Abstract
Multi-modal medical image fusion (MMIF) aims to integrate complementary information from different modalities to obtain a fused image that contains more comprehensive details, providing clinicians with a more thorough reference for diagnosis. However, most existing deep learning-based fusion methods predominantly focus on the local statistical features within images, which limits the ability of the model to capture long-range dependencies and correlations within source images, thus compromising fusion performance. To address this issue, we propose an unsupervised image fusion method guided by stochastic structural similarity (S3IMFusion). This method incorporates a multi-scale fusion network based on CNN and Transformer modules to extract complementary information from the images effectively. During the training, a loss function with the ability to interact global contextual information was designed. Specifically, a random sorting index is generated based on the source images, and pixel features are mixed and rearranged between the fused and source images according to this index. The structural similarity loss is then computed by averaging the losses between pixel blocks of the rearranged images. This ensures that the fusion result preserves the globally correlated complementary features from the source images. Experimental results on the Harvard dataset demonstrate that S3IMFusion outperforms existing methods, achieving more accurate fusion of medical images. Additionally, we extend the method to infrared and visible image fusion tasks, with results indicating that S3IMFusion exhibits excellent generalization performance.
Similar content being viewed by others
Introduction
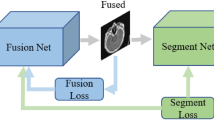
In clinical practice, image scans are essential screening methods to obtain the imaging characteristics of the diseased tissue. For instance, single-photon emission computed tomography (SPECT) effectively detects the activity and metabolic strength of human tissue cells by injecting a radioactive tracer into the patient’s body and analyzing the emitted light. Another well-known technique is magnetic resonance imaging (MRI), which uses electromagnetic waves to characterize the soft tissues of the body. For tissues and organs with higher density, such as bones, computed tomography (CT) is commonly used. CT employs a narrow beam of X-rays to generate cross-sectional images of the body (slices) at a specific thickness. Unfortunately, each of these imaging techniques has its limitations when used independently. SPECT monitors the metabolic activity of tissue cells but provides blurred images, often losing information about tissue structure. Magnetic resonance imaging captures soft tissues, particularly in the brain, with high resolution, but lacks detailed information about the skeleton. CT enhances the details of high-density tissues compared to MRI, but soft tissues are displayed at a lower resolution. These limitations make it tedious and time-consuming for clinicians to switch between different imaging modalities to obtain more comprehensive patient information. As shown in Fig. 1, multi-modal medical image fusion combines complementary information from multiple modalities to create a single image with richer details, providing clinicians with a more accurate imaging basis for patient diagnosis and treatment1,2. This technology is widely applied in intraoperative navigation3, tumor segmentation4 and adjuvant radiotherapy5.
Schematic of multimodal medical image fusion.
Currently, deep learning-based methods are the dominant algorithms for medical image fusion6,7. These methods use convolutional neural networks (CNNs) to extract and fuse image features8,9. The fusion strategies in these methods are typically learned by the network itself, without manual intervention. Coupled with the powerful representational capacity of CNNs, this has led to impressive performance in image fusion tasks. However, convolutional operations inherently limit the receptive field of the network, resulting in insufficient consideration of long-range correlations within the image, thus limiting the fusion performance. To address this, many researchers have introduced transformers to improve the ability of the network to model long-range dependencies, partially alleviating the issue, but a complete solution has not yet been found10. Moreover, since image fusion lacks a ground truth, it is inherently an unsupervised problem. The performance of fusion models largely depends on the design of the loss function. Existing fusion methods typically design the loss function based on local statistical information between pixel points, overlooking the importance of non-local features within the image. This limitation constrains the overall performance of fusion networks.To address this issue, this paper proposes a multi-modal medical image fusion method guided by stochastic structural similarity, called S3IMFusion. First, we design a fusion framework using a convolutional neural network combined with transformer architecture, enabling the model to correlate both local and global features in the image. Then, we propose a stochastic structural similarity loss function that preserves the complementary information from the input images in the fusion result by constructing a stochastic structural feature similarity loss between the fused image and the source images. This results in a fused image with richer information. The main contributions of this paper are summarized as follows:
-
We propose an end-to-end medical image fusion network, termed S3IMFusion, which is capable of extracting and fusing both local detail features and non-local complementary features from the input images, resulting in fused image with richer feature representations.
-
We design a novel loss function that effectively interacts with both local and non-local features. Specifically, the features of the fused result and source images are randomly shuffled along the pixel columns, creating an image that incorporates non-local features. The structural similarity loss is then computed between the shuffled fused result and the source images, thereby enhancing the global feature correlation within the fused image.
-
We evaluate the performance of S3IMFusion on CT-MRI image fusion and SPECT-MRI image fusion tasks. Experimental results demonstrate that the network exhibits excellent fusion performance, preserving significant structural and tissue information from the input images in the fusion results. Furthermore, experiments on the RoadScene dataset show that S3IMFusion can be seamlessly extended to infrared and visible image fusion tasks, yielding satisfactory results.
Related works
In this section, we present related works on traditional medical image fusion methods and deep learning-based medical image fusion methods.
