Abstract
Disease detection plays an important role in shrimp aquaculture to ensure the health and sustainability of farming operations. Specifically, detecting viral infections at early stages can prevent significant losses. Image processing applications have been developed to detect different types of diseases in shrimp. However, theaccuracy of detection models needs improvement to detect various diseases through a single model. Therefore, this research presents a novel disease detection model using an Enhanced Recurrent Capsule Network (ERCN) with a hybrid optimization model for enhanced detection performance. The proposed ERCN utilizes dynamic routing of capsules to extract spatial hierarchies and patterns in shrimp images, while the recurrent layer extracts temporal dependencies. Performance is further improved by incorporating spatial and channel attention models to select optimal regions and features in the images for the fusion process. The dual-level feature fusion procedure combines local and global features, providing a final fused data to classify different types of diseases. Additionally, the proposed work incorporates a hybrid optimization that combines Harris Hawks Optimization (HHO) with the Marine Predator Algorithm (MPA) to fine-tune the classifier model parameters. Experiments evaluate the performance of the proposed disease detection model through various metrics such as accuracy, precision, recall, specificity, Matthews correlation coefficient, and F1-score. The resutls confirms that the performance of the proposed model is superior with precision of 94.9%, recall of 93.5%, F1-score of 94.6% and detection accuracy of 95.2% over conventional Recurrent Neural Network (RNN), Convolutional Neural Network (CNN), Gated Recurrent Unit (GRU), and Long Short Term Memory (LSTM) Networks.
Similar content being viewed by others
Introduction
In the global seafood industry, shrimp aquaculture occupied a significant place in providing nutritious seafood to consumers. The billion-dollar business has a market size valued around 68.40 billion USD in 2022 and it is expected to increase to 110.75 billion USD in 2031. Shrimps has many nutritional health benefits, and it is rich in vitamins, proteins, minerals, and antioxidants. To meet the increasing demand, large scale shrimp cultivation is followed throughout the world. However, maintaining sustainability and increasing profitability while doing large scale farming is threatened due to the infections and diseases that affect shrimp production. The economic losses due to the diseases in shrimps provide serious challenge to food production1,2. Early detection of diseases in shrimp aquaculture will help to reduce the mortality rate of shrimps, help to secure the remaining shrimps in the farms and thus ensure the sustainability of the shrimp farming3. The major factor that affects shrimps are viruses, bacteria, fungi, and parasites. The diseases can cross the shrimp population quickly and lead to significant mortality and reduce overall productivity. Some of the diseases and their symptoms are presented in Table 1 to better understand the diseases factors.
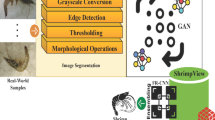
Among all viral diseases, WSSV is the most dangerous disease as these infections cause widespread destruction in shrimp farms worldwide. Bacterial infections caused by vibrio species leads to vibriosis10 and the fungal infections results in Fusariosis. The infections due to the parasites are less but it also brings a threat to shrimp health. Detecting diseases with specific symptoms and making accurate decisions for timely diagnosis is challenging in shrimp aquaculture. The manual inspection and biochemical test are time consuming, and it requires more labor support. To handle these issues, modern farming industries adopt image processing applications for diagnosing shrimp diseases. Advanced machine learning and deep learning techniques are introduced in various domains as efficient image processing applications11,12. Specifically in shrimp aquaculture CNN has been extensively used to learn and recognize disease patterns from the shrimp images13. Deep learning models exhibit a promising result over traditional image processing applications in detecting shrimp diseases14. However, the existing models have limitations in processing complex spatial relationships in the input shrimp images. The limitation is distinct if the symptoms are not noticeable and dispersed across different parts of the shrimp. The performance of these models reduces when the input image has large variations due to the presence of noise, effects due to lighting conditions and image quality.
To overcome the above challenges in shrimp disease detection, this research work presents a novel shrimp disease detection model. The proposed optimized ERCN overcomes the limitations in traditional deep learning models in shrimp disease detection by effectively capturing the spatial and temporal dependencies in shrimp images. The proposed model utilizes the capsule networks dynamic routing mechanism to handle the spatial hierarchies which are essential in recognizing shrimp disease patterns. The recurrent layer in the proposed ERCN analyzes the sequence of image frames and helps to detect the progression of diseases over time. The proposed model additionally employed a hybrid optimization model by combining HHO with MPA to fine tune the ERCN parameters. The hybrid optimization algorithm provides a better balance between exploration and exploitation in search space and provides optimal configurations to the model to detect the diseases with high accuracy. The main contributions to the proposed work are summarized as follows:
-
Presented a novel optimized ERCN to capture the spatial and temporal features in shrimp images. The complex disease patterns in the shrimp images are extracted and classified through the proposed ERCN model to attain better performance in disease detection.
-
The proposed ERCN model incorporates spatial and channel attention mechanisms to enhance the feature extraction performances. The extracted features are fused through dual level feature fusion procedure and then classified to detect different types of diseases.
-
Presented a hybrid optimization algorithm that combines HHO and MPA for optimizing the parameters of proposed ERCN model.
-
Presented a detailed simulation analysis of proposed model using different types of shrimp images and evaluated the performance through metrics like precision, recall, f1-score, specificity, Mathew correlation coefficient and accuracy.
-
A detailed comparative analysis is presented to validate the better performance of proposed model over existing methods like CNN, RNN, LSTM, GRU and VGG16 deep learning algorithms.
The remaining discussions in the research work are arranged in the following order. Section 2 provides a brief discussion on existing algorithms. The research gaps are summarized in Sect. 3 and Sect. 4 presents the proposed ERCN model in detail. Section 5 presents the simulation results and discussion, and Sect. 6 presents the conclusion of research work.
Related works
Recent studies suggest numerous approaches for shrimp disease detection. Machine learning models are used widely in shrimp disease detection research works. A combination of random forest and chi squared automatic interaction detection model was presented in15 to detect white spot diseases in shrimps. The presented ensemble model utilizes the random forest for feature extraction and classification. The CHAD model used in the research work statistically analyzes the detection results and improves the detection accuracy further. However, the presented experimental model results highlight the need for further enhancement of real time efficiency and integrating recent learning technologies to attain better accuracy in disease detection.
A comparative analysis of different machine learning models presented in16 utilizes logistic regression, artificial neural network, decision tree and k-nearest neighbor algorithms for detecting diseases in shrimps. The analysis utilizes different shrimp images and evaluates the machine learning models’ performance. The experimental results highlight the superior detection accuracy of decision tree model over other algorithms in case of training. But in case of validation accuracy, logistic regression model attained better performances over other algorithms. Similar comparative analysis presented in17 utilizes machine learning algorithms like random tree, extra tree and J48 decision tree algorithms to analyze the shrimp images. The presented model includes spatial features in addition to that image features to classify different types of diseases in shrimps. The experimental results highlight the superior performance of extra tree model over other decision tree algorithms. However, it has a limitation of lower variance in results due to the randomization procedure followed in extra tree model.
