Abstract
Bird migration has fascinated natural historians and scientists for centuries. While the timing of migration is known to vary by species, population, sex, and individual, identifying the cause of this variation can be challenging. Here we investigate factors underlying migratory timing in a long-distance migratory bird, the Common Yellowthroat (Geothlypas trichas), using a population genomic approach. We begin by creating a map of genetic variation across geographic space (a “genoscape”) using lcWGS from across the breeding range. We then utilize genetic assays to assign 249 wintering and 1050 northward migrating birds to genetically distinct breeding populations. Additionally, we estimate the expected spring onset date in each predicted breeding region and calculate the remaining migratory distance for northward migrating birds. Our findings indicate that when population genetic structure is not a factor in the analysis, it appears that birds captured early in the season are migrating to breeding grounds where spring arrives later, which contrasts with prior research. However, when we incorporate population structure into our analysis, our results align with predictions, indicating that birds captured earlier in the season are indeed heading to breeding grounds where spring arrives earlier. Further analysis revealed that the disparity between results obtained with and without population genetic structure can be attributed to the fact that individuals from the western genetic group migrate three times the distance to the west, despite breeding at the same latitude. Our findings suggest that categorizing large numbers of migrating birds into genetically distinct groups can reveal population-specific patterns in migratory timing and shed light on the relative contributions of different selective forces responsible for the observed patterns.
Similar content being viewed by others
Introduction
The ability of millions of migratory birds to precisely time their departure from their tropical wintering grounds and journey thousands of miles north to their temperate breeding areas stands as one of the most intriguing mysteries of bird migration. Classic captive breeding experiments, where hybrid offspring display intermediate migratory timing compared to their parents, support the notion that timing is partially genetically controlled1,2,3,4,5,6,7,8. It is also well established that that the failure to synchronize migration with climate and resource peaks can have detrimental effects on an individual’s fitness9,10,11. Currently, our understanding of the role of genetics on migratory timing in birds is based mostly on captive breeding experiments, with limited examples of how these factors operate in the wild. Thus, there is a need for further research in natural settings to comprehensively assess the interplay between genetic, environmental, and demographic factors that shape migratory timing.
While most research to date has concentrated on species-scale patterns in timing, it is important to consider that the evolutionary forces shaping migratory timing, such as genetic drift and natural selection, likely operate at the population level, particularly in species with broad geographic distributions12. Each year, migrants from multiple breeding populations mix extensively on the wintering grounds in a situation akin to a common garden13,14. Despite this, individuals from distinct breeding populations often leave the same wintering areas at different times. Recent research indicates that population-specific variations in migratory timing are regulated in part by genetic alterations in genes responsible for governing biological clocks4,15,16,17. Additional factors that influence migratory timing include sex, social signals, weather, and wind patterns12,18. However, the interaction of these factors with population genetic differences is not yet comprehensively understood.
Despite the importance of considering population-specific differences in migratory timing, research in this domain has faced challenges due to the absence of suitable tools for distinguishing between genetically distinct populations12. Most migratory tracking studies separate individuals into populations based on loosely defined environmental regions rather than using genetics19,20,21. A particularly useful tool in this research has been the use of isotopes to predict breeding location and population; however, this method is typically restricted to coarse predictions along latitudinal bands22,23. Information on population genetic structure provides critical insights into the evolutionary dynamics between groups within a species that cannot be attained from geography alone. Genetic approaches allow for estimates of gene flow supporting the extent to which genetically distinct populations are also demographically independent. While early attempts to use population genetics to define migratory populations faced challenges related to low genetic marker resolution, the recent availability of genomic tools allows for the identification of even subtle patterns of genetic differentiation between populations24. Therefore, by harnessing the power of genome-wide genetic sequencing it is now possible to assess how environmental and demographic factors within genetically distinct populations influence migratory timing.
To implement a genomic framework to study migratory timing, the first step is to construct a map of genetically distinct breeding populations. Such a map has been referred to in previous work as a genoscape13. Once the genoscape has been established, it can be used as a baseline for assigning thousands of individuals from across the annual cycle to their most likely breeding population of origin. In addition to providing one of the most comprehensive assessments of migratory connectivity available, a genoscape-based approach can also be used to identify population-specific differences in migratory timing if samples are collected in a time series from migratory stopover locations. One of the few studies that has implemented this method to assess migration timing in a songbird, the Wilson’s Warbler (Setophaga petechia), found that genetically distinct populations migrate at different times, but the potential evolutionary explanations for these differences was not explored further13.
