustrated in Fig. 3c where we observe that MI can remain close to its maximum value for a broader range of external input strengths than in the case of amplitude encoding. We will discuss later under what conditions MI for frequency encoding can remain relatively constant as Med\((I_{ext})\) is varied, something that is not always satisfied as illustrated in Fig. 3d. We focus in what follows on the reasons that underlie the qualitative differences between amplitude and frequency encoding which limit the information transmission capabilities of the former to a narrower range of stimulus strengths than for the latter.
The qualitative difference between amplitude and frequency encoding derives from the different way in which they map the environment
The different dependence of MI with Med\((I_{ext})\) for amplitude and frequency encoding can be traced back to the different way in which the set of external inputs is “mapped” on the subsequent steps. As we show in what follows, while a mapping in the form of Eqs. (3), (8) only allows to discern a relatively narrow set of external inputs around or below \(EC_{50}\), Eqs. (9) and (10) can map the whole set of external inputs onto a set of discernible interpulse time distributions.
Let us consider two nearby values, \(I_{ext}\) and \(I_{ext}+\Delta _I\). In the case of amplitude encoding (Eqs. 3, 8), these two values will yield two Gaussian distributions of standard deviation, \(\sigma _\zeta\), centered around \(A_{TF}(I_{ext})\) and \(A_{TF}(I_{ext}+\Delta _I)\), respectively. This first step will allow \(I_{ext}\) and \(I_{ext}+\Delta _I\) to be distinguishable if the ratio,
satisfies
In the case of frequency encoding (Eqs. 9 and 10) \(I_{ext}\) and \(I_{ext}+\Delta _I\) will yield two exponential distributions for the “shifted” interpulse time, \(\tau -T_{{\text{min}}}\) of mean \(T_{IP}(I_{ext})-T_{{\text{min}}}\) and \(T_{IP}(I_{ext}+\Delta _I)-T_{{\text{min}}}\), respectively. These two distributions will be distinguishable if the quantiles, p and \(1-p\), of each of them be further apart. This is satisfied if
for some \(p>1/2\) (e.g., \(p=3/4\) guarantees that the overlap of the two distributions does not exceed 1/4 of the total probability).
Equations (18) and (19) are qualitatively different: \(\Delta _A\) is a non-monotone function of \(I_{ext}\), while \(\Delta _F\) does not depend on \(I_{ext}\). For \(h=8\), for example, the term multiplying \({\Delta _I}/{EC_{50}}\) in Eq. (18) attains its maximum value at \(x\equiv I_{ext}/EC_{50}\approx 1\) and decays by 50% at \(x\approx 0.77\) and \(x\approx 1.20\). Thus, the first step of amplitude encoding (Eqs. 3, 8) for \(h=8\) would distinguish values, \(I_{ext}\), that differ among themselves by \(\sim 0.2 EC_{50}\) if \(0.77<I_{ext}/EC_{50}<1.20\). This is consistent with the behavior of MI vs Med\((I_{ext})\) in Fig. 3a. A similar discernment is achieved over the ranges \(I_{ext}/EC_{50}\le 0.4\) for \(h=1\) and \(0.17<I_{ext}/EC_{50}<1.4\) for \(h=2\), consistently with the results of Fig. 3b. Equation (19), on the other hand, shows that for large enough b, the first step of frequency encoding, Eqs. (9) and (10), will allow the discernment of nearby input strengths with the same resolution across the whole range of \(I_{ext}\) values. This qualitative difference is also found in the Kullback-Leibler divergence (KL) (a measure of the statistical distance) between the conditional distributions for \(I_{ext}\) and \(I_{ext}+dI\). Namely, KL depends on \(I_{ext}\) for amplitude encoding (KL=\(2\Delta _A^2\) in this case) while it does not for frequency encoding for which it reads:
Conditional means as functions of \(I_{ext}\) of the TF amplitude, A (Eq. 8), and the interpulse frequency, \(\nu \equiv 1/\tau\), (\(A_{TF}\) and \(\overline{\nu }\), respectively, depicted with solid curves in (a) and (b)), and inverse of the conditional mean of the interpulse time, \(\tau\) (\(1/T_{IP}\), depicted with a dashed curve in (b)) and corresponding CDFs of A (c) and \(\nu\) (d) derived using the \(I_{ext}\) distributions of Fig. 2a with \(\alpha =1.2\) (solid line), \(\alpha =1.4, 2\) (dashed lines), \(\alpha = 2.4\) (dashed-dotted line) and \(\alpha =3\) (dotted line). The parameters used for the \(I_{ext}\)-TF mappings are \(EC_{50}=0.5\) and \(h=8\) in (a) and (c) (same parameters as the curve with solid circles in Fig. 3a) and \(T_{{\text{min}}}=5\,{\text{min}}\), \(b=4\) and \(\kappa =9\) in (b) and (d) (same parameters as the curve with open squares in Fig. 3c) .
