Abstract
The evolution of COVID-19 pandemic has been characterized by the rapid emergence of new SARS-CoV-2 variants, each of which poses unique challenges to public health. This study analyzes the dispersion profiles during the Pre-Omicron and Post-Omicron phases in different epidemiological contexts. The Brazilian state of Espirito Santo, despite its low population density, plays a critical role as a commercial hub due to its intense port activity, which may have contributed to COVID-19 cases and mortality rates being higher than the national average. The state recorded 34,000 confirmed cases and 377 deaths per 100,000 inhabitants. Genomic surveillance revealed that the Pre-Omicron phase was dominated by the B.1.1.33 lineage, characterized by localized intraregional circulation. In contrast, the Post-Omicron phase, dominated by the BQ.1.1 lineage, exhibited greater diversity in circulating lineages, increased international interactions, and rapid viral dissemination, highlighting distinct transmission dynamics between such periods. This study highlights the need for adaptive public health strategies that account for both viral behavior and regional socioeconomic factors, while highlighting the strategic importance of Espirito Santo in monitoring SARS-CoV-2 evolution.
Similar content being viewed by others
Introduction
The evolution of COVID-19 pandemic has been marked by a series of challenges posed by the rapid emergence of new SARS-CoV-2 variants, each with distinct characteristics that significantly impacted disease control strategies1,2.The transition between different phases of the pandemic, particularly concerning the emergence of the Omicron variant and its sublineages, underscores the need to understand not only the virulence of these new lineages, but also their dynamics of spread and prevalence in populations previously exposed to different epidemiological contexts and public health interventions3. In this context, it becomes imperative to explore how these variants, particularly in the early period and throughout the pandemic, following the emergence of the Omicron variant, behaved in terms of dissemination and adaptation. And especially considering changes in the global pandemic landscape, such as the production of vaccines and the relaxation of social measures3,4.
In Brazil, the beginning of the pandemic was marked by the confirmation of the first COVID-19 case in February 2020, followed by the spread of the B.1.1.28 (PANGO lineage, ancestral to Gamma and Zeta variants, GISAID clade GR) and B.1.1.33 (PANGO lineage, GISAID clade GR) lineages, which dominated nearly all Federation States and drove the first wave of cases and deaths5,6. Subsequently, in October 2020, the P.2 (Zeta, PANGO lineage, GISAID clade GR, WHO-classified VOI) lineage emerged in Rio de Janeiro state, spreading rapidly and becoming dominant in several regions by January 20217. Almost simultaneously, the P.1 (Gamma, PANGO lineage, GISAID clade GR/GRS, WHO-classified VOC) variant emerged in Amazonas state and, owing to its high transmissibility and ability to evade the immune system, it rapidly became the dominant variant throughout the country, significantly exacerbating the ongoing health crisis8,9.
Brazil also experienced the introduction of other variants of concern (VOCs) and variants of interest (VOIs), including the Mu variant (B.1.621), which was first detected in Colombia in January 2021 and introduced into Brazil between mid and late 2021, primarily through neighboring countries. Although the Mu variant did not achieve dominance like Gamma or Delta (B.1.617.2, GISAID clade G/478K.V1, WHO-classified VOC), its presence during this period highlighted the importance of Brazil’s geographical and epidemiological dynamics in the dissemination of SARS-CoV-2, especially due to its connections with other South American countries10.
In mid-2021, the Delta variant, originally identified in India, began to compete with Gamma, resulting in the cocirculation of both variants in different regions of Brazil, marking a new phase of the pandemic11. By the end of 2021, the Omicron variant (PANGO lineage B.1.1.529, GISAID clade GRA, WHO-classified VOC) led to an exponential increase in cases, driven by its multiple mutations in the spike protein, although the severity of infections was lower, possibly owing to widespread vaccination. In this context, the Omicron sublineage BQ.1.1 (PANGO lineage, GISAID clade GRA, WHO-classified VOC) emerged as a major variant, standing out owing to its high transmissibility and ability to partially evade previously acquired immunity12.
In this context, characterized by a rising number of positive cases and severe impacts on public health, including overload of healthcare systems and high mortality rates, as well as profound economic losses resulting from necessary containment measures, genomic surveillance has emerged as a critical tool for investigating and tracking the evolution and spread of the virus13.
Accordingly, by the end of July 2024, over 16 million viral genome sequences detected worldwide had been deposited in open-access databases such as GISAID. The United States, Japan, and European countries such as the United Kingdom, Germany, and France stand out with 5.9 million, 682 thousand, 3.2 million, 950 thousand, and 675 thousand deposited sequences, respectively. Brazil, with over 250 thousand deposited sequences, has made significant contributions to genomic surveillance14.
The contributions of genomic surveillance are essential for pandemic management, allowing for real-time monitoring of SARS-CoV-2 evolution, detection of virus dispersal patterns, and identification of new variants with greater pathogenic potential, transmissibility, and ability to evade natural or vaccine-induced immunity, as well as reduced susceptibility to therapeutic agents. This information has supported the development of effective measures to minimize damage and guide pandemic control strategies, as each wave of the epidemic presented unique challenges that required adaptations in public health policies15,16.
The primary objective of this study was to analyze the dispersion profiles of the SARS-CoV-2 lineages B.1.1.33 (Pre-Omicron) and BQ.1.1 (Post-Omicron) across different epidemiological contexts. By comparing these lineages, this study aims to shed light on how the virus evolved in response to various public health interventions and epidemiological conditions. We believe that the analysis conducted here provides valuable insights into the effectiveness of these responses, offering evidence-based recommendations for future strategic decisions and interventions to manage potential.
Understanding the spatial and temporal dynamics of SARS-CoV-2 during the Pre- and Post-Omicron periods provides critical insights into how public health interventions and viral evolution shaped the progression of the pandemic. By selecting lineages based on the prevalence of sequenced samples in each period, this study highlights the key transitions in viral transmissibility and diversity. These findings establish a robust framework for future preparedness, offering important lessons for managing emerging pathogens in a rapidly evolving global health landscape.
Results
Diversity and prevalence of SARS-CoV-2 lineages in local samples
A total of 215 samples were subjected to sequencing, and a minimum coverage of 50% was achieved for approximately 86.5% (186/215) of the samples. Among these samples, 46.8% (n=87) corresponded to samples from the Pre-Omicron group, and 53.2% (n=99) corresponded to those from the Post-Omicron group. Among the sequenced samples, the circulation of multiple SARS-CoV-2 lineages was observed during the studied periods (Fig. 1). The most prevalent lineages during the Pre-Omicron period were B.1.1.33 (52.9%, n=46), B.1.1.28.2 (17.2%, n=15), and B.1.1.28.1 (10.3%, n=9). In the Post-Omicron period, a greater diversity of circulating lineages was observed, with the BQ.1.1 lineage being the most prevalent at 60.6% (n=60), followed by BE.9 at 9.1% (n=9) and BQ.1.18 at 7.1% (n=7).
