Abstract
Multimodal medical image fusion is crucial for enhancing diagnostic accuracy by integrating complementary information from different imaging modalities. Current fusion techniques face challenges in effectively combining heterogeneous features while preserving critical diagnostic information. This paper presents a Temporal Decomposition Network (TDN), a novel deep learning architecture that optimizes multimodal medical image fusion through feature-level temporal analysis and adversarial learning mechanisms. The TDN architecture incorporates two key components: a salient perception model for discriminative feature extraction and a generative adversarial network for temporal feature matching. The salient perception model identifies and classifies distinct pixel distributions across different imaging modalities, while the adversarial component facilitates accurate feature mapping and fusion. This approach enables precise temporal Decomposition of heterogeneous features and robust quality assessment of fused regions. Experimental validation on diverse medical image datasets, encompassing multiple modalities and image dimensions, demonstrates the TDN’s superior performance. Compared to state-of-the-art methods, the framework achieves an 11.378% improvement in fusion accuracy and a 12.441% enhancement in precision. These results indicate significant potential for clinical applications, particularly in radiological diagnosis, surgical planning, and medical image analysis, where multimodal visualization is critical for accurate interpretation and decision-making.
Similar content being viewed by others
Introduction
Medical image fusion is a process that fuses multiple medical images to enhance the performance range of medical diagnosis1. A target information enhancement-based multimodal medical image fusion technique is used in medical applications. The fusion technique uses a feature extraction approach to extract relevant features from medical images2. The extracted features produce the necessary data for image fusion, eliminating unwanted noises from the images.
The fusion technique decreases the saliency and data loss rate while performing the fusion process3. The image fusion technique enhances the quality of images and information, elevating the efficiency range of diagnosis services. An open-source approach is also used for multimodal image fusion4. The approach uses an anatomical and thermal 3D model that gathers optimal medical images for fusion. The 3D models reconstruct the relevant pixels and identify important datasets from input images. The fusion approach minimizes the computational cost of image fusion, which increases the performance level of further disease detection and prediction processes5,6.
Feature-based medical image fusion techniques are also used in healthcare applications. Feature-based techniques elevate the overall quality range of image fusion7. A dual-branch feature-enhanced network model is used for the medical image fusion process. The model uses a convolutional neural network (CNN) algorithm to extract appropriate image details from the medical images8,9. The extracted details are used as a dataset, decreasing the process’s computational complexity. The CNN algorithm reduces the error data during image fusion10. The network model increases the fusion’s precision range, which maximizes the disease diagnosis services’ performance ratio. A multi-features diffusion model is also used as an image fusion model in medical applications11. The diffusion model uses an augmentation strategy that categorizes the features according to importance. It also eliminates the unnecessary noises and patterns from the images12. The model identifies the optimal features that are gathered for image fusion. The diffusion model increases the effectiveness and feasibility range of the image fusion process13.
Machine learning (ML) algorithm-based medical image fusion approaches are also used in healthcare systems, particularly for early Alzheimer’s disease diagnosis14. A hierarchical wavelet generative adversarial network (HW-GAN) enabled model is used for medical image fusion. The HW-GAN model uses a hierarchical wavelet fusion module to construct optimal designs to fuse detailed information15. While traditional fusion methods and recent deep learning models have demonstrated improved accuracy, they often fail to maintain structural fidelity and salient feature representation across diverse modalities. To address these limitations, the proposed TDN framework integrates saliency perception with a generative adversarial paradigm, supported by a decomposition-based architecture designed to enhance spatial coherence and functional integration.
The motivation for TDN research stems from the need to improve multimodal medical image fusion, addressing challenges like feature misalignment, noise interference, and loss of structural integrity. TDN aims to elevate diagnostic capabilities and support precise clinical decision-making by enhancing accuracy and reliability in integrating diverse imaging modalities. Perceptual video quality assessment is vital to multimedia evaluation. In a survey on video quality, subjective and objective evaluation approaches were highlighted16. Examining screen content quality assessment presented innovative standards and methods compared to previous methods17. In audiovisual analysis, quality assessment methodologies were analyzed for audio and visual signals18. The study used multimodal analysis to predict fixation, providing insights into human attention mechanisms19. It created a multimodal saliency model for videos with high audiovisual correlation to better explain perceptual relevance in multimedia processing20.
The Temporal Decomposition Network (TDN) addresses the challenge of high-fidelity feature preservation in multimodal medical image fusion. It separates homogeneous and heterogeneous feature representations, refines feature selection by dynamically identifying key regions, and ensures adaptive feature alignment across different modalities. TDN mitigates limitations in clinical applications like tumor detection from MRI-CT fused images by employing a feature-matching mechanism that enhances discriminative representation while preserving critical anatomical structures. Experimental results show that TDN improves fusion accuracy and precision compared to state-of-the-art approaches, reinforcing its significance in medical imaging applications. Improved extraction and integration of modality-specific structural and functional characteristics are achieved via the TDN’s unique use of saliency perception and generative adversarial learning, a decomposition-driven fusion framework. Using a temporal-frequency decomposition technique, TDN improves the representation of delicate anatomical and pathological information, unlike previous approaches that often depend on manual fusion rules or shallow feature aggregation. Furthermore, adaptive feature learning and structural consistency preservation are enabled by its generative adversarial framework, mapping, and unmapping procedures, which are not effectively captured by classic fusion models or transformer-based architectures. The main contributions of the study are given as:
-
A novel Temporal Decomposition Network (TDN) is developed to enhance multimodal medical image fusion by integrating salient perception and adversarial learning mechanisms for improved feature extraction and temporal analysis.
-
The fusion of heterogeneous medical image features is optimized by implementing a dual-component architecture that combines discriminative feature classification with temporal decomposition for higher precision fusion outputs.
-
The effectiveness of the TDN framework is validated across diverse medical imaging modalities and dimensions, with quantifiable improvements in fusion accuracy (11.378%) and precision (12.441%) demonstrated compared to existing methods, thereby advancing diagnostic visualization capabilities.
The rest of the paper is followed by Section “Related works”, which discusses the latest literature review on the proposed topic. Section “Proposed temporal decomposition network (TDN) for medical image fusion” describes the proposed temporal decomposition network (TDN) for medical image fusion. Section “Results and discussion” gives the results and discussion in detail, while the conclusion of the study is finally drawn in Section “Conclusion”.
Related works
Jiang et al.21 developed a lightweight multimode medical image fusion method for healthcare applications. The developed method uses a fuzzy set theory to analyze the features and patterns of the medical images. The similarity measures presented in the images are also identified, which produce relevant data for image fusion. The developed method enhances the performance range of the image fusion process. He et al.22 introduced a fidelity-driven optimization (FDO) reconstruction and details preserving guided fusion method for medical images. The introduced method uses multi-modality medical images as inputs, fused to get feasible data for disease diagnosis services. The texture and detailed information from the images are calculated, decreasing the latency. The introduced method maximizes the robustness level of the fusion process. Shi et al.23 designed a multilevel bidirectional feature interaction network model for medical image fusion. The designed model extracts specific and shared features from medical images. The extracted features are interacted with to gather necessary information for further disease detection and diagnosis processes. Both features, which are located horizontally and vertically, are interacted. The designed model elevates the performance and efficiency range of medical fusion.
Zhao et al.24 proposed a transformer-based universal fusion algorithm for multimodal images. The proposed algorithm uses a convolutional neural network (CNN) to extract relevant features from the images. The algorithm identifies the noises and eliminates the unwanted noises from the dataset. Spatial and temporal features are identified to form optimal information for the image fusion process. The proposed algorithm enlarges the effectiveness and minimizes the mean error rate. Di et al.25 developed a multimodal medical image fusion method using an attention mechanism and MobileNetV3 (AMMNet). The developed method uses a muti-scale convolutional layer to extract the feature information from the dataset. The attention mechanism is employed here to fuse the detailed information and texture to get optimal images for further processes. Experimental results show that the method developed increases the texture and sharpness of fused images. Fu et al.26 introduced a cascaded dense residual network (CDRNet) for grayscale and pseudocolor medical image fusion. The introduced model extracts the necessary features, and the extracted features are used as a dataset for fusion. The multimodal images are trained to get relevant information, decreasing the process’s computational cost. The introduced model maximizes the overall performance range of medical image fusion.
Song et al.27 proposed a medical image fusion method using a transformer and DenseNet. It uses a deep learning-enabled method that extracts limited features from medical images. The transformer algorithm is used here to train the dataset and get feasible detailed information for image fusion. Compared with others, the proposed method increases the visual quality and metrics of the fused medical images. Zhou et al.28 designed a gradient-intensity oriented model for multimodal Chest tumor image fusion. An intensity attention module (IAM) is utilized here to analyze the actual intensity rate of the features. The analyzed features are used as input for the medical image fusion process. The fusion problems are identified and solved via IAM, eliminating excess energy consumption rate. The designed model improves the accuracy and specificity range of medical images. Ding et al.29 proposed a new multimodal medical image fusion approach using multi-receptive-field and multiscale feature integration (M4FNet). A discrete wavelet packet transformer (DWT) algorithm is employed to extract the depth features from medical images. The DWT algorithm also identifies the textural features used as data for the fusion process. Experimental results show that the proposed approach significantly improves the precision level of medical images.
