Abstract
Abiotic stresses induced by climate change and soil pollution cause significant crop yield loss. Identifying stress-resistant genes for genetic improvement is urgent and necessary. Reticulon (RTN, also called RTNLB in plants) is a widely distributed endoplasmic reticulum-associated protein in eukaryotic cells, playing a crucial role in stress response activities, including autophagy, endoplasmic reticulum stress, and osmotic regulation. However, the diversity of RTNLBs in plants has hindered comprehensive analysis. This study identified 20 OsRTNLBs from Oryza sativa through a comprehensive whole-genome analysis. Detailed information on their chromosomal localization, protein motifs, phylogenetic relationships, and gene collinearity has been provided. The effects of five major abiotic stresses—cold, heat, heavy metals, salt, and drought—on rice seedling development have been assessed. The expression levels of RTNLBs under these abiotic stresses show significant changes, with an overall increase in transcription levels under simulated drought stress and a decrease under the other four stresses. Three OsRTNLBs (OsRTNLB4, OsRTNLB8 and OsRTNLB9) have been identified as key proteins contributing to abiotic stress.
Similar content being viewed by others
Introduction
Reticulon (RTN) is a kind of endoplasmic reticulum (ER)-associated proteins. RTNs are widely spread in eukaryotic organisms, but there no homologous protein has been discovered in prokaryotes1. RTNs includes a conserved reticulon homology domain (RHD) which is the interval of other 2 domains: N-terminal and C-terminal2. RTNs can participate in cell activities, like division and apoptosis, protein transport, membrane formation and stabilization, also ER tube or vesica composition1,3.
Reticulon homologs in eukaryotes are classified into six reticulon-like protein subfamilies (RTNLs), with the plant subfamily referred to as RTNLB4. RTNLBs are found in various membrane organelles and are involved in the composition, alteration, and function of membranes3,5. Some specific RTNLBs, although unable to regulate membrane curvature, exhibit a punctate distribution within cells and possess enzymatic activity related to biosynthesis6. RTNLBs play a significant role in ER stress through selective autophagy7. While the functions of a few RTNLBs have been identified3,5, much about them remains unknown.
Rice (Oryza sativa) is a major food crop cultivated worldwide8. It is highly susceptible to various abiotic stresses such as drought9,10, heat11, cold12, salt13, and heavy metals14. These stresses can induce ER stress, triggering selective autophagy pathways to repair damaged organelles or misfolded proteins15. In Zea mays, selective autophagy is regulated by ZmRTNLB27. Consequently, RTNLB proteins, which regulate autophagy in response to ER stress, may play a crucial role in plant stress resistance7,15. However, systematic studies on the interaction between plant RTNLB proteins and abiotic stress are lacking. A comprehensive analysis of gene families responding to abiotic stress would facilitate the integration of existing research on RTNLBs across different species, aiding in the prediction and screening of key genes. Therefore, research on RTNLB genes related to the abiotic stress response in rice is expected to identify key RTNLBs that contribute to stress resistance.
To investigate the role of OsRTNLBs in abiotic stress, we focus on the following questions: (i) What are the composition, evolutionary relationships, and regulatory networks of OsRTNLBs? (ii) How do OsRTNLBs respond to abiotic stress, and which of them are crucial? Addressing these questions should provide novel insights into the study of abiotic stress responses and RTNLBs in O. sativa.
Result
Identification and system development analysis of OsRTNLBs
The Reticulon domain (PF02453) has been used to perform a hidden Markov model search (HMM) on the whole genome protein sequence of Oryza sativa. 20 OsRTNLBs in O. sativa Japonica Group and 19 in O. sativa Indica Group (ZS97: O. sativa subsp. indica cv. Zhenshan 97) were identified and named according to their chromosomal positions (Table 1). O. sativa Indica, one of the most commonly cultivated sub-species of rice, has a deletion of the OsRTNLB9 gene.
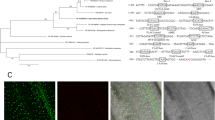
A multiple sequence alignment analysis has been conducted, and conservative segments were analyzed using the Align tool. Furthermore, a comparative analysis combined the amino acid sequences of 20 OsRTNLBs from O. sativa, 21 AtRTNLBs from A. thaliana and 23 OsRTNLBs from Z. mays to construct a Maximum Likelihood (ML) system developmental tree, leading to the construction of a phylogenetic evolutionary tree (Fig. 1A). 7 Clades were delineated based on the distances in the evolutionary tree.
A. Evolutionary tree of RTNLBs in (A) thaliana, O. sativa, and Z. mays. Different species are represented by distinct colors, with the branch lengths proportional to the evolutionary distances. (B) Motif predictions for OsRTNLBs. Different colors indicate distinct motifs. In the left panel, the width of the color blocks corresponds to the length of each motif, while their positions on the black line represent the motif’s location within the protein sequence. The height of the color block reflects the similarity between the actual protein sequence and the motif sequence. In the right panel, the height of the letters indicates the level of conservation of the amino acid residue across all OsRTNLBs, with taller letters indicating higher conservation.
Clade1 exclusively contains RTN proteins from Z. mays and O. sativa. This may be result in the fact that Z. mays and O. sativa both belong to monocotyledonous plants, while A. thaliana is a dicotyledonous plant. ZmRTNLB1 and ZmRTNLB2 in Clade 1 have been suggested their function in endoplasmic reticulum autophagy7. Clade 2 has a cluster of AtRTNLBs, with another part contained only AtRTNLB8. Both AtRTNLB8 and AtRTNLB3, which in Clade 2, are involved in the Agrobacterium Tumefaciens Infection process16. In Clade 4, ZmRTNLB16 is participated in the assembly of vacuolar membrane pumps and response to drought stress17. In Clade 6, A. thaliana RTN19 and RTN20 contain domains which is effected in sterol synthesis6. In Clade 7, there are substantial evidences of AtRTNLB13, suggesting its key role in endoplasmic reticulum morphogenesis18. However, AtRTNLB13 is far evolutionary distance to the OsRTNLBs which in the same clade, indicating potential differences in endoplasmic reticulum morphogenesis mechanisms between A. thaliana and O. sativa. Based on existing research, information is lacked about RTNLBs in Clade 3 and 5. On the phylogenetic tree, RTNLBs are less likely to exhibit species-specific clustering, indicating that the differentiation of RTN proteins is more likely based on functional divergence.