An unsupervised machine learning based segmentation model presented in18 detects white, black, and red spot diseases in aquatic animals. The presented model utilizes k-means clustering for image segmentation and then normalizes the images to enhance the details. Further classification is performed based on the Euclidean and Manhattan distances. The experimental results highlight that the presented model attain better accuracy in detecting spots over existing methods. The shrimp disease detection model presented in19,20,21,22 utilizes canny edge detection with gray level co-occurrence matrix for feature extraction. Further the extracted features are classified through a simple artificial neural network model to detect shrimp diseases. The presented model detects white spot syndrome with better detection accuracy. However, the model requires improved feature extraction procedure to attain better accuracy in detecting shrimp white spot disease.
The shrimp disease detection model presented in23 detects IHHNV infections in shrimp farms using polymerase chain reaction and sequencing analysis tests. The typical analysis utilizes specific primers to detect viral nucleic acids and then perform phylogenetic analysis to detect the strain in the raw shrimp images. The experimental results highlight that the disease detection model has high specificity and sensitivity. However recent advanced image processing applications are simple compared to this sequence analysis and more accurate than the test results. A threshold-based image fusion procedure reported in24 detects WSSV with better detection performance over traditional segmentation procedure. The presented approach utilizes adaptive median filter to eliminate the noise in the input shrimp image. Then using thresholding algorithm, the preprocessed image is segmented based on intra and inter class variances. Finally using expectation maximization algorithm, the detection procedure is fine-tuned and attained better detection performance.
The deep learning model presented in25 for WSSV disease detection preprocess the input image using local binary pattern technique. Then using total variation based fuzzy c-means algorithm the images are segmented and then the features are extracted using probabilistic linear discriminant analysis. The extracted features are then classified using enhanced GRU to attain better detection accuracy. The presented model additionally includes Wild Geese Migration Optimization algorithm to fine the classifier parameters. The presented approach attains better detection performance in shrimp disease analysis compared to traditional GRU and machine learning algorithms.
The disease detection model presented in26 utilizes the deep CNN model, hybrid optimization algorithm and random forest algorithms. The essential features from the images are initially extracted through the CNN model and then to solve the global optimization a hybrid optimization is presented by combining Ant Colony Optimization and genetic algorithm. Finally, the features are classified through random forest and attained better detection performance. However, due to the multiple models, the computational complexity of the presented approach increases compared to traditional methods31,32,33,34,35,36.
Research gaps
From the literature analysis it can be observed that utilizing deep learning algorithms in shrimp disease detection provides enhanced performance over traditional approaches. The incorporation of optimization algorithms in the disease detection model enhances the classification performance by fine tuning the classifier parameters. However, the existing models struggle to capture complex spatial relationships in the shrimp images specifically when the symptoms are spread across different portions of shrimps. The existing methods need to enhance detection accuracy and efficiency by incorporating efficient learning algorithms to detect different types of diseases. Accurate feature extraction methods are required for precise disease detection which highlights the need for improved techniques. Thus, in this research work a novel shrimp disease detection model is presented using advanced image processing technique with hybrid optimization algorithm. The features of ERCN with hybrid optimization are combinedly utilized to detect different types of diseases in shrimps with enhanced detection performance over traditional approaches.
Proposed work
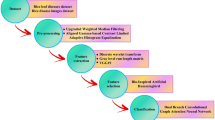
The proposed novel shrimp disease detection model utilizes advanced image processing and machine learning algorithms to attain high detection accuracy in identifying various types of shrimp diseases. The hybrid optimized learning network comprises of An ERCN for disease detection. To fine tune the detection model performance the parameters of the classifier are selected using hybrid optimization algorithm that combines HHO and MPA. The main element of the proposed model is the ERCN classifier which is developed to extract the spatial and temporal dependencies in shrimp images. Unlike traditional CNN that exhibit limited performance in defining the local spatial relationships due to the loss of important contextual information, the proposed ERCN utilizes the capsule network dynamic routing mechanism to preserve the hierarchical spatial relationships. This will help to recognize the complex patterns associated with shrimp diseases. The enhanced model utilizes recurrent layer to capture the temporal dependencies which is further useful in analyzing sequence of image frames so that the progression of disease over time can be identified. The proposed model utilizes the hybrid HHO-MPA that ensures better exploration and exploitation in the search space to provide optimal configurations to the ERCN model. The complete overview of the proposed model is presented in Fig. 1. The proposed shrimp detection model has a preprocessing step that enhances the quality of input images. Normalization, data augmentation and segmentation are performed in the preprocessing step.
Proposed Hybrid optimized Enhanced Recurrent Capsule Network.
Preprocessing
The disease detection model processes the shrimp image through the optimized ERCN model. Before providing the input to the ERCN model, the input image is preprocessed. The input image is normalized and then augmented in the preprocessing step. Since the data used in the proposed work is created by collecting different types of shrimp images available in open-source platforms and to increase the number of samples, data augmentation is performed in the preprocessing step. After that, to remove the unnecessary elements in the input image, the proposed model includes Grab cut segmentation algorithm that isolates the shrimps from the background. The normalization in the preprocessing adjusts the pixel values of input image into standard scale. Mathematically the normalization process is expressed as
where \(\:I\) represents the input image, \(\:\text{min}\left(I\right)\) represents the minimum pixel value in the image and \(\:\text{max}\left(I\right)\) represents the maximum pixel value in the image. \(\:I\left(x,y\right)\) represents the pixel value at position \(\:\left(x,y\right)\) in the input image, and the normalized pixel value is represented as \(\:{I}_{norm}\left(x,y\right)\). Followed by normalization, the input images count is increased through data augmentation. The common data augmentation procedures like rotation, flipping, scaling and translation are employed. In the rotation, the images are rotated at an angle of 90°, 180°, and 270°. While applying flipping, the input images are flipped horizontally and vertically. The images are resized through a scaling factor and using translation the image is shifted either horizontally or vertically by a certain number of pixels. The mathematical formulation for data augmentation is presented as follows.
where \(\:{I}_{rot}\left({x}^{{\prime\:}},{y}^{{\prime\:}}\right)\) represents the rotation, \(\:\left({x}^{{\prime\:}},{y}^{{\prime\:}}\right)\) represents the coordinates of the rotated image. \(\:{I}_{flipH}\left(x,y\right)\) and \(\:{I}_{flipV}\left(x,y\right)\) represents the horizontal and vertical flip respectively. \(\:{I}_{scale}\left({x}^{{\prime\:}},{y}^{{\prime\:}}\right)\) and \(\:{I}_{trans}\left({x}^{{\prime\:}},{y}^{{\prime\:}}\right)\) represents the scaling and translated input image respectively. After performing these preprocessing steps, segmentation is performed using Grab cut segmentation algorithm.