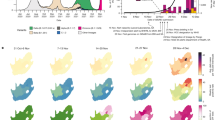
Distribution of locally sequenced SARS-CoV-2 lineages from the Pre and Post-Omicron periods in Espirito Santo state, Brazil. The figure was generated using Python (Version 3.12, https://www.python.org/) with Matplotlib (Version 3.9.1, https://matplotlib.org/) and Pandas (Version 2.2.2, https://pandas.pydata.org/).
The representativeness of the sample collection in this study was assessed by comparing the lineage distributions in our samples with those from sequences of Espirito Santo deposited in GISAID during the same periods. A comprehensive analysis revealed consistent patterns between the datasets, reinforcing the robustness of the findings. Local sequencing of the samples revealed the cocirculation of multiple lineages (Fig. 2B). In the Pre-Omicron group, the same circulation pattern observed in GISAID was found (Fig. 2A,B), with cocirculation becoming more pronounced from January 2021. In the Post-Omicron group, the distribution of lineages deposited in the database in the analyzed period (Fig. 2C) revealed a high diversity of sublineages of the Omicron variant circulating in a short period of time, similar to that observed in the study samples (Fig. 2D). The most prevalent lineages in GISAID were also identified via our sequencing, reinforcing the representativeness and validity of our observations, emphasizing that the variations between the datasets can be attributed to the number of sequenced samples.
Temporal Distribution of Circulating Lineages in Pre- and Post-Omicron Groups (2020-07-09 to 2021-03-29 and 2022-11-14 to 2022-12-28). (A) Distribution of sequences deposited in GISAID for lineages identified in Espirito Santo during the Pre-Omicron period. (B) Distribution profile of lineages identified by sequencing conducted in the present study during the Pre-Omicron period. (C) Distribution of sequences deposited in GISAID for lineages identified in Espirito Santo during the Post-Omicron period. (D) Distribution profile of lineages identified by sequencing conducted in the present study during the Post-Omicron period. *Other (Post-Omicron GISAID sequences): BA.2.75.5, BA.5.2.57, BQ.1.14, EF.1.1.1, BA.4.6, BA.5.3.1, BF.39, BF.41.1, BF.7.5, BN.1.5, BQ.1, BQ.1.1.17, BQ.1.1.18, BQ.1.1.4, BQ.1.1.60, BQ.1.18, BQ.1.2, BQ.1.22, BQ.1.25, BQ.1.3.1, EE.2, XBB.1, XBB.1.18, BN.1.3.1. **Other (Post-Omicron local sequences): BA.4.6, BA.5.2.1, BA.5.2.7 , BN.3.1, BQ.1.1.18, BQ.1.1.60, BQ.1.22, EF.1.1, BQ.1. The figure was generated using Python (Version 3.12, https://www.python.org/) with Matplotlib (Version 3.9.1, https://matplotlib.org/) and Pandas (Version 2.2.2, https://pandas.pydata.org/).
Building a comprehensive dataset for SARS-CoV-2 phylogenetic analysis
The datasets were designed to include the most representative samples from the analyzed periods, prioritizing lineages with higher prevalence as identified through local sequencing. Lineages B.1.1.33 and BQ.1.1 were selected to represent the Pre-Omicron and Post-Omicron groups, respectively, based on their dominance in the corresponding periods. The Pre-Omicron group included 43 samples (coverage> 90%) sequenced in this study, as well as 3879 sequences obtained from GISAID, totaling 3922 sequences. The Post-Omicron group comprised 4627 sequences, of which 21 samples (coverage> 90%) were sequenced in this study, and 4606 sequences were obtained from GISAID, enabling a detailed analysis of the distribution and evolution of these lineages throughout the different periods of the pandemic.
For the final dataset, which integrates sequences generated in this study with those retrieved from GISAID, demographic information such as sex and age was available only for a subset of patients. Among the sequences in the Pre-Omicron dataset with available demographic data, the mean age was \(45.5 \pm 18.1\) years, and 45% (N=1762) of these patients were female. In the Post-Omicron group, the mean age was \(49.8 \pm 21.3\) years, and, similarly, among the patients with available demographic data, 38.8% (N=1803) were female. It is important to note that these percentages reflect only the subset of sequences with linked demographic information and do not represent the entire dataset. Detailed demographic data for both groups are provided in Supplementary Table S2.
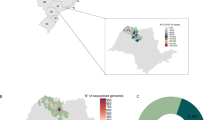
Figure 3 shows the collection locations of the samples that make up the Pre- and Post-Omicron groups worldwide and, in more detail, in the Brazilian context on the basis of the final dataset. Figure 3A shows the origin of the samples in the Pre-Omicron group, highlighting the significant predominance of samples from Brazil, representing 88.2% of the total. Other countries contributed a smaller proportion of samples, with the United States contributing 3.8%, Chile contributing 1.9%, and Uruguay contributing 1.2%. The geographic distribution of the samples shifted significantly from the Pre-Omicron group to the Post-Omicron group (Fig. 3b). While Brazil remained a major contributor (20.21%), the Post-Omicron group presented a more balanced representation of samples from various countries worldwide. The United States led with 29.83% of the samples, followed by France (9.75%), the United Kingdom (5.84%), and Canada (5.58%), indicating widespread viral dissemination in these regions.
Geographic distribution of absolute numbers of SARS-CoV-2 sample sequences from the Pre-Omicron (a) and Post-Omicron (b) groups and the normalized distribution of sequences by Brazilian state per 100,000 inhabitants. In panel (a), the state of Acre (AC) is identified below the respective values, while in both panels (a) and (b), the state of Espirito Santo (ES) is labeled for clarity. The numbers represent the composition of the dataset selection based on the similarity of the sequences from Espirito Santo in the Post-Omicron group and do not reflect the actual number of sequences deposited in GISAID. (The maps were generated using Microsoft Excel 365 (Microsoft® Excel® for Microsoft 365 MSO, Version 2501 Build 16.0.18429.20132, 64-bit, www.microsoft.com) with built-in mapping features.).
When normalizing the SARS-CoV-2 sample distribution in Brazil for the Pre- and Post-Omicron groups, Espirito Santo, despite its smaller population than other states in the federation, had rates of 3.20 and 8.8 samples per 100,000 inhabitants for Pre and Post-Omicron, respectively (Fig. 3A,B). In comparison, Sao Paulo state, with the largest population, presented rates of 1.70 and 0.46 samples per 100,000 inhabitants for the Pre and Post-Omicron groups, respectively. Moreover, Rio de Janeiro state had a significantly higher rate of 4.32 for the Pre-Omicron group. With respect to the Pre-Omicron group, Acre state had the highest rate, with 23.25 samples per 100,000 inhabitants.