In this study we use a genoscape-based approach to investigate factors underlying migratory timing between genetically distinct populations of the Common Yellowthroat (Geothlypas trichas), a widespread and abundant Nearctic-Neotropical migrating bird25. To explore how the inclusion of population genetic structure affects our capability to identify factors influencing migratory timing, we integrate population assignment information with data on average date of spring onset at the predicted breeding site and remaining migratory distance, and approximate wintering area, as well as relevant demographic data (in particular, sex). We then estimate their relative influence on the migratory passage date across the coast of the Gulf of Mexico (i.e. the Gulf Coast). Specifically, we seek to address the following research questions:
1) How are Common Yellowthroat populations connected across time and geographic space? 2) Do genetically distinct populations migrate at different times? 3) How do the date of spring onset on the breeding grounds, the remaining migratory distance, and sex influence migratory timing within genetically distinct populations? 4) Does our interpretation of the factors underlying migratory timing differ with and without including population genetic structure?
Results
Genoscape construction
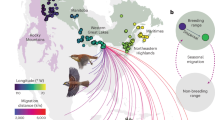
Initial population genomic analysis of 148 breeding individuals after filtering out 2 related individuals at 6,721,176 SNP loci revealed support for genetic differentiation between five groups including: West (green), Atlantic (purple), Midwest (blue), Southwest (orange), and a weak cluster of primarily resident breeders in Central California (red; SI Figure 2). While our STRUCTURE analysis revealed that a K value of 2, 3, 4, and 5 populations were biologically realistic hypotheses for the number of groups within the species (SI Figure 3), the goal of our analysis was not to find the most likely value of K, but to identify spatially explicit genetically distinct groups that could be tracked across the full annual cycle, as in fisheries stock management26. Thus, we set the number of groups to 5 based on concordance between spatially informative genetic clusters identified in the genome wide PCA (SI Figure 2), the STRUCTURE runs based on the 96 SNP loci (Fig. 1), and the power to assign individuals to groups at K = 5 using rubias (Table S1). Evidence of admixture between the Midwest (blue) and the Atlantic Coast (purple) was detected in New York, Ontario, and Quebec samples. Furthermore, genetic differentiation in the resident Central California population (Fig. 1, red) was weak, suggesting high admixture with the surrounding Western migrants. Due to this weak differentiation, a K value of 4 was used in all subsequent assignments of migratory individuals, with the California population being integrated into the broader West (green) population.
The Common Yellowthroat Genoscape. The top panel depicts the STRUCTURE analysis with each line representing the posterior probability of assignment of each individual belonging to one five genetic groups. Numbers indicate the locations of sampling sites on the map. The bottom panel depicts a spatially explicit map of population structure clipped to the breeding range (eBird 2021), with the intensity of color representing the certainty in the assignment of individuals from a particular region belonging to a genetic group. Circles on the wintering represent sampled individuals color coded by the genetic assignments according to rubius. The wintering dots are jittered to facilitate visualization.
Genetic assignments of population, breeding location, and sex
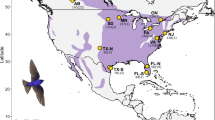
Assignment of wintering individuals to breeding groups using rubias suggested extensive mixing among breeding populations on the wintering grounds. Western breeding birds (Southwest and West) were found wintering primarily from Central Mexico, west to Southern Baja, while eastern breeding birds (Midwest and Atlantic Coast) were primarily restricted to the eastern part of the wintering range (from Central Mexico to Puerto Rico; Fig. 1). Population-level assignments of 509 migrant individuals from Johnson’s Bayou, Louisiana, identified 81% from the Midwestern, 16% from the Atlantic, 3% from the Western, and none from the Southwestern population (n = 412, 81, 16, 0, respectively). Of the 541 migrants sampled from Mad Island, Texas, 79% were from the Midwestern, 9% from the Atlantic, 10% from the Western, and 1% were from the Southwestern populations (n = 428, 51, 56, 6, respectively). All individuals included in these totals were assigned to their genetic population with 80% or greater confidence by OriGen (see Methods) and the vast majority of these predictions (96.5%) fell within (coincided with) the broader population-specific geographic region determined by rubias, giving additional confidence in the results from both assignment methods (Fig. 2A; SI Figure 4). The 3.5% that were not concordant were largely due to misassignments between the closely related Atlantic and Midwest populations. Of migrants sexed in the field at Johnson’s Bayou, LA, 352 were male, 129 female, and 28 undetermined. Migrants from Mad Island, TX, were genetically sexed and determined to consist of 209 males, 279 females, and 47 of undetermined sex. Sex breakdown by population is summarized in Table S1, and roughly aligns with species-wide sexual ratios. In total, 969 migrant individuals had confident assignment of both genetic population and sex; these individuals were used in all subsequent analyses.
(A) Predicted breeding locations of each captured migrant. Lines are drawn between the capture location and the location with the highest probability of breeding as calculated by OriGen (note that this single point is not what is used in analyses). Line color shows population assignment by rubius. (B) Time series of the proportion of migrants through each capture site, separated by week and population. At both the Louisiana (top) and Texas (bottom) capture sites, the Atlantic population showed peak migration roughly two weeks earlier than the Midwest population’s peak. Western individuals are not included in the Louisiana chart due to low sample size. (C) Spring onset dates across the ranges of each respective population, with mean spring onset indicated by dotted vertical line. The breeding ranges of the Atlantic and Midwest populations experience nearly identical spring onset, while spring arrives much later across the Western breeding range.