Segmentation using grab cut algorithm
The Grab cut segmentation algorithm19 is an advanced segmentation procedure that utilizes graph cuts to iteratively enhance the image segmentation. The proposed model utilizes different color images thus color image segmentation is performed. The unnecessary backgrounds in the input image are removed and the exact portion that is occupied by shrimps are segmented through the Grab cut model. The initialization phase of the Grab cut use a bounding box around the object. In this initial segmentation process, the pixels inside the box are marked as foreground and the other are considered as background. Basically, the likelihood of pixels is measured to differentiate the foreground and background. Mathematically it is expressed as
where the probability of pixel \(\:{I}_{ij}\) belongs to foreground or background is represented as \(\:P\left({I}_{ij}|{M}_{ij},Z\right)\). The binary mask is represented as \(\:M\) and the data is indicated as \(\:U\). Further a smoothness factor \(\:V\) is used to encourage neighbor pixels to have similar labels. This process provides smoothness in the segmentation, and it is mathematically formulated as
where \(\:\mathcal{N}\) represents the set of neighbor pixel pairs, \(\:{K}_{ij,kl}\) represents the penalty term based on color difference and it is mathematically expressed as
where \(\:{\upgamma\:}\) and \(\:{\upbeta\:}\) are the constant parameters. The grab cut model utilizes Gaussian Mixture Models (GMM) for foreground and background. A graph is constructed considering each pixel as a node that connects two points. The edge weights of the nodes are obtained from the GMM probabilities data \(\:\left(U\right)\) and smoothness \(\:\left(V\right)\) factors. The graph cut procedure utilizes min-cut and max flow algorithm to segment the graph from background. Based on the graph cut results the mask M is updated, and this process continues till there is no change between successive iterations. Compared to other segmentation algorithms, the accuracy of segmentation is better in grabcut algorithm due to the integration of data \(\:\left(U\right)\) and smoothness \(\:\left(V\right)\) factors. The segmented images are further fed into ERCN for feature extraction and classification.
Enhanced recurrent capsule network for feature extraction and classification
The proposed ERCN model extracts the features from the segmented shrimp image to detect different types of diseases. The proposed model utilizes the features of capsule networks28, recurrent layers29, and attention mechanism12 to capture the spatial and temporal features in the segmented images. The capsule network comprises of small group of neurons called capsule which provides an output vector for the given input. The vector summarizes various properties of the object like position, orientation, scale. This provides a better representation of spatial hierarchies in the data. The capsule network represents each capsule output as an activity vector instead of a single scalar value. Consider the output vector of capsule \(\:\left(i\right)\) in each layer is \(\:{u}_{i}\) and this vector dimensionality will vary depending on the complexity. Using a transformation matrix, each capsule in the lower layers predicts the output of subsequent higher layer capsules.
The ERCN model is depicted in Fig. 2.
ERCN layer visualization.
This transformation helps to learn the relationship between the capsules. A learned weight matrix is used to calculate the predicted output vector for capsule in the next layer. Mathematically the process is formulated as
where \(\:{W}_{ij}\) represents the weight matrix represents the output of capsule \(\:i\) in the lower layer to capsule \(\:j\) in the higher layer, \(\:{\widehat{u}}_{j|i}\) represents the predicted output vector for capsule \(\:j\) in the next layer. Through this, spatial relationships are learned and modelled by the network. The main feature of the capsule network is its dynamic routing mechanism. This procedure defines how the lower layer capsule outputs are routed to the higher layer capsules. This iterative procedure refines the coupling coefficient that represents the degree of agreement between lower layer capsules and higher layer capsules. The coupling coefficient is mathematically formulated as
where \(\:{c}_{ij}\) represents the coupling coefficient, \(\:{b}_{ij}\) represents the logits. These logits start as zero and updates iteratively based on the predicted output \(\:{\widehat{u}}_{j|i}\) and actual output \(\:{s}_{j}\) of the higher layer capsule. Mathematically the process of calculating total input to capsule is defined as a weighted sum of the predicted outputs as follows.
The actual output vector of a capsule is obtained by applying a non-linear squashing function to \(\:{s}_{j}\). This process ensures that the vector length remains in range 0 and 1, which is essential to represent the probability of the object’s presence. Mathematically it is given as
where \(\:{v}_{j}\) represents the actual output vector of a capsule \(\:j\). Further the agreement between predicted output and actual output is measured through dot product which is expressed as follows.
where \(\:{a}_{ij}\) represents the agreement that made between predicted \(\:{\widehat{u}}_{j|i}\) and actual output \(\:{v}_{j}\). The logits are updated based on the agreement and it influences the coupling coefficient in the subsequent iterations. mathematically the update process is formulated as \(\:{b}_{ij}\leftarrow\:{b}_{ij}+{a}_{ij}\). Due to this update procedure the routing process continuously refines the coupling coefficient and improves the connection between capsules. The iterative routing procedure allows the network to dynamically adjust the routing paths to provide more accurate and robust representation of the input data.
In the proposed work a recurrent layer is included to learn the temporal dependencies in the input image. The incorporated recurrent layer allows the network to process the sequence of feature vectors over time. The information from the previous time steps can be retained and utilized to enhance the ability to recognize the changes over time. The recurrent layer in the proposed work utilizes LSTM network which is suitable to process complex input images. The gates in the LSTM network relation or forget the information based on the necessity to ensure its higher performance in capturing the long-term dependencies. The input, output and forget gate in the LSTM is controlled by a set of parameters and sigmoid activation function. Consider \(\:{h}_{t}\) is the hidden state, \(\:{c}_{t}\) is the cell state, \(\:{v}_{t}\) is the feature vector at time step \(\:t\) the update equations are formulated as follows.
where the input gate is indicated as \(\:{i}_{t}\), forget gate is indicated as \(\:{f}_{t}\), output gate is indicated as \(\:{o}_{t}\), sigmoid function is indicated as \(\:\sigma\:\). The weights and biases of input, forget, output and cell state are indicated as \(\:({W}_{i},{W}_{f},{W}_{o},{W}_{c})\), \(\:({U}_{i},{U}_{f},{U}_{o},{U}_{c})\) and \(\:({b}_{i},{b}_{f},{b}_{o},{b}_{c})\) respectively. The input gate defines the quantity of new input \(\:{v}_{t}\) into the cell state. The forget gate defines which previous cell state \(\:\left({c}_{t-1}\right)\) information must be retained for further process. The output gate control how the cell state is exposed to the hidden state. The element wise multiplication \(\:\odot\:\) ensures that the gates can control the information flow. The information is accumulated over time in cell state \(\:{c}_{t}\) and the output that includes the temporal dependencies at each time step is provided by the hidden state \(\:{h}_{t}\). The recurrent layer in the proposed work processes the feature vectors \(\:{v}_{t}\:\)from the capsule network. The information about the previous state is retained in the recurrent layer to recognize the temporal patterns and changes which indicate the presence of disease in shrimps.
Spatial and channel attention mechanisms
The proposed model includes a spatial and channel attention mechanism to enhance the feature extraction performances. The attention mechanism allows the ERCN model to focus on the most relevant features in the input image which is essential for accurate disease detection. The spatial attention mechanism identifies and highlights the most important portions in an image. A spatial attention map is generated in which a weight is assigned to each spatial location in the feature map to represent its importance. The spatial attention mechanism is mathematically formulated as
where input feature map is indicated as \(\:F\), the spatial coordinates are indicated as \(\:\left(x,y\right)\) and the channel index is indicated as \(\:c\). The dimension of feature map is given as \(\:H\times\:W\times\:C\) in which \(\:H\) indicates the height, \(\:W\) indicated the width and \(\:C\) indicates the number of channels. The spatial attention map is computed by applying convolution to the input feature map. This convolution process collects information across the channels and generates a two-dimensional feature map \(\:{F}_{spatial}\). Then by applying an activation function \(\:\sigma\:\) the spatial attention maps \(\:{M}_{s}\) are generated. The spatial locations which are most essential are highlighted through the attention map. Finally, the spatially attended feature map \(\:{F}_{sa}\) is obtained by performing element wise multiplication of input feature map \(\:F\) and spatial attention maps \(\:{M}_{s}\).