Traditional medical image fusion methods
Many multi-modal medical image fusion methods have been proposed in the last decade or more11. Multi-scale decomposition-based ideas have historically been predominant in traditional fusion methods, in which source images are decomposed into different components, a fusion strategy is manually set to combine these components, and the fused image is reconstructed by a corresponding inverse transformation12. Such fusion methods include pyramid transform13; wavelet-based methods14, including shearlet transform15, discrete wavelet16, and stationary wavelet17; and other transform methods18. Harmanpreet et al.19 proposed a fusion framework based on multi-scale edge-preserving filters and visual saliency detection, which effectively solves the problem of high computational complexity of fusion algorithms. To alleviate the problem of losing critical information in original image, Harmanpreet et al.20 decompose the image into detail and base layers based on an anisotropic diffusion filter, and then fuse the different feature components so as to effectively retain the critical information in the source image. However, a common problem with these methods is that they use the same decomposition method for images of different modalities. Note that a single decomposition is usually unable to represent the whole image feature distribution. In addition, the manually designed fusion strategies are also not generalizable due to the incomplete fusion of complementary information in different modalities medical images. Moreover, it is computationally infeasible to implement these traditional methods for a large image dataset.
Deep learning-based medical image fusion methods
In recent years, deep learning-based medical image fusion methods have been proposed21. Lui et al.22 introduced deep learning into the field of medical image fusion for the first time, which computed the weight map for image fusion by CNN for fusing CT and MR images. However, stacking multiple convolution layers resulted in the algorithm losing the underlying information in the images and increased the computational burden. To alleviate the problem, Zhang et al.23 proposed a fast unified image fusion network called PMGI, which models the image fusion problem uniformly as a texture and intensity preservation problem of an image, and then exchanges intensity and texture information in the image using CNN with different channels. This method achieves a unified image fusion task, but the algorithm tends to over-fuse redundant features in the image, resulting in artifacts that adversely affect the accuracy of the fusion. Xu et al.24 proposed an unsupervised, unified end-to-end image fusion network called U2Fusion by utilizing CNN for feature extraction and information metrics on images. This method achieves unified processing by preserving adaptive similarity between the fused image and the source images. A notable disadvantage is that the simplicity of the fusion rule leads to the omission of important information from the original image in the fused images. Zhang et al.25 proposed the IFCNN fusion method, which involves extracting salient features from the source images using CNN, followed by fusing these features using element-wise maximum and minimum operations, and finally reconstructing the fused image through a reconstruction network. In this method, a unified image fusion framework is achieved by combining deep learning with manual fusion strategies. However, the use of an element-level fusion strategy makes the network susceptible to noise. Wang et al.26 proposed a generalized fusion framework based on the mask attention mechanism, which incorporates information filtering and fusion control strategies to enhance the retention of complementary information while eliminating redundant features in the fusion result. Although existing deep learning-based medical image fusion methods can achieve good performance in many tasks, there are still some shortcomings. Since existing deep learning-based methods typically rely on a convolutional operation which only extract local features in images, and do not make good use of global features, which means that global semantic information is often ignored. In addition, the aforementioned methods rely on loss functions primarily derived from pixel-level features, making them susceptible to noise interference.
Transformer is first applied in the field of natural language processing (NLP) and achieved great success27. It has also found applications in the field of computer vision for tasks28,29,30,31. In image fusion task, Ma et al.32 proposed a universal image fusion framework based on Swin Transformer. By modeling long-range dependencies in the source images, the network can fully achieve domain specific information extraction and cross domain complementary information integration, while maintaining appropriate apparent strength from a global perspective. However, a significant drawback of this method is its reliance solely on the Transformer for fusion strategy design, which results in limited extraction of global contextual features within the image. Tang et al.33 proposed an unsupervised multi-modal medical image fusion method by introducing adaptive convolution and multi-scale adaptive Transformer to model long-range dependencies. This method can effectively extend to infrared and visible image fusion tasks, but it is challenging to generalize this method to CT and MRI image fusion tasks. Rao et al.34 proposed a fusion method for infrared and visible images based on Transformer. By designing different attention modules, the fusion performance of the network is improved by interacting the attention module with the Transformer fusion module, and at the same time refining the fusion relationship in the spatial and cross-channel ranges. While this method achieves good results above the retention of global information, it looses portion local information, resulting in fused images with artifacts. Yang et al.35proposed a generalized image fusion network that combines Transformer and diffusion models. The image is first compressed into low-resolution latent features through encoder downsampling, which are then decoded by a decoder to preserve the high-resolution information. Finally, a Transformer-based denoising network and fusion network are employed to ensure the fusion produces highly detailed images. Liu et al.36 proposed a multi-scale feature fusion network based on MixFormer, which enhances scale diversity in the fusion results by utilizing MixFormer as the backbone for feature extraction. A feature fusion module based on multi-source spatial attention is then designed to perform multi-scale fusion of features from the source image. Although this method demonstrates excellent fusion performance, the network architecture exhibits high complexity and substantial computational overhead. Moreover, the above Transformer-based methods do not make full use of the complementariness of global and local information. While it is important to consider global information, local features cannot be ignored as they carry local complementary information. It turns out that these Transformer-based methods use loss functions at the pixel level, which makes it difficult to measure the complementariness of global and local information.
Proposed method
This section first presents the framework of the proposed end to end multi-modal medical image fusion network. Then, we present the details of the structure containing local and non-local feature extraction modules. Finally, the design of the loss function is presented.
Network architecture
The framework of the proposed S3IMFusion.
