Abstract
In chronic infections caused by Staphylococcus aureus, biofilm is a major virulence factor. In Staphylococcus aureus biofilms, bacteria are embedded in a matrix of extracellular polymeric substances and are highly tolerant to antimicrobial drugs. However, the lack of effective solutions to inhibit biofilm formation remains a challenge, and the mechanism of inhibition of biofilm formation targeting extracellular polymeric substances is unclear. The aim of the present study was to investigate the inhibitory mechanisms of Plumbagin against Staphylococcus aureus biofilms formation by affecting secretion of extracellular polymeric substances using the high-content screening. Our results showed Plumbagin (16 µg/mL) inhibited biofilm formation, revealing a significant reduction in both biomass and bacterial metabolic activity, and disrupted the biofilm structure, leading to a significant decrease in both biological volume and average thickness (P ≤ 0.01). High-content screening imaging indicated that the Plumbagin treatment induced alterations in the extracellular polymeric substances of Staphylococcus aureus biofilm, significantly reducing the quantities of extracellular polysaccharide, proteins and extracellular DNA. Interestingly, extracellular DNA within the matrix was found to be the most sensitive to Plumbagin treatment. Extracellular DNA formation was significantly inhibited at a concentration of 4 µg/mL, whereas the inhibition of extracellular polysaccharide and proteins required a higher concentration of 8 µg/mL. Overall, these results demonstrated the inhibitory effects of Plumbagin on Staphylococcus aureus biofilm formation and extracellular polymeric substances secretion, suggesting that extracellular DNA may be a potential target for the anti-biofilm activity of Plumbagin. These findings will provide new insights into the mode of action of Plumbagin in treating infections caused by Staphylococcus aureus biofilms.
Similar content being viewed by others
Introduction
Chronic wounds are a growing medical problem, resulting in high morbidity and mortality. Staphylococcus aureus (S. aureus) is one of the predominant causes of persistent infections in chronic wounds, mainly due to its remarkable ability to adapt to different environments and to acquire genes associated with antimicrobial resistance1,2. Bacteria of the Staphylococcus genus are highly efficient in establishing biofilms resulting in persistent infections that are difficult to treat3.
Biofilms are complex microbial architectures that attach to surfaces and encase microorganisms in a matrix composed of self-produced hydrated extracellular polymeric substances (EPS). In S. aureus biofilms, poly-N-acetyl-β-(1–6)-glucosamine (PNAG; also known as polysaccharide intercellular adhesin PIA), proteins and extracellular DNA (eDNA) are broadly viewed as the main components of the EPS matrix4. Previous studies suggested that removing the residual EPS matrix could be at least as crucial as killing bacteria in the management of biofilm infections5. Targeting the EPS may be an effective strategy to remove biofilm, disaggregate bacteria and disrupt the pathogenic environment6. Additionally, due to the variability in the composition of S. aureus EPS matrix and the interaction between their multiple components, the strategies to disrupt the matrix should ideally target several constituents of the EPS matrix simultaneously7. Currently, it is believed over 80% of chronic infectious diseases are mediated by biofilms, and it is known that conventional antibiotic medications are inadequate at eradicating these biofilm-mediated infections8. Therefore, there is an urgent need for alternative compounds capable to treat S. aureus biofilms.
Natural product therapeutics are currently receiving significant attention due to their antimicrobial efficacy and their ability to avoid inducing drug resistance. Examples of such natural products include polyphenols, flavonoids, quinones, terpenoids, alkaloids, and tannins, all of which have been demonstrated to possess antimicrobial properties9,10,11,12,13. Plumbagin (5-hydroxy-2-methyl-1,4-napthoquinone, PLB) is a naturally occurring naphthoquinone isolated from the roots of the medicinal plant Plumbago zeylanica L. PLB has been documented to exhibit a wide range of biological activities including anti-inflammatory, anticancer, antidiabetic, antioxidant, antibacterial, antifungal, and anti-atherosclerotic properties14. It is considered a natural agent with the potential to control chronic diseases and has shown in vitro antibacterial activity against S. aureus and Candida albicans15,16.
In this study, S. aureus biofilm was used as a research model for chronic wound infection in vitro to examine the antibiofilm activity of PLB against S. aureus and to explore the mechanism of inhibiting the formation of S. aureus biofilm by influencing EPS secretion.
Results
MIC determination and growth curve construction
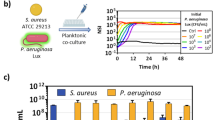
To evaluated the sensitivity of PLB to S. aureus, the MIC of PLB was determined. Figure 1A illustrated that the MIC of PLB against planktonic S. aureus was 4 µg/mL. At a PLB concentration of 4 µg/mL, the growth of S. aureus planktonic cells was markedly inhibited (Fig. 1B). Given that biofilm formation significantly enhanced antimicrobial resistance, we constructed a straightforward biofilm growth curve to delineate the growth characteristics of the biofilm and pinpoint the optimal time for antibiofilm research. According to Fig. 1C, the biofilm formation of S. aureus reached its maximum at 24 h, after which a decline in growth was noted. Consequently, we selected the 24-hour-old biofilm as an in vitro model to evaluate the inhibitory effect of PLB on biofilm formation.
MIC determination and growth curve construction. (A) The MIC of PLB against planktonic S. aureus. (B) The effect of PLB on growth of S. aureus. (C) The growth curve of the biofilm from 0 h to 48 h. Data were presented as the mean ± standard deviation.