The spatial attention mechanism highlights the important region in an image while the channel attention mechanism identifies the most informative features across different channels. The channel attention mechanism generates a channel attention map as follows.
\(\:{F}_{ca}\left(x,y,c\right)={M}_{c}\left(c\right)\cdot\:F\left(x,y,c\right)\) (24) \(\:{M}_{c}=\sigma\:\left(\text{FC}\left(G\right)\right)\) (25)
where \(\:F\) indicates the input feature map and it utilizes global average pooling to summarize the spatial information into channel descriptor. The global average pooling provides a one-dimensional feature vector F with dimensions of \(\:(1\times\:1\times\:C)\). The channel attention map \(\:{M}_{c}\) is obtained by applying a sigmoid activation function to the channel descriptor \(\:G\). Finally, the channel-attended feature map \(\:{F}_{ca}\) is obtained by performing element wise multiplication with input feature map \(\:F\) and channel attention map \(\:{M}_{c}\).
The proposed model utilizes the spatially attended feature map \(\:{F}_{sa}\) and channel attention feature map \(\:{F}_{ca}\) to produce the final feature map as follows.
where \(\:{F}_{final}\) indicates the final feature map that includes spatial and channel attention features.
To enhance interpretability and user trust in the proposed Enhanced Recurrent Capsule Network (ERCN), we have included visualizations of the spatial and channel attention maps alongside capsule activation patterns. These visualizations offer insights into the regions and features within shrimp images that the model emphasizes during the disease detection process. The spatial attention maps highlight disease-relevant areas in the input images, such as blackened gills or white spots, while disregarding irrelevant backgrounds. For instance, brighter regions in the maps correspond to areas with significant disease features, confirming that the model effectively identifies critical regions for classification. Similarly, the channel attention mechanism highlights the most informative features across different channels, showcasing how the model differentiates between diseases based on these features.
The capsule layers provide a hierarchical representation of spatial features, with activity vectors summarizing properties such as position and orientation. Visualizations of capsule activation patterns demonstrate how the dynamic routing mechanism selectively routes disease-relevant features to higher-level capsules, which reinforces the model’s decision-making process. Additionally, the recurrent layers analyze temporal dependencies in sequences of shrimp images, tracking the progression of disease symptoms over time. Visualizations of sequential attention patterns further highlight how the model integrates spatial, temporal, and feature-based information to make accurate predictions. By demonstrating the reasoning behind its predictions, the proposed ERCN model not only achieves high performance but also enhances user trust, offering actionable insights for stakeholders in shrimp aquaculture. This additional layer of explainability contributes to the reliability and practical applicability of the model in real-world settings.
Dual-Level feature fusion
In the next stage of proposed ERCN, dual level feature fusion is employed to combine the local and global features which are extracted from the input image. This feature fusion ensures the utilization of detailed and contextual information in the shrimp disease detection process. The features obtained through the capsule network, recurrent layer and attention mechanism are integrated in the fusion process. The local feature indicates the details and specific patterns in the small regions of the image. In the proposed work capsule network, captures the spatial hierarchies and its output vectors encodes the properties like position, scale and orientation to provide details about the local features. While the global features are obtained from the recurrent layers. The sequence of feature vectors \(\:{v}_{t}\) at each time step \(\:t\) is processed by the recurrent layer as follows.
where the hidden state is indicated as \(\:{h}_{t}\). From this hidden layer, the final global features are obtained. Further the local and global features are fine-tuned through spatial and channel attention mechanisms. Finally, the dual level feature fusion combines the fine-tuned local and global features into a single feature by concatenating the weighted local and global features as follows.
where the local features are indicated as \(\:{F}_{local}\), global features are indicated as \(\:{F}_{global}\), spatial attention map is indicated as \(\:{M}_{s}\) and channel attention map is indicated as \(\:{M}_{c}\). The element wise multiplication is indicated through \(\:\odot\:\) operator. The final fused vector is then passed into the classification layer to detect different types of diseases in shrimps.
Classification
The final layer in the proposed model is the classification layer which classifies the feature representations into specific class probabilities that indicate the presence or absence of diseases in shrimp images. The fused feature is processed by a fully connected layer which transforms the high dimensional feature vector into vector of logits. The logit vector obtained through the fully connected layer is formulated as
where \(\:{W}_{f}\) and \(\:{b}_{f}\) indicates the fully connected layer weights and biases respectively. \(\:z\) indicates the logits vector. The logits are then converted into class probabilities by applying a SoftMax activation function. The SoftMax function normalizes the logits and makes them interpretable as probabilities. Mathematically it is expressed as
where \(\:{z}_{k}\) indicates the logit for class \(\:k\). The proposed ERCN model is trained and the discrepancy between the true class labels and predicted probabilities are obtained through cross entropy loss function. The cross-entropy loss function used in the proposed work is formulated as
where the true label for class \(\:k\) is indicated as \(\:{y}_{true,k}\). Generally, model parameters like weight and biases are obtained through backpropagation during the training process. The gradient procedure indicates the direction of parameters to be adjusted to minimize the loss. To avoid this and to provide optimal parameters for better classification performance a hybrid optimization model is proposed in the research work.
Hybrid optimization model for ERCN parameter optimization
The proposed hybrid optimization18 for ERCN parameter optimization incorporates HHO and MPA. The proposed hybrid optimization algorithm provides a better balance between exploration and exploitation in search space to find the optimal solutions for the given problem.
The cooperative and chasing strategies of Harris Hawks are formulated in HHO optimization. The optimization begins with initializing the Hawks population that represents a potential solution. Consider \(\:X=\{{X}_{1},{X}_{2},\dots\:,{X}_{N}\}\) be the population of \(\:N\) hawks in which \(\:{X}_{i}\) indicates the vector of parameter for \(\:{i}^{th}\) hawk. After initialization, the fitness of each hawk \(\:f\left({X}_{i}\right)\) is evaluated through an objective function. In the proposed work, the entropy loss in the ERCN training process is considered as an objective function. After fitness evaluation, the process guiding the search process towards optimal solution is formulated considering prey as optimal solution. Mathematically the position of prey is given as
where \(\:{X}_{i}\) indicates the \(\:{i}^{th}\) hawk position, \(\:{X}_{prey}\) indicates the prey position. The \(\:arg\) function indicates that minimizes the fitness function \(\:\left(f\right)\) identifying the hawk with the best performance. The position of the prey is considered as reference point so that other hawks update their position based on this reference point. While the hawk trying to attack the prey, the escaping energy of the prey is formulated
where the initial escaping energy is indicated as \(\:{E}_{0}\), the current iteration is indicated as \(\:t\) and the total number of iterations are indicated as \(\:T\). The escaping energy indicated as \(\:E\) decreases over time as the prey gets tired which switches the algorithm from exploration to exploitation phase. The decrease of energy is linear from \(\:2{E}_{0}\) to 0. When the escaping energy is high the algorithm highlights the exploration and allows hacks to search in diverse regions. When the energy decreases the algorithm shifts towards exploitation and focuses on refining the solution around the best prey position.