Evolution of SARS-CoV-2 from local clusters to global dissemination
After phylogenetic analysis of the Pre-Omicron group (Fig. 4A), we selected clusters comprising two sequences from the Espirito Santo state.
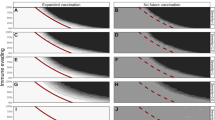
Phylogenetic analysis of Pre-Omicron SARS-CoV-2 sequences via two distinct methods. (A) Phylogenetic tree generated via the maximum likelihood (ML) method, showing the evolutionary relationships among the samples on the basis of their genomes. This tree can be accessed as an interactive version via the link: https://itol.embl.de/shared/MHnpDKTypmbn. (B) Phylogenetic tree generated via Bayesian inference, highlighting the posterior probability of the phylogenetic relationships. Both trees illustrate the genetic diversity and relationships among sequences before the introduction of the Omicron variant.
In the Pre-Omicron group, 12 monophyletic clusters (bootstrap/SH-aLRT > 96/90%) were observed, composed predominantly or exclusively of samples from Espirito Santo state, highlighting the widespread circulation of the virus within the state. With the exception of clusters 6, 9, and 10, the other clusters were composed entirely of samples from Espirito Santo, indicating intraregional transmission. On the other hand, clusters such as clusters 6 and 10 presented heterogeneous compositions, with samples from several Brazilian states. Cluster 6 was composed of 73.7% of the samples from Espirito Santo and 26.3% from other states, specifically Sao Paulo (3), Parana (1), and Rio de Janeiro (1). Similarly, cluster 10 also presented a considerable composition of samples, with 26.5% of the sequences coming from other Brazilian states, including Alagoas, Amazonas, Bahia, and Ceara, as well as a sample from Paraguay, and several from Rio de Janeiro and Sao Paulo. Cluster 9 was particularly diverse, with 45.5% distributed among the following Brazilian states: Amazonas, Bahia, Ceara, Maranhao, Minas Gerais, Rio de Janeiro, Santa Catarina, Sergipe, and Sao Paulo, as well as including international samples from the United States.
Following the phylogenetic analysis and the definition of the Pre-Omicron group clusters (Supplementary Material), three monophyletic groups were selected for the Bayesian analysis and inference of the origin and dispersion of this variant (Fig. 4B). Clusters 6, 9, and 10 were evaluated individually, totaling 143 sequences by grouping each cluster with the sequences of their respective ancestral nodes. Among these sequences, the majority (68.5%) originated from Espirito Santo state.
The Bayesian analysis revealed that Cluster 6, with posterior support of 1.00, had its most recent common ancestor (tMRCA) estimated to have emerged in mid-August 2020, with a high posterior density (HPD) interval between July and late August 2020. This clade appears to have originated in Espirito Santo, and subsequently exported the virus to other Brazilian states, including Parana (PR), Rio de Janeiro (RJ), and Sao Paulo (SP), covering the South and Southeast Regions.
The tMRCA of Cluster 9 (posterior support of 0.99) was estimated until June 2021, with an HPD interval between May and August 2021. This clade originated in Espirito Santo and was responsible for spreading the virus to several states, including Sao Paulo (SP), Maranhao (MA), Amazonas (AM), Sergipe (SE), Minas Gerais (MG), Bahia (BA), and Rio de Janeiro (RJ), covering Northeast, North, and Southeast Regions, as well as international exportation to the United States (USA). Finally, cluster 10, with support from cluster 1, had Espirito Santo as its origin and exported to several states, including Sao Paulo (SP), Alagoas (AL), Amazonas (AM), Bahia (BA), Ceará (CE) and Rio de Janeiro (RJ), in addition to international exports to Paraguay. This cluster probably began to circulate in June 2021, with a confidence interval between May and July 2021.
These results demonstrate that in the analyzed groups, the branches are longer, suggesting a dispersion pattern characterized by viral introduction in Espirito Santo state that occurred some time ago, followed by slower spread. This pattern indicates that the virus circulated gradually, resulting in a more prolonged and less accelerated transmission chain.
The phylogenetic profile in the Post-Omicron period (Fig. 5A), compared to the Pre-Omicron period, reveals a more complex and internationalized pattern of viral dispersion. While 12 clusters, predominantly composed of samples from Espirito Santo state, were observed during the Pre-Omicron period, with some mixed clusters, the new analysis shows an increase in the total number of clusters (25), and a greater diversity of external samples.
Phylogenetic analysis of Post-Omicron SARS-CoV-2 sequences via two distinct methods. (A) Phylogenetic tree generated via the Maximum Likelihood (ML) method, which illustrates the evolutionary relationships among the samples on the basis of their genomes. This tree can be accessed as an interactive version via the link: https://itol.embl.de/shared/M38MGliM59CO. (B) Phylogenetic tree generated via Bayesian inference, highlighting the posterior probability of the phylogenetic relationships. Both trees depict the genetic diversity and relationships among sequences after the introduction of the Omicron variant.
In the Pre-Omicron period, most clusters were exclusively composed of samples from Espirito Santo state, with the exceptions of a mixed composition of local and external samples. However, in the Post-Omicron period, there was a significant increase in the presence of clusters with mixed compositions, including international samples and those from other regions of Brazil. Clusters such as 7, 8, and 9 stand out for their diverse external samples, including sequences from different countries and continents.
Cluster 8, in the Post-Omicron period, comprises 86.6% of the external samples, whereas in the Pre-Omicron period, the most diverse cluster (Cluster 9) comprises 45.5% of the external samples, suggesting the introduction of the virus into Espirito Santo state from diverse global and national sources, which contrasts with the more balanced composition observed in the Pre-Omicron period.
In contrast, when we analyzed the Post-Omicron period through the Bayesian approach (Figure 05-B), and focused on the transmission chains within the Espirito Santo state, we observed a distinct profile of viral dissemination. The tMRCAs of the main clades of the BQ.1 lineage indicate recent dates, with Cluster 1 presenting the tMRCA on November 3, 2022 (HPD: October 9 to November 29, 2022); Cluster 5 with the tMRCA on October 5, 2022 (HPD: September 28 to November 21, 2022); and Cluster 23 with the tMRCA on October 31, 2022 (HPD: October 16 to November 11, 2022).
Discussion
In this study, we presented the molecular epidemiology of SARS-CoV-2 in two distinct periods: Pre-Omicron, which analyzes the dispersion profile of the B.1.1.33 lineage, and Post-Omicron, which focuses on the BQ.1.1 (Omicron, VOC) lineage. By understanding the dispersion dynamics of these lineages at different times, we highlight the main characteristics of each period and how implemented measures influenced pandemic outcomes.