Models of migratory timing
Two linear mixed models of migratory timing, one without genetic population data and one with, were created, summarized in Tables S2 and S3. Each included sex, estimated spring onset, latitude-corrected migration distance (hereafter referred to as LCMD; see methods), and capture site as predictor variables. Both models showed a significant difference between sexes and a weak effect of capture site. Specifically, in both models, males migrated approximately four days before females (no-pop model: estimate = − 3.51, SE = 0.74, t = − 4.74, p < < 0.001; pop model: estimate = − 4.13, SE = 0.73, t = − 5.65, p < < 0.001; Tables S2 and S3), and migrants arrived in Texas a quarter of a day later than in Louisiana (no-pop model: estimate = 0.3, SE = 0.73, t = 0.42, p = 0.68; pop-model: estimate = 0.19 SE = 0.72, t = 0.26, p = 0.79; Tables S2 and S3). In the model with no population data (R2 = 0.27; df = 964; Table S3), both spring onset and LCMD were predicted to have weak (but significant) negative relationships with migration date. In particular, migrants were predicted to migrate a day earlier for every fourteen days spring arrived later at their predicted breeding site (estimate = − 0.069, SE = 0.024, t = − 2.88, p = 0.004), and to migrate a day earlier for each additional 62.5 km of LCMD they had remaining to travel (estimate = − 0.016, SE = 9.2e−4, t = − 17, p < < 0.001).
In the full model (R2 = 0.31, df = 958; Table S2) there was a significant negative relationship between timing and LCMD in both the Midwest and West populations. Specifically, birds in these regions migrated a day earlier for each additional 81.3 and 97.1 km traveled, respectively (Table S2). There were significant differences in the timing of migration between the Midwest (mean migration ordinal date 113.5 ± 10.59 [sd]) and Atlantic (99.14 ± 11.83 [sd]) populations, with the Atlantic population migrating approximately two weeks earlier than the Midwest (Fig. 2B; p < < 0.001). This pattern was consistent between migrating birds sampled at the Louisiana and Texas sites and across years (SI Figure 5). The Western population also migrated significantly earlier (96.83 ± 14.89 [sd]) than the Midwest population with much greater variation in migration dates relative to the number of samples (Fig. 2B; p < < 0.001). The Western population did not differ significantly in timing from the Atlantic (p = 0.99). In contrast to migration passage times, the average date of spring onset (as determined by random sampling of spring onset across the population ranges; Fig. 2C) was not significantly different between the Atlantic and the Midwest (p = 0.95, t = 0.276, df = 225.95), while average date of spring in the West was significantly later and more variable (t = − 15.778, df = 702.94, p < 0.001). Mean LCMD differed significantly between populations, with individuals from the Atlantic (1012 ± 215 km) and Western (993 ± 434 km) breeding populations traveling approximately three times the distance of Midwestern breeding birds (335 ± 267 km; compared to Atlantic: p < < 0.001; compared to West: p < < 0.001; West compared to Atlantic: p = 0.35; Fig. 4).
Discussion
Here, we built a map of population genetic structure across the breeding and wintering range of the Common Yellowthroat to identify the movement patterns of genetically distinct groups across time and space. We find that genetically distinct groups migrate across the Gulf Coast at different times and that the influence of sex, breeding ground phenology, and migration distance on migratory passage dates vary when accounting for or excluding population genetic structure. These findings emphasize the importance of considering population genetic structure when investigating the factors that influence migratory timing, particularly in species with extensive geographical distributions that experience diverse selective pressures.
Here, we mapped the breeding and wintering areas for each genetically distinct population of the Common Yellowthroat to contextualize our data on migratory passage dates across the Gulf Coast within the species’ full annual cycle. We found that the strongest genetic differentiation on the breeding grounds existed between eastern and western populations, consistent with earlier findings27. The observed weaker genetic differentiation within eastern and western groups confirmed some previous research but also uncovered previously unknown population subdivisions. Specifically, in the west, we confirmed the previously recognized split between the Southwest and West populations. Within the east we identified previously unrecognized genetic differentiation between Atlantic Coast and Midwest populations. We also identified a new genetic split between resident populations in California and other western migratory populations. Of the 5 genetic groups identified by our analysis of population structure across the breeding range, only the Atlantic, Midwest, and Western groups passed through the Gulf Coast during migration in large numbers, and as such our conclusions about migratory timing are limited to those populations.