The position update of hawk is performed based on the escaping energy and position of prey. Different strategies are followed based on the escaping energy to update the hawk position such as soft besiege, hard besiege, and surprise pounce. When the escaping energy is less than 1 \(\:\left|E\right|\ge\:1\), soft besiege is performed which is formulated as
In the hard besiege strategy, hawks are more aggressive and perform a direct approach in this stage the escaping energy will be \(\:\left|E\right|<1\) and it is formulated as
In the surprise pounce or rapid dive strategy the hawks perform rapid dives to surprise the prey which is formulated as
where \(\:{X}_{i}^{old}\) indicates the position of \(\:{i}^{th}\) hawk before update, and \(\:{X}_{i}^{new}\) indicates the position of \(\:{i}^{th}\) hawk after update. In order avoid local minima a random perturbation \(\:R\) is introduced in the position update process. When the escaping energy is high, the optimization model exploration phase searches the solution while in the exploitation phase, the optimization model finds the global optimum.
The hybrid model includes MPA in addition to HHO to obtain the optimal parameters. The foraging strategies of marine predators are formulated by combining the levy flight and Brownian motion so that a better balance between exploration and exploitation are obtained in the optimization process. The initialization of MPA includes \(\:P=\{{P}_{1},{P}_{2},\dots\:,{P}_{N}\}\) as the initial predator population in which \(\:{P}_{i}\) indicates the predator parameter vector. After evaluating the fitness of each predator, the optimization model utilizes levy flight and Brownian motion to model the marine predator movement. This combination provides better exploration and exploitation by obtaining different search patterns and effectively navigates the predators into solution space.
The levy flight formulates the series of step with length based on levy distribution and it allows the algorithm to explore the search space extensively. Levy provides the advantage that the search agent can move over large distance so that the optimization model avoids local optima and discovers the new search space. The flight of the levy is mathematically formulated as
where the step size is indicated as \(\:{\Delta\:}X\), search space scaling factor is indicated as \(\:S\) and it controls the step size. The levy distribution \(\:\text{Levy}\left({\uplambda\:}\right)\) is formulated as
where \(\:{\uplambda\:}\) is the distribution parameter. The levy distribution in the MPA produces a higher probability of large steps compared to normal distribution. This ensures the predators’ significant movement which improves the exploration ability.
The Brownian motion used in the MPA takes small successive steps in random directions. This small step movement is used to refine the local search and allows the predators to exploit for promising solution. The Brownian motion is mathematically formulated as
where normal distribution with mean zero and variance \(\:{{\upsigma\:}}^{2}\) is indicated as \(\:N\left(0,{{\upsigma\:}}^{2}\right)\). The MPA utilizes both levy and Brownian motion to provide better exploration and exploitation. Further the position update of predator is defined based on the step size \(\:{\Delta\:}X\) as follows.
where \(\:{P}_{i}^{old}\) indicates the old position and \(\:{P}_{i}^{new}\) indicates the predator new position. The step size \(\:{\Delta\:}X\) is obtained either by using levy flight or Brownian motion based on the strategy selected for the current iteration. The movement of predators based on levy and Brownian motion is formulated as
The position update through the combined strategy in MPA for exploration and exploitation phase is formulated as
The levy flight in the MPA provides wide exploration features while the Brownian motion focuses on exploitation to optimize the search process. The adaptive transition between these phases enhances the ability to solve complex optimization problems. The final solution of hybrid optimization combines the solutions of HHO and MPA through weighted averaging procedure. Mathematically it is formulated as
where \(\:\alpha\:\) indicates the weighting factor. The HHO best solution is indicted as \(\:{X}_{HHO}\) and the best solutions of MPA is indicated as \(\:{P}_{MPA}\). This combined solution is used to update the ERCN model parameters to minimize the loss and improve the detection performances. The summarized pseudocode for the proposed optimized ERCN model is presented as follows.

The hybrid Harris Hawks Optimization (HHO) and Marine Predator Algorithm (MPA) method is employed to optimize key parameters of the Enhanced Recurrent Capsule Network (ERCN), significantly enhancing its performance in shrimp disease detection. The parameters optimized include those related to the capsule network, such as the number of capsules and their dimensionality, which influence the network’s ability to encode spatial hierarchies effectively. In the recurrent layers, the number of LSTM units is optimized to ensure accurate modeling of temporal dependencies in sequential data. The spatial and channel attention mechanisms are fine-tuned by optimizing the spatial attention kernel size and the channel attention reduction ratio, enabling the model to focus on the most relevant regions and feature channels. Additionally, regularization parameters like the dropout rate and regularization weight are optimized to mitigate overfitting, while training parameters such as the batch size and learning rate are fine-tuned to balance computational efficiency and convergence. The cross-entropy weight in the loss function is also adjusted to achieve an optimal balance in training. By combining the exploration capabilities of HHO and the exploitation strengths of MPA, this hybrid optimization ensures optimal tuning of these parameters, resulting in improved accuracy, precision, recall, and other performance metrics, thereby enhancing the model’s reliability in detecting shrimp diseases.
Results and discussion
The proposed model experimentation analysis for shrimp disease detection is performed in python tool using our own dataset. The simulation environment includes essential python libraries like TensorFlow and Keras for deep learning model implementation. To perform image preprocessing OpenCV and scikit-image libraries are used in the proposed model experimentation. The proposed model is aimed at detecting different types of shrimp diseases thus the data set includes different diseased shrimp images. The dataset consists of shrimp images collected from various open-source platforms to ensure diversity in environmental conditions, lighting, and disease symptoms. The dataset includes a total of 1,599 images, distributed across six disease categories: Black Gill (320 images), Infectious Hypodermal and Hematopoietic Necrosis Virus (IHHNV; 280 images), White Spot Syndrome Virus (WSSV; 300 images), Yellow Head Virus (YHV; 180 images), Taura Syndrome Virus (TSV; 245 images), and Vibrios (274 images). The sample images of shrimps are presented in Fig. 3. To enhance the generalization ability of the model, data augmentation techniques such as rotation, flipping, scaling, and translation were applied to increase the number and variability of samples during preprocessing.
Additionally, the images were annotated using expert-based manual labeling to ensure high-quality annotations. To further improve dataset representativeness, segmentation was performed using the Grab Cut algorithm to isolate shrimp features from irrelevant backgrounds. The dataset distribution across categories has been clarified in Table 2 of the manuscript to highlight its representativeness. These steps ensure diversity and robustness, which contribute to improving the model’s generalization ability. From this, the entire dataset is divided into the ratio of 80:20 for training and testing. The details about the dataset are presented in Table 2. For training the proposed optimized ERCN model, 1279 sample images are used and to test the model 320 sample images are used.