The comparative analysis of the Pre-Omicron and Post-Omicron periods not only documents distinct epidemiological patterns but also underscores the importance of integrating molecular epidemiology with public health strategies. By focusing on transitions driven by strict interventions and vaccination, our findings provide a model for understanding the interplay between viral behavior and sociopolitical dynamics, which is essential for future pandemic preparedness.
Additionally, the dispersion patterns of these lineages from the Brazilian state of Espirito Santo, a significant commercial hub, due to its intense port activity, were analyzed. Despite being a state with one of the lowest population densities in the country, COVID-19 cases and mortality rates in Espirito Santo exceeded the national average (18,000 confirmed cases and 339 deaths per 100,000 inhabitants), with Espirito Santo reporting 34,000 confirmed cases and 377 deaths per 100,000 inhabitants17,18.
The analysis of Pre and Post-Omicron datasets revealed distinct patterns in SARS-CoV-2 evolution as well as dissemination throughout the pandemic. As discussed by Candido et al.19, the Pre-Omicron group, dominated by the B.1.1.33 lineage and with a concentration of samples within Brazil, indicates a lower diversity of circulating lineages in the early phase of the pandemic19,20. This pattern can be attributed to the strict containment measures implemented, which limited mobility and, consequently, the introduction of new variants into the country, as well as slower viral evolution at that time10,21.
In contrast, the Post-Omicron group showed greater lineage diversity, driven by both the virus’s natural evolution and advancements in sequencing technologies, which allowed for more accurate detection and characterization of circulating viruses10,21. Parther et al.22 noted that the selective pressure exerted by vaccination contributed to the emergence of Omicron sublineages classified as Variants of Concern (VOC), including BA.4, BA.5, BQ.1, and XBB. These sublineages harbor mutations in the spike protein, such as L452R, F486V, and E484K, which hinder antibody recognition and reduce vaccine and prior infection efficacy22,23. Ramirez et al.24 further supported this, showing that the Mu variant, carrying mutations like N501Y and E484K, not only increased transmissibility but also enabled immune evasion, facilitating reinfection and escape from vaccine-induced immunity. This behavior is similar to that of the Omicron variant, where mutations such as E484K play a key role in immune escape. These findings highlight the significant impact of these mutations on both viral spread and vaccine effectiveness. However, immunization remains crucial for reducing disease severity, leading to fewer severe cases22,23,24. Additionally, the relaxation of containment measures, such as reopening borders and easing mobility restrictions, significantly influenced the spread of these new variants. This trend, observed globally, coincided with the increased transmissibility of the Omicron (BQ.1.1, VOC) variant24. In Colombia, the emergence and global spread of variants like Omicron and Mu were facilitated by changes in epidemiological and mobility patterns25 further supporting this observation. These results align with global trends, illustrating the complex relationship between viral evolution, immune escape, and public health interventions26.
In addition to vaccination interventions, continuous monitoring of viral mutations and evolution is crucial for understanding changes in the epidemiological landscape. In this context, pandemic control and management have been implemented in a manner that relies heavily on global genomic surveillance of SARS-CoV-2, as the rapid sharing of data on viral evolution and the identification of new variants have been essential for decision-making and vaccine development27.
Phylogenetic analysis of the Pre-Omicron group revealed strong evidence of intraregional circulation of SARS-CoV-2 in Espirito Santo, with monophyletic clusters composed predominantly of samples from this state. This pattern of internal transmission, with no detectable external influences in most clusters, reflects the situation described in the study by Giovanetti et al.15,28, which reported that cocirculation of multiple lineages was initially driven by international introductions, but rapidly evolved into sustained local transmissions within Brazilian regions. On the other hand, the presence of clusters with a heterogeneous composition, comprising samples from other Brazilian states and foreign countries, underscores the intricate nature of interregional interactions and viral mobility, as supported by previous research15,29.
The Bayesian analysis of the Pre-Omicron period, particularly the emergence of Cluster 6 between July and late August 2020, highlighted Espirito Santos pivotal role in viral dissemination. Despite being a state of the southeast region of Brazil with a smaller population, its intense port activity, strategic geographic location, and above-average case and mortality rates underscore its importance as a model for understanding viral dispersion.
Thus, our results revealed that Espirito Santo served as a point of origin for viral spread to regions such as Parana, Rio de Janeiro, and Sao Paulo, driven by regional factors like mobility and socioeconomic activities. This pattern was particularly evident during periods of relaxed containment measures, further emphasizing the states integration potential in the broader pandemic context. Bayesian analysis relied on assumptions, including a relaxed log-normal molecular clock and non-parametric coalescent models, which enabled robust reconstructions of lineage origins and dispersal patterns. However, these findings must be interpreted in light of the models limitations, as the assumptions directly influence the inferences drawn. These findings reinforce Espirito Santos relevance as a model for analyzing transmission dynamics during distinct pandemic phases, offering insights into the interplay between regional factors and viral mobility.
In turn, the Bayesian analysis during the Post-Omicron period suggests that the transmission of the BQ.1.1 lineage in Espirito Santo occurred over a relatively short time frame, reflecting a rapid spread of the virus within the state, as indicated by the tMRCAs, which point to recent dates in the main clades. The emergence of Clusters 1, 5, and 23, between October and November 2022, reinforces this conclusion. These origin dates coincide with a period of relaxed pandemic containment measures, during which the circulation of new variants, such as BQ.1.1, may have been facilitated30,31. The rapid dissemination characteristic of the BQ.1.1 lineage is further supported by the observed genetic homogeneity among sequences, which significantly influences phylogenetic analyses. The low SH-aLRT values observed in the Post-Omicron group can be attributed to the high genetic similarity of the BQ.1.1 lineage, reflecting its rapid and widespread transmission during this period. This genetic homogeneity, marked by minimal evolutionary divergence, results in a “star-like” topology in phylogenetic trees, where the lack of sufficient genetic differences between sequences undermines statistical support for specific branching patterns. Such topologies are a common feature in lineages with rapid global spread and limited time for significant mutational accumulation, as observed in the BQ.1.1 lineage. These factors collectively weaken the ability to resolve distinct clusters robustly, as described in recent studies32.
This pattern aligns with findings from other studies on Omicron sublineages, such as BA.1 in the Brazilian state of Mato Grosso do Sul32. Similar challenges in resolving distinct clusters due to reduced genetic diversity were reported, underscoring the broader implications of accelerated viral transmission in shaping phylogenetic topologies. The rapid transmission in susceptible populations inherently limits genetic variability, affecting statistical metrics like SH-aLRT. Despite these limitations, high bootstrap values across the phylogenetic tree reinforce the robustness of the analysis, providing a more reliable measure of branch support in low-variability datasets. Furthermore, the clustering patterns observed in this study align with known epidemiological data, further validating the tree topology. These findings underscore the evolutionary dynamics of the BQ.1.1 lineage, emphasizing its efficiency in spreading with minimal mutations, a trait that has been consistently noted across various Omicron sublineages33.