According to relevant published reports and our results, we have highlighted Clades 1, 4, and 6, as well as OsRTNLB4, 8, 9, and 20 from these clades, for increased attention in subsequent studies. This focus is due to the presence of homologous RTN proteins from other species, which provide a basis for predicting the functions of OsRTNLBs based on existing research. However, this does not imply that other clades are less important. We have merely marked these clades to facilitate further investigation, without using this as a criterion for selection.
Characterization of OsRTNLBs
An analysis of the motifs of the 20 RTN proteins (Fig. 1B) revealed that the RTN protein motifs are mostly identical among most members. Among the six conserved motifs identified, the red, yellow, and blue motifs are highly conserved in 15 members (OsRTNLB1-6, OsRTNLB8, OsRTNLB10-17) (Fig. 1B). These motifs constitute the conserved RHD and APH domains18,19, indicating that these members likely retain the ability to localize to the endoplasmic reticulum membrane and regulate membrane curvature changes. Analysis with the evolutionary tree together, it is observed that motifs in Clade 1 and Clade 4 are relatively conservative, while in Clade 6, OsRTNLB9 and OsRTNLB20 share similarities but differ from OsRTNLB18. Additionally, the predictions indicate that the tertiary structures of various members within this family are highly diverse (Figure S1).
To investigate the chromosomal positions of OsRTNLBs genes in the genome, we conducted computer-aided mapping analysis of genetic loci. The results indicate that the 20 OsRTNLBs are only unevenly distributed across chromosomes 1 to 9 (Fig. 2A), while 12 chromosomes in rice. We have done a cross-species collinearity analysis (Fig. 2B), and the substantial collinearity events between Z. mays and O. sativa suggest extensive duplication events in the evolutionary history of the RTN family. This may contribute to the relatively large size (relative to animal20) of the RTN family in plants. Meanwhile, the fewer collinear events between A. thaliana and O. sativa might be due to their distant evolutionary relationship, which illustrate the remarkably high diversity of the RTN family among species with distant phylogenetic ties.
(A) Chromosomal localization of the rice RTN gene family. This panel shows the distribution of OsRTNLB genes across the rice chromosomes, with their positions labeled on each chromosome. (B) Synteny analysis of the OsRTNLBs with AtRTNLBs and ZmRTNLBs. Chromosomes from O. sativa, A. thaliana, and Z. mays are represented by green, blue, and purple rectangles, respectively. Curved lines between the chromosomes illustrate synteny relationships, indicating homologous gene pairs across the species.
Previous reports suggested that OsRTNLBs comprise 18 members7, but in this study, we identified 20 RTN family members in the rice genome. This discrepancy may be due to updates and improvements in the Rice Genome Database, emphasizing the importance of utilizing the latest resources for gene analysis. The newly identified members, OsRTNLB9 and OsRTNLB20, exhibit substantial differences in their protein sequences compared to other family members, while still retaining the conserved Reticulon Homology Domain (RHD), a hallmark of RTN proteins. This highlights the potential functional diversification of these newly identified genes within the RTNLB family. Collinearity analysis revealed significant collinear events between Z. mays and OsRTNLBs, implying that segmental duplication is crucial for the extensive size of OsRTNLBs in higher plants (Fig. 2B). In contrast, minimal collinearity between rice and A. thaliana suggests substantial differences in RTNLB proteins among evolutionarily distant plant species (Fig. 2B), indicating diverse functions and mechanisms among species.
CREs analysis of OsRTNLBs
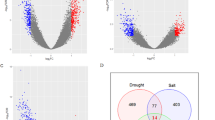
CREs (cis-regulatory elements) are non-coding DNA sequences in the gene promoter region, typically located within the 2000 bp region preceding the promoter21. They are essential for gene expression and widely participate in the regulation of plant growth, development, and stress responses22. We analyzed the CREs of OsRTNLBs, predicting the transcription factors (TFs) regulating OsRTNLBs (Figure S2A). A total of 4935 TF binding sites were annotated, belonging to 36 TF families, and a regulatory network between RTNLBs and TFs is depicted (Figure S3A, Data S1). The top 10 TF families with the highest binding site counts were selected to construct the core regulatory network (Fig. 3A). The top 10 TF families are ERF, Dof, MYB, bZIP, C2H2, BBR-BPC, TCP, MIKC_MADS, NAC, and bHLH. These gene families are significantly enriched in the promoter region of OsRTNLBs, suggesting their crucial role in regulating OsRTNLBs expression levels.
(A) Putative transcription factor (TF) regulatory network analysis of OsRTNLBs. The network diagram displays interactions between the OsRTNLB genes and the top 10 enriched TF families. Each TF family is represented by a node, with node color intensity reflecting the enrichment rank (darker colors indicate higher enrichment). Lines represent regulatory interactions between TF families and OsRTNLBs. (B) Number and proportion of Top 10 TFs targeting OsRTNLB genes. Pie charts illustrate the distribution of TF families regulating specific OsRTNLB genes. The total number of TF binding sites is indicated below each pie chart. (C) Word cloud of CREs function prediction. The word cloud highlights predicted functions of cis-regulatory elements (CREs) in the promoters of OsRTNLB genes. The font size corresponds to the frequency of CREs associated with specific functions, with larger fonts indicating higher counts.
We found that OsRTNLB4 is controlled by all top 10 transcription factor families with a balanced distribution of binding sites. ERF, Dof, MYB, and C2H2 have a major impact on OsRTNLB8. The main TFs for OsRTNLB9 are ERF and Dof. However, ERF has a minimal effect on OsRTNLB20, implying the specificity of OsRTNLB20 within the RTN family (Fig. 3B). Comparing the transcriptional regulation of each member of OsRTNLBs, we observed that the transcriptional regulation of this family does not have a unified master regulator or core, suggesting the functional diversity of the RTN family.