By employing genetic assignments that link wintering birds to their breeding populations, we also gained some limited insight on how approximate wintering location may have affected our results. We found that Midwest and Atlantic Coast breeders predominantly winter in eastern wintering sites, while Southwest, West, and California breeders’ winter in western sites. In the eastern wintering region, some mixture occurred between Atlantic and Midwest breeders in places like eastern Mexico and Jamaica, with a greater number of Atlantic breeders overwintering in the Southeastern United States. Similarly, in the western region, substantial mixing between West and Southwest genetic groups took place in locations such as Baja, western Mexico, and Central Mexico. A major limitation of our dataset is that we do not know the specific departure times of our wintering birds or the exact wintering location of birds passing through the Gulf Coast. However, the extensive mixing between populations, particularly the Midwest and Atlantic, on the wintering grounds suggests that distinct migratory passage dates along the Gulf Coast are likely not entirely due to these populations wintering in separate regions. Future research combining a genoscape-based approach with detailed information on timing and location from external tracking devices like GPS tags would greatly enhance the ability to understand the combined role of population structure and wintering location on the timing of spring passage dates.
Ornithologists and students of natural history have long observed that unique waves of migratory birds come through migration monitoring stations at varying times9, yet the underlying reasons for these timing disparities is usually unclear. In this study, we employ genetic assays to identify the breeding population of origin for > 1000 birds migrating through the Gulf Coast and, in so doing, uncover clear and distinct patterns of migratory timing within each group. Specifically, we show that Western, Midwest, and Atlantic experience peak migration at different times. Western and Atlantic breeders are the first to migrate through the Gulf Coast, followed by Midwestern breeders approximately 14 days later (Fig. 2B). This trend remains consistent across different sample locations and over multiple years (SI Figure 5). Additionally, in line with previous migratory bird research28, we observed that males migrate before females (SI Figure 6). Collectively, these findings provide evidence that distinct waves of migrants observed at bird monitoring stations can be attributed, at least partially, to genetically distinct populations migrating at different times.
Population-specific differences in migratory passage dates are thought to result from a combination of genetic, environmental, and demographic factors, but disentangling the relative contribution of each can be difficult without information on population genetic structure. Here we show that the crucial determinants of migratory passage dates across the Gulf Coast vary when accounting for or excluding population genetic structure. Notably, when considering all individuals within the species, our results suggested that those individuals captured earlier in the season were heading to breeding grounds where spring onset occurred later (Fig. 3). This outcome was confusing because it challenges the widely held notion that spring migration timing is driven partly by the need to align arrival dates with spring’s onset on the breeding grounds9,11. However, when we analyzed genetically distinct populations separately, our results aligned with expectations in two of the three populations, indicating that early migrants travel to areas where spring arrives earlier (Fig. 3). Overall, the ability to include population genetic structure into our analysis helped clarify factors underlying the timing of migratory passage.
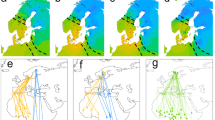
The relationship between migration date and spring onset (A and B) and latitude-corrected migration distance (C and D), with and without population as a factor. Trendlines represent linear regression predicted using full models of timing with 95% confidence intervals. Without considering population, spring onset and latitude-corrected migration distance were strong predictors of timing with negative relationships (p < < 0.001 in both cases; A and C), with earlier migrants traveling to locations with later spring onset and a much greater distance east/west of the capture site. When the data was broken up by population however, the relationship between spring onset and timing changed to a positive one in both the Midwest and Atlantic populations (B). Similarly, at the population scale, latitude-corrected migration distance flipped its relationship in the Atlantic population.
Further analysis within genetically distinct groups revealed that latitude-corrected migration distance also significantly influences spring passage dates. Importantly, our joint analysis revealed that the observed counterintuitive relationships between spring onset and migratory passage dates in part result from the confounding influence of the longitudinal (rather than latitudinal) travel distance remaining for birds from different genetic groups. Further, LCMD appears to explain the consistent difference in timing between the Atlantic and Midwest populations, where Atlantic migrants each year migrate roughly two weeks earlier. Atlantic breeders migrate to locations where the average date of spring is not significantly different from that of Midwestern breeders (Fig. 4B), but Atlantic breeders travel approximately three times further east (Fig. 4A). Similarly, the fact that birds from the West migrate through first even though the average date of spring across the West occurs significantly later than in either the Midwest or Atlantic (Fig. 4B) is likely explained by the fact that the average Western migrants must travel significantly farther to the west relative to Midwestern birds.
Average migration date within each genetically distinct population versus (A) the latitude-corrected migration distance and (B) the predicted spring onset on the breeding grounds. Dots represent mean values, while bars represent one standard deviation. Both plots show that the Atlantic and West migrate roughly two weeks earlier, on average, than the Midwest population. However, while the Atlantic and Midwest largely overlap in their spring onset distribution (with spring arriving earlier in these breeding ranges than the more northerly West), there is a significant difference in remaining migration distance between the two, as the Atlantic population must migrate nearly double the latitude corrected migration distance of the Midwest.