Figure 4 illustrates the different behavior of amplitude and frequency encoding. We have plotted in Fig. 4a the conditional mean, \(A_{TF}\), of the TF amplitude, A (Eq. 8), as a function of \(I_{ext}\) and, in Fig. 4b, the inverse of the conditional mean, \(T_{IP}\), of the interpulse time, \(\tau\) (Eq. 9) and the conditional mean, \(\overline{\nu }\), of the interpulse frequency, \(\nu \equiv 1/\tau\), with dashed and solid curves, respectively, for the parameters \(h=8\) and \(EC_{50}=0.5\) in (a) and \(T_{{\text{min}}}=5\,{\text{min}}\), \(b=4\) and \(\kappa =9\) in (b). We observe that \(\overline{\nu }\) and \(1/T_{IP}\) map the whole range of \(I_{ext}\) values in an approximately uniform manner while \(A_{TF}\) is only sensitive to variations of \(I_{ext}\) around \(I_{ext} =0.5\) (the value of \(EC_{50}\) in this example). Not only the means behave differently, but the whole distribution of the random variables, A and \(\nu\), do, as illustrated in Fig. 4c and d where we have plotted their CDFs (A in (c) and \(\nu\) in (b)) computed for each encoding type as explained in Methods using the same parameters as in (a) and (b) and the \(I_{ext}\) distributions of Fig. 2a with medians 0.26 (solid curve), 0.32 (long-dashed curve), 0.5 (short-dashed curve), 0.62 (dashed-dotted curve) and 0.8 (dotted curve). Analyzing Fig. 4c in terms of the MI that is eventually conveyed for amplitude encoding with \(h=8\) and \(EC_{50}=0.5\) (solid circles in Fig. 3a) we conclude that the \(I_{ext}\) distribution that yields maximum MI for these parameters (the one with Med\(I_{ext}=\) 0.5) corresponds to the amplitude CDF which is closest to that of a uniform distribution (short-dashed curve in Fig. 4c). In the case of frequency encoding, the analogous comparison should be made with the curve depicted with open squares in Fig. 3c (which was obtained using \(T_{{\text{min}}}=5\,{\text{min}}\), \(b=4\) and \(\kappa =9\)). In this case, almost all the CDFs depicted in Fig. 4d are similarly close to that of a uniform distribution over a certain support. This could explain the weak dependence of MI with Med\((I_{ext})\) for the curve depicted with open squares in Fig. 3c. The fact that the distribution of the intermediary of the response which yields maximum MI is almost uniform ressembles the optimal input/output relation derived for cases with small, independent of the mean, noise50,51. This description, however, corresponds to the first step in the generation of the response and other uncertainties are subsequently added which further degrade the information. In particular, this is very relevant in the case of frequency encoding as we explain in the following Section.
The invariance of MI with stimulus strength in frequency encoding is limited by two key timescales
The examples of Fig. 3c,d illustrate that MI for frequency encoding can remain relatively invariant as Med(\(I_{ext}\)) varies (c) but that it can also decrease for small or large values of the median (d). This different behavior depends on some of the timescales of the transcription step. On one hand, the finite observation time imposes a limit on the minimum frequency that will likely lead to mRNA production. This limitation is also relevant in physiological situations, due to the finite turn over time of proteins and the need to generate responses within a time frame. In fact, MI decreases with Med(\(I_{ext}\)) if the probability of eliciting at least one pulse during the observation time (\(T=100\,{\text{min}}\) in our simulations) becomes too small. This is particularly noticeable in the examples of Fig. 3d. A simple calculation (see Supplementary note) yields an estimate of this probability at Med\((I_{ext})\)=0.27 of \(\sim 0.21\) and \(\sim 0.64\) for the cases depicted, respectively, with circles and squares in Fig. 3d and of \(\sim 0.83\) and \(\sim 1\) for those depicted with circles and squares, respectively, in Fig. 3c. On the other hand, too large input strengths can become indistinguishable for the mRNA production if the difference, \(\Delta T_{IP}\), between their corresponding mean interpulse times, \(T_{IP}\) (Eq. 10), is so small that it is filtered out by some of the slower processes of the transcription step. This can cause a decrease in MI with increasing Med\((I_{ext})\), as observed in Fig. 3c. The analysis of the examples of Fig. 3c (see Supplementary note) shows that the minimum discernible \(\Delta T_{IP}\) is \(\sim 7-8\, {\text{min}}\). This timescale is approximately equal to the characteristic mRNA degradation time of the simulations (\(\sim 8 \,{\text{min}}\)) which, in turn, agrees with the fastest mRNA turnover times determined in yeast52. We have previously observed that this timescale is key in limiting the information transmitted through the transcription step (see Fig. 3D in Ref.16). The observation of indistinguishable mean interpulse times in experiments (e.g., Fig. 4B in Ref.31 or Fig. 3C in Ref.28) can then be used to determine the range of inputs for which frequency encoding can work.
