Abstract
Staphylococcus aureus subsp. anaerobius (SAAN) causes abscesses in small ruminants, known as Morel’s Disease. This study describes the presence of SAAN for the first time in Germany and Austria and examines the phylogenetic relationship among these isolates and previously described European and Sudanese ones. A total of 35 sheep and 10 goat isolates from 12 herds in Germany were available for analysis. SAAN isolates from four Polish goats and three Austrian sheep from different herds were included. Genome comparisons and phylogenetic analyses were conducted using core genome multilocus sequence typing. The comparison of the 52 SAAN core genomes revealed a close phylogenetic relationship among most German isolates (n = 38), with allelic differences ≤ 6 in two clusters associated with ST4581. In contrast, distinct clusters of the same ST included the four Polish goat isolates and two ovine isolates from Austria, respectively. A fifth cluster of ST3756 strains was identified on three German farms (six sheep, one goat) and an Austrian sheep. Tight phylogenetic relationships were observed irrespective of the host species. All isolates shared a common set of virulence genes and few known antimicrobial resistance determinants. The introduction of SAAN into herds is mostly unknown, but purchases appear to play a critical role.
Similar content being viewed by others
Introduction
The bacterium Staphylococcus (S.) aureus subsp. anaerobius (SAAN) was first described in France by Morel in 19111. Bacteriological identification of SAAN is delicate; it typically does not sprout under aerobic conditions but grows slowly in microaerophilic or anaerobic atmospheres. SAAN causes abscesses, primarily in close association with superficial lymph nodes in sheep and goats2,3. This disease is known as Morel’s Disease (MD) or abscess disease and has been reported in small ruminant populations in many African and Middle Eastern countries4,5,6,7,8,9. Since 2000, reports of MD in small ruminants have increased throughout Europe, including countries such as Denmark2, Croatia10, Poland3, Switzerland11, and the Czech Republic12. In Spain, SAAN was even identified as the most frequent causative agent of abscesses in small ruminants13. To date, SAAN has not been reported in sheep and goats from Germany and Austria14.
MD is endemic and contagious and is mainly encountered in young animals within the first 4–10 months of life15. However, adult animals are also severely affected, especially when the pathogen enters a herd for the first time3. Transmission occurs through direct or indirect contact with pus from spontaneously drained abscesses. International trade with sheep and goats is a significant risk factor for introducing SAAN into naïve herds2,3,12. MD causes financial losses due to rejection of shipments or condemnation of sheep carcasses during meat inspection in slaughterhouses. Nevertheless, the exact economic losses due to MD have not yet been computed16. The zoonotic potential of SAAN has been suspected, but final proof thereof is still pending17.
Recently, a detailed study of the SAAN genome revealed a highly conserved clone, tracing its lineage back to a Staphylococcus (S.) aureus progenitor that existed approximately 1,000 years ago18. The biochemical identification of SAAN is very limited due to lack of growth under aerobic conditions. Additionally, commercially available kits, such as API 20 A, API rapid ID 32 A, or VITEK ANI card (all BioMerieux, Nuertingen, Germany) do not contain anaerobic staphylococci in their databases17. In biochemical identification algorithms, the presence of catalase is listed as a characteristic of S. aureus19. This virulence factor degrades hydrogen peroxide to water and oxygen. Produced in vivo by bacteria, the enzyme prevents chemical damage, for example, after phagocytosis by neutrophils. However, SAAN is a catalase negative organism, and its virulence potential should not be underestimated. It is assumed that this microaerophilic metabolism is because of the loss of catalase function and other oxidoreductase genes, resulting in higher susceptibility to oxidative free radicals18. In contrast to the sometimes misleading results of microbiological or biochemical approaches, a reliable identification at species level is possible using the molecular approach based on the 16SrRNA gene sequence analysis17 or by MALDI-TOF analysis. A further differentiation below species level can be realized by different molecular typing tools. Genetic analyses of SAAN strains from several countries using different methods such as pulsed-field gel electrophoresis (PFGE), multilocus sequence typing (MLST), and random amplified polymorphic DNA (RAPD) have revealed very high homogeneity among these isolates14,20. Nevertheless, two distinct lineages of SAAN containing African and European isolates exist18.