Assessment of the inhibitory effect of PLB on biofilm formation
Effect of PLB on the biofilm biomass
Based on the susceptibility of planktonic S. aureus cells to PLB, we evaluated the impact of PLB on biofilm formation by measuring the binding of CV to the biofilms. The antibiofilm activity of PLB at concentrations ranging from 0.5 to 64 µg/mL was shown in (Fig. 2A). In our biofilm biomass quantification assay, we observed a concentration-dependent increase in the antibiofilm activity of PLB. Specifically, biofilm formation was reduced by 2, 43, and 92% at concentrations of 4, 8, and 16 µg/mL, respectively, as shown in (Fig. 2A). However, no significant increase in antibiofilm activity was observed when the PLB concentration surpassed 16 µg/mL. Consequently, this concentration was identified as the minimum biofilm inhibitory concentration (MBIC) for PLB.
Effect of PLB on the metabolic activity
To assess the effect of PLB on bacterial metabolic activity within biofilms, we performed an XTT reduction assay on the biofilms. The XTT assay showed that the metabolic activity of bacteria within the biofilms decreased in a concentration-dependent manner with PLB treatment. The findings from the XTT reduction assay provided clear evidence that PLB, at its MBIC, specifically targeted biofilm formation without compromising the viability of S. aureus (Fig. 2B). Specifically, the metabolic activity of the S. aureus biofilm was found to be reduced by only 64% at the MBIC. Furthermore, when the concentration of PLB was elevated to 32 µg/mL, the inhibition rate of PLB on the metabolic activity within S. aureus biofilms increased to 90%.
Light microscope and HCS analysis
The impact on biofilm architecture is a crucial criterion for evaluating the efficacy of any antibiofilm agent. The S. aureus biofilm was visualized using a light microscope after staining with CV (Fig. 2C). The untreated wells exhibited a clumped structure. In contrast, samples treated with PLB displayed a significant reduction in biofilm formation, suggesting that PLB effectively inhibited both biofilm formation and adhesion.
To substantiate our findings, the HCS system was employed to assess changes in biofilm architecture between PLB-treated and untreated biofilms. Analysis of the z-stack images, captured using this microscopy technique and processed with COMSTAT software, clearly demonstrated that PLB treatment disrupted biofilm development. This was evidenced by a decrease in the average biofilm biovolume and thickness, as well as an increase in the roughness coefficient post-treatment, as illustrated in (Fig. 2D, E).
Effect of PLB on S. aureus biofilm formation. (A) Effect of PLB on the biofilm biomass of S. aureus as quantified by CV staining. (B) The viability of biofilm cells of S. aureus was determined by XTT assay. (C) Light microscope images at 40 times magnification. (D) The HCS Z-stack maximum projections, 3D reconstructions and merge images of S. aureus biofilm stained with the Film Tracer™ LIVE/DEAD biofilm kit. (E) COMSTAT image analysis. Data were presented as the mean ± standard deviation. **Indicated P ≤ 0.01 compared to control.
Effect of PLB on biofilm matrix components
EPS is crucial for maintaining biofilm architecture. To elucidate the potential mechanisms underlying the inhibition of biofilm formation, we investigated the effect of PLB on EPS secretion. Treatment of S. aureus biofilms with PLB led to a significant reduction in EPS production (Fig. 3). FITC-WGA staining revealed a marked decrease in PIA production in S. aureus biofilms treated with 8 and 16 µg/mL of PLB when compared to the untreated control group (Fig. 3B). Additionally, the protein content within the biofilm was significantly diminished at a concentration of 8 µg/mL of PLB (Fig. 3C). The presence of eDNA in the biofilms was assessed using PicoGreen, a novel method that involves fluorescence intensity measurement. When the stain was added, PicoGreen did not penetrate the cells but bound exclusively to eDNA. The inhibitory effect of PLB on eDNA was particularly pronounced, with a concentration of 4 µg/mL significantly suppressing eDNA formation (Fig. 3D, E). The fluorescence intensity quantified by HCS images also observed the same trend which showed a significant change in fluorescence intensity during biofilm formation after PLB treatment (Fig. 3A).
Effect of PLB on the EPS components of S.aureus biofilms. (A) HCS images of PIA, protein and eDNA in the biofilm formation assay. (B) The quantitative analysis of PIA production in each sample was determined by measuring the intensity of green fluorescence using Image J software. The date was presented as the PIA production in each layer of the biofilm (1 μm sections). Total PIA production in biofilms was presented as the area under the curve. (C) The quantitative analysis of protein production in each sample was determined by measuring the intensity of red fluorescence using Image J software. The data are presented as the protein production in each layer of the biofilm (1 μm sections). Total protein production in biofilms was presented as the area under the curve. (D) The concentration of eDNA was assessed using the Quant-iT™ PicoGreen® dsDNA Assay Kit. (E) The percentage of EPS components (PIA, protein, eDNA) in biofilms treated with various concentrations of PLB compared to the untreated control was calculated. Data were expressed as mean ± standard deviation. **Indicates a significant difference P ≤ 0.01 compared to control.
Discussion
The presence of chronic wound infection significantly hampers the process of normal wound healing and typically necessitates antibiotic treatment. However, the excessive use of antibiotics contributes to the development of bacterial resistance against these drugs17. Biofilm formation is a bacterial resistance mechanism that acts as a physical barrier to antibiotics and leads to metabolic differences, limiting antibiotic effectiveness18,19.