The framework of the S3IMFusion network is illustrated in Fig. 2. Initially, the input images from different modalities are concatenated along the channel dimension, and a single-layer convolutional network is employed to perform hybrid feature extraction. The feature extraction process is then divided into two primary branches: salient feature extraction and multi-scale feature extraction. The salient feature extraction branch is responsible for capturing high-level features, such as contours and object boundaries, which are crucial for identifying anatomical structures in medical images. The multi-scale feature extraction branch, on the other hand, is designed to extract complementary information across various scales, enhancing the network’s ability to capture both fine-grained and global details. Subsequently, a stacked Transformer block is incorporated to model non-local dependencies and long-range correlations within the image, further improving the fusion process. Finally, a weighted fusion strategy is applied to combine the feature components from each branch, followed by a convolutional layer to reconstruct the fused image. The detailed structures of each branch are described in the following sections.
Salient feature extraction subnetwork
Salient information, such as contours and targets within the input image, constitutes crucial feature components. Efficient extraction of this information is essential for the subsequent reconstruction of high-quality fused images. Typically, this salient information manifests as higher pixel intensity values within the image. As shown in Fig. 2, we propose a salient feature extraction sub-network based on convolution and maximum pooling operations. The process begins with the combination of features extracted from the source images via a CNN. We then apply a convolutional layer with a kernel size of \(3 \times 3\), followed by a max-pooling layer. Next, attention weights are computed using a sigmoid function, generating a saliency weight map. Finally, the output of the subnetwork is obtained by performing element-wise multiplication between the results of the Relu activation function and the saliency weight map. The formula is expressed in Eq. (1).
where \(F_{in}\), \(F_{out}\) and \(\otimes \) denote the hybrid features extracted via the convolutional layer, salient feature and the product of their corresponding elements. Meanwhile, \(Conv(\cdot )\), \(BN(\cdot )\), \(MP(\cdot )\), sigmoid and \(Relu(\cdot )\) denote the convolutional operation, Batch normalization, Max-pooling, activation function and Rectified linear unit, respectively.
Complementary detail feature extraction subnetwork
The hybrid features extracted from the input image after the initial convolutional block contain complex complementary features across different channels, which are essential for enhancing the detail clarity of the fused image. To further capture these complementary details, we developed a complementary detail feature extraction subnetwork. This subnetwork, depicted as the second and third branches in Fig. 2, is constructed using skip connections between convolutional layers. Additionally, to improve feature reuse across different branches, multi-scale features are cascaded between branches and re-fed into the next convolutional layer of both branches. This approach ensures that information at multiple scales can effectively interact between the branches, facilitating better feature fusion.
Global correlation feature extraction module
In the previous subnetwork design, a series of convolutional operations are employed to extract multi-scale and salient features from the image. However, CNN are limited by their restricted receptive field, which hampers the network’s ability to capture global correlation information in the image. In contrast, the Transformer module mitigates this issue through its self-attention mechanism and positional encoding, which effectively preserve global correlation information37. To address the receptive field limitation of the subnetwork, we designed a Transformer-based global correlation feature extraction module. As shown in Fig. 2, the global correlation feature extraction module, located at the upper right, is constructed by connecting multiple Transformer units. This module operates in two stages: the first is the multi-head attention calculation, expressed in Eq. (2).
where \(MSA(\cdot )\), \(LN(\cdot )\), \(F_{s1}^{in}\) and \(F_{s1}^{out}\) denote the multi-head attention computation, layer normalization operation, the input and output features of the module, respectively.
The second stage is multi-layer perceptual computation, which is represented by Eq. (3).
where \( F_{s2}^{Out}\) denotes the output of the global feature extraction block, \(LN(\cdot )\) denotes the layer normalization operation and \(MLP(\cdot )\) represents the multi-layer perception. The outputs of the global feature extraction module are then integrated through a weighted combination of features. Finally, the fused image is reconstructed by applying a convolutional layer followed by a tanh activation function.
Loss function
Stochastic structural similarity loss
Existing learning-based medical image fusion methods usually rely on the pixel-level structural similarity in the images to design the loss function38. The formula for calculating the pixel level structural similarity between images is shown in Eq. (4).
where (x, y) denotes the corresponding pixel position indexes of different images; \(\mu _{x}\), \(\mu _{y}\) ,\(\sigma _{x}\) ,\(\sigma _{y}\) and \(\sigma _{xy}\) denote the expectation, variance and covariance of pixels in a localized region in image respectively, \(c_{1}\) and \(c_{2}\) are hyperparameters.
Although pixel-level loss functions effectively preserve local structural and luminance information in the input images, they typically fail to capture long-range dependencies and neglect non-local correlation information. Based on this consideration and motivated by this article39, we propose a novel loss function that effectively preserves non-local correlation information in the fused image. Specifically, during the computation of the structural similarity loss, we first introduce a random permutation of pixel orderings in both the fusion result and the input images to disrupt the pixel-localized statistical properties. This random shuffling step removes any spatial correlation between pixels across the images. The permuted images are then divided into equally sized image blocks. For each corresponding pair of blocks from the fusion result and the input image, the structural similarity is computed separately. The average structural similarity loss across all image blocks is then calculated, yielding the final loss. This process effectively computes a random structural similarity loss for the fusion result and the input image pair. The mathematical formulation is described in Eq. (5).
where SSIM denotes the similarity computed in an image region with a window size of \(k \times k\) and a stride of k, \(I_{1}\), \(I_{2}\) and \(I_{fus}\) denote the input images and the fused image respectively; \(P^{(m)}(I_{1})\), \(P^{(m)}(I_{2})\) and \(P^{(m)}(I_{fus})\) denote a randomly selected pixel blocks of size 64 \(\times \) 64 in the input images and the fused image respectively.