Interest has been steadily growing in the remarkably high number of toxins and other virulence factors produced by S. aureus and their impact on disease, as recently reviewed by Cheung and colleagues21. For SAAN, the vwb gene, encoded on one of the two mobile genetic elements (MGEs) in the accessory genome, is considered highly relevant for forming classical abscesses in MD18. Moreover, antimicrobial resistance (AMR) is very prominent in many S. aureus strains, especially those that pose a major health threat in humans and animals, like methicillin-resistant S. aureus, and may be conferred by gene modifications or resistance genes that are encoded chromosomally or on MGE22. In contrast, AMR determinants in SAAN are still largely unknown.
Due to the emerging numbers of SAAN isolation in small ruminant herds in Germany, a study was performed to describe the presence of MD, to characterize them, and to examine the phylogenetic relationship among SAAN isolates from different German regions. In addition, samples from Poland and Austria as well as genomic sequences from public databases were included in the genetic analysis to investigate a possible connection. To obtain a complete picture of the Central European SAAN isolates, virulence factors and AMR determinants were assessed. Furthermore, the zoonotic potential of SAAN was discussed due to carcass condemnations caused by abscesses in a significant number of lambs within the herds presented here.
Material and methods
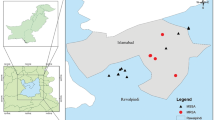
Samples were collected from sheep and goats from 17 small ruminant farms, with 12 herds located in Germany, three sheep herds in Austria, and two dairy goat herds in Poland (Fig. 1). In total, 35 sheep and 10 goats were sampled between December 2018 and September 2022 in Germany. These animals had developed abscesses in the region of the superficial lymph nodes and were sampled during controlled abscess lancing by the authors. Moreover, the farmers were concerned about carcass condemnations at the slaughterhouse due to abscesses in several lambs, which seemed to increase their willingness to engage with local veterinarians to identify the cause of the disease. In addition to the German specimens, four samples from two Polish dairy goat farms, obtained from a previous study in 2006–20073 were included, along with three samples from Austrian sheep obtained from routine diagnostics. Detailed information about the analyzed samples is given in Table 1 and in Supplementary Table 1.
The study was approved as it complied with the animal welfare guidelines of the University of Veterinary Medicine Hannover, the German Animal Welfare Act, and Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the protection of animals used for scientific purposes. All animal procedures were reviewed and approved by the Ethics Committee of the University of Veterinary Medicine Hannover (approval code: TiHo_EA_14_24–24) and were conducted in accordance with the ARRIVE guidelines. Animal-derived data were obtained exclusively for diagnostic purposes. Consequently, the use of these data did not require additional authorization from the IACUC or the competent authority under Directive 2010/63/EU.
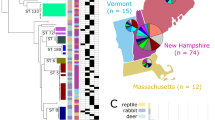
Location of the Staphylococcus aureus subsp. anaerobius (SAAN)-positive sheep and goat herds in Germany, Austria, and Poland. Different colors indicate different genomic clusters of SAAN based on cgMLST analysis. © GeoBasis-DE / BKG (2023).
Isolation of SAAN
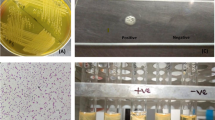
The abscess samples were examined applying classic cultural bacteriologic methods. In brief, swabs were plated onto Columbia Agar with 5% sheep blood and incubated under aerobic, microaerobic, and anaerobic conditions for 24, 48, and 72 h at 37 °C. No growth was observed on all plates after 24 h. After 48 h, small white colonies were visible on plates incubated under microaerobic and anaerobic conditions, while there was still no growth on aerobic plates. Presumptive S. aureus colonies were subcultured and confirmed by MALDI-TOF MS. Apart from different ambient conditions, SAAN can be differentiated from S. aureus subsp. aureus by a negative catalase test.