Zhang et al.30 suggested a dynamic residual focal transformer network for multimodal medical picture fusion. The context dynamic extraction module (CDEM) is used to create high-quality fused images for medical purposes. The module improves feature representation and fusion by extracting semantic information from images. The model uses dynamic residual learning to reduce noise and artifacts while retaining important information. This method outperforms traditional methods by improving medical image fusion for diagnostic and analytical purposes.
Safari et al.31 pioneered multimodal medical picture fusion using spatial and temporal feature extraction to maximize fusion. This method extracts important medical image attributes using an unsupervised deep generative adversarial network (GAN). GAN improves feature discrimination and preserves essential information during fusion. The study reveals that this method increases picture precision and anatomical visualization. This method improves medical diagnostics by merging multimodal data, making it a viable advanced medical image processing tool.
Transfer learning (TL)-based medical image fusion was developed by Dinh and Giang32. Integrating pertinent medical data with a coupled neural P system (CNPS) improves fused pictures in their model. CNPS learns from pre-trained models to improve feature selection and fusion while reducing computational complexity and preserving accuracy. Results show this strategy improves image fusion efficiency and performance. The method has great potential in medical imaging, especially for high-quality picture integration for illness diagnosis and therapy planning.
Zhang et al.33 developed SS-SSAN for multimodal medical picture fusion and image reconstruction. The system can learn representations without labeled training data since the SS-SSAN model evaluates optimal fusion data. The network emphasizes essential features and reduces redundancy to strengthen fused pictures. This strategy improves fusion efficiency and maintains diagnostic information through considerable experimentation. This model’s self-supervised learning methodology allows it to adapt to medical imaging applications and produce high-quality images for clinical interpretation.
Pan et al.34 presented self-supervised medical image fusion to reduce picture fusion delay. Medical photos are processed using a weighted autoencoder and tensor low-rank decomposition network to extract important textural information. This method preserves anatomical features while reducing computational overhead in existing fusion methods. The suggested method improves medical picture fusion by improving feature representation and lowering processing latency. This study advances real-time medical imaging applications that need fast and precise fusion.
MIn et al.35 introduced a high-audio-visual correspondence multimodal saliency (MMS) model for videos. Through a novel free energy concept and cross-modal kernel canonical correlation analysis, the model recognizes spatial and temporal saliency maps from the visual modality and audio saliency maps from both. A two-stage adaptive audiovisual saliency fusion approach is developed to merge these maps. MMS models account for audio, which deep learning-based saliency models ignore. Combining deep saliency models and MMS improves performance by 5%. Results from audiovisual attention databases reveal that models including audio cues outperform state-of-the-art image and video saliency models that use a single visual modality.
Khan et al.36 developed a multimodal medical image fusion (MMIF) model to improve picture structure. The model analyzes low- and high-frequency picture components with a non-subsampled Shearlet transformer (NSST), boosting fusion accuracy. This method keeps precise details and reduces fused output distortions and artifacts. Experimental results suggest that the proposed technique improves medical picture fusion, making it appropriate for high-quality integrated image applications. The model’s structural stability makes it attractive for improved medical picture analysis.
Min et al.37 implemented a blind IQA (BIQA) framework that relies on PRI and a novel reference known as pseudo-reference image (PRI). Depending on the application, PRI is created from skewed photographs and takes on significant distortion. The BIQA framework based on PRI creates measures particular to distortion to measure blockiness, sharpness, and noisiness. Blind PRI-based (BPRI) is a general-purpose BIQA method that incorporates the PRI-based metrics. Compared to cutting-edge opinion-aware and OU-BIQA models, the BPRI measure is nearly training-free and opinion-unaware. It works admirably with both screenshots and photographs of real-life scenes.
Min et al.38 introduced MPRIs to improve blind image quality assessment (IQA) by degrading the distorted image and comparing it to MPRIs. The full-reference IQA framework calculates quality using four types and five levels of distortion aggravation to handle typical distortions. Similarity scores measure quality; local binary pattern features describe similarities between the deformed image and MPRIs. Gooder quality means closer to a pseudo reference image (PRI). Having numerous PRIs minimizes image content’s impact, making image quality inference more accurate and consistent. The suggested method outperforms state-of-the-art blind IQA measures in four mainstream natural scene picture and screen content image quality assessment databases.
Min et al.39 suggested RichIQA, a subjective evaluation system beyond mean opinion score (MOS) to forecast wild image quality. A multi-label training strategy that uses rich subjective quality information like MOS, SOS, and DOS simultaneously and a three-stage image quality prediction network that uses the Convolutional vision Transformer (CvT) to mimic the brain’s short-term and long-term memory mechanisms define RichIQA. Three-stage network predicts rich quality information, whereas multi-label training maximizes subjective quality rating. RichIQA surpasses state-of-the-art competitors in wild IQA databases with rich subjective rating labels in large-scale experiments. RichIQA code will be on GitHub.
Huang et al.40 proposed an end-to-end feature disentangled Transformer (FDT) for entanglement-free and semantic feature representation for accurate pathology grading of squamous cell carcinoma (SCC). The FDTs use a semantic instance-based feature, a disentangled learning framework, and an instance attention block to divide images into multiscale instances for parallel optimization and human-interpretable semantics. Experiments on real-world SCC datasets show the proposed FDTs outperform state-of-the-art methods for trustworthy SCC grading.
Huang et al.41 proposed an end-to-end network with transformers constrained by learned-parameter-free attention (LA-ViT) to improve the accuracy and interpretability of grading laryngeal squamous cell carcinoma (LSCC) based on histopathological images. The LA-ViT network uses learned-parameter-free attention to learn high-effect target semantic information and reduce background semantics. The first-type LA transformer block uses a feature map position subspace for the query and feature channel subspace for the key, while the second-type LA transformer block uses model probability matrix information and decision level weight information for the key and query. The LA-ViT model outperforms other state-of-the-art models. Huang et al.42 introduced MamlFormer, a transformer network utilizing manifold adversarial multi-modal learning and prior-experience guidance for improved histopathological grading of laryngeal tissue, enhancing diagnostic accuracy and model generalization.
Kanika Bhalla et al.43 suggested artificial intelligence-assisted radiologic imaging biomarkers in triple-negative breast cancer. The present research compiles the most cutting-edge AI radiologic imaging applications to facilitate the diagnosis, treatment, and prognosis of TNBC. From 2013 to 2024, the author aims to thoroughly review the advancements in radiomic and deep learning-based AI and how they have advanced TNBC management. It will briefly introduce AI, radiomics, and deep learning to ensure the review is comprehensive. After that, the author focuses on AI-based models useful for diagnosis, prediction, and prognosis in radiologic breast images assessed in TNBC. The discussion concludes by outlining potential avenues for further research into AI that might improve TNBC diagnosis, treatment response prediction, and prognostic outcomes.
Santanu Roy et al.44 proposed the Mathematical analysis of histogram equalization techniques for medical image enhancement. Conventional Global Histogram Equalization (GHE), Histogram Specification (HS), and Brightness Preserving Dynamic Histogram Equalization (BPDHE) are re-investigated in this study using a fresh mathematical technique. Even though researchers in image processing and medical image diagnostics often use these HE approaches, it has been noted that they have substantial drawbacks when it comes to data loss. Since every Histogram Equalization (HE) approach is inherently non-linear, this work provides mathematical evidence that all HE methods always result in data loss. Two datasets were used to apply these histogram equalization methods: one containing MRI images of brain tumors and the other containing histopathology images of colorectal cancer that had been H and E-stained. All HE approaches fall within the range of 0.6–0.95 for the Structural Similarity Index Matrix (SSIM) and Pearson Correlation Coefficient (PCC). A linear contrast enhancement approach called the Reinhard method is also evaluated with these findings.
Yo-Ping Huang et al.45 recommended the Wavelet K-Means Clustering (KMC) and Fuzzy-Based Method for Segmenting MRI Images Depicting Parkinson’s Disease. The MRI pictures were upgraded after increasing the pixel values, and detailed information was recovered using discrete wavelet transform. The segmentation process was started by feeding the improved pictures into the KMC model. The last step was to use the FMF to filter out noise and uncertainty from the segmented pictures. You may also use this approach to segment MRI images of disorders like atrial fibrillation (AF) and tuberous sclerosis (TBS). The author offered the findings of a quantitative and qualitative assessment that the author ran on a clinical dataset and an open-source benchmark dataset. By comparing the suggested technique to existing segmentation methods, the author determined its efficiency based on structural similarity, peak signal-to-noise ratio, and mean-squared error. These findings demonstrate that the suggested approach may be useful for detecting PD, TBS, and AF in MRI scans at an early stage.
Kanika Bhalla et al.46 discussed the fuzzy convolutional neural network for enhancing multi-focus image fusion. Images taken by the cameras include blurriness, pixels that have been incorrectly categorized, uncertainty, and low contrast. Hence, MFIF can fuse an image’s in-focus objects into a unified picture by combining input image attributes. Nevertheless, a lack of consistency is caused by its computational complexity. Consequently, the fused picture is created using the MFIF approach, which combines the FS and CNN to identify focused and unfocused areas in the two original images. We also compare it against six competing MFIF approaches, including guided filters, CNN, ensemble CNN, image fusion-based CNN, and deep regression pair learning (DRPL). Reference metrics like Peak signal-to-noise ratio (57.23) and root mean square error (1.814) are used, along with four non-reference assessment measures such as mutual information (1.1678), edge information (0.7281), structural similarity (0.9850), and human perception (0.8020), to validate the superiority of the proposed FCNN method on benchmark datasets.