The changes in the profile of SARS-CoV-2 clusters between the Pre- and Post-Omicron periods in Espirito Santo reveal a significant transformation in the dynamics of transmission, marked by an increase in the diversity and in the presence of external samples in the clusters of the Post-Omicron period. This pattern aligns with the findings of Ramírez and collaborators30, who highlighted the rapid global dissemination of the Omicron variant, resulting in significant diversification of the viral genomes in several regions, such as New York, as well as in United Kingdom and South Africa, and intensified the interactions between different regions and the diversification of the viral clusters, both in the Brazilian state of Espirito Santo and in other parts of the world30,31.
The analysis of viral dispersion observed in the Maximum Likelihood analyses reinforces the idea of significant transmission of SARS-CoV-2, both within Brazil and on an international scale, especially in the Post-Omicron period. This pattern is similar to that observed in a study conducted in Germany, which highlighted the importance of detailed data on social contacts to understand the dynamics of virus transmission34. In Brazil, the lower geographic dispersion in the Pre-Omicron period may be associated with more localized transmission control, similar to the early phases of the pandemic in Germany, when strict contact restriction measures were in place, limiting the spread of the virus34,35.
The intensification of regional and international interactions in Espirito Santo after Omicron, facilitated by increased population mobility and global exchanges, contributed to the diversification and expansion of viral clusters, reflecting the globalization of the pandemic. Previous studies conducted in the region, specifically in Colombia, where the Mu variant of interest (VOI) (B.1.621/B.1.621.1) was identified, suggest that the increase in cases associated with this variant may have facilitated its spread across various cities in the country. This phenomenon could have been exacerbated by factors specific to Colombia at the time, such as the social protests against the government that occurred in the Pre-Omicron period, as well as national festivities, which promoted gatherings and increased social contact, thus enhancing the likelihood of virus transmission25.
On the other hand, data from a study on the incidence of COVID-19 in the United States during the Omicron variant outbreak revealed that mobility was a significant determinant of viral spread, with reductions in mobility associated with declines in case incidence. This suggests that the increase in mobility reported in the present study during the Post-Omicron period was a key factor in the rapid spread of the virus. These studies reinforce the importance of integrating mobility control into mitigation strategies during periods of high transmission35,36.
The transmission chains demonstrated in the Bayesian analysis of the analyzed clusters from the Pre-Omicron period indicate that the clusters were already established in the Espirito Santo state, during a period of closed borders and movement restrictions37,38. The presence of longer branches reinforces the idea that viral dispersion during this period occurred in a more controlled manner, with a lower transmission speed, reflecting the characteristics of the epidemiological environment at the time and the response of the control measures in force.
In contrast, the transmission chains of the Post-Omicron group, in which we chose to cut the clades in Espirito Santo, owing to the large number of sequences and to facilitate visualization, revealed that the Espirito Santo sequences were predominantly located at the end of the transmission chain (Supplementary material). These results suggest that the origin of the clades is international, with progressive dissemination until reaching the Brazilian state of Espirito Santo. Unlike the previous period, where we observed longer branches and slower propagation, the Post-Omicron period was characterized by rapid dispersion of the clades once they were introduced into the state, reflecting the high transmissibility of the Omicron variant.
Some limitations should be considered, as the analysis may have been influenced by uncontrolled factors, such as regional variations in public health policies and social behaviors. Furthermore, sequencing coverage was uneven in some geographic areas, potentially leading to underrepresentation of certain regions. While the dataset construction methodologies aimed to minimize biases, the representativeness of sequences from specific regions may still be affected by these uneven sampling efforts. Nevertheless, the combined use of local and global data mitigates this limitation, providing a more balanced perspective on SARS-CoV-2 dispersion patterns.
This study revealed a distinct dynamic in the transmission of SARS-CoV-2 in Espirito Santo, highlighting how the relaxation of containment measures and the evolution of the virus transformed predominantly intraregional circulation during the Pre-Omicron period into a more complex and globally connected spread in the Post-Omicron period. Espirito Santo emerged as a strategic region for genomic surveillance, with high SARS-CoV-2 infection rates in both periods analyzed. The data showed that, despite its small size, the state has a significant representation of viral lineages comparable to those found in GISAID. The diversification of Post-Omicron clusters, driven by increased mobility and international exchanges, underscores the importance of adaptive strategies that consider both viral behavior and socioeconomic and mobility conditions. Although the focus of this study has been on the Brazilian state of Espirito Santo, the results point to broader trends that are relevant for health policy formulation in other contexts facing similar challenges.
The historical insights provided by this study remain relevant for understanding the evolution of SARS-CoV-2 and preparing for future pandemics. By integrating innovative sequence selection strategies and focusing on critical pandemic transitions, this work offers valuable contributions to the field of molecular epidemiology and public health.
Methods
Sample collection for group determination and RNA extraction
Samples from patients with positive nasopharyngeal/oropharyngeal swab RT-qPCR results for SARS-CoV-2 were collected during two distinct periods in the Brazilian state of Espirito Santo, forming two separate groups. The first group, Pre-Omicron, is composed of patients with positive samples, which covers the first phase of the pandemic, before the emergence of the Omicron variant, with collections performed between 07/09/2020 and 03/29/2021. The second group, Post-Omicron, includes samples with collections carried out between 11/14/2022 and 12/28/2022, corresponding to the period after the emergence of the referred variant.
These samples were originally collected by the Laboratório Central de Saúde Pública do Estado do Espirito Santo (Lacen/ES) for routine COVID-19 diagnostic purposes and stored in accordance with institutional and regulatory protocols. The use of these retrospectively collected samples for research purposes was formally approved by the UFES Ethics Committee (Approval Number: 6.173.702—CCS/UFES/Brazil), following ethical and legal guidelines. Since these samples were already available in the Lacen/ES biorepository, obtaining individual informed consent was not feasible, and a waiver of informed consent was granted by the Ethics Committee.
After approval, the samples were processed for RNA extraction using the Bio Gene Viral DNA/RNA Extraction Kit (Bioclin, Brazil) following the manufacturer’s guidelines. Subsequently, real-time reverse transcription-quantitative polymerase chain reaction (RT-qPCR) for SARS-CoV-2 detection was performed using a \(\text {TaqPath}^\text {TM}\) 1-Step RT-qPCR Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) in a \(\text {StepOnePlus}^\text {TM}\) PCR system (Applied Biosystems, Clarsbad, CA, USA). Specific primers and hydrolysis probes targeting two regions of the nucleocapsid gene (N1 and N2), along with an internal control (RNase P), were used, as recommended by the Centers for Disease Control and Prevention (CDC). Negative and positive controls (2019-nCoV RUO Kit, Integrated DNA Technologies, IDT, Iowa, USA) were included in all runs for assay validation. The positive samples with a Ct value below 27 were selected and subjected to SARS-CoV-2.