Combining amplitude and frequency encoding to expand the range of distinguishable stimuli
The different way in which the two types of strategies encode external stimuli might serve to enlarge the range of distinguishable stimulus strengths in cell types that use the two encodings to respond to the same type of stimulus. This could happen in the yeast mating response in which the two types of encodings have been observed28,53,54. Analyzing the combined use of the two codification strategies in this system, even within the framework of our simple model, would require the quantification of several parameters and this goes beyond the scope of the present paper. Yet, there is room for an analysis as the one that we follow in this Section. Namely, we keep the parameters of the transcription step in the values that we have used so far because they allow a good information transmission for frequency encoding (within a frequency range that, as shown in what follows, overlaps with the one observed in the yeast mating response pathway) and that, by changing them, MI would not vary significantly for amplitude encoding16. Then, we focus on the parameters of the \(I_{ext}\)-TF mapping. Relating them to experimental observations, we study whether the range of distinguishable stimuli can be expanded through the combination of frequency and amplitude encoding. We present this analysis in what follows.
The canonical response of haploid mating type a S. cerevisiae cells to mating pheromone (\(\alpha\)-factor) secreted by their potential partners, involves amplitude encoding as in Eq. (3) with \(h\sim 1\) and dimensional \(EC_{50}\approx 3-5nM\)53,54,55. Let us then consider that the curve with \(h=1\) in Fig. 3b (asterisks) represents this situation. Given that for this curve the dimensionless \(EC_{50}\) is 0.5, we need to introduce the transformation [\(\alpha\)-factor]= 6-10\(nM\, I_{ext}\) to make the equivalence between our results and the experiments. We show in Fig. 5 with crosses the plot of MI vs [\(\alpha\)-factor] that is obtained from this curve by using the transformation [\(\alpha\)-factor]= 8\(nM\, I_{ext}\). We see that MI decreases by \(\sim 1\) bit (\(\sim 40\)%) as the median [\(\alpha\)-factor] increases from \(\sim 2\) to \(\sim 5.4nM\) and by \(\sim 1.5\) bit (\(\sim 60\)%) if it increases up to 7nM.
Estimate of how MI could vary as a function of [\(\alpha\)-factor] for amplitude and frequency encoding in yeast. This rough estimate was derived from the curves depicted with asterisks in Figs. (b) and (c) (here plotted with crosses and asterisks, respectively) by using the transformation [\(\alpha\)-factor]= 8\(nM\, I_{ext}\) .
These cells also display intracellular Ca2+ pulses of increasing frequency with increasing pheromone concentration. Ca2+ pulses occur very rarely for [\(\alpha\)-factor]=0 and their mean frequency increases with [\(\alpha\)-factor] to values that become indistinguishable for [\(\alpha\)-factor]\(>10nM\)28. Although the role of these pulses in the pheromone response pathway is not clear yet, it is conceivable that the nuclear localization of some of the TFs involved in the response be pulsatile as well as it has been observed in the response to Ca2+ stress in yeast which involves intracellular Ca2+ pulses and the pulsatile nuclear localization of the TF, Crz16. A rough estimate in the form of Eq. (10) derived from Fig. 3C of28 gives \(T_{{\text{min}}}\approx (10-12)\)min, \(\kappa \approx 8-9\) and a dimensional \(b\sim (0.4-0.5)/nM\). Let us consider that the curve plotted with asterisks in Fig. 3c (\(T_{{\text{min}}}=10\,{\text{min}}\), \(\kappa =9\), \(b=4\)) corresponds to this situation. Given that the dimensionless b for this curve is 4 we need to introduce the transformation [\(\alpha\)-factor]= (8-10)\(nM\, I_{ext}\) to make the equivalence between our results and the experiments. We show in Fig. 5 with asterisks the plot of MI vs [\(\alpha\)-factor] that is obtained from this curve by using the transformation [\(\alpha\)-factor]= 8\(nM\, I_{ext}\). In this case we observe that MI varies between 1.7 and 2 as the median [\(\alpha\)-factor] is varied between \(\sim 2\) and \(\sim 5.4nM\) and it differs from its maximum by less than 25% for the whole support of the [\(\alpha\)-factor] distribution (\([0,8-10nM]\) had we used other transformation from \(I_{ext}\) to[\(\alpha\)-factor]).