Pan Huang et al.47 deliberated the end-to-end transformer network with manifold adversarial multi-modal learning (MamlFormer) for laryngeal histopathological grading. Using Hoeffding’s inequality and multimodal co-regularization, the author first shows that the modal fusion of low and high-magnification LSCC pictures is feasible and meets all the requirements. The second step is to create a brand-new manifold block. Following a set of rules, it builds the manifold subspace. These concepts pertain to the feature matrix’s divisibility, recoverability, and the local distance closest to each picture modality before and after LSCC mapping. At the same time, it’s a great way to fix issues like weak modal semantic consistency and redundant feature matrix information that arise after multimodal learning. Thirdly, the adversarial block is implemented using the encoder and the adversarial loss function. In both low- and high-magnification LSCC pictures, it can learn the latent metrics of the modal distributions adaptively. Hence, it complements LSCC low and high-magnification imaging modalities even better. Then, in terms of grading performance and interpretability, MamlFormer beats other SOTA models, according to many trials.
Pan Huang and Xin Luo48 presented the feature disentangled transformer (FDT) for interpretable squamous cell carcinoma grading. This study presents a novel approach to FDTs by combining two concepts: (1) Constructing a SIFDL framework that properly partitions the image into multiscale instances for multi-objective optimization in parallel, then aggregating those instances to make the feature representations human-interpretable; (2) Adding an IAB to find the connection between semantic instances and grading patterns at the instance level, so that low-effect instances (like lymphatic or muscle) are less entangled with the feature representations that are being grasped. Compared to state-of-the-art methods, the suggested FDTs perform better on two real-world SCC datasets, demonstrating their ability to handle the challenge of reliable SCC grading.
Pan Huang et al.49 introduced the end-to-end network with transformers constrained by learned-parameter-free attention (LA-ViT) for Interpretable Grading in a New Laryngeal Histopathology Image Dataset. First, the author shows that learned-parameter-free attention (LA) is better than parameter attention in learning highly effective target semantic information using the generalized linear model and probabilistic. Additionally, the query is realized using the feature map position subspace employing the first-type LA transformer block of LA-ViT. After realizing the key in the feature channel subspace, an average convergence approach is used to get a value. Those are the building blocks of the LA mechanism. As a result, the feature maps and channels include less background semantics. Thirdly, to accomplish key and query, the second-type LA transformer block of LA-ViT utilizes the information from the model probability matrix and the decision level weight information, respectively. They, therefore, realize the LA mechanism. As a result, class activation maps rely less on background semantics.
Yingbo Qu et al.50 examined the end-to-end causality graph attention mamba network (CGAM) for esophageal pathology grading. To increase the graph neural network’s capacity to learn relational representations, the first step is to use the graph attention mechanism as a graph-building technique for learning topological connection representations. Second, to improve the model’s receptive field and direct the graph neural network to capture the tumor-TME link, we leverage the causality of Mamba’s selected state space method. In the third place, inconsistent two-branch choices may be resolved using the entropy-constrained decision consistent optimization approach. On two pathology datasets, we examined the grading performance of nineteen SOTA models; our grading accuracy was 2.72% and 2.90% higher than the top-performing SOTA model, respectively.
Proposed temporal decomposition network (TDN) for medical image fusion
The saliency perceptron of the image with multimodal medical image fusion occurs by matching similar features. The relevant features are extracted from the medical image, which can be classified as homogeneous or heterogeneous. The quality of the image is extracted, and the intensity and contrast are considered for the CT image. The Decomposition is used to divide the image into smaller regions that are used for the pixel distribution. The dataset for the chest X-ray and Brain MRI datasets have been updated to include preprocessing steps, augmentation techniques, and potential biases. Preprocessing includes grayscale normalization, contrast enhancement, and noise reduction for consistency. Data augmentation techniques like rotation, flipping, scaling, and histogram equalization improve generalization and prevent overfitting. Potential biases like class imbalance and scanner variability have been analyzed, and mitigation strategies have been implemented.
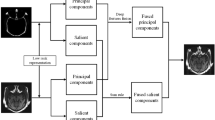
The input CT image is considered for the multimodal medical image and examined. The saliency perceptron extracts the necessary information from the input image. The medical image is taken as the input, and the saliency perceptron’s analysis regarding the quality of the image and the intensity is validated. Figure 1 depicts the proposed TDN model for image fusion.
Proposed temporal decomposition network (TDN) for medical image fusion.
A temporal decomposition step, implemented by the framework, uses multiscale filtering or transform-based algorithms to separate inherent structural and temporal information from input medical pictures acquired from various modalities (e.g., CT, MRI, PET). The emerging features are then classified as either homogeneous (representing traits independent of modality) or heterogeneous (encapsulating discriminative information particular to modality). The Generative Attention Learning (GAL) module uses these attributes to create deep generative frameworks with attention processes. The result is feature maps that are better at recognizing saliency. To help with feature alignment and cross-modality correlation, this module uses mapping operations to project the learned feature distributions into a uniform latent representation space. An unmapping mechanism checks the accuracy of the mapping and sends out a signal to indicate any possible feature discrepancies to ensure the data is accurate and may be undone. Finally, an adaptive fusion operation is carried out to combine the feature maps that have been structurally conserved and those that have been semantically enriched. This creates a composite picture that may be used to retain important diagnostic information from different modalities while reducing data redundancy and loss.
The multimodal medical image is fused to provide visually salient region extraction. The saliency is used to find whether the diseases on the CT image are normal or abnormal. The feature extraction is done on the input region, which is associated with the pixel-based representation. The scope of this paper is to reduce the errors that occur due to unmatching in a fusion process. GAN is proposed for the saliency perceptron of the image where it embarks the homogeneous and heterogeneous feature maps to identify Decomposition matching regions. The preliminary step discussed here is to analyze the saliency perceptron of medical images, and it is formulated as follows:
In Eq. (1):\(\:\varDelta\:\::\:\)Resultant function representing the computed transformation or fusion metric.\(\:{I}_{g}\): Initial input image intensity or grayscale value.\(\:{\varvec{U}}_{\varvec{f}}\): Fusion factor representing the weight applied to image features during processing.\(\:{y}_{c}\) : Coefficients associated with contrast or intensity modulation.\(\:{x}_{t}\): Transformation parameter influencing spatial or intensity adjustments.\(\:{i}_{x}\): Feature index representing pixel-wise interactions.\(\:r{\prime\:}\): Refined residual component related to feature extraction.\(\:r{\prime\:}\left({I}_{g}\right)\): Residual function applied to the input intensity \(\:{I}_{g}\).\(\:{y}_{q}\): Quality factor, determining perceptual enhancement.\(\:{n}_{y}\): Normalization parameter for intensity correction.\(\:{o}_{r}\): Offset factor for pixel intensity calibration.\(\:{i}_{n}\): Noise factor, representing unwanted distortions in image fusion.
The image fusion with saliency feature detection takes place in the above equation, where multimodal image extraction is opted for. The image fusion is represented as\(\:\:F{\prime\:}\), and the examination is described as\(\:\:{N}_{0}\), and the saliency perception is labeled as\(\:\:{S}_{p}\). The features of the image are extracted and provide an efficient feature based on the pixel distribution. The pixel distribution is accounted for the medical image, and detection is done and it is described as\(\:\:{d}_{c}\). For the varying images, the feature extraction is carried out where it estimates the saliency perception for the input image. The input image indicates the necessary features and emphasizes the image fusion technique. The image fusion benchmarks the quality and ensures the pixel distribution with medical images. This examines the saliency feature extraction with necessary information. The perceptron representation before Decomposition is illustrated in Fig. 2.
Decomposition Process Prior to Perceptron Representation.
The perceptron learning illustration is designed with 1 input, 2 conditional, 1 classification, and 1 output layer. The pairs of \(\:\left({o}_{r}\:and\:{i}_{y}\right)\forall\:\left({r}^{{\prime\:}},{p}_{c},\:and\:\varDelta\:\right)\) are used to define the conditional and classification layers. From the conditional layers, \(\:{y}_{q}\) based\(\:\:r{\prime\:}\) and \(\:{i}_{x}\) based \(\:{p}_{c}\) are extracted using\(\:\:\left[\left({U}_{f}*{y}_{c}\right)-{r}^{{\prime\:}}\left({I}_{g}\right)\right]>0\) and \(\:\left[\left({n}_{y}+{o}_{r}\right)-{i}_{n}\le\:1but\ne\:0\right]\) conditions. These outputs (mediate)\(\:\:r{\prime\:}\) and \(\:{p}_{c}\) are used to classify the\(\:\:\left[{U}_{f}\in\:{I}_{g}\right]\forall\:(\varDelta\:\in\:{d}_{c})\). Using this differentiation, the \(\:{U}_{f}\in\:\varDelta\:\) and \(\:{U}_{f}\in\:{i}_{x}\) are classified. The \(\:\varDelta\:\) based features are categorized as similar (homogeneous), and the rest are heterogeneous. Therefore, the Decomposition of the \(\:{I}_{g}\) relies on the \(\:\left({i}_{n}={y}_{c}\right)\) condition using\(\:\:{U}_{f}\) (Refer to Fig. 2). The key factor is the F1-score analyzed for \(\:{o}_{r}\) and \(\:{i}_{y}\) concentration, as presented in Fig. 3 below. Figure 3, the F1 score is estimated for both homogeneous and heterogeneous.\(\:\:{U}_{f}\). This function takes place for the saliency perception, where the necessary information is attained from the noise.\(\:\:{i}_{n}\left({d}_{c}+{S}_{p}\right)*{I}_{g}\). The medical image acquisition is carried out for the feature mapping.\(\:\:{a}_{m}\left({r}^{{\prime\:}}+{n}_{y}\right)*\sum\:_{{S}_{p}}D{e}^{{\prime\:}}\). From this observation step, pixel distribution is examined based on the GAN. This is evaluated for realistic image acquisition and ensures quality.\(\:\:{y}_{q}\left({I}_{g}+{S}_{p}\right)*{U}_{f}\). The processing step is examined for the Decomposition of input images, which is done with feature mapping. The enhancement is done for the reward function attained from the discriminator and generator. This process improves the F1 score based on the feature mapping and evaluates the saliency perception. On this basis, the image fusion relates to the necessary information of the image and finds whether it is normal or abnormal. Based on this detection phase, an image with relevant features is mapped and is said to be fusion.