Detailed texture loss
To ensure that the fusion result retains sufficient texture details, we employ a gradient distribution to capture the texture information within the image. Additionally, we design a gradient loss function, as expressed in Eq. (6).
where \(\nabla \) and \(\Vert \cdot \Vert _1\) denote the Sobel gradient operator and \(l_{1}\) norm. By enforcing the gradient distribution of the fused image to align with the image exhibiting the largest gradient magnitude in the input, we obtain a fusion result with enhanced clarity. To prevent the fusion result from becoming excessively sharp or introducing artifacts such as halos or ringing, we introduce a smoothing loss function, which is defined as Eq. (7).
By implementing consistency between the pixel intensities of the fusion result and those of the input images, the network generates a fused image that is both sharper and smoother. Ultimately, the total loss function is formulated as a weighted combination of the individual loss terms.
Where \(\gamma _{1}\), \(\gamma _{2}\) and \(\gamma _{3}\) are weight hyperparameters.
Experiments and analysis of results
In this section, we experimentally validate the fusion performance of S3IMFusion on two datasets. These include CT and MRI image fusion, SPECT and MRI image fusion and IR and visible image fusion.
Datasets and training details
In this paper, two datasets are utilized. The first is a publicly available multi-modal medical image dataset sourced from the Harvard database, which contains 350 pairs of CT/SPECT and MRI images, each with a resolution of 256 \(\times \) 256. This dataset is widely used in medical image fusion research and provides an effective benchmark for evaluating the performance of fusion models. The second dataset, RoadScene40, is employed for the task of infrared and visible image fusion. It consists primarily of pairs of infrared and visible images depicting various scenes, including streets, pedestrians, vehicles, and buildings.
The experiments are implemented via the PyTorch framework on an NVIDIA GeForce RTX 3090 GPU. During training process, model parameters are updated using an Adam optimizer with a learning rate of 0.001, the batch size of 16, and the number of epochs is 100. The hyperparameters in the loss function are set as \(\gamma _1\) = 10, \(\gamma _2\) = 5, and \(\gamma _3\) = 1.
Comparison methods and evaluation metrics
In this section, we evaluate the fusion performance of the proposed S3IMFusion by comparing it with six state-of-the-art methods: EMFusion41, IFCNN25, MATR33, MUFusion42, U2Fusion24, and DGcGAN43. IFCNN and MUFusion represent medical image fusion methods that rely exclusively on convolutional neural networks (CNNs). U2Fusion is a versatile image fusion approach that demonstrates exceptional performance not only in medical image fusion but also in infrared-visible image fusion, as well as in multi-focus and multi-exposure scenarios. MATR combines CNN and transformer architectures for image fusion, while DGcGAN leverages generative adversarial networks (GANs) to perform image fusion. EMMA44 is a self-supervised fusion method with a priori knowledge of the principles of optical imaging. INet45 is a medical image fusion method that combines discrete wavelet transform with reversible networks.
To thoroughly evaluate the fusion performance of S3IMFusion, we employ eight widely recognized image quality assessment metrics: entropy (EN)46, average gradient (AG)47, mutual information (MI)48, structural similarity index (SSIM)49, peak signal-to-noise ratio (PSNR)50, Qabf51, sum of the correlations of differences (SCD)52, and spatial frequency (SF)53. EN quantifies the information content within an image, providing insights into its richness. AG measures the average local pixel value variations and is commonly used to assess texture and detail preservation. MI evaluates the capacity of fusion methods to retain original information, with higher values indicating better information preservation. SSIM offers a holistic assessment by evaluating brightness, contrast, and structural similarity between images. PSNR quantifies the signal-to-noise ratio between the original and fused images, offering a measure of image fidelity. Qabf, based on the Bandlet transform, emphasizes spectral and spatial fidelity, as well as global consistency in image fusion evaluation. SCD analyzes pixel differences across multiple scales, providing an assessment of information retention. SF reflects the retention of fine image details, such as texture and edges, and assesses the ability of the fusion model to preserve these features. By employing these metrics, a comprehensive and objective evaluation of the fusion performance of S3IMFusion is achieved.
CT and MRI image fusion
The experimental results of our proposed S3IMFusion on the Harvard dataset are shown in Fig. 3.
Results of CT and MRI image fusion on Harvard.