Sample images in the dataset.
In order to attain the best performance in disease detection, the parameters of the proposed ERCN are optimized through the hybrid optimization algorithm that combines HHO with MPA. The simulation parameters used in the proposed model experimentation are listed in Table 3.
In the proposed work, input image is initially normalized and then data augmented to increase the number of samples. Further the images are segmented using Grab cut segmentation algorithm. This segmentation provides shrimp images and removes the external backgrounds in the input image. Due to this, the computation complexity of the proposed classifier is reduced.
The performance of the proposed model is evaluated through metrics like accuracy, precision, recall, f1-score, specificity, and Mathew correlation coefficient. Table 4 provides the performance results of the proposed model for the training and testing process. From table it can be observed that the proposed model achieved the highest accuracy of 95.8% in the training process and in the testing process the proposed model attained 95.2% of accuracy. The precision obtained in the training and testing process is 95.4% and 94.9% while the recall in the training and testing process is 93.8% and 93.5%. The F1-score of the proposed model for the training and testing process is 95.0% and 94.6%. The proposed model attained specificity of 98.0% and 97.8% for the training and testing process. In case of MCC, the proposed model exhibited 92.4% and 92.1% in the training and testing process. These results indicate the proposed model effectiveness in disease detection during the training and testing process.
The accuracy and loss curves of proposed model for training and testing process is presented in Figs. 4 and 5 respectively. The analysis given in Fig. 4 demonstrate the proposed model better accuracy through the stable line after 15 epochs. The accuracy increases from the beginning and after continuous update and fine tuning of parameters the proposed model exhibit accuracy of 95.8% in the training process and 95.2% average in the testing process. Similarly, the loss analysis given in Fig. 5 indicates the minimum loss of proposed model. During the initial stage of epoch, the loss is high, and it gradually reduces and reaches to minimum after 20th epoch. The minimum error continued till the last with slight variations which indicates the stable performance of the proposed model.
Accuracy vs. epoch.
Loss vs. epoch.
The precision recall analysis of proposed model is given in Fig. 6 for different diseases classes. It can be observed from the results, the average precision (AP) value of all the disease classes is above 0.99 which indicates the better performance of the proposed model. Specifically, the proposed model attained better AP for BG and YHV classes as 0.9948 and 0.9949 respectively. The minimum AP score 0.9876 of WSSV class indicates that the proposed model showed slightly reduced performance in detecting WSSV. However, the consistent performance of proposed model for all the other classes indicates the model reliability in multi-class classification.
Precision-Recall analysis.
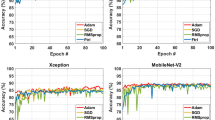
To evaluate the superior performance of the proposed model, the performance metrics are comparatively analysed with existing deep learning algorithms. As the dataset was created on our own, standard deep learning models are considered as existing methods and compared with the proposed model performance. Algorithms like CNN27, RNN28, LSTM29, GRU30 and VGG1631 are used for comparative analysis. Each model is implemented individually using the parameters listed in Table 5 and then the results are used to plot the comparative analysis graphs. For all the algorithms, the epoch is commonly selected, Adam optimizer is used for LSTM and VGG16, Nadam optimizer is used for GRU, RMS prop optimizer is used for RNN and CNN has SGD as optimizer.
The comparative analysis given in Fig. 7 depicts precision analysis of proposed and existing deep learning algorithms in shrimp disease detection. The result depicts the superior performance of proposed model reaching maximum of 94.9%, but the existing models RNN and VGG16 exhibits around 90% as precision which is 5% lesser than the proposed. Similarly, the CNN and GRU exhibits precision of around 88% which is 7% lesser than the proposed model. The precision of LSTM model is around 92% which is 3% lesser than the proposed model. This indicates the proposed model better learning ability and providing reliable results over different types of inputs in shrimp disease detection. Table 6 depicts the numerical values of precision attained by the proposed and existing deep learning algorithms.
Precision comparative analysis.
Recall comparative analysis.
The comparative analysis given in Fig. 8. depicts recall analysis of proposed and existing deep learning algorithms in shrimp disease detection. The result depicts the superior performance of proposed model reaching maximum of 93.5%, but the existing models RNN and VGG16 exhibits around 90% as recall which is 4% lesser than the proposed. Similarly, the CNN and GRU exhibits recall of around 87% which is 6% lesser than the proposed model. The precision of LSTM model is around 91% which is 3% lesser than the proposed model. Table 7 depicts the numerical values of recall attained by the proposed and existing deep learning algorithms. The superior recall value of the proposed model highlights the model ability in identifying relevant instances for disease detection compared to existing deep learning models.
F1-score comparative analysis.
The comparative analysis given in Fig. 9 depicts F1-score analysis of proposed and existing deep learning algorithms. The result depicts the proposed model superior performance with maximum f1-score of 94.6%. The existing models RNN and VGG16 exhibits around 90% as F1-score which is around 5% lesser than the proposed. Similarly, the CNN and GRU exhibits F1-score of around 88% which is 7% lesser than the proposed model. The F1-score of LSTM model is around 91% which is 4% lesser than the proposed model. The numerical values of F1-score attained by the proposed and existing deep learning algorithms are comparatively presented in Table 8 to better understand the superior performance of the proposed model.
The specificity comparative analysis given in Fig. 10 for the proposed and existing deep learning algorithms depicts the proposed model superior performance with maximum specificity of 97.8%. The existing models RNN and VGG16 exhibits specific of around 94% which is 4% lesser than the proposed. Similarly, the CNN and GRU exhibits specificity of around 93% which is 5% lesser than the proposed model. The LSTM model exhibit specificity of around 95% which is 2% lesser than the proposed model. Table 9 depicts the results of proposed and existing models for different epochs, and it highlights the superior performance of the proposed model.
Specificity comparative analysis.
Figure 11 depicts Mathew correlation coefficient analysis of proposed and existing deep learning algorithms. The better performance of proposed model is depicted in the comparative results with maximum MCC of 92.1% whereas the existing RNN and VGG16 models exhibits around 88% specificity which is 3% lesser than the proposed. Similarly, the specificity of CNN and GRU exhibits is around 86% which is 6% lesser than the proposed model. The specificity of LSTM model is around 89% which is 3% lesser than the proposed model. Table 10 depicts the numerical values of the proposed model and existing models for specificity metric.
MCC comparative analysis.
Accuracy comparative analysis.
The accuracy comparative analysis of proposed and existing model is given in Fig. 12 and the results depicts the maximum accuracy of proposed ERCN model over existing deep learning algorithms. The maximum accuracy of 95.2% is 4% higher than the RNN and VGG16 models. when compared to CNN and GRU, the proposed model exhibited 5% higher accuracy for 50th epoch. The performance of LSTM is comparatively less with accuracy of 92.3% which is 3% lesser than the proposed model. From the results the proposed ERCN model superior performance can be obtained and numerically it can be observed in Table 11.