A variety of plant-derived compounds have been demonstrated to exhibit potential antimicrobial and antibiofilm actions, including cell membrane rupture; blockade of QS activity; inhibition of bacterial motility, cell adhesion, EPS formation, and cell proliferation; efflux pump downregulation; and synergism with other antimicrobials20. PLB is a naturally occurring naphthoquinone that has been reported to exhibit antibiofilm effects. Previous studies have demonstrated that Plumbagin exhibits anti-biofilm activity against S. aureus, enhances the antibacterial efficacy of antibiotics, and inhibits biofilm formation by S. aureus21,22,23. Our study confirms these findings and further elucidates the underlying mechanism by which PLB interferes with biofilm formation, specifically focusing on its effects on the biofilm extracellular matrix. Guo et al.24 demonstrated that PLB inhibited the expression of PIA and the release of eDNA in S. aureus biofilms using extraction and isolation techniques. To further investigate the effects of PLB on the biofilm extracellular matrix, we employed molecular probe labeling to selectively label PIA, proteins, and eDNA. These components were then quantitatively analyzed to identify potential targets through which PLB may inhibit biofilm formation. However, its effects on biofilm formation, as well as the related molecular mechanisms, particularly against S. aureus, remain largely unknown. In this study, the antibacterial and antibiofilm activities of PLB were evaluated. The results demonstrated that PLB exhibited strong antibacterial activity against S. aureus, with a MIC of 4 µg/mL. This antibacterial activity may be related to the naphthoquinone structure of PLB. It has been reported that shikonin and luteolin, which also have the naphthoquinone structure, also have antibacterial activity against S. aureus25,26. As demonstrated by CV staining, PLB significantly inhibited biofilm formation at a concentration of 16 µg/mL. However, the results from the XTT reduction experiment revealed that PLB had limited inhibition effects on the metabolic activity of bacteria within biofilms at this concentration. Hence, it is reasonable to state that the anti-biofilm potential of PLB in MBIC does not arise from its ability to inhibit bacterial growth. Biofilms are more resistant to antimicrobials compared to planktonic cells. The most prominent characteristic of a biofilm is the formation of a three-dimensional structure through colony aggregation and stacking27. S. aureus biofilm is often encased in a self-produced extracellular matrix which is pivotal for the three-dimensional architecture of the biofilm, facilitating its adherence to various surfaces and ensuring the cohesion among the bacterial communities within the biofilm28. The complex nature of the matrix represents a diffusion barrier for antimicrobial agents, since it limits their penetration into deeper layers of the biofilm29. This indicates that the matrix components are a crucial focus in the investigation of the anti-biofilm activity exerted by plant-derived bioactive compounds (PLB) against S. aureus.
The antibiofilm effect of PLB on S. aureus was assessed using CV staining, a widely used method for high-throughput quantification of biofilm biomass30,31. However, the use of CV has limitations including toxicity, unspecific binding to negatively charged molecules, and low reproducibility due to uneven dye extraction or differential removal of biofilm biomass throughout the washing steps32,33,34. Traditional microscopy approaches suffer from low throughput and subjectivity in image acquisition35. In this study, we utilized a novel high-throughput screening platform based on high-content screening that provided rapid, reliable, quantitative assessment of biofilm formation. SYTO9 and PI fluorescent dyes in combination with COMSTAT software were utilized to further analyze the impact of PLB on biofilm formation. The findings demonstrated that PLB disrupted the biofilm structure, leading to a significant decrease in both biological volume and average thickness. MatthewV et al.35 also employed high-content screening system to assess the influence of inhibitors on C. jejuni biofilms viability and structural integrity. They conducted initial screening for the metabolically active living bacterial population as well as the eDNA of the biofilm using TMAMR and SytoX fluorescent dyes. Powell et al.36 utilized COMSTAT image-analysis software of CLSM z-stack images to quantify the structural changes in Pseudomonas aeruginosa biofilms treated with oligosaccharides, indicating a disruption of the biofilm through reduction in polysaccharides and extracellular DNA components of the matrix. Although the composition of Pseudomonas aeruginosa biofilms is different compared to S. aureus biofilms, the paradigm of targeting EPS and disrupting biofilms in S. aureus needs to be explored. Thus, to clarify possible mechanism of inhibiting biofilm formation, the influence of PLB on the secretion of EPS was investigated.
The EPS plays a crucial role in protecting bacteria within biofilms by forming multiple layers around the cells and facilitating attachment to surfaces37,38. Biochanin A39 and phenyllactic acid40 effectively prevent S. aureus biofilm formation through the reduction of EPS production. These polymeric substrates have been widely recognized as prominent molecular targets for the development of anti-biofilm agents.
Exopolysaccharides serve as the primary constituents of the EPS matrix, facilitating adhesion and maintaining structural integrity to form bacterial aggregates39. Numerous phytochemicals have been discovered to effectively reduce biofilm formation by directly targeting exopolysaccharides within the EPS41,42,43. Aloe-emodin41 and Quercetin44 have demonstrated their efficacy in reducing biofilm formation by inhibiting protein synthesis. As expected, the HCS images confirmed that PLB effectively inhibited the formation of biofilm EPS.
FITC-WGA and SYPRO Ruby staining, along with fluorescence intensity measurements, revealed that PLB caused changes in the distribution of PIA and proteins during biofilm formation. At a concentration of 8 µg/mL, there was a significant reduction in PIA and protein content. For eDNA content determination, we utilized a rapid and direct method involving staining with PicoGreen followed by measuring fluorescence intensity45. Remarkably, at a concentration of 4 µg/mL, the release of eDNA was significantly suppressed. Therefore, it can be concluded that eDNA is more susceptible to PLB. It has been demonstrated that eDNA plays an important role in the S. aureus biofilm matrix by acting as an electrostatic net binding cells together through the proteinaceous layer of the biofilm matrix. Given its prevalence in S. aureus biofilms, eDNA holds potential as a primary target for anti-biofilm therapeutics46,47. This paper proposes that targeting this vital structural matrix component can be achieved through the use of a plant-derived active ingredient PLB. Since eDNA plays a crucial role in initial bacterial attachment to a surface and subsequent biofilm growth, including cell aggregation, the absence of eDNA or its inability to form a structural framework hinders the biofilm from establishing a strong foundation and maturing, ultimately inhibiting its growth48,49.