To provide a more detailed and illustrative evaluation, we select two local regions (indicated by red-boxed areas) for zoomed-in visual comparisons of the fused images. Each of the compared methods exhibits distinct strengths and limitations. The DDcGAN method enhances the brightness of fused images; however, it introduces significant artifacts, leading to pronounced blurring and reduced structural integrity. EMFusion effectively integrates salient features from CT images into the fused outputs, though at the cost of losing texture details from MRI images. A similar compromise is observed in the IFCNN method. MATR demonstrates a notable ability to combine detailed texture information and salient features, yet suffers from visible blurring and reduced brightness in the fused images, particularly in the CT-MRI fusion context. MUFusion introduces undesirable noise artifacts, severely compromising the visual quality of the fusion results. U2Fusion excels at incorporating intricate details from MRI images into the fused outputs but neglects critical complementary information from CT images, resulting in a loss of balance between modalities. EMMA effectively preserves the salient features of the original image; however, the fusion result suffers from a lack of fine-grained detail, leading to insufficient representation of intricate information. INet achieves a more complete preservation of the mutual information from the original image in the fused output, owing to the reversibility of the network, which effectively mitigates information loss. Nevertheless, this approach is plagued by the issue of color distortion. In contrast, the proposed S3IMFusion method demonstrates superior performance by effectively preserving salient features from CT images while maintaining the intricate texture details from MRI images. Moreover, it achieves an optimal balance between brightness and detail preservation, resulting in fused images with enhanced visual clarity and overall quality. This capability underscores the robustness and effectiveness of S3IMFusion in handling multi-modal medical image fusion tasks.
Table 1 presents the evaluation results derived from the eight metrics mentioned earlier. This evaluation is conducted using 21 pairs of CT and MRI images. For each metric, the final score is calculated by averaging the assessment scores of the 21 test samples. From Table 1, it can be seen that the proposed S3IMFusion method performs well in EN, AG and MI. INet achieves excellent results on SSIM, Qabf and SCD metrics, due to the information lossless extraction capability of the invertible network, which allows it to retain more structural information in the image. S3IMFusion also demonstrates relatively sub-optimal results in metrics such as PSNR and SSIM. Both U2Fusion and MUFusion demonstrate superior performance in terms of PSNR and SF metrics. The comprehensive analysis underscores the stability of S3IMFusion in producing fused images and its capability to achieve higher-quality outputs by effectively integrating both global and local features from the source images.
SPECT and MRI image fusion
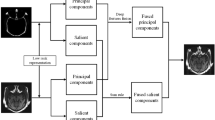
When fusing SPECT and MRI images, the SPECT image is initially transformed from the RGB color space to the YUV color space. In this representation, the U and V channels capture the chromaticity information of the image, while the Y channel encapsulates the luminance information. To leverage the luminance details for fusion, the Y-channel features are directly utilized in combination with the MRI image to generate the grayscale fusion result. Subsequently, the RGB fusion result is reconstructed by reintegrating the chromaticity information preserved in the U and V channels. The detailed workflow of this process is illustrated in Fig. 4.
SPECT and MRI image fusion process.
Similarly, we conducted experiments using the Harvard dataset, and the experimental comparison results are shown in Fig. 5, where the local features of the fusion results are zoomed in and labeled with green and red rectangular boxes for comparison purposes.
Results of SPECT and MRI image fusion.
As illustrated in Fig. 5, for images containing rich and intricate features, the existing methods fail to achieve a satisfactory fusion of SPECT and MRI images. The EMFusion method effectively preserves texture details from MRI images; however, it tends to lose critical structural information, particularly in organ structures such as the human eye. In contrast, the DDcGAN method excels at fusing contour information from both modalities but compromises the preservation of texture details from MRI images, thus negatively impacting the overall clarity of the fused image. Additionally, significant color distortion occurs when fusing the chromaticity information from the SPECT images. The fused images generated by IFCNN appear excessively smoothed, lacking adequate preservation of texture details from the source images. The MATR method, while successful in fusing detailed texture and salient features, suffers from over-fusion, retaining excessive chromaticity information, and neglecting important texture features from MRI images. MUFusion struggles to harmoniously integrate complementary information, resulting in fused images with low clarity. Similarly, while U2Fusion manages to retain complementary information, it introduces artifacts that degrade the overall image quality. EMMA effectively preserves contour and target information within an image; however, it is less effective at retaining edge intensity information, as exemplified by the region of the eyeball highlighted in the green box of the third result. This leads to blurring in the fused image. In contrast, INet excels at preserving detailed texture information and produces high-definition fusion results. Nevertheless, it tends to lose some intensity information, as indicated by the red rectangular box in the first fusion result, which results in the loss of edge features. In contrast, our proposed S3IMFusion method effectively preserves complementary information from both modalities, seamlessly integrating salient features from SPECT images with texture information from MRI images. Moreover, S3IMFusion generates fused images with superior clarity, retaining more chromaticity information and texture details compared to existing methods. To further assess the performance, we conduct objective evaluations of the fused images using the eight metrics previously mentioned.
As shown in Table 2, the quantitative evaluation results for S3IMFusion demonstrate its superior performance across seven metrics, including EN, AG, MI, SSIM and SF. EMMA achieves optimal performance in terms of PSNR and Qabf metrics, owing to the network training process being aligned with the principles of optical imaging. This alignment enables the network to adhere to the iso-realistic a priori, resulting in fusion outputs that are clearer and richer in detail. The superior performance of INet in SCD metrics can be attributed to its multichannel lossless feature extraction method, which enhances the consistency of the fusion results. These results align with the findings in Table 1, further highlighting the ability of S3IMFusion to maintain exceptional fusion quality, even when dealing with more complex image features. This underscores the enhanced generalization capability of S3IMFusion in comparison to other fusion methods.