F1-Score Determination for\(\:\:\left({\varvec{o}}_{\varvec{r}},\varDelta\:,\:{\varvec{y}}_{\varvec{c}}\right)\) and\(\:\:\left({\varvec{i}}_{\varvec{y}},\:\varDelta\:,\:{\varvec{y}}_{\varvec{c}}\right)\)
To maintain modality-specific structural and textural information during fusion, the generator architecture was purposefully designed to facilitate progressive feature extraction and noise suppression. This architecture consists of numerous convolutional layers with ReLU activation and batch normalization. Improving spatial resolution and guaranteeing smooth reconstruction of fused outputs were the goals of including upsampling layers. To make the generator generate more realistic outputs, the discriminator was fine-tuned to discriminate between actual and fused pictures using LeakyReLU activations and a fully linked classification layer. To ensure that both networks received consistent gradients during stable adversarial training, the Binary Cross-Entropy (BCE) loss function was used.
Table 1 details the GAN’s architecture, hyperparameters, and training. The generator and discriminator structure, activation functions, optimization methods, and loss functions are described. Training parameters such as batch size, learning rate, regularization methods, evaluation metrics, and data augmentation procedures were defined for clarity. The exact hyperparameters for the proposed GAN architecture were chosen based on empirical testing and acknowledged best practices in medical image synthesis and fusion tasks. To achieve seamless convergence across complicated feature distributions, we opted for the Adam optimizer, which has a learning rate of 0.0002. It has a history of success in adversarial training. The batch size 32 was chosen to achieve the sweet spot between computational efficiency and training stability, guaranteeing sufficient gradient updates free of memory constraints. Because the discriminator tends to remember training samples in small datasets, we used dropout regularization (0.3) to prevent overfitting. Experimental validation was conducted on an image patch size of 256 × 256 pixels to ensure enough structural features were preserved while tolerating the computational burden. Furthermore, validation-based monitoring helped define the training epochs (1200) with early exit to avoid overtraining and guarantee generalizability.
By processing this, computation time and detection time decrease. The contrast and intensity are considered for the saliency perception for varying input images. From this observation step, the noise is detected and reduced, and the analysis is followed up with the image fusion method. The image fusion is processed to maintain the quality for the saliency perception. So, based on this section, saliency perception is done using the image fusion mechanism for the multimodal input image. This is further examined using the GAN method discussed in the topics below. From this study of saliency perception for multimodal image fusion, the Decomposition for medical images is executed and performed in the steps below.
Decomposition
Decomposition reduces the number of unnecessary image features according to the pixel distribution. This mechanism relates to the medical image and ensures the quality with contrast and intensity bases. This reduction of unnecessary features is observed in the steps below.
Step 1: Feature extraction and enhancement
The image splitting takes place on boundaries and detects the image which is having the high contrast and intensity bases\(\:\:a{\prime\:}\left({I}_{g}\right)\to\:{d}_{c}\left({y}_{c}*{r}^{{\prime\:}}\right)\).
Step 2: Noise Reduction
The noise in the image occurs due to overlapping or blurriness. Based on the components, the noise detection takes place using Decomposition \(\:D{e}^{{\prime\:}}\left({i}_{n}+{y}_{q}\right)-r{\prime\:}\)
Step 3: Image fusion
The relevant images are fused to reduce the computation time, so multimodal image fusion takes place after the noise reduction step,\(\:\:{d}_{c}\left({i}_{x}*{a}^{{\prime\:}}\right)+{N}_{0}\)
Step 4: Matching and Un-Matching Detection
The matching and un-matching estimation is performed by detecting the boundary and fixing the region for the detection phase, \(\:{d}_{c}\left({I}_{g}*{a}^{{\prime\:}}\right)+{S}_{p}\left({y}_{c}\right)\).
The above four steps are required to perform decomposition for the relevant image extraction, where saliency perception takes place based on decomposition, as formulated in the equation below.
The Decomposition is described as\(\:\:De{\prime\:}\), the splitting is represented as\(\:\:a{\prime\:}\), and the examination states the Decomposition of an image based on saliency perception. The above four steps indicate the working flow of the Decomposition of the image that divides the image input into smaller, whereas the\(\:\:r{\prime\:}\) detection of the image is based on the pixel. Here, pixel distribution takes place for the saliency perception, where it estimates that the smaller image will decompose. This part illustrates the medical image processing. The analysis for saliency feature detection is executed in the following equation.
The analysis is done for saliency perception, where the necessary feature extraction is done for the input image, which takes place from the decomposition steps. This analysis is done for saliency perception, examining the detection of features and the intensity and contrast basis. The noise is detected using the decomposition method, where the extraction takes place. The image splitting is done for the input image, whereas the image fusion is done for the saliency perception of feature extraction. This analysis is used to reduce the region, and pixel distribution is accounted for saliency perception. The classification of features is performed as described in the equation below.
The classification is done for the homogeneous and heterogeneous method, and it is represented as\(\:\:\beta\:\). The heterogeneous feature is to maximize the fusion precision, whereas homogeneous refers to the image pixel with the same intensity and contrast. Thus, the analysis done for the image fusion examines the salient features of the input image. This analysis provides the appropriate features for the Decomposition. The steps discussed above are diagrammatically illustrated for the\(\:\:\beta\:\) process.
Graphical Representation of the of the\(\:\:\varvec{D}\varvec{e}{\prime\:}\) Steps for\(\:\:\varvec{\beta\:}\)
The \(\:De{\prime\:}\) process is performed to generate feature maps using heterogeneous and homogeneous. \(\:{U}_{f}\) identified from \(\:\left({i}_{x}+{i}_{y}\right)\) of any\(\:\:{I}_{g}\). Depending on the \(\:({y}_{c}+{i}_{n})\:\)and\(\:\:\left({y}_{c}-{i}_{n}\right)\) the \(\:{U}_{f}\) classification \(\:\forall\:\left[{a}^{{\prime\:}}+\left({r}^{{\prime\:}}*{x}_{t}\right)\right]\) and\(\:\:\left({r}^{{\prime\:}}-{y}_{q}\right)\) is performed. Using this classification, the successive decomposition instances are defined as separate. \(\:\left({i}_{x}+{i}_{y}\right)\) and\(\:\:{d}_{c}\). Thus, \(\:\varDelta\:\) it serves as the intermediate point of \(\:{y}_{c}\) and \(\:r{\prime\:}\) differentiation using\(\:\:{S}_{p}\). Therefore the \(\:{D}_{e}{\prime\:}\) process enclosures are directly used for \(\:\beta\:\) if a new\(\:\:r{\prime\:}\) is identified (Refer to Fig. 4). This classification reduces the false rate for any range of fusion using\(\:\:{U}_{f}\). The false rate analysis is presented in Fig. 5 below.
False Rate Analysis for \(\:\left(\varvec{\beta\:},\:{\varvec{i}}_{\varvec{y}},\:{\varvec{y}}_{\varvec{c}}\right)\) and \(\:(\varDelta\:,\:{\varvec{i}}_{\varvec{y}},\:{\varvec{y}}_{\varvec{c}})\)
The false rate in this TDN is less for varying input images where it estimates the discriminator and generator that attains reward and penalty. Both are acquired from the discriminator and generator process and maintain the quality.\(\:\:{y}_{q}*\sum\:_{S{\prime\:}}(\varDelta\:*{r}^{{\prime\:}})\). The quality is observed based on the decomposition methodology for the saliency perception of medical images. The image features are extracted on the pixel distribution method, where the matching is attained from the image fusion.\(\:\:{A}_{t}+F{\prime\:}\left({I}_{g}\right)\). From this step, region-based segmentation is followed up for the intensity and contrast of the medical image. This false rate is reduced based on the detection of noise where the intensity and contrast.\(\:\:\left({n}_{y}+{o}_{r}\right)*\prod\:_{{I}_{g}}\left({U}_{f}+{i}_{n}\right)\). On executing this process, the false rate is addressed and reduced and maintains the higher resolution from the image fusion technique (Fig. 5). This indicates homogeneous and heterogeneous image processing. The upcoming process is the pixel distribution for the homogeneous image, which is derived as follows:
The pixel distribution is done from the region extracted from the input image, which indicates the homogeneity associated with the pixel representation. From this mechanism, intensity and contrast refer to the necessary information of the image. This process suggests splitting the image and estimating saliency perception based on the detection. For every set of input images, the pixel distribution is carried out, which indicates the decomposition of the image. It is performed from the classification step, which provides the distribution of the image and applies the pixel-based distribution for saliency perception. Generative adversarial learning embarks on homogeneous and heterogeneous feature maps to identify decomposition-matching regions.