Whole-genome sequencing and bioinformatics analyses
Presumptive SAAN isolates and the DSM 20714 type strain were inoculated in 5 mL brain heart infusion broth and anaerobically incubated at 37 °C for 24 h. DNA of 1 mL culture was extracted using the Qiagen DNeasy Blood and Tissue Kit (Qiagen GmbH, Hilden, Germany) according to the manufacturer’s protocol, modified by adding 10 µL lysostaphin (0.1 mg/mL; Sigma-Aldrich Chemie GmbH, Taufkirchen, Germany) to the lysis buffer.
The DNA library was prepared using an Illumina Nextera DNA Prep kit (Illumina Inc., San Diego, CA, USA), and the 150 bp paired-end sequencing run was performed on an Illumina NextSeq 500 instrument. Raw Illumina reads were trimmed and de novo assembled using a SPAdes algorithm with the in-house developed AQUAMIS pipeline23. Bacterial characterization was conducted with the in-house developed BakCharak pipeline (https://gitlab.com/bfr_bioinformatics/bakcharak, accessed on February 8, 2023) using the NCBI AMRfinder database24 for screening of AMR genes, the virulence factor database (VFDB) SetB (http://www.mgc.ac.cn/VFs/, accessed August 26, 2022) for the detection of virulence genes, and MLST based on the scheme of Enright and colleagues25. For the presence of virulence and resistance genes, a 50% sequence coverage and 50% identity were considered. Genome comparisons and phylogenetic analyses were conducted by core genome multilocus sequence typing (cgMLST) using Ridom SeqSphere + version 7.0.4 (Ridom GmbH, Münster, Germany), and the integrated 1861 loci scheme was used26.
Additionally, raw paired-end reads of six publicly available genomes at the European Nucleotide Archive (ENA) from published strains18 that were isolated in Denmark, Italy, Poland, Spain, and the Sudan were assembled as described above and included in the cgMLST comparison. The respective strain IDs were complemented with ‘*_ENA’ and are listed in Supplementary Table 2.
Results
Phylogenetic analysis
The comparison of the 52 SAAN core genomes from this study, the core genome of DSM 20714 (Ref-DSMZ), and those from the further international SAAN strains available at the ENA database revealed a close phylogenetic relationship of most German isolates (n = 34) with allelic differences (AD) ≤ 6 in Cluster 1 (Fig. 2). In contrast, we found further distinct clusters of the same ST4581 with the goat isolates and ENA strain from Poland (P1, P2, and P_ENA, Cluster 3), sheep isolates (n = 4) from Bavaria (G5, Cluster 4), two sheep isolates from Austria (A2, A3, Cluster 2), and singletons like the Ref-DSMZ and the Spanish ENA strain that were more distantly related. Furthermore, we identified a fifth cluster of ST3756 strains that were assigned to two Bavarian farms (G2, G3), one farm in Baden-Wuerttemberg (G8), and a sheep farm from Austria (A1) as well as two singletons with the ENA strains from the Sudan that belonged to ST5843.
Interestingly, the four isolates from Poland (Cluster 3) and two isolates from Austria (Cluster 2) each assigned to ST4581 were phylogenetically highly related with AD ≤ 4, whereas a relationship between isolates of different countries could only be assumed in the case of the German and Austrian ST3756 isolates (G8, A1, Cluster 5), with the caveat that a clustering cut-off in that case was hard to define due to the lack of definitively epidemiologically unrelated ST3756 strains.
Furthermore, tight phylogenetic relationships were observed irrespective of the host species (sheep, goat) from which they originated, indicating no association between the sequence type and a certain host species.
Minimum spanning tree based on comparative cgMLST analyses of SAAN isolates from this study, the type strain DSM 20714 and published genome sequences from the ENA database. The ENA strains and the DSM strain are marked in gray, whereas the isolates of this study are colored according to clusters 1–5. The allelic differences (AD) between strains are indicated as numbers between the nodes. The branch length is not proportionate to the allelic differences. Clusters are defined by strains with less than 10 ADs.