RNA extraction and RT-qPCR for sample selection
RNA from the selected samples was reverse-transcribed into cDNA using the Luna Script RT SuperMix. The cDNA was then amplified in two pools of primers following the Midnight RT PCR Expansion Kit (EXP-MRT001) protocol. This protocol generates overlapping 1200 bp amplicons covering the SARS-CoV-2 genome. Quantification of the pooled amplicons was performed using the \(\text {Qubit}^\text {TM}\) dsDNA HS Assay Kit, ensuring concentrations above 20 ng/\(\upmu\)L for each pool before proceeding to sequencing.
Additionally, to improve sequencing of Omicron variant samples, the following additional primers were added to the pools: in Pool 1, primers Pool1_1200_23_Left_Omicron (ACTTTAGAGTTCAACCAACAGAATCT) and Pool1_1200_21_Right_Omicron (GTGTATGATTGAGTTCTGGTTGTAAG); and in Pool 2, primers Pool2_1200_22_Right_Omicron (AACAGATGCAAATTTGGTGGCG), Pool2_1200_24_Left_Omicron (GCTGAATATGTCAACAACTCATATGA), and Pool2_1200_28_Left_Omicron (TTTGTGCTTTTTAGCCTTTCTGTT). Amplicons from both primer pools were combined and purified with a 1\(\times\) volume of Ampure XP beads.
Sequencing and data processing
Sequencing was performed on the Oxford Nanopore Technologies (ONT) platform. MinION library preparation was performed using the Ligation Sequencing Kit SQK-RBK-110.96. The resulting library was loaded onto Oxford MinION R9.4 flow cells (FLO-MIN106) and sequenced using the MinION Mk1C device. ONT MinKNOW software was used to collect raw data. Quality control and high-precision base-calling analyses were performed using Guppy (v6.0.1). High-precision database assembly called fastq files was performed using the 2019-nCoV-2 novel coronavirus bioinformatics protocol to generate consensus sequence39. All sequences were deposited in the GISAID database (EPI_SET_240818zu; doi:10.55876/gis8.240818zu).
Comparative genome analysis
Sequences were analyzed with Nextclade v3.7.440 to determine the clustering and the number of lacunar regions. Lineages were assigned to each genome using Pangolin v3.1.1741. Subsequently, the sequences corresponding to the predominant lineages in the Pre and Post-Omicron sequencing periods were grouped into group Pre and Post-Omicron respectively.
Dataset construction
Espirito Santo was selected as the focal point of this study due to its unique epidemiological profile and strategic importance for understanding viral dispersion. Despite its low population density, COVID-19 case and mortality rates in the state significantly exceeded the national average, highlighting its critical role in the pandemic’s trajectory. Additionally, its position as a commercial hub with intense port activity and geographical connectivity to major Brazilian states made it an ideal region to examine SARS-CoV-2 dispersion patterns in both regional and global contexts. To minimize potential regional biases, the inclusion of global datasets in both Pre- and Post-Omicron analyses ensures that the findings reflect a broader viral dispersion pattern while maintaining a strong regional foundation.
Based on Espirito Santo’s unique context and the predominant lineages identified through local sequencing, two groups were defined for comparative analysis: the Pre-Omicron Group (B.1.1.33 lineage) and the Post-Omicron Group (BQ.1.1 lineage). Specific and common procedures were followed for dataset construction to ensure consistency and representativeness.
For the Pre-Omicron Group, the sequences obtained in this study were combined with all sequences of the same lineage deposited in GISAID up to March 5, 2024, to evaluate the phylogenetic relationships of viruses circulating in a global context. The filters ’Complete,’ ’Low Coverage Excluded,’ and ’Collection Date Complete’ were applied (GISAID access: EPI_SET_240818ds, doi: 10.55876/gis8.240818ds).
In the Post-Omicron Group, due to the large number of available BQ.1.1 sequences, a different approach was adopted. All sequences of this lineage collected in Espirito Santo were downloaded, and using the ’Audacity GISAID tool’, the 150 most similar sequences from all regions of the world were selected. The threshold of 150 sequences was chosen to balance dataset manageability with global representativeness, ensuring that the selected sequences capture the diversity of the BQ.1.1 lineage while avoiding potential overrepresentation of specific regions. Redundant sequences were excluded, and to ensure high quality, a 20% cutoff was applied, eliminating sequences with more than 2500 gaps. Throughout the dataset construction process, quality control measures were applied, including manual inspection of alignments and validation of sequence metadata, to ensure the reliability of the datasets used for phylogenetic and Bayesian analyses. To confirm that all sequences belonged to the BQ.1.1 lineage, the Pangolin tool (v3.1.17) was used for classification, and those that did not match the desired lineage were removed from our database. This approach ensured that the dataset was both manageable and representative of the global dynamics of the BQ.1.1 lineage, offering a robust framework for studying viral evolution and dispersion patterns. The GISAID access number (EPI_SET_240818xv, doi: 10.55876/gis8.240818xv) corresponds to the dataset of sequences deposited for this analysis.
Phylogenetic analysis
Multiple sequence alignments were performed using MUSCLE in MEGA 7.0 software, aligning the dataset sequences with the Wuhan reference sequence42(GenBank accession number: NC_045512.2). The Wuhan sequence served as the outgroup, providing a reference point for rooting the phylogenetic trees. Default settings were applied during the alignment process, and the aligned sequences were manually inspected to ensure accuracy and quality.
Maximum likelihood (ML) phylogenetic trees were constructed using IQ-TREE (v2.3.6). The Transitional (TIM) model of nucleotide substitution, with empirical base frequencies (+F) and a FreeRate model with two categories (+R2), was selected using the ModelFinder tool integrated into IQ-TREE. To validate the tree topology, 1000 bootstrap replicates and SH-aLRT branch tests were applied. The GTR+I+G4 substitution model and the relaxed log-normal molecular clock were chosen due to their demonstrated ability to model viral evolutionary dynamics, particularly in datasets with rapid mutation rates like SARS-CoV-2. The trees were visualized and analyzed using Figtree (v1.4.4), and branch support values (bootstrap and SH-aLRT) were calculated for each cluster.
Clusters were defined when at least two samples from Espirito Santo grouped together with a support value greater than 0.8 on the phylogenetic tree. Subsequently, all samples descending from the ancestral node of the defined cluster were considered part of the same transmission cluster. To enhance the visualization of the temporal and geographic dispersion of the clusters, all trees were calibrated to the same time frame, and color codes were used to represent sequences from different states. Additionally, a local BLAST database was built to identify the 20 most similar sequences for each clade observed in Espirito Santo, ensuring both local representativeness and the preservation of phylogenetic relationships. Support values, along with the geographic composition and number of sequences in each cluster, are provided in the supplementary table for transparency.