Analysis of loss function
To evaluate the efficacy of the global similarity loss and random region pixel intensity loss functions proposed in this study, we conduct an ablation experiment on the loss functions. In this experiment, the proposed loss functions are replaced with traditional structural similarity loss and pixel intensity loss functions, while all other conditions are kept consistent. This approach aims to isolate and assess the specific impact of these novel loss functions on the overall performance. The \(L_{ab}\) is described in Eq. (9).
where SSIM, \(L_{\textrm{grad}}\) and \(L_{smooth}\) denote structure similarity index, gradient loss and smoothing loss, and \(\gamma _1\), \(\gamma _2\) and \(\gamma _3\) are the corresponding weighting parameters.
Results of loss functions ablation experiments.
The experimental results are presented in Fig. 6. The fusion network trained exclusively with the general similarity loss function fails to adequately preserve global complementary information during the fusion process, as evidenced by the blurred texture details and poorly preserved salient features in the fused images. In contrast, the network guided by the proposed loss function demonstrates a significant improvement. It effectively integrates complementary features from the source images, resulting in a fused image with sharper definition and richer texture details. Similarly, the results across the eight evaluation metrics, as shown in Table 3, further corroborate these findings. From a comprehensive perspective, S3IMFusion with \(L_{total}\) exhibits substantial advantages in visual perception indices. When \(L_{total}\) is replaced with \(L_{ab}\), a notable decline is observed in the indices related to both image features and image structure in the fused images. This indicates that \(L_{total}\) plays a crucial role in enhancing edge information and preserving fine texture details in the fused image.
Extension to infrared and visible image fusion
In general, RGB camera imaging offers the advantages of rich texture and high clarity. However, in extreme weather conditions or low-light environments, a single RGB camera struggles to effectively capture the external world. In contrast, infrared cameras leverage thermal radiation to image objects, offering superior stability and reliability under challenging conditions. Therefore, the fusion of infrared and visible light images can capitalize on the complementary strengths of both camera types, resulting in fused images of higher quality. Infrared and visible light image fusion has thus emerged as a crucial subfield within multi-modal image fusion. In this work, we extend the proposed S3IMFusion framework to infrared and visible light image fusion, evaluating the generalizability of the algorithm through experiments conducted on the RoadScene dataset. Consistent with the fusion of SPECT and MRI images, we first convert the visible image from the RGB color space to the YUV color model. Fusion is then performed on the Y-channel of the visible image with the infrared image. Finally, the fusion result is transformed back to the RGB color space to reconstruct the fused image. The experimental results are shown in Fig. 6.
Results of visible and infrared image fusion.
We compare the experimental results with six existing methods. Among them, CDDFuse produces fusion results that are closest to our S3IMFusion; however, its performance suffers a reduction in clarity when fusing images with richer edge information, as seen in the region highlighted by the red rectangular box in the third set of images in Fig. 7. DATFuse and U2Fusion fail to adequately preserve the detailed texture information from the input images, resulting in blurred fusion outputs. Although DDcGAN performs well in fusing prominent features such as pedestrians, it suffers from significant color distortion and blurring, leading to substantial information loss. IFCNN and SwinFuse experience feature loss when fusing images with weak texture features, such as the streetlights marked by the green rectangular boxes in the fourth set of images. In contrast, our proposed S3IMFusion effectively addresses the limitations observed in the aforementioned methods. It successfully retains the rich texture information from the visible image while preserving the salient features from the infrared image. When confronted with targets exhibiting distinct edge distributions, such as streetlights and buildings, S3IMFusion produces clear and precise fusion results, avoiding color distortions and maintaining high image clarity.
The quantitative evaluation results are presented in Table 4, which demonstrates that the performance of S3IMFusion across the reevaluated metrics aligns well with the results shown in Fig. 7. Notably, S3IMFusion achieves optimal performance on the EN, AG, MI, PSNR, Qabf, and SF metrics. Both subjective visual assessment and objective quantitative metrics indicate that S3IMFusion performs exceptionally well, exhibiting strong scalability in the context of infrared and visible image fusion tasks.
Conclusion
In this paper, we proposed a multi-modal medical image fusion method guided by stochastic structural similarity, termed S3IMFusion. This method leverages CNN and Transformer modules to design a multi-channel sub-network that extracts global correlation information from the input images, enabling the precise fusion of complementary features. By incorporating a salient attention module, the network effectively preserves the most informative regions of the images. Furthermore, a novel loss function is designed, addressing both global and local aspects of image fusion. The global correlation features are retained by constructing a stochastic structural similarity loss between the fusion result and the input images. The texture loss function, which is based on gradient modeling, ensures the preservation of rich texture and fine-grained details in the fusion result. Experimental results demonstrate that the proposed method outperforms existing fusion methods in terms of both visual quality and quantitative assessment, achieving accurate fusion of the input images. Additionally, we extend S3IMFusion to the task of infrared and visible image fusion, where it produces promising results, indicating the robust generalization ability of our method.
Data Availability
The Harvard dataset underlying this article are available in https://www.med.harvard.edu/aanlib/home.htm. The another dataset underlying this article are available in https://github.com/hanna-xu/RoadScene.
References
Liu, R. S., Liu, J., Jiang, Z., Fan, X. & Luo, Z. A bilevel integrated model with data-driven layer ensemble for multi-modality image fusion. IEEE Trans. Image Process. 30, 1261–1274 (2020).