Results in Table 12 indicate that the incorporation of HHO-MPA optimization significantly enhances the performance of the ERCN model. For instance, accuracy improves from 89.5 to 95.2%, resulting in an enhancement rate of 6.38%. Similarly, the precision, recall, and F1-score were improved by 6.99%, 7.23%, and 7.63%, respectively. These improvements demonstrate that the hybrid optimization method effectively fine-tunes the model parameters, enabling it to capture spatial and temporal features more accurately and enhance its classification capabilities. The specificity, which measures the ability to correctly identify negative cases, also shows a notable improvement of 3.71%, indicating the model’s robustness in detecting different types of shrimp diseases.
To evaluate the individual contributions of the Spatial Attention and Channel Attention modules to the performance of the proposed Enhanced Recurrent Capsule Network (ERCN), a series of ablation experiments were conducted. The model was tested under four configurations: (1) without any attention mechanism (baseline), (2) incorporating only the Spatial Attention module, (3) incorporating only the Channel Attention module, and (4) utilizing both modules (full model). For each configuration, the model was trained and tested using the same dataset and experimental setup as described in the manuscript. The evaluation metrics, including accuracy, precision, recall, F1-score, and specificity, were used to assess the impact of each attention module. This approach allowed us to isolate and quantify the performance improvements introduced by the Spatial and Channel Attention modules, as well as their combined effect. The results, summarized in Table 13, provide a comprehensive understanding of how these attention mechanisms enhance the model’s detection capabilities.
The results from the ablation analysis shown in Table 13 clearly demonstrate the contributions of the Spatial and Channel Attention modules to the performance of the proposed ERCN model. It is found that incorporating only the Spatial Attention module improved the model’s ability to focus on critical spatial regions within the input images, resulting in a notable increase in precision and accuracy compared to the baseline. Similarly, using only the Channel Attention module enhanced the selection of discriminative features across channels, leading to a significant improvement in recall and F1-score over the baseline. The full model, combining both modules, achieved the highest performance across all metrics, emphasizing the complementary effects of these attention mechanisms. The accuracy improved by 6.1% over the baseline, while the F1-score and specificity improved by 6.7% and 3.5%, respectively, highlighting the synergistic benefits of integrating both modules.
The computational complexity of the model was evaluated by analyzing the number of parameters, floating-point operations per second (FLOPs), and memory requirements. The proposed ERCN model, which incorporates capsule networks, recurrent layers, and attention mechanisms, contains approximately 2.5 million trainable parameters and requires 4.1 GFLOPs for processing an input image. While this complexity is higher than traditional CNN models, it is justified by the enhanced detection accuracy and robustness achieved by the proposed architecture. The real-time performance was measured in terms of inference time per image using a standard system configuration with an NVIDIA RTX 3060 GPU and TensorFlow framework. The average detection time was found to be 58 ms per image, which translates to approximately 17 frames per second (FPS). This performance demonstrates the model’s suitability for real-time disease detection in practical aquaculture scenarios.
Conclusion
This research work presents an optimized ERCN model for detecting different types of shrimp diseases. The proposed optimized ERCN model effectively captures the spatial and temporal features from the segmented images. The features are then combined through dual level feature fusion procedure and then classified to detect different types of diseases. The parameters of the classifier are optimized through a hybrid optimization model that combines HHO with MPA. The detection accuracy of the proposed model is enhanced due to the optimal selection of parameters using the hybrid optimization algorithm. The experimental results demonstrate the superior performance of proposed model with detection accuracy of 95.2% over existing CNN, RNN, LSTM, GRU, and VGG16 models which attain accuracy of 90.1%, 91.3%, 92.3%, 89.8% and 91.7%. Though the proposed model has better detection performance, it has a minor limitation in terms of its computation complexity due to multiple algorithms. However, it can be neglected due to its higher detection performance. In future, this research work can be extended to optimize the computational efficiency of the proposed model considering quantization techniques.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Patil, P. K., Geetha, R., Ravisankar, T., Avunje, S. & Solanki, H. G. Thangapalam Jawahar Abraham, Sakkarai Pandiyan Vinoth, Karingalakkandy Poochirian Jithendran, Shankar Vinayakarao Alavandi, Koyadan Kizhakedath Vijayan (2021), economic loss due to diseases in Indian shrimp farming with special reference to enterocytozoon hepatopenaei (EHP) and white spot syndrome virus (WSSV). Aquaculture, 533, pp. 1–10 .
Frank Asche, J. L. et al. The economics of shrimp disease. J. Invertebr. Pathol. 186, 1–34 (2021).
Riya Rajeev, S. P. S., Kiran, G. S. & Joseph Selvin Shrimp disease management for sustainable aquaculture: innovations from nanotechnology and biotechnology. Aquacult. Int. 29, 1591–1620 (2021).
Troy, D. et al. Penaeid shrimp in Chesapeake Bay: population growth and black gill disease syndrome. Mar. Coastal. Fisheries. 13, 159–173 (2021).
Prochaska, J., Poompuang, S., Koonawootrittriron, S., Sukhavachana, S. & Uthairat Na-Nakorn Evaluation of a commercial SPF Litopenaeus vannamei shrimp breeding program: resistance to infectious myonecrosis virus (IMNV), Taura syndrome virus (TSV), and white spot syndrome virus (WSSV) from laboratory challenges. Aquaculture 554, 1–12 (2022).
Islam, S. I., Mou, M. J., Sanjida, S. & Sarower Mahfuj A review on molecular detection techniques of white spot syndrome virus: perspectives of problems and solutions in shrimp farming. Veterinary Med. Sci. 9, 778–801 (2023).
P, A. M., Anandajothi, E., Vasudevan, S. & Suresh, E. Major viral diseases in culturable Penaeid shrimps: a review. Aquacult. Int. 28, 1939–1967 (2020).
Aly, S. M., Mansour, S. M., Thabet, R. Y. & Mabrok, M. Studies on infectious myonecrosis virus (IMNV) and infectious hypodermal and hematopoietic necrosis virus (IHHNV) in cultured Penaeid shrimp in Egypt. Dis. Aquat. Organ. 143, 57–67 (2021).
Navaneeth, K. A. et al. Characterization of Vibrio parahaemolyticus isolates from shrimp farms of Southeast coast of India with special reference to Acute Hepatopancreatic Necrosis Disease (AHPND) status, Aquaculture, p. 513:1–39. (2020).
Baskaran Babu, G. et al. Surveillance of disease incidence in shrimp farms located in the East coastal region of India and in vitro antibacterial efficacy of probiotics against vibrio parahaemolyticus. J. Invertebr. Pathol. pp. 179, 1–13 (2021).
Samuel Manoharan, J., Gopal, S. T., Muthu, S. & Selvaraju, S. & Md. Saad Patel. (2022). A hybrid fuzzy based cross neighbor filtering (HF-CNF) for image enhancement of fine and coarse powder scanned electron microscopy (SEM) images. J. Intell. Fuzzy Syst., 42, 6, pp. 6159–6169
Pandurangan, R., Samuel Manoharan, J., Rajalingam, S., Angelo, M. & K A novel hybrid machine learning approach for traffic sign detection using CNN-GRNN. J. Intell. Fuzzy Syst. 44, 1283–1303 (2023).
Quach, N. D. T. L. D. & Chi-Ngon Nguyen Towards classification of shrimp diseases using transferred convolutional neural networks. Adv. Sci. Technol. Eng. Syst. J. (ASTESJ). 5 (4), 724–732 (2020).