Additionally, we obtained annotation information for CREs from the Plant Care database, which annotates CREs’ functions from three perspectives: plant hormones, growth and development, and stress responses. The results show (Fig. 3C, Figure S2B): (i) OsRTNLBs may be significantly influenced by Jasmonic acid and Abscisic acid. (ii) Stimuli such as cold, drought, injury, and hypoxia may affect the expression of OsRTNLBs. (iii) OsRTNLBs may be associated with the cell cycle. To provide an intuitive representation of the transcriptional regulation of OsRTNLB4, OsRTNLB8, OsRTNLB9, and OsRTNLB20, word clouds were generated based on cis-regulatory element (CRE) annotation (Figure S3B). For OsRTNLB4, CREs associated with light, abscisic acid, and MeJA signaling were predominantly enriched. OsRTNLB8 showed a strong association with CREs linked to hypoxia and abscisic acid responses. A broader range of regulatory elements, including stress responsiveness and MeJA, was observed for OsRTNLB9. In contrast, OsRTNLB20 exhibited enrichment for CREs related to drought and salicylic acid signaling.
Modeling and phenotypic detection
This research simultaneously conducted various experiments on Jiahe102 and Nipponbare. Compared to Nipponbare, Jiahe102 exhibits higher yield and stronger heat resistance.
This study used 5 types and 2 gradients of stress, totaling 11 groups each breed including Control (Table 2). The Control groups of both varieties grew well under suitable conditions, with Jiahe102 showing unaffected growth rates in cold or heat stress. The aboveground lengths of Nipponbare in various stress groups were lower than the control group. Cd ions, Na ions, and simulated drought stress greatly inhibited the growth of rice, with Cd ions having the most significant impact (Figure S4). Measurements of the aboveground height, the length of the first and second tillers of seedlings revealed that the growth inhibition in Jiahe102 is correlated with Cd ion concentration and simulated drought stress intensity. It’s noteworthy that Cd ions seem to have a smaller impact on the second tiller (Fig. 4).
Growth records for Jiahe102 under various treatments. The top-left panel displays the overall growth condition of control plants. Bar graphs represent the lengths (cm) of the whole shoot, first tiller, and second tiller under different stress treatments (Cd_1, Cd_10, DR_2, DR_5). Measurements are based on three randomly selected plants, with statistical significance determined by a T-test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001). The lower panels show photographs of plants under control conditions and stress treatments (Cd_1, Cd_10, DR_2, DR_5), with a 10 cm scale bar for reference. Note: DR indicates mannitol used to simulate drought stress.
Generally, plants subjected to external stress exhibit the classical phenotype of elevated MDA (malondialdehyde), H2O2 (hydrogen peroxide), T-AOC (total antioxidant capacity), and simultaneously decreased chlorophyll content due to oxidative stress. We tested the leaves of Jiahe102 for MDA, H2O2, T-AOC, and chlorophyll content (Fig. 5A, Table S1). Usually, in variety stresses, MDA increased, and chlorophyll decreased. H2O2 and T-AOC often simultaneously increased or decreased under different stresses. H2O2 levels did not increase under extremely low temperature (4 °C), moderate-high temperature (37 °C), and salt stress, accompanied by a slight decrease in T-AOC, possibly due to the normal operation of the antioxidant system consuming reactive oxygen species. We found that under Cd ion and simulated drought stress, MDA, H2O2, and T-AOC significantly increased, while chlorophyll significantly decreased (Fig. 5B-C). Additionally, the detection results for Cd ion and simulated drought stress were same between Jiahe102 and Nipponbare (Figure S5A, Table S2), indicating significant oxidative stress in rice. Furthermore, under cold, heat, and salt stress, Nipponbare exhibited an increase in chlorophyll content. Considering its reduced growth rate, we believe that the abundant chlorophyll under stress may be detrimental to growth.
(A) Heatmap of biochemical test results under different stress treatments. The heatmap displays log-normalized values for malondialdehyde (MDA), hydrogen peroxide (H₂O₂), total antioxidant capacity (T-AOC), and chlorophyll content across treatments. The control group is normalized to a value of 1, with varying colors indicating relative changes in biochemical levels under different stress conditions. (B) Biochemical responses under Cd_10 treatment. Bar plots represent the relative content of MDA, H₂O₂, T-AOC, and chlorophyll under Cd_10 stress compared to the control group. Statistical significance is indicated with T-test P-values displayed above each group. (C) Biochemical responses under DR_5 treatment. Similar to panel B, bar plots show the relative biochemical levels under DR_5 stress, with P-values displayed above the bars. Statistical analysis was performed using a T-test. Note: DR indicates mannitol used to simulate drought stress.
Tissue-level analysis has been performed by slicing leaves under various stresses (Jiahe102). Only Cd ion and simulated drought stress caused significant changes in leaf tissue (Figs. 6 and 7). In simulated drought stress, the lignification of rice seedling leaves increased, accompanied by an enlargement of mechanical tissue (Fig. 6B). Ventilation tissue in rice seedlings almost disappeared under Cd ion stress (Fig. 7B), suggesting that the normal growth of seedlings has been hindered, leading to slow development, or abnormal programmed cell death might have occurred. The mechanisms behind these abnormal phenotypes require further exploration. The leaf aeration tissue disappears in rice leaves under Cd ion stress (Fig. 7), possibly due to the collective decrease in RTN protein expression and suppression of programmed cell death in thin-walled tissues23. However, further research is needed.
(A) Microscopic observations of rice first tiller cross-sections under control and DR_5 conditions. The sections were stained with saffranine solid green, highlighting key tissues: VB (vascular bundle), Mec (mechanical tissue), Aer (aerenchyma), and Mes (mesophyll tissue). Images on the top row show lower magnification (scale bar = 100 μm), while the bottom row presents higher magnification (scale bar = 40 μm). (B) Quantitative analysis of tissue proportions in the main vein. Bar plots compare the proportions of lignified tissue, vascular bundles, mechanical tissue, aerenchyma, and mesophyll tissue between control and DR_5 treated plants. Statistical significance was evaluated using a T-test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns = not significant). Note: DR indicates mannitol used to simulate drought stress.