Genotypic characterization: virulence-associated genes
The highly clonal structure and conserved genome could also be confirmed by analyzing genes that are associated with virulence and antimicrobial resistance. All isolates from this study independent of their ST harbored the same classical S. aureus virulence genes with relevance for adhesion like cna (collagen binding), eap (extracellular adherence protein), emp (extracellular matrix protein), cflB, efb and fnbA (fibronectin-binding), sas (S. aureus surface protein), srtB (sortase B), the expression of a type VII secretion system like the esa/ess/esx-operon, and the production of exotoxins (hemolysins: hlb, hld, hlgB, hlgC, hly; staphylococcal superantigen-like proteins: set25, set30, set31; leukotoxins: lukD, lukE, lukG, lukH) as well as exoenzymes (aureolysin: aur; hyaluronate lyase: hysA; lipase: geh, lip; coagulase: coa; serine proteases: ssp., spl; Von Willebrand factor-binding protein (vWbp): vwb). Furthermore, genes that affect immune evasion, like adsA (Adenosine synthase A), capsule-related genes (cap8B, cap8C, cap8E, cap8F, cap8H to cap8M, cap8O, cap8P, capA, capN), and sbi (staphylococcal binder of immunoglobulin), or are important for biofilm synthesis like the ica-operon, the sdrC gene, and the autolysin gene atl were present in all genomes. Iron acquisition is crucial for host infection and associated gene operons like sbn, sfa, and sir could be identified. All of these virulence genes were chromosomally encoded.
Genotypic characterization: antimicrobial resistance genes
The number of AMR associated genes or genetic modifications in SAAN were very limited and the same for all isolates in this study. We identified point mutations in the genes glpT_A100V, murA_E291D, and murA_T396N that are associated with fosfomycin resistance, and detected the mepA and tet(38) genes which confer resistance to tetracyclines. Furthermore, the lmrS gene of the lincomycin resistance protein, an efflux pump that belongs to the major facilitator superfamily27, was present. All genes were encoded chromosomally, plasmids could not be identified.
Discussion
To the best of the authors’ knowledge, this study describes the presence of SAAN in small ruminants located in Germany and Austria for the first time. However, it is noteworthy that a German breeding buck potentially introduced SAAN to a naïve Polish dairy goat herd in 20053. Therefore, MD appears to have been present in the German small ruminant population before. The reason for the sudden emergence in sheep herds in southern Germany is still unclear and might be associated with increased awareness of farmers and veterinarians. Besides the already described likely transmission from a German buck to a Polish goat herd, two SAAN outbreaks were reported in Denmark2 and the Czech Republic12, caused by the importation of sheep from France. However, current information about the presence of SAAN in France exists only for goat herds28. This emphasizes the need to implement quarantine measures, such as isolation and palpation of animals’ superficial lymph nodes, even for small ruminants from countries where SAAN has not yet been reported. In the present study, limited information was available regarding the source of SAAN introduction to small ruminant herds, though some farmers suggested that animal purchases may be responsible. Additionally, shearing has been reported as a risk factor for SAAN transmission to naïve herds15. Only one shearing contractor (Contractor A) sheared three of the sheep flocks included in this study (flocks G1, G3, and G7; see Supplementary Table 1), while other flocks were sheared by different contractors. However, SAAN transmission from flock G1 to flock G3 appears unlikely, as different MLST types were detected in the two flocks. Furthermore, SAAN was introduced to flock G7 due to the purchase of lambs from flock G1. Thus, shearing seems to play a less significant role in the transmission of SAAN among the herds studied here, but the small sample size limits the epidemiological traceability of SAAN transmission among sheep farms.
Interestingly, even though herds G2 and G3 were located in the same district and belong to the same cluster, we can exclude any animal movements between both farms, as they are pedigree breeders of different sheep breeds, and cross-breeding did not occur. Although the owners of herds G2 and G3 hired different shearing contractors (Supplementary Table 1), an indirect contact, e.g., by vehicles, cannot be completely ruled out. It is known that S. aureus can survive for up to seven months on inanimate surfaces29, but information about the longevity of SAAN is lacking. Other anaerobic bacteria, such as Bacteroides melaninogenicus and Fusobacteria nucleatum, can tolerate air for up to 72 h30. Such a short oxygen tolerance has to be proven for SAAN in the future. Moreover, ticks, in combination with the tick-borne bacteria Anaplasma (A.) phagocytophilum, favor tick pyemia caused by S. aureus in lambs31. This combination of tick infestation and A. phagocytophilum might also trigger the onset of clinical MD cases, as A. phagocytophilum is very common in the German sheep population and the pathogen has an immune-suppressive effect on small ruminants32.