Kumar, A., Fulham, M., Feng, D. & Kim, J. Co-learning feature fusion maps from PET-CT images of lung cancer. IEEE Trans. Med. Imaging 39, 204–217. https://doi.org/10.1109/TMI.2019.2923601 (2019).
Guo, P. et al. Nuclei-based features for uterine cervical cancer histology image analysis with fusion-based classification. IEEE J. Biomed. Health Inform. 20, 1595–1607 (2016).
Catana, C., Drzezga, A., Heiss, W.-D. & Rosen, B. R. PET/MRI for neurologic applications. J. Nucl. Med. 53, 1916–1925 (2012).
Li, Z. et al. Image fusion technique for target volume delineation in 125I seed implant brachytherapy for parotid gland cancers. J. Cancer Res. Ther. 18, 470–475 (2022).
Zhang, H., Xu, H., Tian, X., Jiang, J. & Ma, J. Image fusion meets deep learning: A survey and perspective. Inf. Fusion 76, 323–336 (2021).
Azam, M., Khan, K., Salahuddin, S. & Rehman, E. A review on multimodal medical image fusion: Compendious analysis of medical modalities, multimodal databases, fusion techniques and quality metrics. Comput. Biol. Med. 144, 105253 (2022).
Zhou, T. et al. Deep learning methods for medical image fusion: A review. Comput. Biol. Med. 160, 106959 (2023).
Li, Y., Zhao, J., Lv, Z. & Li, J. Medical image fusion method by deep learning. Int. J. Cognit. Comput. Eng. 2, 21–29 (2021).
Shamshad, F. et al. Transformers in medical imaging: A survey. Med. Image Anal. 88, 102802 (2023).
Hermessi, H., Mourali, O. & Zagrouba, E. Multimodal medical image fusion review: Theoretical background and recent advances. Signal Process. 183, 108036 (2021).
Karim, S. et al. Current advances and future perspectives of image fusion: A comprehensive review. Inf. Fusion 90, 185–217 (2023).
Zhang, Q. et al. Similarity-based multimodality image fusion with shiftable complex directional pyramid. Pattern Recognit. Lett. 32, 1544–1553 (2011).
Ma, J. & Zhou, Y. Infrared and visible image fusion via gradientlet filter. Comput. Vis. Image Understand. 197, 103016 (2020).
Geng, P. et al. Image fusion by pulse couple neural network with shearlet. Opt. Eng. 51, 067005 (2012).
Bhavana, V. & Krishnappa, H. Multi-modality medical image fusion using discrete wavelet transform. Procedia Comput. Sci. 70, 625–631 (2015).
Ganasala, P. & Prasad, A. Medical image fusion based on laws of texture energy measures in stationary wavelet transform domain. Int. J. Imaging Syst. Technol. 30, 544–557 (2020).
Yan, J. et al. Multispectral and hyperspectral image fusion based on low-rank unfolding network. Signal Process. 213, 109223 (2023).
Kaur, H. et al. Multimodal medical image fusion utilizing two-scale image decomposition via saliency detection. Curr. Med. Imaging 20, e15734056260083 (2024).
Kaur, H., Vig, R., & Kumar, N. et al. Fusion of multimodal medical images based on fine-grained saliency and anisotropic diffusion filter. Curr. Med. Imaging 20, (2024).
Wang, W., He, J. & Liu, H. EMOST: A dual-branch hybrid network for medical image fusion via efficient model module and sparse transformer. Comput. Biol. Med. 179, 108771 (2024).
Liu, Y., Chen, X. & Cheng, J. A medical image fusion method based on convolutional neural networks. in Proceedings of the 20th International Conference on Information Fusion (IEEE, 2017), pp. 1-7.
Zhang, H., Xu, H., Xiao, Y. et al. Rethinking the image fusion: A fast unified image fusion network based on proportional maintenance of gradient and intensity. In Proceedings of the AAAI Conference on Artificial Intelligence, pp. 12797–12804 (2020).
Xu, H. et al. U2Fusion: A unified unsupervised image fusion network. IEEE Trans. Pattern Anal. Mach. Intell. 44, 502–518 (2020).
Zhang, Y. et al. IFCNN: A general image fusion framework based on convolutional neural network. Inf. Fusion 54, 99–118 (2020).
Wang, X. et al. MMAE: A universal image fusion method via mask attention mechanism. Pattern Recognit. 158, 111041 (2025).
Vaswani, A., Shazeer, N., & Parmar, N. et al. Attention is all you need. Adv. Neural Inf. Process. Syst. 30 (2017).
Srinivas, A., Lin, T., & Parmar, N. et al. Bottleneck transformers for visual recognition. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 16519-16529 (2021).
Sun, Z., Cao, S., Yang, Y. et al. Rethinking transformer-based set prediction for object detection. In Proceedings of the IEEE/CVF International Conference on Computer Vision, pp. 3611-3620 (2021).
Chen, J., Lu, Y., & Yu, Q. et al. Transunet: Transformers make strong encoders for medical image segmentation. arXiv preprint arXiv:2102.04306 (2021).
Sun, L. et al. MCnet: Multiscale visible image and infrared image fusion network. Signal Process. 208, 108996 (2023).
Ma, J. et al. SwinFusion: Cross-domain long-range learning for general image fusion via swin transformer. IEEE/CAA J. Autom. Sin. 9, 1200–1217 (2022).