Our work highlights the importance of a genoscape-based approach for understanding factors underlying migratory timing. Our research revealed consistent differences in migratory timing between genetically distinct populations and demonstrates how the inclusion of information on population genetic structure clarifies the importance of spring onset in determining migratory passage date. Future work linking specific genes, their expression, and the regulatory pathways involved with the observed population specific patterns would help improve our understanding of the ultimate mechanisms controlling migratory timing within genetically distinct groups. In addition, landscape genomic analysis may help reveal the links between observed genetic difference between populations and natural selection across the annual cycle. In sum, we believe this study provides a significant step forward in our ability to distinguish factors regulating migratory timing, relationships that are likely to become increasingly dynamic under future climatic change.
Methods
Ethics
Sampling of live specimens was conducted using CSU Veterinary approved IUCAC protocols (#5449) and were carried out following ARRIVE guidelines.
Sampling and DNA extraction
Breeding and wintering birds were captured with mist-nets at 37 sites across the breeding range (Table 1) and 41 sites across the wintering range (SI Table 1), while migrants were captured throughout spring migration at two long-term migration monitoring stations on the Gulf Coast–Johnson’s Bayou, Louisiana (Johnson’s Bayou, 29° 45′ N, 93° 37′ W) during 2004–2012 and Clive Runnells Family Mad Island Marsh Preserve, Texas (Mad Island, 28° 37′ N, 96° 6′ W) during 2013–2019 (Fig. 2A) 29,30. In total, 271 breeding, 249 wintering, and 1050 migrating birds were sampled. Upon capture, 20 uL of blood from the sub-brachial wing vein or one tail feather from each bird was collected for sexing and genotyping. Blood samples were suspended in Queen’s lysis buffer and stored at − 20 °C until processing. Feather samples were similarly stored in coin envelopes at − 20 °C until extraction. DNA was extracted from migrant feathers and blood using Qiagen DNeasy Blood and Tissue extraction kits (Qiagen Inc., Valencia, CA, USA) following the standard tissue extraction protocol. DNA quantification was done using a Qubit dsDNA HS Assay kit (Thermo Fisher Scientific) prior to genomic library preparation.
Genoscape construction
We used a modified version of Illumina’s Nextera Library Preparation protocol to prepare whole-genome sequencing libraries and pooled the libraries prior to sequencing on two Illumina HiSeq4000 (Illumina) lanes (see Supplementary Methods). Reads were processed using Genome Analysis Toolkit (GATK) following the Best Practices Guide31 and aligned to a Common Yellowthroat reference genome (accession number: GCA_009764595.1) using hisat232. Ninety-six of the top 300 most highly differentiated SNPs between the 5 major groups identified in a preliminary population structure analysis were converted into SNP Assays for subsequent population assignment (see Supplementary Methods). We then tested the effectiveness of our markers for population assignment and filled in sampling gaps across the breeding distribution by genotyping all individuals in our original genomic panel as well as an additional 121 breeding individuals across the range using the resulting 96-SNP panel. Samples with missing genotypes at more than 10% of SNP assays were filtered out from subsequent population structure analysis.
To assess population structure across the breeding region, we used the admixture model in structure33, a model-based clustering method. Using the locprior model with sampling locations as prior information, uncorrelated allele frequencies, a burn-in period of 50,000, and total run length of 150,000 we ran 5 iterations of each assumed number of genetic clusters (K), where K ranged from 1 to 633. To understand how patterns of population structure were distributed across geographic space, we visualized the posterior probability of group membership estimates from structure as transparency levels of different colors overlaid upon a base map from Natural Earth (naturalearthdata.com) and clipped to a map of the Common Yellowthroat breeding range34, making use of the R packages sp, RGDAL, and raster35,36,37. We scaled the transparency of colors within each distinguishable group, so that the highest posterior probability of membership in the group according to structure is opaque and the smallest is transparent.
Population and individual-based assignments
To assign wintering and migrating individuals to breeding areas, we used two different methods—one to assign individuals to genetically distinct breeding populations as determined by the genoscape and the other to make specific predictions of breeding location for each individual. We leveraged the genoscape to assign individuals sampled during winter and migration to genetically distinct populations using the R package rubias38. The reference breeding bird panel excluded the southern California populations due to the ambiguous assignment resulting from admixed ancestry between the West and Central California breeding units. Wintering and migrating individuals with posterior probability of assignment less than 80% were considered as uncertain and were not used for the remaining analyses. To give a more precise prediction of the probable breeding destinations of migrating individuals within genetically distinct groups, we then used the R package OriGen39. We first created a continuous allele frequency surface across the breeding range using allele frequencies from 271 individuals of known breeding location and the program-specific parameters used in related studies (MaxGridLength = 70, RhoParameter = 10)39,40. We then estimated the probability that each migrant individual belonged to a 1.5° longitude and 0.5° latitude grid cell based upon the given allele frequency distributions across the breeding range. The grid cells were trimmed to the breeding range of the Common Yellowthroat and probabilities were rescaled to sum to 1. The resulting rasters of the probability of breeding in each cell of the range were used to weight individual measures of spring onset and migration distance (see Spring onset and migration distance).