(A) Microscopic observations of rice first tiller cross-sections under control and Cd_10 conditions. The sections highlight key tissues using saffranine solid green staining: VB (vascular bundle), Mec (mechanical tissue), Aer (aerenchyma), and Mes (mesophyll tissue). The scale bar represents 100 μm for both panels. (B) Quantitative analysis of tissue proportions in the main vein. Bar plots compare the proportions of lignified tissue, vascular bundles, mechanical tissue, aerenchyma, and mesophyll tissue between control and Cd_10-treated plants. Statistical analysis was conducted using a T-test (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001; ns = not significant).
Although the stress analysis presented in this study aligns with findings from prior studies, its novelty lies in the comparative approach between Jiahe102 and Nipponbare under identical stress conditions. By systematically examining five types of stress with two gradient levels, the study provides a unique dataset that integrates physiological, biochemical, and tissue-level analyses. These findings contribute to the broader understanding of stress tolerance mechanisms in rice and provide a basis for future studies to explore the genetic and molecular underpinnings of these traits.
Expression level detection
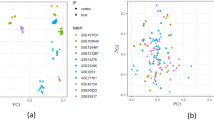
Taking leaf tissues of Jiahe102, we measured the expression levels of 20 OsRTNLBs under various stresses (Fig. 8, Table S3). Due to very low expression levels in leaves, OsRTNLB13 and OsRTNLB14 were excluded. We observed an increase in expression levels of OsRTNLBs under simulated drought stress. The expression levels under the other four stresses showed a decrease, with the most significant lower expression observed under heat stress. Among them, OsRTNLB8, OsRTNLB17, and OsRTNLB19 showed extreme changes in expression levels under various stresses. OsRTNLB2 exhibited lower expression in expression levels under all stresses. OsRTNLB1, OsRTNLB4, OsRTNLB6, OsRTNLB16, and OsRTNLB18 had relatively high average expression levels under various stresses. The expression levels of OsRTNLB20 showed relatively small variations under all stresses.
Changes in OsRTNLBs expression levels under different stress conditions. The heatmap illustrates the log(2−△△Ct) values for OsRTNLB gene expression in JH102 rice under various stresses, including heat, cold, Cd²⁺, Na⁺, and mannitol. Each row represents an OsRTNLB gene, and the color scale indicates the relative expression levels, with red representing upregulation and blue representing downregulation compared to the control group. Genes are hierarchically clustered based on their expression patterns across stress conditions.
Additionally, we collected OsRTNLBs expression data for Nipponbare (Figure S5B, Table S4). We found that the overall trend of OsRTNLBs expression is same as Jiahe102, but there were more OsRTNLBs with increased expression levels under stress in Nipponbare. Compared to Jiahe102, OsRTNLB1, OsRTNLB6, and OsRTNLB16 showed higher expression levels under various stresses in Nipponbare. Interestingly, OsRTNLB20 in Nipponbare exhibited a significant upregulation under various stresses (Figure S5C).
Function speculates
Endoplasmic reticulum (ER) stress and oxidative stress are intricately intertwined. Indicators such as H2O2, MDA, and T-AOC reflect ER stress occurrence24,25. In plant cells, chlorophyll content is also affected25. Under various abiotic stress conditions, MDA levels in rice leaves are consistently elevated (Fig. 5A, Figure S5A), indicating that these stressors disrupt redox balance. ER stress occurs under drought and Cd ion stress (Fig. 5B-C). Some RTN family members participate in ER stress-induced selective autophagy7, which is crucial for plant resistance. ZmRTNLB2 promotes autophagy during ER stress7. Members in Clade 1, including OsRTNLB2, OsRTNLB4, OsRTNLB6, and OsRTNLB16, may have similar functions (Fig. 1A). The expression of these OsRTNLBs in Nipponbare is upregulated under drought and Cd ion stress (Figure S5B). However, in Jiahe102, two interesting phenomena were observed: (i) Lower expression of OsRTNLB2 under all stress conditions. (ii) Lower expression of Clade-1-OsRTNLBs under Cd ion stress (Fig. 8). These observations will be the focus of our subsequent research. OsRTNLB4, the closest rice RTN to ZmRTNLB2 in the evolutionary tree, shows smaller expression changes between the two varieties under drought and Cd ion stress compared to other Clade-1-OsRTNLBs (Fig. 8, Figure S5B). With 31 transcription factor families regulating OsRTNLB4 (Table 1), it is the most regulated among OsRTNLBs, with targets evenly distributed across families (Fig. 3B). Therefore, OsRTNLB4 is likely a key molecule in Clade 1. Additionally, OsRTNLB1 in Clade 3 has a similar expression pattern under various stresses to OsRTNLB6 and OsRTNLB16, suggesting its involvement in RTN protein-mediated selective autophagy (Figs. 1A and 8, Figure S5B).
In Z. mays, ZmRTNLB16 enhances drought resistance by promoting the localization of V-H+-ATPase and its E3 subunit on vacuolar membranes17. OsRTNLB8, the rice homolog of ZmRTNLB16 (Fig. 1A), shows significantly increased expression under drought conditions (Fig. 8, Figure S5B). OsRTNLB8 target transcription factors, including ERF, Dof, MYB, and C2H226, are associated with drought response (Fig. 3B). OsRTNLB8 may play a crucial role in assembling the vacuolar ATP pump, promoting rice resistance to drought. Furthermore, the high expression of OsRTNLB8 in Nipponbare under cold stress, despite Nipponbare’s relatively poor cold tolerance compared to Jiahe102 (Figure S5B), raises questions about a possible correlation between these events.