Bayesian analysis
Bayesian analyses were conducted to infer the spatiotemporal spread of the virus using the BEAST v1.10 package43 with the BEAGLE44 library to accelerate computations. The analyses included basal samples from selected clusters, along with all sequences between the first and second ancestral nodes. The inclusion of two ancestral nodes ensures that the temporal and phylogenetic breadth of the clusters is adequately represented. In cases where the tree topology prevented the inclusion of two ancestral nodes, sequences from the nearest ancestral node were selected to maintain consistency.
The GTR+I+G4 nucleotide substitution model, a relaxed log-normal molecular clock, calibrated with a normal prior based on previous estimates6,45 (mean: \(8 \times 10^{-4}\) substitutions/site/year; SD: \(2 \times 10^{-4}\)), and a non-parametric Bayesian skyline coalescent model were used. This molecular clock model assumes rate variation among lineages, which is critical for capturing the rapid evolutionary dynamics of SARS-CoV-2. We employed a non-parametric Bayesian skyline coalescent model due to its flexibility in reconstructing changes in effective population size over time without requiring prior assumptions about demographic history.
Discrete asymmetric (nonreversible) phylogeographic models46 were employed to reconstruct the geographic spread of the virus, allowing directional migration rates between regions to be estimated. The model incorporated location states as discrete traits, reflecting the sampling regions, and enabled the inference of lineage movements across geographical locations.
Markov-chain Monte Carlo (MCMC) sampling was performed for \(10^8\) generations, with convergence assessed using TRACER v1.7.447. Adequate effective sample sizes (ESS> 200) were confirmed after discarding 10% of the chains as burn-in. Results from Bayesian analyses, including the phylogeographic reconstruction, were used to interpret the temporal and spatial dynamics of SARS-CoV-2 within the context of the dataset.
Data Availability
The datasets generated and/or analyzed during the current study are available in the GISAID database under the following accession codes: Post-Omicron Group: EPI_SET_240912dp (https://doi.org/10.55876/gis8.240912dp); Pre-Omicron Group: EPI_SET_240818ds (https://doi.org/10.55876/gis8.240818ds); Sequences Generated in This Study: EPI_SET_240818zu (https://doi.org/10.55876/gis8.240818zu).
References
Chan, J.F.-W. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. The Lancet 395, 514–523. https://doi.org/10.1016/s0140-6736(20)30154-9 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273. https://doi.org/10.1038/s41586-020-2012-7 (2020).
Fan, Y. et al. SARS-CoV-2 Omicron variant: Recent progress and future perspectives. Signal Transduct. Target. Therapy 7, 66. https://doi.org/10.1038/s41392-022-00997-x (2022).
Ramesh, S. et al. Emerging SARS-CoV-2 variants: A review of its mutations, its implications and vaccine efficacy. Vaccines 9, 1195. https://doi.org/10.3390/vaccines9101195 (2021).
da Saúde, M. Brasil confirma primeiro caso de infecção pelo novo coronavírus.) OPAS. figshare https://www.paho.org/pt/node/69303 (2020).
Resende, P. C. et al. Evolutionary dynamics and dissemination pattern of the SARS-CoV-2 lineage B.1.1.33 during the early pandemic phase in Brazil. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.615280 (2021).
Voloch, C. M. et al. Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. J. Virol. https://doi.org/10.1128/jvi.00119-21 (2021).
dos Santos, C. A. et al. SARS-CoV-2 genomic surveillance in Northeast Brazil: Timing of emergence of the Brazilian variant of concern P1. J. Travel Med. https://doi.org/10.1093/jtm/taab066 (2021).
Martins, A. F. et al. Detection of SARS-CoV-2 lineage P. 1 in patients from a region with exponentially increasing hospitalisation rate, February 2021, Rio Grande do Sul, Southern Brazil. Eurosurveillance https://doi.org/10.2807/1560-7917.es.2021.26.12.2100276 (2021).
Alcantara, L. C. J. et al. SARS-CoV-2 epidemic in Brazil: How the displacement of variants has driven distinct epidemic waves. Virus Res. 315, 198785. https://doi.org/10.1016/j.virusres.2022.198785 (2022).
Kupferschmidt, K. & Wadman, M. Delta variant triggers new phase in the pandemic. Science 372, 1375–1376. https://doi.org/10.1126/science.372.6549.1375 (2021).
Stefanelli, P. et al. Tracking the progressive spread of the SARS-CoV-2 Omicron variant in Italy, December 2021 to January 2022. Eurosurveillance https://doi.org/10.2807/1560-7917.es.2022.27.45.2200125 (2022).
Gardy, J. L. & Loman, N. J. Towards a genomics-informed, real-time, global pathogen surveillance system. Nat. Rev. Genet. 19, 9–20. https://doi.org/10.1038/nrg.2017.88 (2017).
Khare, S. et al. Gisaid’s role in pandemic response. China CDC Wkly. 3, 1049–1051. https://doi.org/10.46234/ccdcw2021.255 (2021).
Giovanetti, M. et al. Genomic epidemiology of the SARS-CoV-2 epidemic in Brazil. Nat. Microbiol. 7, 1490–1500 (2022).
Koelle, K., Martin, M. A., Antia, R., Lopman, B. & Dean, N. E. The changing epidemiology of SARS-CoV-2. Science 375, 1116–1121. https://doi.org/10.1126/science.abm4915 (2022).
de Geografia e Estatística, I. B. Espírito Santo: cidades e estados) IBGE (2024). figshare https://www.ibge.gov.br/cidades-e-estados/es.html (2024).
da Saúde, M. Painel Coronavírus) (2024). figshare https://covid.saude.gov.br/ (2024).
Candido, D. S. et al. Evolution and epidemic spread of SARS-CoV-2 in Brazil. Science 369, 1255–1260. https://doi.org/10.1126/science.abd2161 (2020).
Bergeri, I. et al. Global SARS-CoV-2 seroprevalence from January 2020 to April 2022: A systematic review and meta-analysis of standardized population-based studies. PLOS Med. 19, e1004107. https://doi.org/10.1371/journal.pmed.1004107 (2022).
Cascini, F. et al. A cross-country comparison of Covid-19 containment measures and their effects on the epidemic curves. BMC Public Health https://doi.org/10.1186/s12889-022-14088-7 (2022).
Pather, S. et al. SARS-CoV-2 Omicron variants: Burden of disease, impact on vaccine effectiveness and need for variant-adapted vaccines. Front. Immunol. 14, 66. https://doi.org/10.3389/fimmu.2023.1130539 (2023).