Genetic identification of sex
This species, like many Nearctic-Neotropical migrants, exhibits sex specific migratory timing, with males typically migrating earlier than females41,42. To account for this difference in migration timing, we sexed birds based on morphological features in the hand (samples from Johnson’s Bayou) or using genetic markers to sex birds in the lab (samples from Mad Island). Genetic sexing was performed using methods outlined in Fridolfsson and Ellegren43 and elaborated on in Sehgal et al.44. The CDH1 gene on the avian Z chromosome was amplified using the primers 2550F and MSZ1R. Fifteen microliter reaction mixtures were created containing 2.74 μL of sample DNA, 1.36 μL of each primer at 10 μM, 0.68 μL of 10 mg/mL BSA, 1.36 μL of Q solution (Qiagen), and 7.5 μL of Qiagen Master Mix. The thermocycling profile consisted of 15 min of initial denaturation at 94° C, followed by 43 cycles of 30 s denaturation at 94°, 30 s of annealing at 50°, 45 s of extension at 72° and a final extension period of 10 min at 72°. The results were visualized using Invitrogen 2% Agarose E-gels, with two DNA bands representing females and a single band representing males.
Spring onset and migration distance
To determine the average date of spring onset across recent history, we used the First Leaf Historical Spring Indices generated by the National Phenology Network45. This dataset uses historical leaf-out data to model the “start of spring” for all points in the US and Canada between 1880 and 201346. A 20-year average of spring onset (ordinal date) for each point in North America was created by averaging across rasters from between 1994 and 2013. Additionally, the average spring onset across the breeding range of each genetically distinct population was estimated by randomly sampling 1000 points in each region and averaging across the 20-year spring onset for each grid cell. We created histograms visualizing the distribution of average leaf out date over time by population using density plots in ggplot247.
To determine the average timing of spring onset at the breeding destination and the approximate migration distance for each migrant, we multiplied the probability of breeding in each cell (as determined by OriGen) by the average spring onset in that cell and these values were summed across the range to create a weighted average for each individual. The migration distance was then also calculated using the individual rasters of breeding probability by weighting the great circle distances48 between the capture location and each grid cell by the probability of breeding in that cell. Because spring phenology and latitude are highly correlated and our objective was to assess the relative influence of migration distance and spring onset, we further transformed the data to focus on the component of distance remaining after accounting for travel in the latitudinal direction. We refer to the resulting value as the latitude-corrected migration distance (LCMD), which is calculated by subtracting the distance between the capture site and the predicted breeding latitude at a point due north of the capture site from the total remaining migration distance (SI Figure 1). This essentially represents the additional distance that birds are traveling in an eastern or western direction. All distances were estimated using the R package geosphere49.
Statistical analyses
To assess the relative influence of each predictor variable (including genetic population, estimated spring phenology, LCMD, sex, and capture site) on migratory timing, a linear model was created with capture date as the response variable. To allow for differential relationships with spring phenology and LCMD for each genetic population, an interaction term was included for each. A second linear model without genetic population data was created to assess the importance of including genetic data on the resulting model. The means of key variables (migration date, spring onset, and latitude-corrected migration distance) between pairs of populations were compared using pairwise Wilcoxon tests (pairwise.wilcox.test() in base R) with a Bonferroni correction to p values due to the number of variables in the model. Trend lines were fitted to graphs of spring onset and LCMD versus capture date by using the predict() function of base R in conjunction with our models.
Data availability
All data used in this manuscript, as well as all code for data analysis and figure creation, is publicly available on Dryad at the following link: http://datadryad.org/stash/share/xmCoXi1USvuuM5-83fhuRsI3XF84k43qv1jxi5PBoa0.
References
Berthold, P., Wiltschko, W., Miltenberger, H. & Querner, U. Genetic transmission of migratory behavior into a nonmigratory bird population. Experientia 46, 107–108 (1990).
Gwinner, E. Circadian and circannual programmes in avian migration. J. Exp. Biol. 199, 39–48 (1996).
Moller, A. P. Heritability of arrival date in a migratory bird. Proc. R. Soc. Lond. B Biol. Sci. 268, 203–206 (2001).
Berthold, P. Genetic basis and evolutionary aspects of bird migration. in Advances in the Study of Behavior vol. 33 175–229 (Elsevier, 2003).
Pulido, F. Phenotypic changes in spring arrival: Evolution, phenotypic plasticity, effects of weather and condition. Clim. Res. 35, 5–23 (2007).
Hedenström, A. Adaptations to migration in birds: Behavioural strategies, morphology and scaling effects. Philos. Trans. R. Soc. B Biol. Sci. 363, 287–299 (2008).
Robson, D. & Barriocanal, C. Ecological conditions in wintering and passage areas as determinants of timing of spring migration in trans-Saharan migratory birds: Ecological conditions and timing of spring migration. J. Anim. Ecol. 80, 320–331 (2011).
Ramenofsky, M., Cornelius, J. M. & Helm, B. Physiological and behavioral responses of migrants to environmental cues. J. Ornithol. 153, 181–191 (2012).
