Abstract
The co-detection of SARS-CoV-2 variant with other respiratory virus has been extensively investigated. However, conclusive evidence remains elusive and conflicted. This study investigated the source- and age-dependent prevalence, incidence, and co-detection of multiple viral infections in children younger than 18 years old who presented with clinical symptoms indicative of respiratory infections during SARS-CoV-2 pandemic. We retrospectively obtained and analyzed pediatric patients admitted to the institution and underwent Film Array Respiratory Panel (BIOFIRE® RP2.1) testing between January 2021 and December 2022. This encompassed pre-Delta, Delta and omicron periods, evaluating Film Array results for singular and co-detections. The overall detection rate was 84.2% (1670/1983) among 1,983 pediatric patients. Of these, 106 were SARS-CoV-2 positive. Notably, 45 patients (42%) harbored SARS-CoV-2 as the sole pathogen. Co-detection was significant; 32 cases (30.2%) involved Human Rhinovirus/Enterovirus, 29 (24.5%) involved parainfluenza 3, and 26 (24.5%) involved Respiratory syncytial virus. Peaks of co-detected parainfluenza 3 and Respiratory syncytial virus were evident in the winter of 2022 and absent in 2021. The top three viral strain for co-detection was HRV/EV, PIV-3, and RSV. The emergence order of co-detection strain was HRV/EV ◊PIV-3◊ RSV during the Omicron period in Taiwan. We identified characteristic patterns of SARS-CoV-2-associated co-detections, with a notable emphasis on the co-detection of HRV/EV, PIV3, and RSV alongside SARS-CoV-2. This association appears to hold heightened significance during the Omicron variant era compared to earlier SARS-CoV-2 variants, which contrast with previous studies. The simultaneous circulation of different variants may contribute to variations in viral co-detection, particularly in young children, warranting further investigation.
Similar content being viewed by others
Introduction
The relationship between acute illness and postacute sequelae and SARS-CoV-2 has been extensively investigated by researchers aiming to understand its underlying nature1,2,3,4. Studies conducted during the omicron period revealed a substantial high risk of postacute sequealae even among vaccinated subjects5. Notably, patients with chronic diseases, particularly neurological conditions, showed an elevated risk of severe SARS-CoV-2 during distinct periods dominated by different variants6. Correspondingly, hospitalizations due to SARS-CoV-2 have increased following the emergence of the omicron variant7,8. In line with these findings, a previous study reported a significant increase in the frequency of this clinical presentation during the high transmission period of omicron compared to the delta variant9,10.
Previous studies showed co-detection of respiratory viruses among adult patient with SARS-CoV-2 is common and clinical correlated significantly, thus routine testing should be advised11 Although Agathis, N.T., et al. reported that the co-detection of non-SARS-CoV-2 viral infections in pediatric cases with SARS-CoV-2 infection may increase illness severity, the details of the co-detected pathogens were lacking, and further investigation was limited12.
Thus, considering the existing knowledge gaps, utilizing a multiplex PCR Film Array, a diagnostic tool capable of systemically detecting multiple pathogens with standardized methodology, presents a promising tool for examining the potential association between co-detection pathogen and SARS-CoV-2. This study aimed to investigate the prevalence, incidence, and co-detection of multiple viral infections in children younger than 18 years old during SARS-CoV-2 pandemic.
Methods
Study design
This study retrospectively conducted pediatric patients admitted to the China Medical University (CMU) Children’s Hospital between January 2021 and December 2022, and performed molecular analysis of data obtained from the Film Array Respiratory Panel (BIOFIRE® RP2.1) testing, reviewed data (FA-RP). Inclusion criteria: Pediatric patient presented to outpatient department (OPD), Emergency Department (ED) or impatient (AD) of the institution with symptoms included but were not limited to cough, fever, sore throat, nasal congestion, and difficulty breathing and received the exam of FA-RP was enrolled. Exclusion criteria: Patient older than 18-year-old was excluded from the study. OPD refers to a section staffed to provide healthcare for limited hours to pediatric patients with minor illnesses, while the ED provides 24-hour services and typically treats pediatric patients presenting with more critical conditions.
Data collection and analysis
Deidentified FA-RP full data abstraction was conducted by the laboratory medicine department of the institution, capturing variables such as age, sex, date of FA-RP examination, and the corresponding FA-RP results. The FA-RP assay, a multiplex polymerase chain reaction (PCR) panel, was used to detect multiple respiratory pathogens simultaneously. FA-RP testing was performed as part of the standard clinical care at the CMU Children’s Hospital. The assay employed the BIOFIRE® RP2.1 panel, which encompasses 22 respiratory pathogens, including SARS-CoV-2, influenza, parainfluenza (PIV), coronal viral strain, respiratory syncytial virus (RSV), human rhinovirus/enterovirus (HRV/EV), human metapneumovirus (HMPV), adenovirus, as well as bacterial targets like Bordetella pertussis and Mycoplasma pneumoniae.