Therefore, the anti-biofilm effect of PLB targeting eDNA opens up new avenues for understanding and treating S. aureus biofilm infections. Although our study highlights eDNA as the most sensitive EPS component to PLB, the precise mechanism by which PLB inhibits eDNA release remains to be elucidated. Based on PLB’s chemical structure and prior studies on analogous compounds, we propose two non-exclusive pathways: First, PLB may directly bind to eDNA via electrostatic or intercalative interactions. The compound’s fused aromatic rings resemble classical DNA intercalators such as Doxorubicin50, while its hydroxyl and ketone groups could form hydrogen bonds with DNA bases51. Furthermore, the partial positive charge of PLB at neutral pH (predicted pKa ~ 8.2) may enhance its affinity for the negatively charged DNA backbone, enabling it to sequester eDNA and impede its integration into the biofilm matrix. Second, PLB might indirectly suppress eDNA release by targeting bacterial autolysis. Biofilm-associated eDNA is often derived from cell lysis mediated by autolysins52. PLB could inhibit autolysin activity or downregulate their expression, thereby reducing eDNA liberation. This hypothesis aligns with studies showing that natural compounds (e.g., Geraniol) inhibit autolysis by downregulating autolysin genes53. While the current study did not resolve whether PLB acts directly on DNA or through bacterial pathways, its structural features (e.g., planarity, charge distribution) and phenotypic effects provide a foundation for future mechanistic work.
Our previous investigations explored the interactions between PLB and various antibiotics, including nitrofurantoin, ciprofloxacin, mecillinam, and chloramphenicol. PLB demonstrated synergistic effects with these antibiotics, significantly enhancing their growth-inhibitory activity against S. aureus21. These findings suggest that PLB could serve as a valuable adjunctive therapy in combinatorial strategies targeting biofilm-associated infections. Research indicates that PLB enhances the anti-biofilm activity of chlorhexidine (CHX) against Klebsiella pneumoniae. This combination reduces the MIC of CHX and significantly diminishes bacterial load in biofilms. The synergistic effect is attributed to PLB’s ability to increase membrane permeability and inhibit efflux pump expression, facilitating better antibiotic penetration and efficacy54. Integrating PLB into conventional antibiotic regimens presents a promising strategy to combat biofilm-related infections, particularly those caused by multidrug-resistant pathogens. By improving antibiotic penetration and potency, PLB may reduce required antibiotic dosages, potentially mitigating side effects and delaying the emergence of resistance.
As a natural product, PLB exhibits broad-spectrum antimicrobial and antibiofilm activities against diverse bacterial species, highlighting its potential as a novel anti-infective agent. However, its achievable concentrations in the body and potential toxicity have not been thoroughly studied. Future research should focus on the pharmacokinetic properties of PLB, assessing its absorption, distribution, metabolism, and excretion in the human body to determine its feasibility for clinical application. Additionally, preclinical and clinical trials are necessary to evaluate the safety and efficacy of PLB, providing scientific evidence for its development as an anti-infection drug.
Conclusion
Overall, this study demonstrated that PLB is an effective anti-biofilm agent. Moreover, its anti-biofilm does not arise from its ability to inhibit bacterial growth, so we investigated the mechanism by which PLB inhibits biofilm formation, focusing on its influence on EPS secretion. We used a high-content system to confirm that PLB inhibits EPS formation during biofilm development, and that PLB is more sensitive to eDNA of biofilm, suggesting that eDNA is a target for the observed PLB anti-biofilm activity. Given its enhanced antibacterial efficacy, PLB represents a promising alternative for biofilm inhibition and offers a novel strategy for treating S. aureus biofilm infections. However, further research is necessary to elucidate the precise mechanisms of action of PLB in biofilm inhibition and to evaluate its clinical potential as an anti-biofilm agent.
Materials and methods
Chemicals
PLB was purchased from the Aladdin Industrial Corporation (Shanghai, China). PLB was dissolved in dimethyl sulfoxide (DMSO) at a final stock concentration of 50 mg/mL. Film Tracer™ LIVE/DEAD biofilm kit, Film Tracer™ SYPRO Ruby, and the Quant-iT™ PicoGreen® dsDNA Assay Kit were purchased from Invitrogen (Thermo Fisher Scientific, USA). Fluorescein iso-thiocyanate-conjugate wheat germ agglutinin from Triticum vulgaris (FITC-WGA) was purchased from Sigma-Aldrich. All other chemicals were of analytical grade.
Bacterial strains and growth conditions
S. aureus CMCC(B) 26,003 was purchased from China Institute of Food and Drug Control and stored at -80℃ with 25% glycerol (Aladdin Reagent Ltd., Shanghai, China). The bacteria were inoculated in Luria-Bertani (LB) medium and incubated at 37 °C for 18–22 h. The culture was then adjusted to an optical density (OD) of 0.1 at 625 nm, corresponding to approximately 108 CFU/mL, which represents the mid-logarithmic growth phase of S. aureus55. This cell suspension was used for subsequent experiments. The bacteria were inoculated in LB supplemented with at 37℃ for 16 h to obtain logarithmic phase cells for biofilm assays. Subsequently, the bacterial suspensions were incubated at 37℃ for 24 h in LB medium containing 3% glucose to promote biofilm formation, as previously described with slight modifications56,57.