Generative adversarial learning
The input image features are identified based on the feasibility of two methods, a discriminator and a generator, which optimize saliency perception.
Generative
It converts the noise to complex data that refers to the image. Here, the training takes place using backpropagation to generate high-quality images. The loss function is reduced by using this generative method.
Discriminative
The differentiation of generated and input data and allocating the pixel based on the region-based fusion. This plays a role in identifying the pixel distribution by classifying homogeneous and heterogeneous features.
The following steps are included for the GAN method, and the feature mapping is indicated to identify the decomposition of region matching.
Step 1: Initialization: Both the generator and discriminator are observed for the input image.
The initialization is labeled as\(\:\:{z}_{t}\), the generator and discriminator are symbolized as\(\:\:{G}_{r}\:\text{a}\text{n}\text{d}\:{C}_{r}\). This is carried out for varying input medical, where the image fusion is done to ensure quality. The generator creates a new image that resembles the input image, whereas the discriminator identifies the input image, and the generator creates a new image.
Step 2: First move of generator: The noise of the image is taken as the input that acts as the initial point for the generator creating the image. Based on the image patterns, the generator changes the noise to the newly sampled data, which resembles the image created by the generator.
The generator is processed in the equation where the feature mapping is followed up, and it is represented as\(\:\:{a}_{m}\), the new input generated is symbolized as\(\:\:{w}_{n}\),\(\:\:{p}_{t}\) is defined as a pattern.
Step 3: Discriminator exertion: Two input patterns are extracted, such as original input from the training dataset and generator-created input. So, the role of the discriminator is to find the original input, and it is based on either 0 or 1; based on this, 1 refers to the original input, whereas 0 is not the real input.
The discriminator exertion is done where it refers to the if and otherwise condition and finds the real and not-real input image.
Step 4: Learning method: The learning process indicates the training set for the generator and discriminator. It is lined in simple terms.\(\:\:{\varvec{C}}_{\varvec{r}}\) correctly identifies the fake input, then a reward is generated for both\(\:\:{\varvec{G}}_{\varvec{r}}\:\text{a}\text{n}\text{d}\:{\varvec{C}}_{\varvec{r}}\). The learning process is trained based on the output and works accordingly to find the correct input, which improves feature mapping with image fusion methodology.
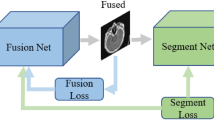
The learning process takes place to observe discrimination, whether it finds the real input or not, based on the training dataset and attains the reward for every step of detection of image fusion. The learning process is represented as\(\:\:L{\prime\:}\). The GAN functions discussed in Steps 1 to 4 are portrayed in Fig. 6.
For detecting matching \(\:({\varvec{C}}_{\varvec{r}}\:output=1)\) and un-matching \(\:\left({\varvec{C}}_{\varvec{r}}output=0\right)\) the GAN model is designed as presented in Fig. 6. The \(\:{C}_{r}\) and \(\:{G}_{r}\) are responsible for \(\:{P}_{t}\) and\(\:\:{a}_{m}\) mapping verification for which\(\:\:\left({w}_{n}+{P}_{t}\right)\to\:{I}_{g}\) is evaluated. Depending on the number of training images available, the \(\:{\varvec{U}}_{\varvec{f}}\) maps between\(\:\:\left({F}^{{\prime\:}},{w}_{n}=0,\:{S}_{p}\right),\:\left(F,{w}_{n}=1,\:{S}_{p}\right),\:and\:\left({F}^{{\prime\:}},{p}_{t},{S}_{p}\right)\) are generated. If either of the above maps are conjoined to form\(\:\:{\varvec{C}}_{\varvec{r}}\), then \(\:\left({y}_{c}-{d}_{c}\right)\) (for homogeneous\(\:\:{\varvec{U}}_{\varvec{f}}\)) and \(\:\left(\varDelta\:-{y}_{q}\right)\) for (for heterogeneous\(\:\:{U}_{f}\)) \(\:{G}_{r}\) is trained using \(\:{\varvec{C}}_{\varvec{r}}=1\) output else new\(\:\:{U}_{f}\) are used to classify the input features through\(\:\:\beta\:\). Thus, the learning is a two-way process for the GAN, where\(\:\:{z}_{t}=\left({y}_{c},{\varvec{U}}_{\varvec{f}}\right)\in\:{D}_{e}{\prime\:}\) is the input.
Step 5: Generator enhancement: If \(\:{C}_{r}\) does not detect the real input generator created, then a penalty is given to the discriminator, whereas the generator improves its mechanism.
The generator enhancement is examined in the above equation, and it is represented as\(\:\:{e}_{n}\). The feature mapping is done to find the real input, and it is not the penalty generated for the discriminator. It is labeledas\(\:\:{l}^{y}.\:\)
Illustrations of GAN Functions from Steps 1 to 4.
Step 6: Discriminator variation: In other cases,\(\:\:{\varvec{C}}_{\varvec{r}}\) Correctly finds the real input the reward is given to \(\:{\varvec{C}}_{\varvec{r}}\) where\(\:\:{\varvec{G}}_{\varvec{r}}\) is given with a penalty. This working process built the generator to perform the realistic input in further computation.
The discriminator variation is\(\:\:{v}_{a}\), as discussed above, the discriminator finds the correct input reward is given; in other cases, the generator is assigned a penalty.
This is how GAN works. Lastly, the generator is improved with new image creation that the discriminator cannot find. So, the realistic image is generated as a well-trained model with input data. From this aspect, the feature map is done for the matching and un-matching process, which is examined in the following steps.
Feature map generation
This represents similar features and input patterns, which are fused with saliency perception. This execution step illustrates the medical image processing that estimates the pixel distribution process to identify similar features, as discussed in the steps below.
Step 1
The initial step is to acquire the input image and perform the matching and un-matching processes. This execution phase defines the pixel distribution based on the salient perception that illustrates the GAN concept.\(\:\:{i}_{x}+\left({I}_{g}\text{*}{x}_{t}\right)+\sum\:_{{y}_{q}}d{w}_{0}.\)
Step 2
This observes the GAN for feature mapping for the input image and analyzes the similarity and dissimilarity. From this methodology, the decomposition of the input image and feature mapping are taken into consideration.\(\:\:{a}_{m}+{F}^{{\prime\:}}\left({I}_{g}\text{*}{r}^{{\prime\:}}\right)+{i}_{x}\).
Step 3
From step 2, the mapping of similar images is done where it refers to the homogenous image processing\(\:\:{a}_{m}\left({H}_{s}+{I}_{g}\right)\text{*}\prod\:_{{N}_{0}}\left({x}_{t}+{d}_{c}\right)\approx\:{A}_{t}\), homogenous is\(\:\:{H}_{s}\), \(\:{A}_{t}\) is matching. This step defines the mapping of similar features that evaluate the detection of images from the medical image dataset.\(\:\:{a}_{m}+\left({F}^{{\prime\:}}\text{*}{y}_{c}\right)\).
Step 4
This refers to the un-matching process, where the differences in features are detected that might be variations in color and intensity\(\:\:\left({o}_{r}+{n}_{y}\right)\text{*}{S}_{p}+{F}^{{\prime\:}}\left({I}_{g}\right)-{E}_{g}\approx\:{H}_{u}\), \(\:{E}_{g}\:\)is the heterogeneous image, \(\:{H}_{u}\) is un-matching.
From the above steps, the matching and un-matching are detected from the input medical image, and it is integrated to observe the feature mapping of matching and un-matching images in the equation below.
The mapping is represented as\(\:\:{m}_{p}\), where it indicates the analysis of matching and un-matching images that state image fusion. Based on feature mapping, the matching forwards the image fusion discussed below. The un-matching image states the error in the computation step.
Image fusion is done through the matching process, which defines the necessary feature extraction. A similar image is\(\:\:S{\prime\:}\), which leads to the saliency perception method and describes the feature mapping. This phase of detection plays an efficient role in image fusion and addresses the error of un-matching images. From the above feature map discussion, the matching and un-matching rates are analyzed using Fig. 7 for the differences.\(\:\:{m}_{p}\) rates.