Smith, R. J. & Moore, F. R. Arrival timing and seasonal reproductive performance in a long-distance migratory landbird. Behav. Ecol. Sociobiol. 57, 231–239 (2005).
Møller, A. P., Arriero, E., Lobato, E. & Merino, S. A meta-analysis of parasite virulence in nestling birds. Biol. Rev. 84, 567–588 (2009).
Gienapp, P. & Bregnballe, T. Fitness consequences of timing of migration and breeding in cormorants. PLoS ONE 7, e46165 (2012).
Pulido, F. The genetics and evolution of avian migration. BioScience 57, 165–174 (2007).
Ruegg, K. C. et al. Mapping migration in a songbird using high-resolution genetic markers. Mol. Ecol. 23, 5726–5739 (2014).
Dossman, B. C., Rodewald, A. D., Studds, C. E. & Marra, P. P. Migratory birds with delayed spring departure migrate faster but pay the costs. Ecology https://doi.org/10.1002/ecy.3938 (2022).
Bazzi, G. et al. Clock gene polymorphism and scheduling of migration: a geolocator study of the barn swallow Hirundo rustica. Sci. Rep. 5, 12443 (2015).
De Greef, E., Suh, A., Thorstensen, M. J., Delmore, K. E. & Fraser, K. C. Genomic architecture of migration timing in a long-distance migratory songbird. Sci. Rep. 13, 2437 (2023).
Le Clercq, L. et al. Time trees and clock genes: A systematic review and comparative analysis of contemporary avian migration genetics. Biol. Rev. 98, 1051–1080 (2023).
Cooper, N. W. et al. Atmospheric pressure predicts probability of departure for migratory songbirds. Mov. Ecol. 11, 23 (2023).
Stanley, C. Q. et al. Connectivity of wood thrush breeding, wintering, and migration sites based on range-wide tracking. Conserv. Biol. 29, 164–174 (2015).
Taylor, C. M. & Stutchbury, B. J. M. Effects of breeding versus winter habitat loss and fragmentation on the population dynamics of a migratory songbird. Ecol. Appl. 26, 424–437 (2016).
Knight, S. M. et al. Constructing and evaluating a continet-wide migratory songbird network across the annual cycle. Ecol. Monogr. 88, 445–460 (2018).
Bowen, G. J., Wassenaar, L. I. & Hobson, K. A. Global application of stable hydrogen and oxygen isotopes to wildlife forensics. Oecologia 143, 337–348 (2005).
Studds, C. E., Wunderle, J. M. & Marra, P. P. Strong differences in migratory connectivity patterns among species of Neotropical-Nearctic migratory birds revealed by combining stable isotopes and abundance in a Bayesian assignment analysis. J. Biogeogr. 48, 1746–1757 (2021).
Savolainen, O., Lascoux, M. & Merilä, J. Ecological genomics of local adaptation. Nat. Rev. Genet. 14, 807–820 (2013).
Guzy, M. J., Ritchison, G. & Rodewald, P. G. Common Yellowthroat (Geothlypis trichas). In Birds of the world (eds Billerman, S. M. et al.) (Cornell Lab of Ornithology, 2020). https://doi.org/10.2173/bow.comyel.01.
McKinney, G. J. et al. Dense SNP panels resolve closely related Chinook salmon populations. Can. J. Fish. Aquat. Sci. 77, 451–461 (2020).
Escalante, P., Márquez-Valdelamar, L., de la Torre, P., Laclette, J. P. & Klicka, J. Evolutionary history of a prominent North American warbler clade: The Oporornis-Geothlypis complex. Mol. Phylogenet. Evol. 53, 668–678 (2009).
Morris, S. R., Pusateri, C. R. & Battaglia, K. A. Spring migration and stopover ecology of common Yellowthroats on Appledore Island, Maine. Wilson Bull. 115, 64–72 (2003).
Cohen, E. B. et al. Spring resource phenology and timing of songbird migration across the gulf of Mexico. In Phenological synchrony and bird migration: changing climate and seasonal resources in North America (eds Wood, E. M. & Kellermann, J. L.) 78–97 (CRC Press, 2015). https://doi.org/10.1201/b18011-8.
Cohen, E. B. et al. The strength of migratory connectivity for birds en route to breeding through the Gulf of Mexico. Ecography 42, 658–669 (2019).
Van der Auwera, G. A. et al. From FastQ to high-confidence variant calls: The genome analysis toolkit best practices pipeline. Curr. Protoc. Bioinforma. https://doi.org/10.1002/0471250953.bi1110s43 (2013).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pritchard, J. K. & Wen, W. Documentation for structure software: Version 2. (2002).
Sullivan, B. L. et al. eBird: A citizen-based bird observation network in the biological sciences. Biol. Conserv. 142, 2282–2292 (2009).
Bivand, R. S., Pebesma, E. & Gomez-Rubio, V. Applied spatial data analysis with R (Springer, 2013).