Under physiological conditions, distinguishing relatively subtle differences in [\(\alpha\)-factor] is important for the cell to grow towards the largest [\(\alpha\)-factor] regions to encounter its potential partner. If the partners are apart from one another, we can expect that amplitude encoding be used at the earliest stages of the detection, on one hand, because, as illustrated by the curve with \(h=1\) of Fig. 3b, it works correctly for relatively small median concentrations. On the other hand, because if [\(\alpha\)-factor] is too low it will take a relatively long time for an individual cell to collect enough statistics and “respond” correctly using frequency encoding (our estimate of the mean interpulse time derived from28 yields \(\sim 70\,{\text{min}}\) at [\(\alpha\)-factor]=1nM). Furthermore, there is a time lag between exposing the cells to \(\alpha\)-factor and the occurrence of Ca2+ pulses which, at the saturating level [\(\alpha\)-factor]=100nM, is of 30 min on average28. As cells change their form, getting closer to their partners, the pheromone concentration around the growing mating projection gets larger. Amplitude encoding might then cease to discriminate [\(\alpha\)-factor] values, but frequency encoding could still do its job. Therefore, the apparently redundant use of amplitude and frequency encoding to mount the pheromone response might serve the purpose of allowing the cell to detect differences in [\(\alpha\)-factor] across different concentration ranges.
Summary, discussion and conclusions
In this paper we compared two strategies commonly used by cells to encode changes in the environment and generate responses: amplitude and frequency encoding. While in the former increasing stimulus intensities are transduced into increasing concentrations or activation levels of the intermediaries of the pathway, in the latter, pulsatile behaviors are induced in which the frequency increases with the stimulus strength. We had previously studied the information capabilities of the transcription step when the Transcription Factor’s (TF) nuclear fraction displayed one or the other dynamics16. In the present paper we broadened our approach and focused on the effect that the transduction of the external stimulus into the TF’s nuclear fraction had on the information transmission capabilities for each codification strategy. To this end we assumed that amplitude encoding entailed a mapping from the external stimulus, \(I_{ext}\), to the TFs mean concentration, \(A_{TF}\), in the form of a Hill curve (Eq. 3), an expression that describes many gene input functions56 including those involved in the canonical pheromone response pathway in yeast53,54,55. For frequency encoding, we assumed that the interpulse time, \(\tau\), was the sum of a constant, \(T_{{\text{min}}}\), and an exponentially distributed variable, \(\eta\), (Eq. 9) with a mean, \(T_{IP},\) that depended exponentially on \(I_{ext}\) (Eq. 10), as observed experimentally in sequences of intracellular Ca2+ pulses31 and derived theoretically for different classes of noisy driven excitable systems42,44. Screening a set of \(I_{ext}\) distributions defined over the same compact support, of similar variance but different medians, we studied how the mutual information, MI, between the mRNA produced over a finite time, O (Eq. 7), and the stimulus strength, \(I_{ext}\), varied with the median of the distribution, Med\((I_{ext})\). We performed this analysis under the assumption that Med\((I_{ext})\) was representative of the stimulus strengths that constituted the bulk of each \(I_{ext}\) distribution.