We have described the relationship of isolates from different herds within Germany and strains from neighboring and distant countries, which allows us to draw conclusions regarding potential transmission pathways. However, we have not found any clear indications of an international transmission of strains within the last years. The isolates analyzed in this study mostly revealed a distant relationship across country borders, but the conclusion is limited due to the lack of complete surveillance data. Moreover, in the case of potential introduction into a Polish dairy goat herd, the time difference between the sampling of the goat herd and the German small ruminant herds is approximately 12 years, which additionally hampers the comparison. Nevertheless, single transmission events cannot be excluded and could be exemplarily present in the relationship of isolates G8 and A1 as part of Cluster 2, with the limitation of an uncertain cut-off between related and unrelated ST3756 strains. In addition, the same strain was also identified in sheep from the Czech Republic12, but a French origin of SAAN was suspected, which indicates a pathogen transmission among four countries. However, transmission events between herds and regions in Germany are more frequent than between countries, which results in the formation of endemic SAAN populations that can be phylogenetically distinguished using highly discriminatory molecular methods like single nucleotide polymorphism or cgMLST analyses.
Most of the virulence genes detected in the SAAN isolates occur ubiquitously in S. aureus. However, Yebra et al.18 described the frequent occurrence of pseudogenes in SAAN isolates with approximately 8% of the genome in contrast to S. aureus subsp. aureus isolates due to point mutations that affect the transcription of these genes. This indicates that beyond the pure detection of virulence and resistance genes in our study, statements regarding their functionality are very limited. Insofar some of the above-mentioned virulence genes might be probably inactive, namely capH8, cflB, emp, fnbA, hly, lip, lukD/E, and sbi which were described as pseudogenes and seem not relevant or beneficial regarding the adaptation to ruminants. In contrast, others do not show signs of impairment so far and are also reported to be relevant for attachment (cna, eap, efb, sas, srtB) or cytotoxicity (hlb, hld, hlgB, hlgC). Especially those encoding the Type VII Secretion System (esa/ess/esx-operon)33, which is important for pathogenicity by secreting toxins and other virulence factors as well as the interbacterial competition do not include pseudogenes. Further virulence genes that are similarly generalist and ensure the iron acquisition (sbn, sfa, sir), or confer advantages in the environment by enabling the production of biofilms (atl, ica, sdrC) have not been evolutionary pseudogenized.
The results reported here and elsewhere have shown the fundamental reduction of virulence genes in contrast to S. aureus subsp. aureus. On the other hand, a few virulence genes are much more prevalent in SAAN than in S. aureus subsp. aureus isolates like vwb or cna. Coagulase production is a prerequisite for coagulase-positive Staphylococci like S. aureus, but in contrast to coa, which is the classical mediator in virtually all S. aureus strains, vwb is only present in certain S. aureus strains. Both factors have been proven to synergistically affect the abscess formation in mice34 although each protein is able to coagulate plasma. The presence of the vwb gene in contrast to coa is, according to Pickering and colleagues35, the result of a host-dependent evolution, with a host-specific functionality especially with regard to ruminants and equines36. This was concurrently confirmed for SAAN by Yebra and colleagues18 who highlighted vWbp as an important virulence factor, which is encoded on one of only two MGEs in the accessory genome of SAAN strain MVF7, the SAAN pathogenicity island SaaPIMVF7, and hypothesized this particular vwb variant to be highly relevant for the formation of classical abscesses in MD18.