Among the 20 members of the RTN family, 5 members (OsRTNLB7, OsRTNLB9, OsRTNLB18-20) unique distinctive motifs, suggesting that they may have unique functions (Fig. 1B). The functions of AtRTNLB17-18 and AtRTNLB21 are not yet clear, and the rice RTN proteins most homologous to them are OsRTNLB7 and OsRTNLB19 (Fig. 1A). OsRTNLB9, OsRTNLB18, and OsRTNLB20 are closely related to A. thaliana AtRTNLB19 and AtRTNLB20 (Fig. 1A). AtRTNLB20 and AtRTNLB19 exhibit 3Beta_HSD domains and enzymatic activities related to sterol biosynthesis, displaying a distinct punctate expression pattern in cells compared to other RTN proteins6. We found that OsRTNLB9 and OsRTNLB20 also possess predicted 3Beta_HSD domains (Figure S6). Brassinosteroids (BRs) are sterol compounds that play a crucial role in plant growth and development27. BRs are involved in various responses to abiotic stresses, but the expression changes of OsRTNLB9 and OsRTNLB20 differ under different stress conditions (Fig. 8). Comparing Japonica Group (Nipponbare) and Indica Group (Jiahe102), the expression of OsRTNLB20 in Indica Group under various stresses is the same as that of OsRTNLB9 in Japonica Group but different from that of OsRTNLB20 in Japonica Group (Figure S5B). BRs can promote the synthesis of lignin in plant cells28. Under simulated drought stress: (i) The lignification of leaves increases in Japonica Group29; (ii) The degree of leaf lignification in Indica Group increases, and mechanical tissues enlarge (Fig. 6); (iii) OsRTNLB9 in Japonica Group and OsRTNLB20 in Indica Group are upregulated only under simulated drought stress (Fig. 8, Figure S5B), along with the observed lignification increased. The TCP transcription factor family plays a crucial regulatory role in BR biosynthesis in A. thaliana30. CRE analysis indicates that OsRTNLB20 is not regulated by TCPs, while OsRTNLB9 may be one of the targets of TCP regulation (Fig. 3B). In summary, OsRTNLB9 may play a crucial role in the regulation of BR biosynthesis and influence lignin synthesis in rice, while OsRTNLB20 may have an auxiliary role in Japonica Group and provide compensation in Indica Group.
Based on existing literature and our data, OsRTNLB4, OsRTNLB8, and OsRTNLB9 are marked as key players in the stress response of rice.
Discussion
The OsRTNLB gene family in Oryza sativa has been updated and systematically analyzed in this study, encompassing evolutionary relationships, protein motifs, collinearity, and cis-regulatory elements (CREs). A scientifically rigorous framework for understanding the OsRTNLB family has been constructed by integrating comprehensive genomic data with insights from existing research. Expression analyses under various abiotic stresses, combined with phenotypic observations, have provided a detailed perspective on the functional roles of individual OsRTNLB members. This analysis prioritized OsRTNLB members closely homologous to functionally characterized RTNs from other species, identifying three key genes (OsRTNLB4, OsRTNLB8, and OsRTNLB9) for further investigation.
The upstream regulation of OsRTNLB4 does not appear to directly involve elements associated with abiotic stress, suggesting that its role in stress responses may be indirect (Figure S3B). Studies have shown that the autophagy marker protein ATG8 can be regulated by transcription factors that respond to environmental stressors31. Moreover, ZmRTNLB2, a maize homolog closely related to OsRTNLB4, has been reported to interact with ATG87. RTN proteins act as autophagy receptors, facilitating the recruitment of ATG8 to initiate autophagy7. Additionally, the endoplasmic reticulum (ER) is widely recognized as playing a central role in cellular responses to stress conditions, and the association between RTN proteins and ER stress has been well established2,32. These findings imply that the involvement of OsRTNLB4 in stress responses may be primarily triggered by ER stress rather than direct regulation by stress-specific transcription factors. Further investigation is required to clarify the mechanisms underlying this regulatory relationship and the specific role of OsRTNLB4 in stress adaptation.
Interestingly, upstream regulatory elements related to light were widely identified among OsRTNLBs, suggesting a potential role in light-responsive pathways (Fig. 3C, Figure S2B, Figure S3B). For instance, Arabidopsis thaliana AtRTNLB4 forms a basket-like structure around chloroplasts under fluorescence microscopy, indicating its involvement in ER-chloroplast trafficking20,33,34. Similarly, MpRTNLB1 in Marchantia polymorpha has been shown to interact with photoreceptors, reshaping the ER and promoting chloroplast migration3. These findings highlight the potential participation of RTNLBs in signaling and metabolic networks within plant cells, particularly in adapting to changing light intensities—a process essential for photosynthetic efficiency35.
Compared to other eukaryotes, the RTN family in plants exhibits greater diversity and complexity, particularly in higher plants. This can be attributed to several factors: (1) the partitioning of the endoplasmic reticulum (ER) into diverse subdomains in plants36, with RTNLBs localized in structures such as chloroplasts3, vacuoles37, and plasmodesmata38; (2) the specialized functions of plant RTN proteins, often requiring synergistic or antagonistic interactions among multiple RTNLBs within the same biological processes6,38,39,40; and (3) the unique roles plant RTN proteins may play, extending beyond the ER41. Plant RTN proteins exhibit a broader range of functions; these distinctions highlight the necessity of gene family analyses to optimize research resource utilization by identifying key targets for further study.
Bioinformatics tools and computational approaches are increasingly recognized as effective methods for analyzing biological data and synthesizing insights from existing research42. Gene family studies, such as this one, demonstrate how these methods provide a data-driven framework for bridging established knowledge with future research. By evaluating protein homology, phylogenetic relationships, and transcriptional regulation, this approach facilitates the identification of functionally relevant homologous proteins and their applicability in rice. The systematic characterization of the OsRTNLB family not only completes but also enriches the understanding of this gene family, providing a foundation for future functional and applied research.
Despite these advancements, certain limitations remain. The scarcity of studies on plant RTN proteins has restricted the ability to predict rice RTN protein functions solely through phylogenetic grouping. Moreover, the specific mechanisms and functional roles of the identified OsRTNLB proteins require experimental validation, which is expected to be addressed in future investigations.