Tang, W. et al. MATR: Multimodal medical image fusion via multiscale adaptive transformer. IEEE Trans. Image Process. 31, 5134–5149 (2022).
Rao, D., Xu, T. & Wu, X. TGFuse: An infrared and visible image fusion approach based on transformer and generative adversarial network (IEEE Trans, Image Process, 2023).
Yang, B. et al. LFDT-Fusion: A latent feature-guided diffusion Transformer model for general image fusion. Inf. Fusion 113, 102639 (2025).
Liu, Y. et al. MM-Net: A Mixformer-based multi-scale network for anatomical and functional image fusion. IEEE Trans. Image Process. 33, 2197–2212 (2024).
Xiao, T. et al. Early convolutions help transformers see better. Adv. Neural. Inf. Process. Syst. 34, 30392–30400 (2021).
Ding, K. et al. Image quality assessment: Unifying structure and texture similarity. IEEE Trans. Pattern Anal. Mach. Intell. 44, 2567–2581 (2020).
Xie, Z., Yang, X., Yang, Y., et al. S3im: Stochastic structural similarity and its unreasonable effectiveness for neural fields. In Proceedings of the IEEE/CVF International Conference on Computer Vision, pp. 18024–18034 (2023).
Xu, H. et al. FusionDN: A unified densely connected network for image fusion. Proc. AAAI Conf. Artif. Intell. 34, 12484–12491 (2020).
Xu, H. & Ma, J. EMFusion: An unsupervised enhanced medical image fusion network. Inf. Fusion 76, 177–186 (2021).
Cheng, C., Xu, T. & Wu, X. MUFusion: A general unsupervised image fusion network based on memory unit. Inf. Fusion 92, 80–92 (2023).
Ma, J. et al. DDcGAN: A dual-discriminator conditional generative adversarial network for multi-resolution image fusion. IEEE Trans. Image Process. 29, 4980–4995 (2020).
Zhao Z., Bai H., Zhang J. et al. Equivariant multi-modality image fusion. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 25912-25921 (2024).
He, D. et al. MMIF-INet: Multimodal medical image fusion by invertible network. Inf. Fusion 114, 102666 (2025).
Roberts, J., Van Aardt, J. & Ahmed, F. Assessment of image fusion procedures using entropy, image quality, and multispectral classification. J. Appl. Remote Sens. 2, 023522 (2008).
Cui, G. et al. Detail preserved fusion of visible and infrared images using regional saliency extraction and multi-scale image decomposition. Opt. Commun. 341, 199–209 (2015).
Qu, G., Zhang, D. & Yan, P. Information measure for performance of image fusion. Electron. Lett. 38, 1 (2002).
Kaur H., Vig R., Kumar N. et al. Objective image quality assessment of pixel level image fusion algorithms for medical imaging. In 2023 Second International Conference on Electrical, Electronics, Information and Communication Technologies (ICEEICT), IEEE 2023, pp. 01-08.
Hore, A., Ziou, D. Image quality metrics: PSNR vs. SSIM. In Proceedings of the 2010 20th International Conference on Pattern Recognition, IEEE, pp. 2366-2369 (2010).
Xydeas, C. & Petrovic, V. Objective image fusion performance measure. Electron. Lett. 36, 308–309 (2000).
Aslantas, V. & Bendes, E. A new image quality metric for image fusion: The sum of the correlations of differences. AEU-Int. J. Electron. Commun. 69, 1890–1896 (2015).
Eskicioglu, A. & Fisher, P. Image quality measures and their performance. IEEE Trans. Commun. 43, 2959–2965 (1995).
Zhao, Z., Bai, H., Zhang, J., et al. Cddfuse: Correlation-driven dual-branch feature decomposition for multi-modality image fusion. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 5906–5916 (2023).
DATFuse: Infrared and visible image fusion via dual attention transformer. IEEE Trans. Circuits Syst. Video Technol. 33, 3159–3172 (2023).
Wang, Z. et al. SwinFuse: A residual swin transformer fusion network for infrared and visible images. IEEE Trans. Instrum. Meas. 71, 1–12 (2022).
Acknowledgements
This research is supported by the Foundation of Medical Science and Technology Project of Zhejiang Province of China under Grant 2023RC189, the Zhejiang Province Leading Geese Plan of China under Grant 2024C04032, and the Zhejiang Provincial Outstanding Youth Science Foundation of China under Grant LR20F030004.
Author information
Authors and Affiliations
Contributions
Conceptualization: Junhui Lv; Methodology: Junhui Lv, Xiangzhi Zeng; Writing - Review and Editing: Xiangzhi Zeng, Junhui Lv; Validation: Junhui Lv, Xiangzhi Zeng; Funding acquisition: Bo Chen, Junhui Lv, Zheming Wang; Software: Xiangzhi Zeng, Mingnan Hu; Writing - Original Draft: Xiangzhi Zeng, Junhui Lv; Investigation: Bo Chen, Mingnan Hu, Xiang Qiu, Zheming Wang; Supervision: Junhui Lv, Zheming Wang, Xiang Qiu, Shuxu Yang.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lv, J., Zeng, X., Chen, B. et al. A stochastic structural similarity guided approach for multi-modal medical image fusion. Sci Rep 15, 8792 (2025). https://doi.org/10.1038/s41598-025-93662-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-93662-6