The demographic and clinical characteristics of the study population were summarized using descriptive statistics. The prevalence of single viral infections and co-detection of multiple viral pathogens were determined based on the results obtained from FA-RP. Furthermore, multi-viral co-detection was compared considering the pathogen, timing, and key events. We defined different variants according to the pandemic period reported by Taiwan CDC database because not all the sample was send for identification of variants. (Taiwan CDC data: Pre-delta : April – June 2021, Delta : July 2021 – January 2022, Omicron : since February 2022)13,14 Continuous variables were expressed as mean (SD), and differences between groups were assessed using analysis of variance (ANOVA) or the Mann–Whitney U test based on the normality assumption. Discrete variables are presented as N (%), and differences between groups were evaluated using the chi-square test or Fisher’s exact test. Statistical analyses were performed using appropriate software, with p-values < 0.05 indicating statistical significance.
Ethical considerations
This study complied with the ethical principles outlined in the Declaration of Helsinki. The research protocol was approved by the Institutional Review Board (IRB: CMUH110-REC3-083) of the CMU Children’s Hospital. All methods were performed in accordance with relevant guidelines and regulations. Data were collected, reviewed, deidentified, and anonymized before analysis. The ethics committee of China Medical University Hospital waived the requirement for informed consent because of the anonymous nature of the data and the scientific purpose of the study.
Results
Demographic data and prevalence
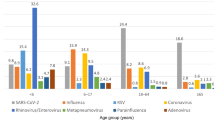
A total of 1983 pediatric patients under the age of 18 years were collected in this study. The overall positivity rate for respiratory infections was 84.2% (1670/1983) after excluding duplicated subjects. Table 1 showed an enumeration of the 10 most prevalent viruses detected. Among these, the top three aligned closely with the most recently published data15,16. Within this cohort, several noteworthy discoveries have emerged concerning the prevalence of various viruses in pediatric patients across distinct healthcare settings. Specifically, the admitted children were notably younger than their counterparts seeking treatment in the OPD or ED. In the group of admitted children, the rates of detection for HRV/EV was significantly lower (p = 0.004) than in ED and OPD. RSV has the highest detection rates at ED compared with OPD and AD. Notably, PIV 3 has emerged as the second most prevalent virus detected in this cohort, displayed no significant variance across the different patient source settings.
Furthermore, SARS-CoV-2 was detected more frequently in pediatric patients who visited the ED (p = 0.019). However, the incidence rates of PIV 4 and coronavirus 229E were higher in the OPD group (p = 0.005 and 0.002, respectively) (Fig. 1).
The viral co-detection associated with SARS-CoV-2 analysis showed Human Rhinovirus/enterovirus to be the most prevalent one, parainfluenza virus 3 the second, and respiratory syncytial virus (RSV) the third.
SARS-CoV-2 co-detection strains and their trends
Over a 2-year retrospective observational study, HRV/EV presented as the predominant detected strain, displaying a pronounced seasonal decline, particularly evident in July 2021. Furthermore, in 2022, this decline occurred earlier and shifted to June (Fig. 2, label 1). Concurrently, a noticeable increase in SARS-CoV-2 detection rate was observed during the same period (Fig. 2, label 2).
“The line chart (above) illustrates the seasonal correlation change between SARS-CoV-2 and the co-detected viruses during October 2021 to December 2021 when human rhinoviral prevalence was observed without SARS-CoV-2 or other viral detections. With the presence of SARS-CoV-2 since April 2022 (label 3), both Parainfluenza-3 and RSV became more prevalent than during the same period last year (2021). Labels: 1: Decline in Human rhinovirus. 2: Increase in SARS-CoV-2 detection rate. 3: Beginning of SARS-CoV-2 detection. 4: Parainfluenza-3 detection as a single pathogen. 5: RSV detection as a single pathogen. 6: Co-detection of Human rhinovirus with SARS-CoV-2. 7: Co-detection of Parainfluenza-3 with SARS-CoV-2. 8: Co-detection of RSV with SARS-CoV-2.
Among the 106 patients who tested positive for SARS-CoV-2, 45 (42%) had SARS-CoV-2 as the sole pathogen. The top three viral strain for co-detection was HRV/EV, PIV-3, and RSV, with 32 (30.2%) had co-detection of HRV/EV, 29 (24.5%) had co-detection of PIV 3, and 26 (24.5%) had co-detection of RSV (Fig. 1). Additionally, 5 cases had co-detection with adenovirus, and only 1 showed co-detection with influenza B. The peak emergence order of co-detection pairs was observed (HRV/EV ◊PIV-3◊ RSV) after July 2022, the Omicron period in Taiwan17,18.