Another example of a gene which was prevalent in all SAAN isolates in this study but not very common in other S. aureus is cna, which encodes an adhesin that enables the binding to collagen37 and thereby resists clearance by the host defense mechanisms. It was observed that cna is especially prevalent in certain S. aureus clonal complexes (CC) like CC1, 12, 22, 30, 45, 51, and 23938, and that this might be linked to a colonizing rather than an invasive lifestyle. Furthermore, this gene was recently reported to be prevalent in atypical CC133 S. aureus from waterfowl39. However, it is yet not clear what drives this uneven distribution of cna between different S. aureus types and how far it could be linked to the adaptation to ruminants in SAAN or the progress of MD. Furthermore, it must be constrained that the cna reference sequence in the VFDB Set B was only covered by 61% by the assembled SAAN sequences. Thus, it is not yet ruled out that this gene is not functional in SAAN or is just a pseudogene.
Similar to the virulence genes, the few AMR determinants found in the SAAN isolates are very common in S. aureus and have been described for S. aureus subsp. aureus strains of different origin40,41,42. We observed point mutations in genes that encode for relevant antimicrobial targets and genes encoding for efflux pumps that limit the exposure to antimicrobial agents. Point mutations in antimicrobial target genes can confer a certain degree of resistance. For example, fosfomycin interferes with the function of the UDP-N-acetylglucosamine enolpyruvyl transferase (MurA), which is relevant for cell wall synthesis43. A point mutation leading to an amino acid substitution as is described here for murA can decrease the affinity for fosfomycin. GlpT is a transporter protein relevant for fosfomycin transport into the bacteria cell. Point mutations in glpT therefore affect the cell wall permeability for fosfomycin and act synergistically with murA mutations.
MepA, Tet(38), and LmrS are multi-drug efflux pumps belonging to different families and confer cross resistance to different antibiotics and biocides44. How far they are expressed in SAAN and biologically active compared to S. aureus subsp. aureus still has to be determined. At least mepA was reported to be impaired by a point-mutation18.
The current findings of low genetic diversity obtained with NGS align with other studies that used different molecular techniques14,18. This homogeneity may be utilized to develop vaccines that can be broadly applied and reduce the likelihood of pathogen evasion mechanisms. The success of vaccination programs in controlling SAAN in sheep herds was already demonstrated in Spain and the Czech Republic12,45 and is currently being investigated by the authors in SAAN-infected small ruminant herds in Germany46.
On the farms presented here, carcass condemnations occurred at slaughterhouses due to the presence of abscesses in a significant number of lambs. Therefore, it is important to discuss reports regarding the zoonotic potential of SAAN. Generally, transmission of S. aureus from livestock to humans and vice versa occurs, and depending on the lineages of S. aureus, the bacteria can cause severe diseases in both animals and humans47. A review of the literature reveals only a few reports of SAAN (catalase-negative S. aureus) infections in patients from several countries48,49,50. In most described cases, the strains were isolated from patients with a previous medical history. Although the pathogens were identified as catalase-negative S. aureus, and SAAN as a causative agent cannot be ruled out, there was no explicit evidence of this pathogen. To our knowledge, the only comprehensive case report of a human with a plausible SAAN infection came from Australia17. A farmhand showed severe clinical signs, including alveolar infiltrates, pain, and induration of the abdomen, phlebitis of superficial veins in one leg, and respective inguinal nodes. An obligately anaerobic S. aureus was isolated, and epidemiologic circumstances revealed that the man had handled sheep manure without wearing protective gloves. However, MD has not yet been diagnosed in Australian sheep flocks (personal communication Peter Windsor, The University of Sydney, Australia). Due to the limited differentiation of the subspecies anaerobius from other S. aureus strains, accurately determining the actual occurrence of the pathogen from case reports is challenging. More recent findings suggest a probable human-to-ruminant host transfer in the past and indicate a highly niche-specific ecology of SAAN adapted to sheep and goats18. While SAAN infection in patients other than sheep and goats is extremely rare, it cannot be completely ruled out. Considering the diagnostic difficulties, it remains unknown whether SAAN infection might be underdiagnosed in human medicine.
We are aware of the limitations of the present study. The analyzed isolates were collected using a convenience sampling approach and do not represent the prevalence of SAAN in Germany. Details about the probable introduction of SAAN are based on anecdotal reports from farmers due to the lack of surveillance data. Therefore, this information must be interpreted with caution.