Conclusion
This study has provided a prediction of the possible functions and molecular mechanisms for OsRTNLBs. OsRTNLB4, OsRTNLB8, OsRTNLB9 has been marked as key protein in abiotic stress response. The research on the rice RTN gene family has significant importance, laying the foundation for functional verification and potential applications in future genetic engineering projects.
Method
Collection of data resources
The genome-wide information of Arabidopsis thaliana, Oryza sativa, and Zea mays genome data was downloaded from the ensemble database (http://plants.ensembl.org/index.html).
Genome-wide identification of OsRTNLBs
To perform a comprehensive identification of the OsRTNLBs in O. sativa, the following steps were then taken: (1) The Hidden Markov Model (HMM) profile of the Reticulon domain (PF02453) was downloaded from the protein family database (Pfam: http://pfam.xfam.org/) with an E-value of 1*10− 5. (2) The amino acid sequences of A. thaliana, O. sativa, and Z. mays RTN (AtRTNLB/OsRTNLB/ZmRTNLB) proteins were retrieved from the Rice Gene Index (RGI: https://riceome.hzau.edu.cn/) and the Phytozome (https://phytozome-next.jgi.doe.gov/) respectively, which were then used to map the phylogenetic tree. (3) The confirmed OsRTNLBs genes were renamed according to their positions on O. sativa chromosomes.
Multiple sequence alignment and phylogenetic tree
To observe the evolutionary relationship of the RTN gene family, we constructed a phylogenetic tree with AtRTNLB, OsRTNLB, and ZmRTNLB protein sequences. Multiple sequence alignment of the protein sequences was conducted by MUSCLE43. For the alignment, the BLOSUM62 substitution matrix was used, with an open gap penalty = 2 and extension gap penalty = 0.2 set. The maximum likelihood (ML) method phylogenetic tree based on the protein sequence alignment was constructed according to the Whelan and Goldman substitution model (WAG + G + F) by MEGA 11 software44. Relative branch support was evaluated by 1000 bootstrap replicates, the branch lengths were calculated by pairwise comparisons of genetic distances, and missing data were treated by pairwise deletions of gaps. For better visualization, the phylogenetic tree was visualized using the online tool iTOL v6 (https://itol.embl.de/). The tertiary structure of OsRTNLBs have been obtained from SWISS-MODEL (https://swissmodel.expasy.org/).
Conserved protein motifs
To identify conserved motifs in the OsRTNLBs proteins, we utilized the Multiple Em for Motif Elicitation (MEME v5.5.0: https://meme-suite.org/meme/) online program. We set the ideal breadth of each motif to range from 6 to 50 residues. The number of repetitions was set arbitrarily, and the maximum number of motifs considered was limited to 6.
Chromosomal distribution and gene duplication analysis
All OsRTNLBs were anchored to their corresponding chromosomes using TBtools (v1.100)45. To demonstrate the synteny of orthologous RTN genes obtained from Oryza sativa and other selected species, we analyzed gene duplication events by using the Multicollinearity Scanning Toolkit (MCScanX)46.
CREs (cis-regulatory elements) analysis
The upstream 2000 bp sequence of each OsRTNLBs gene was retrieved by TBtools software45 according to the genomic full length DNA sequences of Oryza sativa. Then, the Plant Care database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) was used to predict putative promoter regions of RTN genes in Oryza sativa CREs. Finally, the results were visualized using TBtools.
A regulation prediction tool, the Plant Transcriptional Regulatory Map (http://plantregmap.gao-lab.org/binding_site_prediction_result.php) had been used to infer potential regulatory interactions of Transcription Factors (TFs) in the upstream (2000-bp) regions of OsRTNLBs with threshold (P-value < 1e− 4). The predicted TFs were visualized in a network using the Cytoscape software (v3.9.1)47. Furthermore, the wordcloud was generated by the wordcloud2 package in R.
Plant materials and stress treatments
Jiahe 102 (provided by State Key Laboratory of Hybrid Rice, Wuhan, China) and Nipponbare, had been used as a model rice variety.
The seed coat had been removed, sterilized, and soaked in a dark room at 37 °C for 24 h. The seeds were allowed to germinate at 25 °C in the dark for 48 h and then transferred to a culture bottle for cultivation at 25 °C for 14 days. The control group had been directly sampled. The growth environments of seedlings in the experimental group were changed 7 or 1 day before sampling. Photographic record.
Physiological measurements
Samples were collected to quantify the Malondialdehyde (MDA) levels, Total antioxidant capacity (T-AOC), Hydrogen peroxide contents and Chlorophyll contented by using Malondialdehyde (MDA) Content Assay Kit (Solarbio: BC0020), Total Antioxidant Capacity (T-AOC) Assay Kit (Solarbio: BC1315), Hydrogen peroxide (H2O2) Content Assay Kit (Solarbio: BC3595), and Chlorophyll Assay Kit (Solarbio: BC0990), respectively, according to the manufacturer’s instructions.
Tissue section
The middle part of the first tiller of the leaves was taken and photographed with a scanning electron microscope (Hitachi SU8100, Japan) after the steps of FAA fixation, paraffin embedding, slicing, and dyeing.
RNA extraction and quantitative real-time PCR
Whole shoot tissues (100 mg) of rice from different treatment groups were weighed and placed in a grinding tube containing steel beads. The grinding tubes were immersed in liquid nitrogen for 10–20 min. Finally, the samples were freeze-ground at − 20 °C for 120 s and returned to liquid nitrogen for storage. RNA extraction had been performed using the FastPure Universal Plant Total RNA Isolation Kit (Vazyme, Nanjing, China: RC411-01).
The cDNA had been synthesized using the Revert Aid First Strand cDNA Synthesis Kit (Thermo Scientific, Watham, MA, USA: K1622). qRT-PCR analysis were performed using a LightCycler 96 (Roche, Basel, Switzerland). eEF1 was used as the reference gene48. Gene sequences were searched using Phytozome (https://phytozome-next.jgi.doe.gov/), and qRT-PCR primer sequences were designed using the primer blast tool of NCBI (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The primers used in this study were listed in Table 2. See Data S2 for the MIQE checklist49.