For amplitude encoding we found, in agreement with previous observations57, that the maximum MI is achieved if Med\((I_{ext})\) approximately matches the Hill function’s \(EC_{50}\) in those cases with cooperativity index, \(h>2\), (Fig. 3a) while it is optimal at small values of Med\((I_{ext})\) if \(h=1\) (Fig. 3b). Although Eqs. (3), (8) define only the first part in the generation of the output (Eq. 7), we see that the properties of the Hill function imprint their mark on the final depedendence of MI with Med\((I_{ext})\). Namely, the values Med\((I_{ext})\) that give maximum MI roughly correspond to those of \(I_{ext}\) that yield maximum variability of the Hill function, which, in turn, satisfy Eq. (18), the condition under which two stimuli that differ by \(\Delta _I\) lead to distinguishable nuclear TF concentrations. We found a similar situation in the case of frequency encoding: the properties of the first part of the response generation (Eqs. 9 and 10) highly influenced the dependence of MI with Med\((I_{ext})\). In this case, two stimulus strengths that differ by \(\Delta _I\) lead to distinguishable interpulse frequency distributions if Eq. (19) is satisfied for some \(p>1/2\). \(\Delta _F\) in Eq. (19) and the Kullback-Leibler divergence (KL in Eq. (20)) are scale invariant in the sense that they do not depend on \(I_{ext}\), but only on \(\Delta _I\). This is the reason that underlies the weak dependence of MI with Med\((I_{ext})\) of Fig. 3c. The timescales of the subsequent steps in the generation of the response put limits to this scale invariance. As illustrated in Fig. 3d, MI can decrease for decreasing Med\((I_{ext})\) if the probability of eliciting one TF pulse during the observation time, T, becomes to small. MI can also decrease for increasing Med\((I_{ext})\). The analysis of the Med\((I_{ext})\) values that yield maximum MI in each of the examples of Fig. 3c led us to conclude that the limiting timescale in the high frequency end is that of mRNA degradation. This coincides with our previous studies which showed that this timescale limits the information transmitted through the transcription step (Fig. 3D of16).
The above discussion shows that the qualitative difference between amplitude and frequency encoding derives from the qualitative differences between the Hill and the exponential functions that characterize the first part in the generation of the response. Here the use of the Hill function is incontestable. In the case of the exponential, we provided experimental evidence6,31 and cited theoretical works that derive this dependence for different types of noise-driven excitable systems42,44. Given the widespread presence of excitable dynamics in biology, including the paradigmatic example in which spikes are used to transmit information (neurons), we can expect a pervasive presence of this dependence in pulse-signaling systems. Another important feature of the exponential dependence is that it can yield relatively large ranges of mean interpulse times42, as large as those observed in experiments6,31. This feature is particularly advantageous for frequency encoding, as previously noticed58.
In the paper we also discussed the pheromone response pathway in S. cerevisiae as a potential example in which the combination of frequency and amplitude encoding could enlarge the range of external stimuli (i.e., pheromone concentrations) over which the cells could reliably distinguish different values. In this case, Ca2+ pulses were observed in mating type a (MATa) cells in the presence of the pheromone, \(\alpha\)-factor. Under physiological conditions, these cells secrete the pheromone a-factor which attracts mating type \(\alpha\) (MAT\(\alpha\)) cells. These two pheromones differ in various properties, among them, their diffusivity, secretion mechanisms and extracellular metabolism, so that their gradients around the secreting cell can be expected to differ as well59. This led to the hypothesis that the two pheromones conveyed different spatial information to their potential partners, hypothesis that was contradicted by recent observations59. Furthermore, experiments in which MATa cells were exposed to different \(\alpha\)-factor gradients showed that they could be decoded for a wide range of mean \(\alpha\)-factor concentrations60,61. The ability of MATa cells to detect gradients, i.e., to distinguish [\(\alpha\)-factor] values, across mean concentrations was further confirmed by experiments in which the gradients were produced with different source strengths62. As previously stated59, it seems that the cells “do not rely on a narrow concentration range of pheromone”. As discussed in this paper, frequency encoding can endow the cells with such scale-invariant discrimination ability. We expect to do experiments to analyze this hypothesis in the future.
Data availibility
The underlying code for this study is available in Git-Hub and can be accessed via this link: https://github.com/alangivre/qualitatively-scripts.
Change history
19 June 2025
The original online version of this Article was revised: In the original version of this Article Affiliation 3 was incorrectly given as ‘Departamento de Fisiología, Departamento de Fisiología, Biología Molecular y Celular, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Buenos Aires, Argentina’. The correct affiliation is listed as ‘Departamento de Fisiología, Biología Molecular y Celular, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Buenos Aires, Argentina’.
References
Ventura, A. C. et al. Utilization of extracellular information before ligand-receptor binding reaches equilibrium expands and shifts the input dynamic range. Proc. Natl. Acad. Sci. 111, E3860–E3869. https://doi.org/10.1073/pnas.1322761111 (2014).
Poritz, M. A., Malmstrom, S., Kim, M. K., Rossmeissl, P. J. & Kamb, A. Graded mode of transcriptional induction in yeast pheromone signalling revealed by single-cell analysis. Yeast 18, 1331–8. https://doi.org/10.1002/yea.777 (2001).