In summary, the present study demonstrates for the first time a wide distribution of SAAN in small ruminant herds, especially in southern Germany. Moreover, it closes the epidemiological gap of missing data on SAAN, as noted by others14. The trading of infected animals may have facilitated transmission among herds both nationally and internationally. The identified SAAN isolates in small ruminant herds in Germany, Poland, and Austria exhibited low genetic diversity. Although SAAN was first described over 100 years ago, information such as environmental survival rates and zoonotic potential remain vague and should be investigated in future studies.
Data availability
Data are available upon request from the corresponding author. The assembled genome datasets generated during the current study can be found in the National Center for Biotechnology Information (NCBI) repository, https://www.ncbi.nlm.nih.gov/bioproject, PRJNA634452.
References
Morel, M. Contribution a L’etude de l’adénite caséeuse du Mouton. J. Méd Vét Zootechnie. 73, 513–520 (1911).
Møller, K., Agerholm, J. S., Ahrens, P., Jensen, N. E. & Nielsen, T. K. Abscess disease, caseous lymphadenitis, and pulmonary adenomatosis in imported sheep. J. Vet. Med. B. 47, 55–62. https://doi.org/10.1046/j.1439-0450.2000.00310.x (2000).
Szaluś-Jordanow, O. et al. Epidemiological features of Morel’s disease in goats. Pol. J. Vet. Sci. 13, 437–445 (2010).
Shirlaw, J. & Ashford, W. The occurrence of caseous lymphadenitis and Morel’s disease in a sheep flock in Kenya. Vet. Rec. 74, 1025–1026 (1962).
El Sanousi, S., Hamad, A. & Gameel, A. Abscess disease in goats in the Sudan. Rev Elev Med Vet Pays Trop. 42, 379–382 (1989).
Ben, M. S. et al. Epidemiological and clinical studies of ovine caseous lymphadenitis. Arch. Inst. Pasteur Tunis. 79, 51–57 (2002).
Alhendi, A. B., El-Sanousi, S. M., Al-Ghasnawi, Y. A. & Madawi, M. An outbreak of abscess disease in goats in Saudi Arabia. J. Vet. Med. A. 40, 646–651 (1993).
Al-Harbi, K. B. Prevalence and etiology of abscess disease of sheep and goats at Qassim region, Saudi Arabia. Vet. World. 4, 495–499. https://doi.org/10.5455/vetworld.2011.495-499 (2011).
Pegram, R. An unusual form of lymphadenitis in sheep and goats in the Somali Democratic Republic. Trop. Anim. Health Prod. 5, 35–39 (1973).
Habuš, J. et al. Comparison of the epizootiological and clinical features of caseous lymphadenitis and Morel’s disease in goats. Veterinarski Arhiv. 85, 163–173 (2015).
Rossetti, B., Regi, G., Röttele, K. & Overesch, G. Outbreak of Morel’s disease in a Swiss goat flock. Vet. Rec. Case Rep. 2, e000084. https://doi.org/10.1136/vetreccr-2014-000084 (2014).
Slosarkova, S. et al. New sequence type ST3756 of Staphylococcus aureus subspecies anaerobius as the causative agent of abscessing lymphadenitis in sheep. Comp. Immunol. Microbiol. Infect. Dis. 63, 112–116. https://doi.org/10.1016/j.cimid.2019.01.008 (2019).
De la Fuente, R. et al. Isolation frequency of bacteria causing lymphadenitis and abscesses in small ruminants in central Spain. Small Rumin Res. 154, 5–8. https://doi.org/10.1016/j.smallrumres.2017.06.022 (2017).
Szaluś-Jordanow, O. et al. MLST and RAPD molecular analysis of Staphylococcus aureus subsp. anaerobius isolated from goats in Poland. Arch. Microbiol. 200, 1407–1410. https://doi.org/10.1007/s00203-018-1568-1 (2018).
De la Fuente, R., Cid, D., Sanz, R. & Ruiz-Santa-Quiteria, J. An outbreak of abscess disease associated with shearing. Small Rumin Res. 26, 283–286. https://doi.org/10.1016/S0921-4488(97)00006-0 (1997).