The FA-RP analysis revealed multiple pathogen co-detection of up to five pathogens within a single patient, predominantly among those younger than 4 years. In children aged 5–9 years old, the maximum number of co-detected pathogens decreased to four, and further declined to three in the age group of 10–18 years.
Discussion
Although the positive rates were high, there is still approximately 16% pediatric patient with negative result among the 22-pathogen collected via FA-RP. The possible reason could be the routine use of FA-RP as an admission screen test to differential whether infected with SARS-CoV-2 or not during Taiwan SARS-CoV-2 pandemic without respiratory infection criteria oversight.
Based on our study, SARS-CoV-2 began to emerge in the results of the FA-RP in April 2022 (Fig. 2, label 3), coinciding with the local Omicron period in Taiwan. During this period, there was a noticeable upward trend in the incidence rates of both PIV 3 and RSV, whether detected as single pathogens or co-detected (Fig. 2, label 4,5,7,8). The occurrence of PIV3 and RSV peaks (Fig. 2, label 4, 5) was not observed during the winter of 2021 as it should be, maybe suggest the correlation with SARS-CoV-2 may be stronger than a mere seasonal factor. Meanwhile, co-detection including RSV and HRV/EV may increase illness severity among children under 5 years of age hospitalized with SARS-CoV-2 infection12. These findings highlight the importance of investigations to understand the co-detection between SARS-CoV-2 and other respiratory viruses and their potential impact on disease severity in pediatric.
The occurrence of SARS-CoV-2 co-detection with HRV/EV, PIV 3, or RSV showed sequential peaks, with HRV/EV peaking in September 2022 (Fig. 2, label 6), PIV 3 reaching its highest point in October 2022 (Fig. 2, label 7), and RSV showing its last peak in November 2022 (Fig. 2, label 8). This temporal relationship may be correlated with the concurrent pandemic pattern. However, the potential insight from this result lies in the observation that the co-detection of SARS-CoV-2 and HRV/EV, PIV3, and RSV was rarely observed or mentioned before the emergence of the omicron variant (since February 2022)17,18.
The hypothesis posited in the preceding document suggests that the viruses exhibited consistent patterns between years, called seasonality19. However, this theory encounters significant challenges when attempting to elucidate why SARS-CoV-2 infected pediatric population exhibit a diverse array of co-detection pattern with different SARS-CoV-2 variant. Although Japan study group (Arimura et al.) has reported the reduced likelihood of SARS-CoV-2 co-detection of parainfluenza virus, the research didn’t report on the basis of variant differential outcome. The present investigation showed different co-detection pattern among different SARS-CoV-2 variants, which predominant over Omicron pandemic. We identified characteristic patterns of SARS-CoV-2-associated co-detections, with a notable emphasis on the co-detection of HRV/EV, PIV3, and RSV alongside SARS-CoV-2. This association appears to hold heightened significance during the Omicron variant era compared to earlier SARS-CoV-2 variants, which contrast with previous studies. The simultaneous circulation of different variants may contribute to variations in viral co-detection, particularly in young children, warranting further investigation.
Limitations
This study had certain limitations that warrant consideration. First, owing to the deidentified nature of the results, it was challenging to comprehensively analyze the clinical outcomes and evaluate the severity of illness associated with multi-viral co-detection. Additionally, the inclusion of SARS-CoV-2 in the FA-RP was implemented only in January 2021 (as the 22nd pathogen in RP2.1), thus limiting the traceability of the data to this time point. Furthermore, the successful control of the SARS-CoV-2 pandemic in Taiwan led to negative results for SARS-CoV-2 in all 941 FA-RP cases in 2021, leaving a smaller dataset for analysis during this period. These constraints highlight the need for cautious interpretation and further investigation in future studies to gain a more comprehensive understanding of this subject.
Conclusion
This study proposes that the co-detection of PIV3 and SARS-CoV-2, as well as HRV and RSV, may play a more important role in the era of omicron than previous variants, such as delta, regarding clinical manifestations.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Park, S., You, J., Lee, J. & Park, E. Two case reports of Life-Threatening croup caused by the SARS-CoV-2 Omicron BA.2 variant in pediatric patients. J. Korean Med. Sci. 37 (24), e192 (2022).
Choi, Y. Y. et al. Croup as a manifestation of SARS-CoV-2 Omicron variant infection in young children. J. Korean Med. Sci. 37 (20), e140 (2022).