Relative expression level of target genes were calculated based on the 2–ΔΔCt method for normalization50. The normalized qRT-PCR data was analyzed using a t-test to determine statistically significant differences in gene expression between the control and experimental groups51. Statistical significance was set at P < 0.05.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Oertle, T. & Schwab, M. E. Nogo and its partners. Trends Cell. Biol. 13, 187–194. https://doi.org/10.1016/s0962-8924(03)00035-7 (2003).
D’Eletto, M. et al. Reticulon homology domain-containing proteins and ER-Phagy. Front. Cell. Dev. Biol. 8, 90. https://doi.org/10.3389/fcell.2020.00090 (2020).
Ishikawa, K. et al. The Endoplasmic reticulum membrane-bending protein RETICULON facilitates Chloroplast relocation movement in Marchantia polymorpha. Plant. J. 111, 205–216. https://doi.org/10.1111/tpj.15787 (2022).
Nziengui, H. et al. Reticulon-like proteins in Arabidopsis Thaliana: structural organization and ER localization. FEBS Lett. 581, 3356–3362. https://doi.org/10.1016/j.febslet.2007.06.032 (2007).
Chen, J. et al. The Endoplasmic reticulum: a social network in plant cells. J. Integr. Plant. Biol. 54, 840–850. https://doi.org/10.1111/j.1744-7909.2012.01176.x (2012).
Kriechbaumer, V. et al. The odd one out: Arabidopsis reticulon 20 does not Bend ER membranes but has a role in lipid regulation. Sci. Rep. 8, 2310. https://doi.org/10.1038/s41598-018-20840-0 (2018).
Zhang, X. et al. Reticulon proteins modulate autophagy of the Endoplasmic reticulum in maize endosperm. Elife 9. https://doi.org/10.7554/eLife.51918 (2020).
Sarma, B. et al. Abiotic stress in rice: Visiting the physiological response and its tolerance mechanisms. Plants (Basel) 12. https://doi.org/10.3390/plants12233948 (2023).
Oladosu, Y. et al. Drought resistance in rice from conventional to molecular breeding: A review. Int. J. Mol. Sci. 20 https://doi.org/10.3390/ijms20143519 (2019).
Jian, L. et al. Mutation of OsMYB60 reduces rice resilience to drought stress by attenuating cuticular wax biosynthesis. Plant. J. 112, 339–351. https://doi.org/10.1111/tpj.15947 (2022).
Ferguson, J. N. et al. Rapid temperature responses of photosystem II efficiency forecast genotypic variation in rice vegetative heat tolerance. Plant. J. 104, 839–855. https://doi.org/10.1111/tpj.14956 (2020).
Lou, Q. et al. Cold-adaptive evolution at the reproductive stage in Geng/japonica subspecies reveals the role of OsMAPK3 and OsLEA9. Plant. J. 111, 1032–1051. https://doi.org/10.1111/tpj.15870 (2022).
Jing, P. et al. OsClo5 functions as a transcriptional co-repressor by interacting with OsDi19-5 to negatively affect salt stress tolerance in rice seedlings. Plant. J. 105, 800–815. https://doi.org/10.1111/tpj.15074 (2021).
Li, L. et al. CF1 reduces grain-cadmium levels in rice (Oryza sativa). Plant. J. 110, 1305–1318. https://doi.org/10.1111/tpj.15736 (2022).
Li, B. et al. Ufmylation reconciles salt stress-induced unfolded protein responses via ER-phagy in Arabidopsis. Proc. Natl. Acad. Sci. U S A. 120, e2208351120. https://doi.org/10.1073/pnas.2208351120 (2023).
Huang, F. C. et al. Arabidopsis RETICULON-LIKE3 (RTNLB3) and RTNLB8 participate in Agrobacterium-Mediated plant transformation. Int. J. Mol. Sci. 19 https://doi.org/10.3390/ijms19020638 (2018).
Tian, T. et al. Genome assembly and genetic dissection of a prominent drought-resistant maize germplasm. Nat. Genet. 55, 496–506. https://doi.org/10.1038/s41588-023-01297-y (2023).
Brooks, R. L. et al. Curvature sensing amphipathic helix in the C-terminus of RTNLB13 is conserved in all Endoplasmic reticulum shaping reticulons in Arabidopsis Thaliana. Sci. Rep. 11, 6326. https://doi.org/10.1038/s41598-021-85866-3 (2021).
Breeze, E. et al. A C-terminal amphipathic helix is necessary for the in vivo tubule-shaping function of a plant reticulon. Proc. Natl. Acad. Sci. U S A. 113, 10902–10907. https://doi.org/10.1073/pnas.1605434113 (2016).
Nziengui, H. & Schoefs, B. Functions of reticulons in plants: what we can learn from animals and yeasts. Cell. Mol. Life Sci. 66, 584–595. https://doi.org/10.1007/s00018-008-8373-y (2009).
Marand, A. P. et al. A cis-regulatory atlas in maize at single-cell resolution. Cell 184 https://doi.org/10.1016/j.cell.2021.04.014 (2021).
Zhao, J. et al. Genome-Wide identification and expression profiling of the TCP family genes in Spike and grain development of wheat (Triticum aestivum L). Front. Plant. Sci. 9, 1282. https://doi.org/10.3389/fpls.2018.01282 (2018).
Joshi, R. & Kumar, P. Lysigenous aerenchyma formation involves non-apoptotic programmed cell death in rice (Oryza sativa L.) roots. Physiol. Mol. Biol. Plants. 18, 1–9. https://doi.org/10.1007/s12298-011-0093-3 (2012).
Wang, H. et al. Characterization of bip genes from pepper (Capsicum annuum L.) and the role of CaBiP1 in response to Endoplasmic reticulum and multiple abiotic stresses. Front. Plant. Sci. 8, 1122. https://doi.org/10.3389/fpls.2017.01122 (2017).