Yu, R. C. et al. Negative feedback that improves information transmission in yeast signalling. Nature 456, 755–761 (2008).
Koninck, P. D. & Schulman, H. Sensitivity of CaM kinase II to the frequency of \(ca^2\) oscillations. Science 279, 227–230. https://doi.org/10.1126/science.279.5348.227 (1998).
Albeck, J. G., Mills, G. B. & Brugge, J. S. Frequency-modulated pulses of ERK activity transmit quantitative proliferation signals. Mol. Cell 49, 249–261 (2013).
Cai, L., Dalal, C. K. & Elowitz, M. B. Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455, 485–490. https://doi.org/10.1038/nature07292 (2008).
Hao, N. & O’Shea, E. K. Signal- dependent dynamics of transcription factor translocation controls gene expression. Nat. Struct. Mol. Biol. 19, 31–39 (2012).
Yissachar, N. et al. Dynamic response diversity of NFAT isoforms in individual living cells. Mol. Cell 49, 322–330 (2012).
Dalal, C., Cai, L., Lin, Y., Rahbar, K. & Elowitz, M. Pulsatile dynamics in the yeast proteome. Curr. Biol. 24, 2189–2194. https://doi.org/10.1016/j.cub.2014.07.076 (2014).
Tostevin, F., de Ronde, W. & ten Wolde, P. R. Reliability of frequency and amplitude decoding in gene regulation. Phys. Rev. Lett. 108, 108104. https://doi.org/10.1103/PhysRevLett.108.108104 (2012).
Micali, G., Aquino, G., Richards, D. M. & Endres, R. G. Accurate encoding and decoding by single cells: Amplitude versus frequency modulation. PLoS Comput. Biol. 11, 1–21. https://doi.org/10.1371/journal.pcbi.1004222 (2015).
Selimkhanov, J. et al. Accurate information transmission through dynamic biochemical signaling networks. Science 346, 1370–1373. https://doi.org/10.1126/science.1254933 (2014).
Hansen, A. S. & O’Shea, E. K. Promoter decoding of transcription factor dynamics involves a trade-off between noise and control of gene expression. Mol. Syst. Biol. 9, 704 (2013).
Hansen, A. S. & O’Shea, E. K. Limits on information transduction through amplitude and frequency regulation of transcription factor activity. Elife 4, e06559. https://doi.org/10.7554/eLife.06559 (2015).
Minas, G. et al. Multiplexing information flow through dynamic signalling systems. PLoS Comput. Biol. 16, 1–18. https://doi.org/10.1371/journal.pcbi.1008076 (2020).
Givré, A., Colman-Lerner, A. & Ponce Dawson, S. Modulation of transcription factor dynamics allows versatile information transmission. Sci. Rep. 13, 2652. https://doi.org/10.1038/s41598-023-29539-3 (2023).
Cover, T. M. & Thomas, J. A. Elements of information theory 2nd edn. (Wiley-Interscience, 2006).
Butte, A. J. & Kohane, I. Mutual information relevance networks: Functional genomic clustering using pairwise entropy measurements. Pac. Symp. Biocomput. 5, 415–426 (2000).
Margolin, A. A. et al. ARACNE: An algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinf. 7, S7. https://doi.org/10.1186/1471-2105-7-S1-S7 (2006).
Quian Quiroga, R. & Panzeri, S. Extracting information from neuronal populations: Information theory and decoding approaches. Nat. Rev. Neurosci. 10, 173–185. https://doi.org/10.1038/nrn2578 (2009).
Paninski, L. Estimation of entropy and mutual information. Neural Comput. 15, 1191–1253. https://doi.org/10.1162/089976603321780272 (2003).
Purvis, J. & Lahav, G. Encoding and decoding cellular information through signaling dynamics. Cell 152, 945–956. https://doi.org/10.1016/j.cell.2013.02.005 (2013).
Nelson, M. T. et al. Relaxation of arterial smooth muscle by calcium sparks. Science 270, 633–637 (1995).
Nishiyama, M., Hong, K., Mikoshiba, K., Poo, M.-M. & Kato, K. Calcium stores regulate the polarity and input specificity of synaptic modification. Nature 408, 584–588. https://doi.org/10.1038/35046067 (2000).
Berridge, M. J., Lipp, P. & Bootman, M. D. The versatility and universality of calcium signalling. Nat. Rev. Mol. Cell Biol. 1, 11–21 (2000).