\(\:\:{\varvec{A}}_{\varvec{t}}\) and\(\:\:{\varvec{H}}_{\varvec{u}}\) rates for the different\(\:\:{\varvec{m}}_{\varvec{p}}\)
In Fig. 7, the \(\:{A}_{t}\) and \(\:{H}_{u}\) variants for \(\:{m}_{p}\) rates are analyzed and depicted. The proposed TDM relies on \(\:{C}_{r}\) and\(\:\:{D}_{r}\) to maximize \(\:S{\prime\:}\) from the inputs for which \(\:L{\prime\:}\) decides \(\:\left({y}_{c}-{d}_{c}\right)\) or \(\:\left(\varDelta\:-{y}_{q}\right)\) for any \(\:{i}_{x}\) and \(\:{i}_{y}\) extracted. The feature mapping rate increases for new\(\:\:{U}_{f}\) identified in \(\:{C}_{r}=0\) output to ensure fewer false rates. Therefore, the classification cases of\(\:\:{A}_{t}\) over\(\:\:{H}_{u}\) is high for different\(\:\:{U}_{f}\). Generative adversarial learning embarks on homogeneous and heterogeneous feature maps to identify decomposition matching regions. The identified regions are fused to leverage the temporal image representations. Based on this representation, the quality degradations at the embarked regions are identified.
A saliency perceptron was a computer model that highlighted key picture features. It selected relevant sections based on intensity, contrast, and geographic distribution to preserve medical features during fusion. Over time, the temporal decomposition procedure broke an image into homogeneous and heterogeneous components to improve feature separation and enhancement. This technique enables adaptive feature refinement, retaining crucial diagnostic features while minimizing unnecessary information. Both techniques helped improve the precision and accuracy of multimodal medical image fusion.
Results and discussion
The first part of Section “Results and discussion” presents the comparative analysis of the metrics’ accuracy, precision, sensitivity, specificity, and mean error. The above metrics are analyzed for Chest and brain images with 256 × 256 and 512 × 512 pixels from the datasets. Besides, the above metrics are comparatively analyzed with the MMIF-NSST36, MedFusionGAN31, and GIAE-Net28 methods discussed in Section “Related works”. Severe motion abnormalities, poor signal-to-noise ratios, or inadequate data capture are common issues in medical pictures used in clinical practice, and they may ruin feature extraction and fusion accuracy. Such settings could reduce the model’s sensitivity to distinguishing important structures, which might provide less-than-ideal fusion results. Consequently, researchers must create adversarial training procedures that mimic real-world degradations while building models, robust normalization layers, and noise-aware feature extraction algorithms to increase noise resistance. Incorporating uncertainty quantification algorithms may improve the system’s decision-support capabilities and clinical dependability by giving doctors confidence levels for fusion results under severe or uncertain settings.
Extensive assessment of fusion quality relies on using datasets that are diverse in imaging modalities, have clinical significance, and have ground truth references available. The input distributions were made uniform, and systematic preprocessing techniques such as intensity normalization, histogram equalization, and modality-specific alignment reduced inconsistencies caused by modality. Aside from using data augmentation methods like flipping and rotation to enhance generalization, we also used them to mimic real-world variability in clinical imaging situations. Cross-validation methods and balanced sample techniques were used to address any dataset-induced bias to prevent performance measurements from being influenced by class imbalance or modality dominance.
Accuracy and precision analysis
The accuracy is high if the detection of medical images has a higher fusion rate. This executes the saliency perception where it estimates the extraction of images with high-quality\(\:\:\left\{\left[\left({r}^{{\prime\:}}+{x}_{t}\right)+\left({U}_{f}\text{*}{I}_{g}\right)\right]+{y}_{c}\right\}\). In this execution phase, the necessary information is retrieved, which reduces the noise. The noise is addressed and decreases, and it is oriented with the intensity and contrast of the input image.\(\:\:\left[\left({y}_{q}\text{*}\left({n}_{y}+{o}_{r}\right)\right)-{i}_{n}\right]\). This execution states the image fusion technique, and for this, GAN is developed to illustrate the patterns and features of the input medical image. On this basis, discriminators and generators play a role in improving realistic image extraction where it opts for the learning patterns. From this processing step, the saliency perception defines the homogeneous and heterogeneous Decomposition of the image.\(\:\:\left(\sum\:_{{i}_{x}}\left({U}_{f}\text{*}{y}_{c}\right)\right)\). The Decomposition divides the bigger image into smaller positions and provides the pixel distribution accordingly. The precision shows a higher value for the featured medical image observed from the decomposition technique. This step defines the feature mapping and provides better region-based segmentation with higher quality. The quality is ensured from region to region and finds for the pixel distribution.\(\:\:\sum\:_{{G}_{r}}\left({I}_{g}-{w}_{n}\right)+{p}_{t}-{a}_{m}\). The pixel distribution is performed for the homogenous and heterogeneous method, which observes saliency perception.\(\:\:\left({w}_{n}+{p}_{t}\right)\to\:{I}_{g}\). The saliency perception defines the new input and finds the actual input that states the feature mapping. The mapping is done on Eq. (8), associated with the classification model.\(\:\:{a}_{m}\left({H}_{s}+{I}_{g}\right)|{F}^{{\prime\:}}\left({I}_{g}\right)-{E}_{g}\). This methodology is designed for the necessary feature mapping, which is similar and ensures higher precision (Table 2). The tabulation for sensitivity and specificity is given below.
The generator and discriminator networks were intentionally subjected to dropout regularization (rate 0.3) to promote greater generalization by disrupting neuronal co-adaptations. After every convolutional layer, batch normalization was added to stabilize the learning process by standardizing feature activations and reducing the impact of internal covariate shifts. There was less chance of oscillations in adversarial loss since the Adam optimizer was used with carefully chosen learning rates (0.0002) to improve convergence stability. As an adaptive control mechanism, early stopping based on validation loss halted training as performance plateaued, limiting the danger of overfitting. In addition, the training dataset became more variable due to data augmentation methods, including random flipping, rotation, and contrast modifications, which compelled the model to acquire more generic properties.
Sensitivity and specificity analysis
The sensitivity is high, estimated based on the intensity and contrast, and it illustrates saliency perception. On this basis, feature mapping is followed up to detect noise in the image.\(\:\:\left({w}_{n}+{p}_{t}\right)\to\:{I}_{g}\:\). The pattern and features of the image are analyzed and found for real and fake input, and the reward function is attained if no penalty is generated. Based on the GAN concept, sensitivity plays a role in detecting noise in the feature; in this case, matching is followed up.\(\:\:{i}_{n}+{d}_{c}\left({I}_{g}\right)\text{*}{S}_{p}\). The saliency perception is done with the decomposition of an image where the heterogeneous and homogeneous work to improve pixel distribution.\(\:\:{I}_{g}\left(D{e}^{{\prime\:}}+{S}_{p}\right)\text{*}{d}_{c}+{i}_{x}\). In executing this step, the output image’s necessary features and quality are extracted. The splitting takes place for the region-based segmentation and shows whether it matches. So, by performing this, sensitivity is maintained and improved. The specificity shows the higher value range, performed with the GAN concept, indicating the discriminator and generator.\(\:\:\left({G}_{r}+{C}_{r}\right)\text{*}\sum\:_{{i}_{x}}{y}_{c}+De{\prime\:}\). In this execution phase, the necessary feature mapping is followed for the saliency perception. This execution states the Decomposition of the image and generates the pixel distribution.\(\:\:{x}_{t}\left({I}_{g}\right)-{G}_{r}\left({l}^{y}\right)+{i}_{x}\). Based on the pixel distribution, the intensity and contrast are considered, and the specificity is improved. This case relies on matching similar images, which takes place using the fusion technique.\(\:\:{S}^{{\prime\:}}\left({A}_{t}\right)+{I}_{g}\). Image fusion is carried out by examining images extracted from medical image processing.\(\:\:{a}_{m}+{F}^{{\prime\:}}\left({I}_{g}\text{*}{r}^{{\prime\:}}\right)\). The image processing is enhanced with saliency perception and provides the Decomposition of the image. Table 3 presents the mean error with the corresponding false rate for different sizes and types.
Mean error analysis
The mean error shown is less based on the extractions associated with the feature mapping. This concept defines the pixel distribution, evaluates the feature, and reduces the noise.\(\:\:\sum\:_{{a}_{m}}\left({i}_{n}-{I}_{g}\right)+{A}_{t}\). The noise in the medical image is addressed using the GAN concept, where the reward and penalty are indicated on the region technique.\(\:\:\left(d{w}_{0}+{l}^{y}\right)\text{*}\prod\:_{{y}_{q}}({a}_{m}+{N}_{0})\). The extraction is done for the varying images, which are used to identify the matching and unmatching processes. Image fusion accounts for similar features in the image where the pixel distribution is carried out. Based on this, the error is reduced if image fusion is attained for a similar feature.\(\:\:{U}_{f}\left({I}_{g}\text{*}{F}^{{\prime\:}}\right)-{i}_{n}\). This mechanism uses feature-based mapping with similar patterns and examines the pixel distribution. The pixel distribution is done for the un-matching process where the error is addressed and reduced (Table 4).
Distribution outputs and fused image output
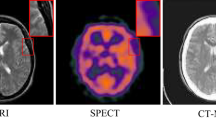
The experimental outcomes are represented in Tables 5 and 6 for the different processes described above. Two types of images, Chest (CT) and brain (MRI) inputs are validated from51] and [52, respectively, where the Chest CT images for pneumonia analysis accounted for 390 infected and 234 normal images for testing and 1341 and 3875 normal and infected images for training. The second dataset boasts 300 glioma tumors for testing and 1321 training images. From both datasets, 256\(\:\times\:\)256 and 512\(\:\times\:\)512-pixel distributed images are selected and validated using MATLAB codes. This setup is executed in a standalone computer with 8GB random access storage, 128GB secondary storage, and a 2.4 GHz 4-core processor. The training iterations are a maximum of 1200 with 4 epochs on average. Some sample results are depicted in the following tables with this experimental environment.