Sharma, S. et al. Croup associated with SARS-CoV-2: pediatric laryngotracheitis during the Omicron surge. J. Pediatr. Infect. Dis. Soc. 11 (8), 371–374 (2022).
Dasdemir, S., Uysal Yazici, M., Gudeloglu, E., Akkuzu, E. & Tezer, H. Croup as a previously unrecognized symptom of COVID-19 in infants. Pediatr. Infect. Dis. J. 41 (8), e332 (2022).
Xie, Y., Choi, T. & Al-Aly, Z. Postacute sequelae of SARS-CoV-2 infection in the Pre-Delta, delta, and Omicron eras. N Engl. J. Med. 391 (6), 515–525 (2024).
Choi, S. H. et al. Clinical characteristics and outcomes of children with SARS-CoV-2 infection during the Delta and Omicron Variant-Dominant periods in Korea. J. Korean Med. Sci. 38 (9), e65 (2023).
Narayanan, N. et al. COVID-19 is observed in older children during the Omicron wave in new York City. J. Emerg. Med. 64 (2), 195–199 (2023).
Acker, K. P. et al. Indications for hospitalization in children with SARS-CoV-2 infection during the Omicron wave in new York City. Child. (Basel) ;9(7), 1043. (2022).
Kenney, P. O., Chang, A. J., Krabill, L. & Hicar, M. D. Decreased clinical severity of pediatric acute COVID-19 and MIS-C and increase of incidental cases during the Omicron wave in comparison to the Delta wave. Viruses ;15(1), 180. (2023).
Lefchak, B. et al. Analysis of COVID-19-Related croup and SARS-CoV-2 variant predominance in the US. JAMA Netw. Open. 5 (7), e2220060 (2022).
Blasco, M. L. et al. Co-detection of respiratory pathogens in patients hospitalized with coronavirus viral disease-2019 pneumonia. J. Med. Virol. 92 (10), 1799–1801 (2020).
Agathis, N. T. et al. Codetections of other respiratory viruses among children hospitalized with COVID-19. Pediatrics ;151(2), e2022059037. (2023).
Shy, C. G. et al. Rapid control of a SARS-CoV-2 B.1.617.2 (Delta) variant COVID-19 community outbreak: the successful experience in Pingtung County of Taiwan. Int. J. Environ. Res. Public. Health ;19(3), 1421. (2022).
Liu, L. T. et al. Identification and analysis of SARS-CoV-2 alpha variants in the largest Taiwan COVID-19 outbreak in 2021. Front. Med. (Lausanne). 9, 869818 (2022).
Lee, J. K., Song, S. H., Ahn, B., Yun, K. W. & Choi, E. H. Etiology and epidemiology of croup before and throughout the COVID-19 pandemic, 2018–2022, South Korea. Child. (Basel) ;9(10). (2022).
Sim, J. Y. et al. Circulating pediatric respiratory pathogens in Taiwan during 2020: Dynamic change under low COVID-19 incidence. J. Microbiol. Immunol. Infect. 55 (6 Pt 2), 1151–1158 (2022).
Chen, Y. C. et al. Risk reduction analysis of mix-and-match vaccination strategy in healthcare workers during SARS-CoV-2 Omicron variant predominant period: A multi-center cohort study in Taiwan. Hum. Vaccin Immunother. 19 (2), 2237387 (2023).
Liu, L. T. et al. The identification and phylogenetic analysis of SARS-CoV-2 delta variants in Taiwan. Kaohsiung J. Med. Sci. 39 (6), 624–636 (2023).
Shirreff, G. et al. Seasonality and Co-Detection of respiratory viral infections among hospitalised patients admitted with acute respiratory Illness-Valencia region, Spain, 2010–2021. Influenza Other Respir Viruses. 18 (10), e70017 (2024).
Acknowledgements
Not applicable.
Funding
No funding was secured for this study.
Author information
Authors and Affiliations
Contributions
I-Hsin Tai and Chiung-Tzu Hsiao reviewed the medical records, analyzed, and interpreted the data, drafted the manuscript, and reviewed the medical. Chi-Hsiang Chu and Wan-Jane Tsai analyzed and interpreted the data. Han-Ping Wu designed and oversaw the study and revised the manuscript. Ying-Ju Chen contributed equally to this work. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Ethical approval and consent to participate
This study was approved by our Institutional Review Board of China Medical University Hospital (CMUH110-REC3-083).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tai, IH., Hsiao, CT., Chu, CH. et al. Co-detection of various viruses in SARS-CoV-2 in children with respiratory infections. Sci Rep 15, 8312 (2025). https://doi.org/10.1038/s41598-025-92878-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-92878-w