Wang, B. et al. ZmNF-YB16 overexpression improves drought resistance and yield by enhancing photosynthesis and the antioxidant capacity of maize plants. Front. Plant. Sci. 9, 709. https://doi.org/10.3389/fpls.2018.00709 (2018).
Hrmova, M. & Hussain, S. S. Plant transcription factors involved in drought and associated stresses. Int. J. Mol. Sci. 22. https://doi.org/10.3390/ijms22115662 (2021).
Sharma, A. et al. Brassinosteroids and metalloids: regulation of plant biology. J. Hazard. Mater. 424, 127518. https://doi.org/10.1016/j.jhazmat.2021.127518 (2022).
Wang, X. et al. Comparative proteomics analysis reveals that lignin biosynthesis contributes to brassinosteroid-mediated response to phytophthora Sojae in soybeans. J. Agric. Food Chem. 68, 5496–5506. https://doi.org/10.1021/acs.jafc.0c00848 (2020).
Bang, S. W. et al. Overexpression of OsTF1L, a rice HD-Zip transcription factor, promotes lignin biosynthesis and stomatal closure that improves drought tolerance. Plant. Biotechnol. J. 17, 118–131. https://doi.org/10.1111/pbi.12951 (2019).
Spears, B. J. et al. Class I TCP transcription factor AtTCP8 modulates key brassinosteroid-responsive genes. Plant. Physiol. 190, 1457–1473. https://doi.org/10.1093/plphys/kiac332 (2022).
Wang, P. et al. Identification of transcription factors that regulate ATG8 expression and autophagy in Arabidopsis. Autophagy 16, 123–139. https://doi.org/10.1080/15548627.2019.1598753 (2020).
Zhu, J. K. Abiotic stress signaling and responses in plants. Cell 167, 313–324. https://doi.org/10.1016/j.cell.2016.08.029 (2016).
Hanson, M. R. & Köhler, R. H. GFP imaging: methodology and application to investigate cellular compartmentation in plants. J. Exp. Bot. 52, 529–539. https://doi.org/10.1093/jexbot/52.356.529 (2001).
Holthuis, J. C. M. & Levine, T. P. Lipid traffic: floppy drives and a superhighway. Nat. Rev. Mol. Cell. Biol. 6, 209–220. https://doi.org/10.1038/nrm1591 (2005).
Wada, M. Chloroplast and nuclear photorelocation movements. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 92, 387–411 (2016).
Stefano, G. & Brandizzi, F. Advances in plant ER architecture and dynamics. Plant. Physiol. 176, 178–186. https://doi.org/10.1104/pp.17.01261 (2018).
Tian, T. et al. Genome assembly and genetic dissection of a prominent drought-resistant maize germplasm. Nat. Genet. https://doi.org/10.1038/s41588-023-01297-y (2023).
Tilsner, J. & Kriechbaumer, V. Reticulons 3 and 6 interact with viral movement proteins. Mol. Plant. Pathol. 23, 1807–1814. https://doi.org/10.1111/mpp.13261 (2022).
Zhang, X. et al. Reticulon proteins modulate autophagy of the Endoplasmic reticulum in maize endosperm. Elife 9, e51918. https://doi.org/10.7554/eLife.51918 (2020).
Huang, F. C. et al. Arabidopsis RAB8A, RAB8B and RAB8D proteins interact with several RTNLB proteins and are involved in the Agrobacterium tumefaciens infection process. Plant. Cell. Physiol. 62, 1572–1588. https://doi.org/10.1093/pcp/pcab112 (2021).
Staehelin, L. A. The plant ER: a dynamic organelle composed of a large number of discrete functional domains. Plant. J. 11, 1151–1165. https://doi.org/10.1046/j.1365-313x.1997.11061151.x (1997).
Cantó-Pastor, A. et al. Arabidopsis bioinformatics: tools and strategies. Plant. J. 108, 1585–1596. https://doi.org/10.1111/tpj.15547 (2021).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 5, 113. https://doi.org/10.1186/1471-2105-5-113 (2004).
Kumar, S. et al. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant. 13, 1194–1202. https://doi.org/10.1016/j.molp.2020.06.009 (2020).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49. https://doi.org/10.1093/nar/gkr1293 (2012).
Kohl, M. et al. Cytoscape: software for visualization and analysis of biological networks. Methods Mol. Biol. 696, 291–303. https://doi.org/10.1007/978-1-60761-987-1_18 (2011).
Ambavaram, M. M. R. & Pereira, A. Setting up reverse transcription quantitative-PCR experiments. Methods Mol. Biol. 678, 45–54. https://doi.org/10.1007/978-1-60761-682-5_4 (2011).
Bustin, S. A. et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622. https://doi.org/10.1373/clinchem.2008.112797 (2009).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta delta C(T)) method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Wilson, E. B. & Worcester, J. Note on the t-Test. Proc. Natl. Acad. Sci. U S A. 28, 297–301. https://doi.org/10.1073/pnas.28.7.297 (1942).
Acknowledgements
The authors acknowledge the assistance of Jun Hu from College of Life Science, Wuhan University, China, who offered great help in providing rice seeds.
Funding
Supported by the Open Research Fund of State Key Laboratory of Hybrid Rice (Wuhan University) (KF202202); the Project for Technology Innovation of Hubei Province (2024BBA005).
Author information
Authors and Affiliations
Contributions
Conceptualization, L.T.; methodology, G.C.; software, G.C.; validation, L.T. and G.C. formal analysis, G.C.; investigation, G.C.; resources, L.T.; data curation, G.C.; writing—original draft preparation, G.C.; writing—review and editing, L.T., X.G., B.X. and Y.Y.; visualization, G.C.; supervision, L.T.; project administration, L.T.; funding acquisition, L.T. and G.C. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cao, GH., Guo, XL., Yang, YJ. et al. RTN family and its response to abiotic stress in Oryza sativa. Sci Rep 15, 13134 (2025). https://doi.org/10.1038/s41598-025-97261-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97261-3