Table 7 compares The Temporal Decomposition Network (TDN) to six cutting-edge medical image fusion algorithms in the Technical Comparison of Multimodal Medical Image Fusion Methods. GAN-based feature matching and temporal Decomposition give the TDN the highest fusion accuracy and precision at a more significant computational cost. Other approaches, including MMIF-NSST, GIAE-Net, TUFusion, AMMNet, and CDRNet, trade computational efficiency, generalization, and feature retention. In medical image fusion tasks, where localized spatial and modality-specific information is crucial, TDN’s higher capacity to extract temporal and frequency-oriented decomposition characteristics proved a deciding factor. However, medical imaging scenarios with limited access to annotated multimodal datasets present unique challenges when trying to achieve optimal performance using transformer-based models, which are powerful in modeling global dependencies but typically involve higher computational complexity and require substantially larger datasets. Also, TDN’s decomposition-guided fusion method takes care of structural alignment and fine-grained saliency preservation, which transformers could have trouble with. Based on the empirical data, TDN is a better and more interpretable option for real-time clinical deployment since it has quicker inference times and improved fusion consistency.
Ablation studies on an independent dataset show the Temporal Decomposition Network (TDN)’s generalization performance. The study uses a varied dataset with various medical imaging modalities and diseases, such as ChestX-ray14 for chest imaging or BraTS for brain tumor imaging. TDN components, including the salient perception model and temporal decomposition processes, are identified and adjusted. FQI, SSIM, and PSNR measure performance. Visualization and statistical analysis evaluate results. TDN’s robustness and potential efficacy in real-world circumstances bolster its status as a multimodal medical picture fusion breakthrough.
TDN’s p-values and confidence intervals are compared to other approaches in Table 8. TDN had statistically significant accuracy, precision, sensitivity, and specificity gains. Lower p-values (< 0.05) indicate that TDN’s improvements are reliable and robust. The table also demonstrates that TDN regularly had the top performance measures with tight confidence intervals, indicating steady improvements. With substantial statistical significance, our analysis supports TDN’s superiority. Further modifications are needed to confirm the results.
Table 9 outlines technical issues in traditional medical image fusion methods and proposes a Temporal Decomposition Network (TDN), highlighting its potential to improve diagnostic accuracy, reliability, and versatility in clinical applications.
Ablation study
An in-depth evaluation of performance measures while removing important modules like the adversarial training component, the temporal decomposition module, or the saliency-guided feature extraction block would show how each subcomponent is related to the others. Excluding the saliency perception module from the fused outputs makes fine-grained anatomical features less clear, and eliminating the adversarial loss leads to decreased structural consistency, according to such an examination. Breaking down the study into its components allows us to understand better how discriminative feature learning affects fusion quality and validate the TDN framework’s design rationale. This, in turn, improves the study’s methodological transparency and technical rigor. Table 10 shows the ablation study of TDN components.
Ablation research has been conducted to determine the specific effects of suggested key architectural elements within the TDN framework. After removing the adversarial training module, lower SSIM and PSNR values were seen, suggesting that the discriminator is critical for improving the fusion output’s structural realism and picture fidelity. Furthermore, fine-detail preservation was further decreased upon removing the saliency perception module, demonstrating its function in directing modalities toward spatially significant feature extraction. The accuracy loss was most pronounced when the temporal decomposition block was removed, proving that it successfully detangled and captured modality-specific temporal-frequency information that strengthened the fusion process. These changes seem essential for feature alignment and domain-level consistency, as their exclusion had a detrimental effect on structural coherence. All factors considered, the findings prove that TDN’s strong performance is the consequence of each component’s distinct and complementary roles, which, when combined, allow for better fusion quality and clinical application.
The performance analysis would be much more rigorous if it included effect size comparisons, confidence intervals, and p-values, greatly improving the evaluation’s statistical reliability. For instance, to better understand the real-world relevance of performance improvements beyond just statistical significance, one may compute Cohen’s d between the suggested TDN and competing approaches like GIAE-Net or MedFusionGAN. In real-world clinical applications, gains made by TDN would be statistically significant and substantively useful if the effect size is substantial (e.g., d > 0.8). This method’s advantage might be better explained using quantitative effect size measurements that better quantify the extent of increased diagnostic reliability and fusion quality.
The implementation across diverse imaging devices is a substantial barrier since model performance and generalizability may be negatively impacted by differences in image quality, acquisition techniques, and device-specific artifacts. Furthermore, the fusion model’s resilience in multiple clinical situations might be compromised by bias introduced by inconsistent and imbalanced training data, especially regarding patient demographics, disease kinds, and imaging modalities. The use of domain adaption methods and meticulous normalization processes is necessary to guarantee uniform performance among institutions with different imaging standards. In addition, complicated deep learning models may not be able to be deployed in clinical settings due to computational restrictions, particularly in low-resource or mobile contexts, unless there is considerable model reduction or hardware optimization. To guarantee clinical reliability and interoperability in real-world healthcare systems, it is necessary to implement stringent validation methods in addition to scalable, adaptive, and hardware-agnostic solutions.
Conclusion
The proposed TDN framework significantly advances multimodal medical image fusion by successfully integrating temporal Decomposition with adversarial learning. The improvements in fusion accuracy (11.378%) and precision (12.441%) validate the effectiveness of combining salient perception models with generative adversarial learning for heterogeneous feature integration. The framework’s performance across different image types and sizes confirms its robustness and practical applicability in clinical settings.
Limitations
The dual-component architecture’s computational complexity could affect real-time processing capabilities, and current validation is limited to specific medical imaging modalities. It relies on high-quality input images for optimal feature extraction and fusion performance and faces challenges in handling extreme image quality variations.
Broader implications
Improved visualization of medical imaging data enhances diagnostic accuracy, reduces errors, and aids clinical decision-making. Advancements in automated medical image analysis techniques contribute to multimodal data fusion and deep learning applications in healthcare.
Future directions
The suggested fusion architecture will have its real-time processing capabilities improved in future work to handle time-critical clinical applications. Additional imaging modalities and higher-dimensional data formats (such as 3D/4D imaging) will be investigated further to enhance diagnostic comprehensiveness. The development of computationally efficient and lightweight designs is fundamental to guarantee the viability of deployment in resource-constrained settings, including mobile health systems and point-of-care systems. This study will use transfer learning and domain adaption approaches to enhance the model’s generalization ability across various datasets and clinical settings. Additional support for verifying the accuracy and precision of the fusion results may be provided by using automated picture quality evaluation criteria. Future studies will look at how to include explainable AI (XAI) components for better model transparency, clinician confidence, and decision-making. Last but not least, research on privacy-preserving picture fusion techniques, such as secure computing methods and federated learning, is increasingly necessary to guarantee data secrecy and conformity with regulations about medical data.
Data availability
The data will be made available by the corresponding author on a request.
References
Zhou, Y., Yang, X., Liu, S. & Yin, J. Multimodal medical image fusion network based on target information enhancement. IEEE Access. 12, 70851–70869 (2024).
Jana, M. & Das, A. Multimodal medical image fusion using a two-stage decomposition technique to combine the significant features of Spatial fuzzy plane and transformed frequency plane. IEEE Trans. Instrum. Meas. 72, 1–10 (2023).
Lopes, C., Vilaca, A., Santos, C. & Mendes, J. An Open-Source software approach to multimodal medical imaging: combination of anatomical and thermal 3D models. Procedia Comput. Sci. 239, 122–129 (2024).
Chen, J. & Chen, J. Multimodal image feature fusion for improving medical ultrasound image segmentation. Biomed. Signal Process. Control. 89, 105705 (2024).
Soltwedel, J. et al. Slice2Volume: fusion of multimodal medical imaging and light microscopy data of irradiation-injured brain tissue in 3D. Radiother. Oncol. 182, 109591 (2023).
Zhu, Z., Liu, Y., Yuan, C. A., Qin, X. & Yang, F. A Diffusion Model Multiscale Feature Fusion Network for Imbalanced Medical Image Classification Research, 256108384 (Computer Methods and Programs in Biomedicine, 2024).
Yang, M. et al. A feature fusion module based on complementary attention for medical image segmentation. Displays 84, 102811 (2024).
Li, W., Zhang, Y., Wang, G., Huang, Y. & Li, R. DFENet: A dual-branch feature enhanced network integrating Transformers and convolutional feature learning for multimodal medical image fusion. Biomed. Signal Process. Control. 80, 104402 (2023).
Xu, S., Chen, Y., Yang, S., Zhang, X. & Sun, F. FCSU-Net: A novel full-scale Cross-dimension Self-attention U-Net with a collaborative fusion of multiscale feature for medical image segmentation. Comput. Biol. Med. 180, 108947 (2024).
Zhu, Z. et al. Lightweight medical image segmentation network with multiscale feature-guided fusion. Comput. Biol. Med. 182, 109204 (2024).
Zhou, Y., Yang, X., Yin, J. & Liu, S. Computers research on multiscale feature fusion network algorithm based on brain tumor medical image classification. Mater. Contin., 79 (3). (2024).