Minimum inhibitory concentrations (MIC) of PLB against S. aureus in planktonic state
The MIC of PLB against S. aureus in planktonic state was determined using the broth microdilution technique, following the guidelines of the Clinical and Laboratory Standards Institute (CLSI), with slight modifications. Briefly, various concentration of PLB ranging from 0 to 128 µg/mL were added into a 96-well microtiter plate with the bacterial suspensions and incubated 37℃ for 24 h. The bacterial growth was measured as optical density by a full-wavelength microplate reader at 625 nm. The MIC of PLB was determined as the concentration that resulted in a 90% inhibition.
Growth curve analysis
The effect of PLB on S. aureus growth was tested as previously described58. To obtain growth curve, 1% inoculum from standard cell suspension (around 1 × 108 CFU/mL) of S. aureus were added to 50 mL sterile LB broth with PLB at 2 and 4 µg/mL and incubated at 37 ℃ with 120 r/min for 24 h. The bacterial cell density (OD at 600 nm) was measured at selected time intervals (0, 2, 4, 6, 8, 10, 12 and 24 h) using a microplate reader (Tecan Spark, Männedorf, Switzerland).
To characterize the temporal progression of S. aureus biofilm formation, we quantified biofilm development at 6, 12, 18, 24, 36, and 48 h using crystal violet staining. The biofilm formation assay was performed in 96-well microtiter plates, as previously described with slight modifications30,31. Biofilms were initiated by inoculating S. aureus into LB medium supplemented with 3% glucose and incubated at 37 °C. Following incubation at each time point, the biofilms were stained with 0.1% crystal violet and the optical density (OD) of the resulting solution was measured at 570 nm using a microplate reader. All assays were performed in triplicate.
Crystal Violet (CV) staining assay
The biofilm formation assay was performed in 96-well microtiter plates, as previously described with slight modifications30,31. Briefly, S. aureus was inoculated into wells containing PLB (0–64 µg/mL) and incubated for 24 h at 37 ℃. After incubation, the planktonic cells were removed and the wells rinsed twice with sterile water. Then, biofilms in the wells were stained with 0.1% crystal violet for 20 min. After removal of unbound dye, the stained biofilms were washed gently with sterile water, and crystal violet dye bound to biofilms was dissolved using 200 µL of ethanol. The optical density (OD) of the ethanol solution described above was measured at 570 nm using a microplate reader. All assays were performed in triplicate. The percentage of inhibition was calculated according to the following formula58 (Formula(1)). Additionally, the stained biofilms were observed by using a light microscope (Olympus IX53, Tokyo, Japan) at 40 times magnification.
Metabolic activity detection by XTT assay
The effect of PLB treatment on S. aureus metabolic activity and proliferation was determined using an XTT assay. The culture method of biofilm and the concentration of Plumbagin were same as above. After 24 h of biofilm culture, the culture medium was removed, and the wells were washed three times with PBS to remove planktonic cells. Samples were then incubated with the solution of 0.5 mg/mL XTT (Sigma-Aldrich, Shanghai, China) and 10mM menadione in the dark for 3 h. The conversion of tetrazolium salt XTT to a colored formazan derivative was measured at 490 nm in a 96-well plate. All assays were performed in triplicate. The percentage of inhibition was calculated according to the following formula58(Formula (2)).
Representation and quantification of biofilm three-dimensional structure
For the representation and quantification of biofilm, the Film Tracer™ LIVE/DEAD biofilm kit, which includes two stains, SYTO 9 dye and propidium iodide (PI) dye, was used. Firstly, the biofilms were cultured on the black 96-well plates as described the above method. After removal of planktonic cells, the biofilm was stained with SYTO9 and PI according to the kit instructions. All samples were stained for 30 min in the dark at room temperature with an appropriate volume of a mixture of the two dyes SYTO9 and PI (3 µL of each component in 1 mL of filter-sterilized water). Furthermore, the images of the stained biofilm were obtained with an ImageXpressmicro high-content screening system (Molecular Devices, Sunnyvale, CA)59. And then, the Imaris software (version 9.0.1, Oxford Instruments, Abingdon, UK) was utilized to represent a three-dimensional structure of the biofilms. For the quantification of biofilm, at least four random optical fields were chosen and observed. Additionally, a series of z-stack images were analyzed using Image J (version 2.14, Wayne Rasband, NIH, Bethesda, MD, USA) and COMSTAT60 image analysis software, and the three-dimensional biofilms were quantified by the measurement of biofilm volume, surface roughness and biofilm depth. All assays were performed in triplicate.
Quantification of biofilm matrix components
To assess the impact of PLB on the extracellular matrix content of biofilms, the biofilms were allowed to form in black 96-well microtiter plates in the absence or presence of PLB for 24 h at 37 ℃.
Quantification of PIA
FITC-WGA was used as a fluorescent probe to evaluate the formation of PIA in biofilms under the influence of PLB61. After removal of planktonic cells, biofilms were stained with a volume of 100 µl FITC-WGA (10 µg/mL) per well and plates were incubated at room temperature in the dark for 2 h. The plates were washed three times with PBS and then 100 µL of PBS were added to each well. Stained biofilms were observed with HCS system. Three-dimensional images of PIA were constructed using Imaris software. At least four random optical fields were captured and analyzed. The production of the PIA was calculated according to the fluorescence intensity using Image J. The date was presented as the PIA production in each layer of biofilm (1 μm). The percentage of PIA in biofilms treated with PLB was presented as area under the curve (AUC) and compared to untreated control. Three separate experiments were conducted, and a single set of representative data was presented.