Bivand, R. S., Keitt, T. & Rowlingson, B. rgdal: Bindings for the ‘Geospatial’ Data Abstraction Library. (2023).
Hijmans, R. J. raster: Geographic Data Analysis and Modeling. (2023).
Moran, B. M. & Anderson, E. C. Bayesian inference from the conditional genetic stock identification model. Can. J. Fish. Aquat. Sci. 76, 551–560 (2019).
Rañola, J. M., Novembre, J. & Lange, K. Fast spatial ancestry via flexible allele frequency surfaces. Bioinformatics 30, 2915–2922 (2014).
Bay, R. A. et al. Genetic variation reveals individual-level climate tracking across the annual cycle of a migratory bird. Ecol. Lett. 24, 819–828 (2021).
Morbey, Y. E. & Ydenberg, R. C. Protandrous arrival timing to breeding areas: A review. Ecol. Lett. 4, 663–673 (2001).
Stewart, R. L. M., Francis, C. M. & Massey, C. Age-related differential timing of spring migration within sexes in passerines. Wilson Bull. 114, 264–271 (2002).
Fridolfsson, A.-K. & Ellegren, H. A simple and universal method for molecular sexing of non-ratite birds. J. Avian Biol. 30, 116 (1999).
Sehgal, R. N. M., Jones, H. I. & Smith, T. B. Molecular evidence for host specificity of parasitic nematode microfilariae in some African rainforest birds. Mol. Ecol. 14, 3977–3988 (2005).
USA National Phenology Network. Historical Annual Spring Indices (1880–2013), First Leaf, Years: 1994–2013. USA-NPN (2017).
Schwartz, M. D., Ault, T. R. & Betancourt, J. L. Spring onset variations and trends in the continental United States: Past and regional assessment using temperature-based indices. Int. J. Climatol. 33, 2917–2922 (2013).
Wickham, H. Ggplot2: Elegant graphics for data analysis (Springer-Verlag, 2016).
Alerstam, T. et al. Great-circle migration of arctic passerines. Auk 125, 831–838 (2008).
Hijmans, R. J. Introduction to the “geosphere” package (Version 1.5-14). 19 (2021).
Acknowledgements
This study was funded by an NSF CAREER award to K.R. (008933-00002), a National Geographic grant to K.R. (WW-202R-170), a UC MEXUS grant to R.F. and K.R. (CN-19-120) and a California Energy Commission grant to T.B.S. and K.R. (EPC-15-043). Sample collection in TX was supported by the Helen DuBois Family and the Conoco Phillips Charitable Investment Global Signature Program and in Louisiana by NSF support to F. Moore (IOS 0078189 and 0844703, DEB 0554754). We thank the Institute for Bird Populations, the many MAPS and MoSI station operators, and individual contributors for dedicating their time and/or donating samples, including, K. Acencio, C. Burelos, A. Ulin, M. Calcao, B. Keith, L. Helton, A. Patterson, K. Slankard, T. Kita, K. Allen, M. Fylling, J., D. Downy, M. Westbrook, C. Hathcock, J. Port, D. Grosshuesch,B. Shepherd, F. Schaffner, M.S. Wilson, C. McLure, A. Finfera, W. Sakai, K. Gordon, M. Voisine, D. Humple, S. Koenig, J.K. Allen, M. Kimura, B. Mila, J. Deppe, C. Arizmendi, F.R. Vasquez, and A.L. Sandoval. We thank the many students and field assistants for their help with bird banding and sample collection, including, C. Crow, Sahid RB, D. Palma, and A. Echeverría. For logistical support on the wintering grounds, we thank S. Arriaga, M. Grosselet, and A. Dorantes. For logistical support at Mad Island, we thank Steven Goertz The Nature Conservancy of Texas. We thank the following individuals for reviewing and providing comments on a previous version of this manuscript: E. Anderson, K. Horton, the Ruegg Lab, the Funk Lab, and the Cohen Lab.
Author information
Authors and Affiliations
Contributions
This project was conceptualized by Taylor Bobowski and Kristen Ruegg. Sample collection was primarily led and conducted by Emily Cohen, Rafael Hernandez, and Ingrid Tello-Lopez. Laboratory work and data analysis for the creation of the genoscape was conducted by Christen Bossu and Teia Schweizer, with input from Kristen Ruegg and Thomas Smith. Laboratory work for migrant and wintering samples was conducted by Taylor Bobowski and Teia Schweizer. Data analysis was conducted by Taylor Bobowski, with significant input from Christen Bossu and Richard Feldman. The manuscript was written by Taylor Bobowski, Kristen Ruegg, and Christen Bossu. All authors contributed to manuscript revision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bobowski, T.J., Bossu, C., Rueda-Hernández, R. et al. Factors underlying migratory timing of a seasonally migrating bird. Sci Rep 15, 8527 (2025). https://doi.org/10.1038/s41598-025-93442-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-93442-2