Ming Sun, X. & Yang, Y. X. Deep learning in aquaculture: A review. J. Computers. 31 (1), 294–319 (2020).
Edeh, M. O., Dalal, S., Obagbuwa, I. C. & Siva Prasad, B. V. V. Shalini Zanzote Ninoria, Mohd Anas Wajid & Ademola Olusola adesina (2022), bootstrapping random forest and CHAID for prediction of white spot disease among shrimp farmers. Sci. Rep., 12, pp. 1–12 .
Khiem, N. M., Takahashi, Y., Oanh, D. T. H., Hai, T. N., Nobuo, K. & Hiroki Yasuma & The use of machine learning to predict acute hepatopancreatic necrosis disease (AHPND) in shrimp farmed on the East Coast of the Mekong delta of Vietnam. Fish. Sci. 86, 673–683 (2020).
Al-Ansari, T. T. T. N. & Nguyen, D. D. Hai Minh Le, thi Nga Quynh Phan, Indra Prakash, Romulus Costache & Binh Thai pham (2023), prediction of white spot disease susceptibility in shrimps using decision trees-based machine learning models. Appl. Water Sci., 14, 2, pp.1–15 .
Barik, R. C. et al. Spot disease identification using unsupervised machine learning based image segmentation with its remedial solution in aquatic fauna. J. Surv. Fisheries Sci. 10 (2), 912–922 (2023).
Ramachandran, L. & Mohan, V. A novel neural network model for shrimp segmentation to detect white spot syndrome. J. Intell. Fuzzy Syst. 43 (1), 1453–1466 (2022).
Ramachandran, L., Mangaiyarkarasi, S. P., Subramanian, A. & Senthilkumar, S. Shrimp classification for white spot syndrome detection through enhanced gated recurrent Unit-based wild geese migration optimization algorithm. Virus Genes. 60 (2), 134–147 (2024).
Ramachandran, L., Mohan, V., Senthilkumar, S. & Ganesh, J. Early detection and identification of white spot syndrome in shrimp using an improved deep convolutional neural network. J. Intell. Fuzzy Syst. 45 (4), 6429–6440 (2023).
Thotakura, K. V. V. P. et al. White spot syndrome detection in shrimp using neural network model Proceedings of the 18th INDIACom; INDIACom-2024; IEEE Conference ID: 57xxx, 2024 11th International Conference on Computing for Sustainable Global Development, 28th Feb-01st March 2024, Bharati Vidyapeeth’s Institute of Computer Applications and Management (BVICAM), New Delhi (INDIA). (2024).
Park, S. C. et al. Jee Eun Han (2020), detection of infectious hypodermal and hematopoietic necrosis virus and white spot syndrome virus in whiteleg shrimp (Penaeus vannamei) imported from Vietnam to South Korea. J. Vet. Sci., 21, 2, pp. 1–5 .
Vijaya Kumar Reddy, L. G. R., Srinivasa Rao, B., Rajesh, G. & Chandra Optimized threshold-based EM fusion technique for shrimp white spot disease detection. Int. J. Adv. Trends Comput. Sci. Eng. 9 (4), 6577–6584 (2020).
Jhansi, G. & Sujatha, K. HRFSVM: identification of fish disease using hybrid random forest and support vector machine. Environ. Monit. Assess. 195, 1–18 (2023).
Visumathi, P. K. J, Hybrid approach of CNN and SVM for shrimp freshness diagnosis in aquaculture monitoring system using IoT based learning support system. J. Internet Technol., 23, 4, pp. 801–810 .
Mercedes, E., Paoletti, J. M., Haut, J. & Plaza, A. P. Scalable recurrent neural network for hyperspectral image classification. J. Supercomputing. 76, 8866–8882 (2020).
Sudipan Saha; Francesca Bovolo; Lorenzo Bruzzone. Change detection in image Time-Series using unsupervised LSTM, IEEE geoscience and remote sensing letters, 19, pp. 1–5. (2022).
Erting Pan, X., Mei, Q., Wang, Y. & Ma, J. M. Spectral-spatial classification for hyperspectral image based on a single GRU. Neurocomputing 387, 150–160 (2020).
Tamnuwat Valeeprakhon, K. & Orkphol and Penpun Chaihuadjaroen Deep Constitutional Neural Networks based on VGG-16 Transfer Learning for Abnormalities Peeled Shrimp Classification, International Scientific Journal of Engineering and Technology, vo. 6, no. 2, pp. 13–23. (2022).
Keshun, Y., Zengwei, L. & Yingkui G. A performance-interpretable intelligent fusion of sound and vibration signals for bearing fault diagnosis via dynamic CAME, Nonlinear Dynamics, 112, 20903–20940. (2024). https://doi.org/10.1007/s11071-024-10157-1
You Keshun and Liu Huizhong. Feature detection of mineral zoning in spiral slope flow under complex conditions based on improved YOLOv5 algorithm. Phys. Scr. 99 (1), 016001. https://doi.org/10.1088/1402-4896/ad0f7d (2023).
Keshun, Y., Puzhou, W. & Peng, H. Gu Yingkui A sound-vibration physical-information fusion constraint-guided deep learning method for rolling bearing fault diagnosis, Reliability Engineering & System Safety, 253, 110556. (2025).
Wang, Y. K., Gu, P. & Yingkui Toward efficient and interpretative rolling bearing fault diagnosis via quadratic neural network with Bi-LSTM. IEEE Internet Things J. 11 (13), 23002–23019 (2024).
Keshun, You & Guangqi Qiu and Yingkui Gu. An efficient lightweight neural network using BiLSTM-SCN-CBAM with PCA-ICEEMDAN for diagnosing rolling bearing faults. Meas. Sci. Technol. 34, 094001. https://doi.org/10.1088/1361–6501/acd5ef (2023).
You Keshun; Qiu Guangqi; Gu Yingkui. A 3-D Attention-Enhanced hybrid neural network for turbofan engine remaining life prediction using CNN and BiLSTM models. IEEE Sens. J. 24 (14), 21893–21905 (2024).
Acknowledgements
Not Applicable.
Funding
No funding was received for this research work.
Author information
Authors and Affiliations
Contributions
All the authors contributed to this research work in terms of concept creation, conduct of the research work, and manuscript preparation.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not Applicable.
Consent for publication
Not Applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Raj, A.S., Senthilkumar, S., Radha, R. et al. Enhanced recurrent capsule network with hyrbid optimization model for shrimp disease detection. Sci Rep 15, 10400 (2025). https://doi.org/10.1038/s41598-025-94413-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-94413-3
Keywords
This article is cited by
-
A Hybrid Self-Attention LSTM-XGBoost Model for Cardiovascular Disease Risk Prediction in Patients with Obstructive Sleep Apnea Using Sleep Heart Rate Variability Analysis
Sleep and Vigilance (2026)
-
A Hybrid Spatial Temporal Transformer Network Optimised for Accurate Disease Identification in Shrimp Producers
Iranian Journal of Science and Technology, Transactions of Electrical Engineering (2025)