Quantification of extracellular proteins
FilmTracer™ SYPRO Ruby (Invitrogen, Molecular Probes) was used as a fluorescent probe to evaluate the formation of biofilms proteins under the influence of PLB62. SYPRO Ruby stain labels most classes of proteins, including glycoproteins, phosphoproteins, lipoproteins, calcium binding proteins, fibrillar proteins, and other proteins that are difficult to stain. The biofilms were stained with a volume of 100 µL SYPRO Ruby per well and plates were incubated at room temperature in the dark for 1 h. Stained biofilms were observed with HCS system. A similar approach to the one described above for PIA was used.
Quantification of eDNA
The eDNA was quantified using the Quant-iT™ PicoGreen® dsDNA Assay Kit to assess the formation of biofilm eDNA under the influence of PLB. PicoGreen® is a fluorescent dye that selectively binds to double-stranded DNA (dsDNA) and exhibits enhanced fluorescence upon binding. Biofilms were cultivated as described in the previous section. After removing planktonic cells, the biofilms were washed twice with PBS. The quantification of eDNA involved adding 100 µL TE buffer followed by 100 µL freshly prepared PicoGreen solution (1 µL PicoGreen dye in 199 µL TE buffer). The wells were mixed and incubated for 2–5 min at room temperature in darkness before measuring fluorescence intensity (excitation 485 nm/emission 535 nm) using a fluorescence microplate reader. Lambda DNA was used to generate a standard curve for each run. Stained biofilms were observed with HCS system. All assays were performed in triplicate.
Statistical analysis
SPSS version 27.0 (SPSS Inc., Chicago, IL, USA) software was used for data quantification and analysis. GraphPad Prism 8.0.2 software was used for mapping. All results were presented as mean ± standard deviation. Analysis of variance (ANOVA) was performed to evaluate significant differences. Statistical significance compared to the untreated control was indicated using double asterisks (**) for P ≤ 0.01.
Data availability
All raw data are available from the corresponding author upon reasonable request.
Abbreviations
- S. aureus :
-
Staphylococcus aureus
- EPS:
-
Extracellular polymeric substances
- PLB:
-
Plumbagin
- PIA:
-
Polysaccharide intercellular adhesin
- eDNA:
-
Extracellular DNA
- DMSO:
-
Dissolved in dimethyl sulfoxide
- FITC-WGA:
-
Fluorescein iso-thiocyanate-conjugate wheat germ agglutinin from Triticum vulgaris
- LB:
-
Luria bertani
- MIC:
-
Minimum inhibitory concentrations
- CLSI:
-
The clinical and laboratory standards institute
- CV:
-
Crystal violet
- PBS:
-
Phosphate-buffered saline
- OD:
-
Optical density
- PI:
-
Propidium iodide
- AUC:
-
Area under the curve
References
James, G. A. et al. Biofilms in chronic wounds. Wound Repair. Regen. 16, 37–44 (2008).
Foster, T. J. Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS Microbiol. Rev. 41, 430–449 (2017).
Götz, F. Staphylococcus and biofilms. Mol. Microbiol. 43, 1367–1378 (2002).
Otto, M. Staphylococcal biofilms. Bacterial Biofilms, 207–228 (2008).
Liu, J., Madec, J. Y., Bousquet-Mélou, A., Haenni, M. & Ferran, A. A. Destruction of Staphylococcus aureus biofilms by combining an antibiotic with subtilisin A or calcium gluconate. Sci. Rep. 11, 6225 (2021).
Gunn, J. S., Bakaletz, L. O. & Wozniak, D. J. What’s on the outside matters: the role of the extracellular polymeric substance of gram-negative biofilms in evading host immunity and as a target for therapeutic intervention. J. Biol. Chem. 291, 12538–12546 (2016).
Hobley, L., Harkins, C. & MacPhee, C. E. Stanley-Wall, N. R. Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes. FEMS Microbiol. Rev. 39, 649–669 (2015).
Li, X. H. & Lee, J. H. Antibiofilm agents: A new perspective for antimicrobial strategy. J. Microbiol. 55, 753–766 (2017).
Wijesundara, N. M. & Rupasinghe, H. P. V. Essential oils from Origanum vulgare and Salvia officinalis exhibit antibacterial and anti-biofilm activities against Streptococcus pyogenes. Microb. Pathog. 117, 118–127. https://doi.org/10.1016/j.micpath.2018.02.026 (2018).
Matilla-Cuenca, L. et al. Antibiofilm activity of flavonoids on Staphylococcal biofilms through targeting BAP amyloids. Sci. Rep. 10, 18968 (2020).
Dahlem Junior, M. A., Edzang, N., Catto, R. W., Raimundo, J. M. & A. L. & Quinones as an efficient molecular scaffold in the antibacterial/antifungal or antitumoral arsenal. Int. J. Mol. Sci. 23, 14108 (2022).
Farha, A. K. et al. Inhibition of multidrug-resistant foodborne Staphylococcus aureus biofilms by a natural terpenoid (+)-nootkatone and related molecular mechanism. Food Control 112, 107154 (2020).
Herrera, K. M. et al. A 3-alkylpyridine-bearing alkaloid exhibits potent antimicrobial activity against methicillin-resistant Staphylococcus aureus (MRSA) with no detectable resistance. Microbiol. Res. 261, 127073 (2022).
Yin, Z. et al. Anticancer effects and mechanisms of action of Plumbagin: Review of research advances. Biomed. Res. Int. 6940953, (2020). https://doi.org/10.1155/2020/6940953 (2020).
de Paiva, S. R., Figueiredo, M. R., Aragão, T. V. & Kaplan, M. A. Antimicrobial activity in vitro of Plumbagin isolated from Plumbago species. Mem. Inst. Oswaldo Cruz 98, 959–961. https://doi.org/10.1590/s0074-02762003000700017 (2003).