Dolmetsch, R. E., Xu, K. & Lewis, R. S. Calcium oscillations increase the efficiency and specificity of gene expression. Nature 392, 933–936. https://doi.org/10.1038/31960 (1998).
West, A. E. et al. Calcium regulation of neuronal gene expression. Proc. Natl. Acad. Sci. 98, 11024–11031. https://doi.org/10.1073/pnas.191352298 (2001).
Carbó, N., Tarkowski, N., Ipiña, E. P., Dawson, S. P. & Aguilar, P. S. Sexual pheromone modulates the frequency of cytosolic \(Ca^2+\) bursts in Saccharomyces cerevisiae. Mol. Biol. Cell 28, 501–510. https://doi.org/10.1091/mbc.E16-07-0481 (2017).
Skupin, A. et al. How does intracellular \(Ca^2+\) oscillate: By chance or by the clock?. Biophys. J . 94, 2404–2411 (2008).
Dragoni, S. et al. Vascular endothelial growth factor stimulates endothelial colony forming cells proliferation and tubulogenesis by inducing oscillations in intracellular \(Ca^2+\) concentration. Stem Cells 29, 1898–1907. https://doi.org/10.1002/stem.734 (2011).
Thurley, K. et al. Reliable encoding of stimulus intensities within random sequences of intracellular Ca2+ spikes. Sci. Signaling 7, ra59. https://doi.org/10.1126/scisignal.2005237 (2014).
Thurley, K. & Falcke, M. Derivation of Ca2+ signals from puff properties reveals that pathway function is robust against cell variability but sensitive for control. Proc. Natl. Acad. Sci. 108, 427–432. https://doi.org/10.1073/pnas.1008435108 (2011).
Lopez, L., Piegari, E., Sigaut, L. & Ponce Dawson, S. Intracellular calcium signals display an avalanche-like behavior over multiple lengthscales. Front. Physiol. https://doi.org/10.3389/fphys.2012.00350 (2012).
Hernández-Navarro, L. et al. Noise-driven amplification mechanisms governing the emergence of coherent extreme events in excitable systems. Phys. Rev. Res. 3, 023133. https://doi.org/10.1103/PhysRevResearch.3.023133 (2021).
Izhikevich, E. M. Neural excitability, spiking and bursting. Int. J. Bifurc. Chaos 10, 1171–1266. https://doi.org/10.1142/S0218127400000840 (2000).
Lechleiter, J., Girard, S., Peralta, E. & Clapham, D. Spiral calcium wave propagation and annihilation in Xenopus laevis oocytes. Science 252, 123–126. https://doi.org/10.1126/science.2011747 (1991).
Lechleiter, J. D. & Clapham, D. E. Molecular mechanisms of intracellular calcium excitability in X. laevis oocytes. Cell 69, 283–294. https://doi.org/10.1016/0092-8674(92)90409-6 (1992).
Li, Y.-X. & Rinzel, J. Equations for insp3 receptor-mediated [Ca2+]i oscillations derived from a detailed kinetic model: A hodgkin-huxley like formalism. J. Theor. Biol. 166, 461–473. https://doi.org/10.1006/jtbi.1994.1041 (1994).
Tang, Y. & Othmer, H. G. Frequency encoding in excitable systems with applications to calcium oscillations. Proc. Natl. Acad. Sci. 92, 7869–7873 (1995).
Batchelor, E., Mock, C. S., Bhan, I., Loewer, A. & Lahav, G. Recurrent initiation: A mechanism for triggering p53 pulses in response to DNA damage. Mol. Cell 30, 277–289. https://doi.org/10.1016/j.molcel.2008.03.016 (2008).
Mönke, G. et al. Excitability in the p53 network mediates robust signaling with tunable activation thresholds in single cells. Sci. Rep. 7, 46571. https://doi.org/10.1038/srep46571 (2017).
Givré, A. & Dawson, S. P. Cell information processing via frequency encoding and excitability. J. Stat. Mech: Theory Exp. 2024, 064002. https://doi.org/10.1088/1742-5468/ad4af8 (2024).
Hänggi, P., Talkner, P. & Borkovec, M. Reaction-rate theory: Fifty years after Kramers. Rev. Mod. Phys. 62, 251–341. https://doi.org/10.1103/RevModPhys.62.251 (1990).
Eguia, M. C. & Mindlin, G. B. Distribution of interspike times in noise-driven excitable systems. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 61, 6490–6499 (2000).