Planas, D. et al. Distinct evolution of SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineages combining increased fitness and antibody evasion. Nat. Commun. https://doi.org/10.1038/s41467-024-46490-7 (2024).
Ramirez, A. L. et al. Impact of SARS-CoV-2 Mu variant on vaccine effectiveness: A comparative genomics study at the peak of the third wave in Bogota, Colombia. J. Med. Virol. 94, 3988–3991 (2022).
Patiño, L. H. et al. Epidemiological dynamics of SARS-CoV-2 variants during social protests in Cali, Colombia. Front. Med. 9, 863911 (2022).
Zhang, N. et al. Explosive household spread of the SARS-CoV-2 Omicron variant in China in late 2022. Build. Environ. 256, 111491. https://doi.org/10.1016/j.buildenv.2024.111491 (2024).
Maxmen, A. One million coronavirus sequences: Popular genome site hits mega milestone. Nature 593, 21–21. https://doi.org/10.1038/d41586-021-01069-w (2021).
Giovanetti, M. et al. Genomic epidemiology reveals the impact of national and international restrictions measures on the SARS-CoV-2 epidemic in Brazil. MedRxiv (2022).
Jorge, D. C. et al. Assessing the nationwide impact of COVID-19 mitigation policies on the transmission rate of SARS-CoV-2 in Brazil. Epidemics 35, 100465. https://doi.org/10.1016/j.epidem.2021.100465 (2021).
Ramírez, J. D. et al. Hotspots for SARS-CoV-2 Omicron variant spread: Lessons from New York City. J. Med. Virol. 94, 2911–2914. https://doi.org/10.1002/jmv.27691 (2022).
Tong, C., Shi, W., Zhang, A. & Shi, Z. Tracking and controlling the spatiotemporal spread of SARS-CoV-2 Omicron variant in South Africa. Travel Med. Infect. Dis. 46, 102252. https://doi.org/10.1016/j.tmaid.2021.102252 (2022).
de Mello Almeida Maziero, L. et al. Unveiling the impact of the Omicron Variant: Insights from Genomic Surveillance in Mato Grosso do Sul, Midwest Brazil. Viruses 15, 1604 (2023).
Arantes, I. et al. Spatiotemporal dynamics and epidemiological impact of SARS-CoV-2 XBB lineage dissemination in Brazil in 2023. Microbiol. Spectr. 12, e03831-23 (2024).
Tomori, D. V. et al. Individual social contact data and population mobility data as early markers of SARS-CoV-2 transmission dynamics during the first wave in Germany—An analysis based on the COVIMOD study. BMC Med. https://doi.org/10.1186/s12916-021-02139-6 (2021).
Fiori, M. et al. Decoupling between SARS-CoV-2 transmissibility and population mobility associated with increasing immunity from vaccination and infection in South America. Sci. Rep. https://doi.org/10.1038/s41598-022-10896-4 (2022).
Harris, J. E. Mobility was a significant determinant of reported COVID-19 incidence during the Omicron Surge in the most populous U.S. Counties. BMC Infect. Dis. https://doi.org/10.1186/s12879-022-07666-y (2022).
Houvèssou, G. M., Souza, T. P. d. & Silveira, M. F. d. Medidas de contenção de tipo lockdown para prevenção e controle da COVID-19: estudo ecológico descritivo, com dados da África do Sul, Alemanha, Brasil, Espanha, Estados Unidos, Itália e Nova Zelândia, fevereiro a agosto de 2020. Epidemiologia e serviços de saúde 30, e2020513. https://doi.org/10.1590/s1679-49742021000100025 (2021).
Liu, Y., Morgenstern, C., Kelly, J., Lowe, R. & Jit, M. The impact of non-pharmaceutical interventions on SARS-CoV-2 transmission across 130 countries and territories. BMC Med. https://doi.org/10.1186/s12916-020-01872-8 (2021).
Network, A. nCoV-2019 novel coronavirus bioinformatics SOP) ARTIC. figshare. https://artic.network/ncov-2019/ncov2019-bioinformatics-sop.html (2023).
Nextstrain. Clade assignment, mutation calling, and sequence quality checks) Nextstrain (2024). figshare https://clades.nextstrain.org/ (2024).
PANGO-Lineages. Latest epidemiological lineages of SARS-CoV-2) PANGO Lineages (2024). figshare https://cov-lineages.org/index.html (2024).
Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature 579, 265–269. https://doi.org/10.1038/s41586-020-2008-3 (2020).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. https://doi.org/10.1093/ve/vey016 (2018).
Ayres, D. L. et al. BEAGLE: An application programming interface and high-performance computing library for statistical phylogenetics. Syst. Biol. 61, 170–173. https://doi.org/10.1093/sysbio/syr100 (2011).
Gräf, T. et al. Phylogenetic-based inference reveals distinct transmission dynamics of SARS-CoV-2 lineages Gamma and P.2 in Brazil. iScience 25, 104156. https://doi.org/10.1016/j.isci.2022.104156 (2022).
Lemey, P., Rambaut, A., Drummond, A. J. & Suchard, M. A. Bayesian phylogeography finds its roots. PLoS Comput. Biol. 5, e1000520. https://doi.org/10.1371/journal.pcbi.1000520 (2009).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using tracer 1.7. Syst. Biol. 67, 901–904. https://doi.org/10.1093/sysbio/syy032 (2018).
Acknowledgements
We would like to acknowledge the financial support provided by the Foundation for Research and Innovation Support of Espirito Santo (FAPES) (Number 283/2020 and 479/2021) and the Coordination for the Improvement of Higher Education Personnel (CAPES) [692/2020], both from Brazil. We also thank the Central Public Health Laboratory of the State of Espirito Santo (LACEN-ES) for providing SARS-CoV-2 samples. We would like to thank GISAID for providing the data and genomic sequences that were essential for this research.
Author information
Authors and Affiliations
Contributions
J.S.A: Sample processing, sequencing, data analysis, and manuscript writing. M.V.M.S: Sample processing, sequencing, data analysis, and manuscript review. B.R.A: Sample processing, sequencing, and manuscript review. A.M.C, R.T, R.R.R, L.C.S, and G.G.P: Data analysis and manuscript review. E.D: Data analysis, manuscript review, and supervision. S.V.V and T.F.B: Data analysis, manuscript review, and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Santa Ardisson, J., Vedovatti Monfardini Sagrillo, M., Ramos Athaydes, B. et al. Comparative spatial–temporal analysis of SARS-CoV-2 lineages B.1.1.33 and BQ.1.1 Omicron variant across pandemic phases. Sci Rep 15, 10319 (2025). https://doi.org/10.1038/s41598-025-95140-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-95140-5