Zhao, C. et al. MHW-GAN: Multidiscriminator Hierarchical Wavelet Generative Adversarial Network for Multimodal Image Fusion (IEEE Transactions on Neural Networks and Learning Systems, 2023).
Wen, J. et al. Msgfusion: medical semantic guided two-branch network for multimodal brain image fusion. IEEE Trans. Multimedia. 26, 944–957 (2023).
Qiu, Z. et al. 3D Multimodal Fusion Network with Disease-induced Joint Learning for Early Alzheimer’s Disease Diagnosis (IEEE Transactions on Medical Imaging, 2024).
Ameen, R. A. & Al Maktoum, L. Machine learning algorithms for emotion recognition using audio and text data. Pattern IQ Min., 1 (4), 1–11. (2024). https://doi.org/10.70023/sahd/241101
Min, X., Duan, H., Sun, W., Zhu, Y. & Zhai, G. Perceptual video quality assessment: A survey. Sci. China Inform. Sci. 67 (11), 211301 (2024).
Min, X. et al. Screen content quality assessment: overview, benchmark, and beyond. ACM Comput. Surv. (CSUR). 54 (9), 1–36 (2021).
Min, X., Zhai, G., Zhou, J., Farias, M. C. & Bovik, A. C. Study of subjective and objective quality assessment of audiovisual signals. IEEE Trans. Image Process. 29, 6054–6068 (2020).
Min, X., Zhai, G., Gu, K. & Yang, X. Fixation prediction through multimodal analysis. ACM Trans. Multimedia Comput. Commun. Appl. (TOMM). 13 (1), 1–23 (2016).
Nair, S. & Kumar, A. Zero-Shot learning algorithms for object recognition in medical and navigation applications. PatternIQ Min. 1 (4), 24–37. https://doi.org/10.70023/sahd/241103 (2024).
Jiang, Q. et al. A lightweight multimode medical image fusion method using similarity measure between intuitionistic fuzzy sets joint laplacian pyramid. IEEE Trans. Emerg. Top. Comput. Intell. 7 (3), 631–647 (2023).
He, K., Zhang, X., Xu, D., Gong, J. & Xie, L. Fidelity-driven optimization reconstruction and details preserving guided fusion for multi-modality medical image. IEEE Trans. Multimedia. 25, 4943–4957 (2022).
Shi, K., Liu, A., Zhang, J., Liu, Y. & Chen, X. Medical image fusion based on multilevel bidirectional feature interaction network. IEEE Sens. J. ( 2024).
Zhao, Y., Zheng, Q., Zhu, P., Zhang, X. & Ma, W. TUFusion: A transformer-based Universal Fusion Algorithm for Multimodal Images (IEEE Transactions on Circuits and Systems for Video Technology, 2023).
Di, J., Guo, W., Liu, J., Ren, L. & Lian, J. AMMNet: A multimodal medical image fusion method based on an attention mechanism and MobileNetV3. Biomed. Signal Process. Control. 96, 106561 (2024).
Fu, J. et al. CDRNet: Cascaded dense residual network for grayscale and pseudocolor medical image fusion. Comput. Methods Prog. Biomed. 234, 107506 (2023).
Song, Y. et al. DesTrans: A medical image fusion method based on transformer and improved densenet. Comput. Biol. Med. 174, 108463 (2024).
Zhou, T. et al. GIAE-Net: A gradient-intensity oriented model for multimodal lung tumor image fusion. Eng. Sci. Technol. Int. J. 54, 101727 (2024).
Ding, Z. et al. M4fnet: multimodal medical image fusion network via multi-receptive-field and multiscale feature integration. Comput. Biol. Med. 159, 106923 (2023).
Zhang, W., Yu, L., Wang, H. & Pedrycz, W. End-to-end dynamic residual focal transformer network for multimodal medical image fusion. Neural Comput. Appl., 1–23. (2024).
Safari, M., Fatemi, A. & Archambault, L. MedFusionGAN: multimodal medical image fusion using an unsupervised deep generative adversarial network. BMC Med. Imaging. 23 (1), 203 (2023).
Dinh, P. H. & Giang, N. L. Medical image fusion based on transfer learning techniques and coupled neural P systems. Neural Comput. Appl. 36 (8), 4325–4347 (2024).
Zhang, Y., Nie, R., Cao, J., Ma, C. & Wang, C. SS-SSAN: a self-supervised subspace attentional network for multimodal medical image fusion. Artif. Intell. Rev. 56 (Suppl 1), 421–443 (2023).
Pan, L., Nie, R., Zhang, G., Cao, J. & Han, Y. WAE-TLDN: self-supervised fusion for multimodal medical images via a weighted autoencoder and a tensor low-rank decomposition network. Appl. Intell. 54 (2), 1656–1671 (2024).
Min, X. et al. A multimodal saliency model for videos with high audiovisual correspondence. IEEE Trans. Image Process. 29, 3805–3819 (2020).
Khan, S. U., Khan, F., Ullah, S. & Lee, B. Multimodal medical image fusion in NSST domain with structural and spectral features enhancement. Heliyon, 9 (6). (2023).
Min, X. et al. Blind quality assessment based on pseudo-reference image. IEEE Trans. Multimedia. 20 (8), 2049–2062 (2017).
Min, X., Zhai, G., Gu, K., Liu, Y. & Yang, X. Blind image quality Estimation via distortion aggravation. IEEE Trans. Broadcast. 64 (2), 508–517 (2018).
Min, X. et al. Exploring rich subjective quality information for image quality assessment in the wild. arXiv:2409.05540. (2024).
Huang, P. & Luo, X. FDTs: A feature disentangled transformer for interpretable squamous cell carcinoma grading. IEEE/CAA J. Autom. Sin. (2025).
Huang, P. et al. LA-ViT: A network with transformers constrained by learned-parameter-free attention for interpretable grading in a new laryngeal histopathology image dataset. IEEE J. Biomed. Health Inform. (2024).
Huang, P. et al. MamlFormer: Priori-experience guiding transformer network via manifold adversarial multi-modal learning for laryngeal histopathological grading. Inform. Fusion. 108, 102333 (2024).
Bhalla, K., Xiao, Q., Luna, J. M., Podany, E., Ahmad, T., Ademuyiwa, F. O., Gastounioti, A. Radiologic imaging biomarkers in triple-negative breast cancer: A literature review about the role of artificial intelligence and the way forward. BJR| Artif. Intell., 1(1), ubae016 (2024).
Roy, S., Bhalla, K. & Patel, R. Mathematical analysis of histogram equalization techniques for medical image enhancement: A tutorial from the perspective of data loss. Multim. Tools Appl., 83 (5), 14363–14392. (2024).
Huang, Y. P. et al. Wavelet k-means clustering and fuzzy-based method for segmenting MRI images depicting Parkinson’s disease. Int. J. Fuzzy Syst., 1–13. (2021).
Bhalla, K., Koundal, D., Sharma, B., Hu, Y. C. & Zaguia, A. A fuzzy convolutional neural network for enhancing multi-focus image fusion. J. Vis. Commun. Image Represent. 84, 103485 (2022).
Huang, P. et al. MamlFormer:Priori-experience guiding transformer network via manifold adversarial multi-modal learning for laryngeal histopathological grading. Inf. Fusion 108, 102333 (2024).
Huang, P. & Luo, X. FDTs: A feature disentangled transformer for interpretable squamous cell carcinoma grading. IEEE/CAA J. Autom. Sin. (2025).
Huang, P. et al. La-vit:A network with transformers constrained by learned-parameter-free attention for interpretable grading in a new laryngeal histopathology image dataset. IEEE J. Biomed. Health Inform. (2024).
Qu, Y. et al. CGAM: an end-to-end causality graph attention Mamba network for esophageal pathology grading. Biomed. Signal Process. Control. 103, 107452 (2025).
Chest X-Ray Images (Pneumonia) Dataset https://www.kaggle.com/datasets/paultimothymooney/chest-xray-pneumonia
Brain Tumor MRI Dataset https://www.kaggle.com/datasets/masoudnickparvar/brain-tumor-mri-dataset
Acknowledgements
This work was funded by the Deanship of Graduate Studies and Scientific Research at Jouf University under grant No. (DGSSR-2023-02-02261).
Author information
Authors and Affiliations
Contributions
Authors Contributions:1. Mohammed Albekairi: Conceptualization, Methodology2. Mohamed vall O. Mohamed: Data curation, Writing–original draft,3. Khaled Kaaniche: Validation, Writing–review and editing4. Ghulam Abbas: Writing–original draft, Writing–review and editing5. Meshari D. Alanazi: Writing–review and editing, Supervision6. Turki M. Alanazi: Formal Analysis, Validation.7. Ahmed Emara: Data curation, Writing–review and editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Albekairi, M., Mohamed, M.v.O., Kaaniche, K. et al. Multimodal medical image fusion combining saliency perception and generative adversarial network. Sci Rep 15, 10609 (2025). https://doi.org/10.1038/s41598-025-95147-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-95147-y
Keywords
This article is cited by
-
Multitask optimization and convergence stability with hierarchical feature learning for self guided optimization
Scientific Reports (2026)
-
Fusion of optimized feature descriptors for medical image retrieval and classification
Iran Journal of Computer Science (2025)
-
A predictive model for differentiating PFA and PFB subtypes of posterior fossa ependymoma using multi-sequence MRI radiomics: a two-center study
Neuroradiology (2025)