Givré, A. & Ponce Dawson, S. Information content in stochastic pulse sequences of intracellular messengers. Front. Phys. https://doi.org/10.3389/fphy.2018.00074 (2018).
Frank, S. A. Input-output relations in biological systems: Measurement, information and the hill equation. Biol. Direct 8, 31. https://doi.org/10.1186/1745-6150-8-31 (2013).
Strong, S., Koberle, R., Steveninck, R. & Bialek, W. Entropy and information in neural spike trains. Phys. Rev. Lett. https://doi.org/10.1103/PhysRevLett.80.197 (1996).
Cheong, R., Rhee, A., Wang, C. J., Nemenman, I. & Levchenko, A. Information transduction capacity of noisy biochemical signaling networks. Science 334, 354–358. https://doi.org/10.1126/science.1204553 (2011).
Gelman, A. et al.Bayesian data analysis. Chapman & Hall/CRC Texts in Statistical Science (CRC Press, 2004).
Laughlin, S. A simple coding procedure enhances a neuron’s information capacity. Zeitschrift fur Naturforschung C 36, 910–912. https://doi.org/10.1515/znc-1981-9-1040 (1981).
Bialek, W. Perspectives on theory at the interface of physics and biology. Rep. Prog. Phys. 81, 012601. https://doi.org/10.1088/1361-6633/aa995b (2017).
Wang, Y. et al. Precision and functional specificity in mRNA decay. Proc. Natl. Acad. Sci. 99, 5860–5865. https://doi.org/10.1073/pnas.092538799 (2002).
Yi, T.-M., Kitano, H. & Simon, M. I. A quantitative characterization of the yeast heterotrimeric G protein cycle. Proc. Natl. Acad. Sci. 100, 10764–10769. https://doi.org/10.1073/pnas.1834247100 (2003).
Bush, A. et al. Yeast GPCR signaling reflects the fraction of occupied receptors, not the number. Mol. Syst. Biol. 12, 898. https://doi.org/10.15252/msb.20166910 (2016).
Colman-Lerner, A. et al. Regulated cell-to-cell variation in a cell-fate decision system. Nature 437, 699–706. https://doi.org/10.1038/nature03998 (2005).
Alon, U. An introduction to systems biology (CRC Press, Boca Raton, 2020).
Walczak, A. M., Mugler, A. & Wiggins, C. H. A stochastic spectral analysis of transcriptional regulatory cascades. Proc. Natl. Acad. Sci. 106, 6529–6534. https://doi.org/10.1073/pnas.0811999106 (2009).
Schuster, S., Marhl, M. & Höfer, T. Modelling of simple and complex calcium oscillations. Eur. J. Biochem. 269, 1333–1355. https://doi.org/10.1046/j.0014-2956.2001.02720.x (2002).
Jacobs, K. C. & Lew, D. J. Pheromone guidance of polarity site movement in yeast. Biomolecules 12, 502. https://doi.org/10.3390/biom12040502 (2022).
Moore, T. I., Chou, C.-S., Nie, Q., Jeon, N. L. & Yi, T.-M. Robust spatial sensing of mating pheromone gradients by yeast cells. PLoS ONE 3, 1–11. https://doi.org/10.1371/journal.pone.0003865 (2008).
Dyer, J. M. et al. Tracking shallow chemical gradients by actin-driven wandering of the polarization site. Curr. Biol. 23, 32–41. https://doi.org/10.1016/j.cub.2012.11.014 (2013).
Jacobs, K. C., Gorman, O. & Lew, D. J. Mechanism of commitment to a mating partner in Saccharomyces cerevisiae. Mol. Biol. Cell 33, ar112. https://doi.org/10.1091/mbc.E22-02-0043 (2022).
Acknowledgements
This research has been supported by UBA (UBACyT 20020170100482BA) and ANPCyT (PICT 2018-02026 and PICT-2021-III-A-00091 to SPD and PICT 2019-1455 to ACL).
Author information
Authors and Affiliations
Contributions
AG and SPD performed numerical and analytical calculations, respectively; AC-L and SPD conceived project; AC-L and SPD wrote paper.
Corresponding author
Ethics declarations
Competing interests
All authors declare no financial or non-financial competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Givré, A., Colman-Lerner, A. & Ponce Dawson, S. Amplitude and frequency encoding result in qualitatively distinct informational landscapes in cell signaling. Sci Rep 15, 8075 (2025). https://doi.org/10.1038/s41598-025-92424-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92424-8
This article is cited by
-
Biological physics to uncover cell signaling
Biophysical Reviews (2025)