Panichayupakaranant, P. & Ahmad, M. I. Plumbagin and its role in chronic diseases. Adv. Exp. Med. Biol. 929, 229–246. https://doi.org/10.1007/978-3-319-41342-6_10 (2016).
Hindy, J. R., Haddad, S. F. & Kanj, S. S. New drugs for methicillin-resistant Staphylococcus aureus skin and soft tissue infections. Curr. Opin. Infect. Dis. 35, 112–119 (2022).
Pontes, J. T. C., d., T., Borges, A. B., Roque-Borda, C. A. & Pavan, F. R. Antimicrobial peptides as an alternative for the eradication of bacterial biofilms of multi-drug resistant bacteria. Pharmaceutics 14, 642 (2022).
Hochbaum, A. I. et al. Inhibitory effects of D-amino acids on Staphylococcus aureus biofilm development. J. Bacteriol. 193, 5616–5622 (2011).
Uddin Mahamud, A. S., Nahar, S., Ashrafudoulla, M., Park, S. H. & Ha, S. D. Insights into antibiofilm mechanisms of phytochemicals: prospects in the food industry. Crit. Rev. Food Sci. Nutr., 1–28 (2022).
Bie, S. et al. Interactions of Plumbagin with five common antibiotics against Staphylococcus aureus in vitro. PLoS One 19, e0297493. https://doi.org/10.1371/journal.pone.0297493 (2024).
Periasamy, H., Iswarya, S., Pavithra, N., Senthilnathan, S. & Gnanamani, A. In vitro antibacterial activity of Plumbagin isolated from Plumbago zeylanica L. against methicillin-resistant Staphylococcus aureus. Lett. Appl. Microbiol. 69, 41–49. https://doi.org/10.1111/lam.13160 (2019).
Alfhili, M. A. et al. Antibacterial and anti-biofilm activity of Plumbagin against multi-drug resistant clinical bacterial isolates. Saudi Med. J. 43, 1224–1233. https://doi.org/10.15537/smj.2022.43.11.20220446 (2022).
Guo, N. et al. Inhibitory effects of Plumbagin on Staphylococcus aureus biofilm in vitro. Adv. Mater. Eng., 683–692 (2015).
Wan, Y. et al. Control of foodborne Staphylococcus aureus by Shikonin, a natural extract. Foods 10, 2954 (2021).
Wan, Y. et al. Antibacterial activity of juglone revealed in a wound model of Staphylococcus aureus infection. Int. J. Mol. Sci. 24, 3931 (2023).
You, J. et al. Inhibition of vibrio biofilm formation by a marine actinomycete strain A66. Appl. Microbiol. Biotechnol. 76, 1137–1144 (2007).
Flemming, H. C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Pinto, R. M., Soares, F. A., Reis, S., Nunes, C. & Van Dijck, P. Innovative strategies toward the disassembly of the EPS matrix in bacterial biofilms. Front. Microbiol. 11, 535344 (2020).
Grossman, A. B., Burgin, D. J. & Rice, K. C. Quantification of Staphylococcus aureus biofilm formation by crystal Violet and confocal microscopy. S. Aureus: Methods Protocols, 69–78 (2021).
Stepanović, S., Vuković, D., Dakić, I. & Savić, B. Švabić-Vlahović, M. A modified microtiter-plate test for quantification of Staphylococcal biofilm formation. J. Microbiol. Methods 40, 175–179 (2000).
Peeters, E., Nelis, H. J. & Coenye, T. Comparison of multiple methods for quantification of microbial biofilms grown in microtiter plates. J. Microbiol. Methods 72, 157–165 (2008).
Kragh, K. N., Alhede, M., Kvich, L. & Bjarnsholt, T. Into the well—A close look at the complex structures of a microtiter biofilm and the crystal Violet assay. Biofilm 1, 100006 (2019).
Amador, C. I., Stannius, R. O., Røder, H. L. & Burmølle, M. High-throughput screening alternative to crystal Violet biofilm assay combining fluorescence quantification and imaging. J. Microbiol. Methods. 190, 106343 (2021).
Whelan, M. V. X., Simpson, J. C. & T, Ó. C. A novel high-content screening approach for the Elucidation of C. jejuni biofilm composition and integrity. BMC Microbiol. 21, 2. https://doi.org/10.1186/s12866-020-02062-5 (2021).
Powell, L. C. et al. Targeted disruption of the extracellular polymeric network of Pseudomonas aeruginosa biofilms by alginate oligosaccharides. NPJ Biofilms Microbiom. 4, 13. https://doi.org/10.1038/s41522-018-0056-3 (2018).
Bridier, A., Briandet, R., Thomas, V. & Dubois-Brissonnet, F. Resistance of bacterial biofilms to disinfectants: a review. Biofouling 27, 1017–1032 (2011).
Richards, J. J. & Melander, C. Controlling bacterial biofilms. ChemBioChem 10, 2287–2294 (2009).
Bai, X. et al. Anti-biofilm activity of Biochanin A against Staphylococcus aureus. Appl. Microbiol. Biotechnol. 107, 867–879 (2023).
Liu, F. et al. Inhibition of biofilm formation and exopolysaccharide synthesis of Enterococcus faecalis by phenyllactic acid. Food Microbiol. 86, 103344 (2020).
Xiang, H. et al. Aloe-emodin inhibits Staphylococcus aureus biofilms and extracellular protein production at the initial adhesion stage of biofilm development. Appl. Microbiol. Biotechnol. 101, 6671–6681 (2017).
Gutierrez-Pacheco, M. et al. Carvacrol inhibits biofilm formation and production of extracellular polymeric substances of Pectobacterium carotovorum subsp. carotovorum. Food Control 89, 210–218 (2018).
Li, C. et al. Separation of phenolics from peony flowers and their inhibitory activities and action mechanism on bacterial biofilm. Appl. Microbiol. Biotechnol. 104, 4321–4332 (2020).
Vazquez-Armenta, F. et al. Quercetin reduces adhesion and inhibits biofilm development by Listeria monocytogenes by reducing the amount of extracellular proteins. Food Control 90, 266–273 (2018).
Tang, L., Schramm, A., Neu, T. R., Revsbech, N. P. & Meyer, R. L. Extracellular DNA in adhesion and biofilm formation of four environmental isolates: a quantitative study. FEMS Microbiol. Ecol. 86, 394–403 (2013).
Dengler, V., Foulston, L., DeFrancesco, A. S. & Losick, R. An electrostatic net model for the role of extracellular DNA in biofilm formation by Staphylococcus aureus. J. Bacteriol. 197, 3779–3787 (2015).
Okshevsky, M., Regina, V. R. & Meyer, R. L. Extracellular DNA as a target for biofilm control. Curr. Opin. Biotechnol. 33, 73–80 (2015).
Das, T., Sharma, P. K., Busscher, H. J., Van Der Mei, H. C. & Krom, B. P. Role of extracellular DNA in initial bacterial adhesion and surface aggregation. Appl. Environ. Microbiol. 76, 3405–3408 (2010).
Dengler, V., Foulston, L., DeFrancesco, A. S. & Losick, R. An electrostatic net model for the role of extracellular DNA in biofilm formation by Staphylococcus aureus. J. Bacteriol. 197, 3779–3787. https://doi.org/10.1128/jb.00726-15 (2015).
Nedělníková, A., Stadlbauer, P., Otyepka, M., Kührová, P. & Paloncýová, M. Atomistic insights into interaction of doxorubicin with DNA: from duplex to nucleosome. J. Comput. Chem. 46, e70035 (2025).
Sánchez-González, Á., Castro, T. G., Melle-Franco, M. & Gil, A. From groove binding to intercalation: unravelling the weak interactions and other factors modulating the modes of interaction between methylated phenanthroline-based drugs and duplex DNA. Phys. Chem. Chem. Phys. 23, 26680–26695 (2021).
Sharma, D. K. & Rajpurohit, Y. S. Multitasking functions of bacterial extracellular DNA in biofilms. J. Bacteriol. 206, e00006–00024 (2024).
Gu, K. et al. Geraniol inhibits biofilm formation of methicillin-resistant Staphylococcus aureus and increase the therapeutic effect of Vancomycin in vivo. Front. Microbiol. 13, 960728 (2022).
Liu, H. et al. Plumbagin enhances antimicrobial and anti-biofilm capacities of chlorhexidine against clinical Klebsiella pneumoniae while reducing resistance mutations. Microbiol. Spectr. 12, e00896–e00824 (2024).
Wiegand, I., Hilpert, K. & Hancock, R. E. Agar and broth Dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Vázquez-Sánchez, D., Habimana, O. & Holck, A. Impact of food-related environmental factors on the adherence and biofilm formation of natural Staphylococcus aureus isolates. Curr. Microbiol. 66, 110–121 (2013).
Jha, S., Bhadani, N. K., Kumar, A. & Sengupta, T. K. Glucose-induced biofilm formation in Bacillus thuringiensis KPWP1 is associated with increased cell surface hydrophobicity and increased production of exopolymeric substances. Curr. Microbiol. 79, 1–10 (2022).
Sivaranjani, M. et al. Morin inhibits biofilm production and reduces the virulence of Listeria monocytogenes—An in vitro and in vivo approach. Int. J. Food Microbiol. 237, 73–82 (2016).
Peng, F., Hoek, E. M. & Damoiseaux, R. High-content screening for biofilm assays. J. BioMol. Screen. 15, 748–754 (2010).
Heydorn, A. et al. Quantification of biofilm structures by the novel computer program COMSTAT. Microbiology 146, 2395–2407 (2000).
Campoccia, D. et al. Exopolysaccharide production by Staphylococcus epidermidis and its relationship with biofilm extracellular DNA. Int. J. Artif. Organs. 34, 832–839. https://doi.org/10.5301/ijao.5000048 (2011).
Ravaioli, S. et al. Various biofilm matrices of the emerging pathogen Staphylococcus lugdunensis: exopolysaccharides, proteins, eDNA and their correlation with biofilm mass. Biofouling 36, 86–100 (2020).
Acknowledgements
The authors acknowledge Zheng Li professor, who offered many helpful comments and criticisms throughout the study and patiently critiqued drafts of the manuscript.
Funding
This research was funded by the Science and Technology Project of Haihe Laboratory of Modern Chinese Medicine (22HHZYSS00003) and Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (ZYYCXTD-D-202002). This research was also funded by the Henan Liwei Biological Pharmaceutical Co., Ltd. (20501/KH2145), and the Anhui Tiancien Biotechnology Co., Ltd. (20501/KH2185).
Author information
Authors and Affiliations
Contributions
SB and HY conceived the study and designed the experiments. HY and CS performed the experiments and handled data curation. HY and SB acquired the funding. CL conducted the investigation. ML, ZY, RL and DL provided critical feedback and helped shape the research, analysis tools, and manuscript. ZC, ML and TM provided the software for visualization. SB and HY wrote the first draft of the manuscript and all authors revised and commented it. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bie, S., Yuan, H., Shi, C. et al. Antibiofilm activity of Plumbagin against Staphylococcus aureus. Sci Rep 15, 7948 (2025). https://doi.org/10.1038/s41598-025-92435-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92435-5