Saeed, E. M. & Alharbi, K. B. Morel’s disease and caseous lymphadenitis: A literature review with special reference to Saudi Arabia. IOSR J. Agric. Vet. Sci. 7, 76–86 (2014).
Peake, S. L. et al. First report of septicemia caused by an obligately anaerobic Staphylococcus aureus infection in a human. J. Clin. Microbiol. 44, 2311–2313. https://doi.org/10.1128/JCM.02125-05 (2006).
Yebra, G. et al. Radical genome remodelling accompanied the emergence of a novel host-restricted bacterial pathogen. PLoS Pathog. 17, e1009606. https://doi.org/10.1371/journal.ppat.1009606 (2021).
Chelikani, P., Fita, I. & Loewen, P. C. Diversity of structures and properties among catalases. Cell. Mol. Life Sci. CMLS. 61, 192–208. https://doi.org/10.1007/s00018-003-3206-5 (2004).
Szaluś-Jordanow, O. et al. PFGE and AFLP genotyping of Staphylococcus aureus subsp. anaerobius isolated from goats with Morel’s disease. Arch. Microbiol. 195, 37–41. https://doi.org/10.1007/s00203-012-0844-8 (2013).
Cheung, G. Y. C., Bae, J. S. & Otto, M. Pathogenicity and virulence of Staphylococcus aureus. Virulence 12, 547–569. https://doi.org/10.1080/21505594.2021.1878688 (2021).
Lakhundi, S. & Zhang, K. Methicillin-Resistant Staphylococcus aureus: Molecular characterization, evolution, and epidemiology. Clin. Microbiol. Rev. 31, e00020-18. https://doi.org/10.1128/cmr.00020-18 (2018).
Deneke, C. et al. Species-specific quality control, assembly and contamination detection in microbial isolate sequences with AQUAMIS. Genes (Basel). 12, 644. https://doi.org/10.3390/genes12050644 (2021).
Feldgarden, M. et al. Validating the AMRFinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, e00483-19. https://doi.org/10.1128/aac.00483-19 (2019).
Enright, M. C., Day, N. P., Davies, C. E., Peacock, S. J. & Spratt, B. G. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 38, 1008–1015. https://doi.org/10.1128/jcm.38.3.1008-1015.2000 (2000).
Leopold, S. R., Goering, R. V., Witten, A., Harmsen, D. & Mellmann, A. Bacterial whole-genome sequencing revisited: portable, scalable, and standardized analysis for typing and detection of virulence and antibiotic resistance genes. J. Clin. Microbiol. 52, 2365–2370. https://doi.org/10.1128/jcm.00262-14 (2014).
Floyd, J. L., Smith, K. P., Kumar, S. H., Floyd, J. T. & Varela, M. F. LmrS is a multidrug efflux pump of the major facilitator superfamily from Staphylococcus aureus. Antimicrob. Agents Chemother. 54, 5406–5412. https://doi.org/10.1128/aac.00580-10 (2010).
Tingaud, R. Étude épidémiologique descriptive de l’infection par le microcoque de Morel (Staphylococcus aureus subsp. anaerobius) en élevage caprin laitier. dumas-03857101 (2022). https://dumas.ccsd.cnrs.fr/dumas-03857101.
Kramer, A., Schwebke, I. & Kampf, G. How long do nosocomial pathogens persist on inanimate surfaces? A systematic review. BMC Infect. Dis. 6, 130. https://doi.org/10.1186/1471-2334-6-130 (2006).
Lu, Z. & Imlay, J. A. When anaerobes encounter oxygen: Mechanisms of oxygen toxicity, tolerance and defence. Nat. Rev. Microbiol. 19, 774–785. https://doi.org/10.1038/s41579-021-00583-y (2021).
Lihou, K., Rose Vineer, H. & Wall, R. Distribution and prevalence of ticks and tick-borne disease on sheep and cattle farms in great Britain. Parasit. Vectors. 13, 406. https://doi.org/10.1186/s13071-020-04287-9 (2020).
Rubel, W., Schoneberg, C., Wolf, A., Ganter, M. & Bauer, B. U. Seroprevalence and risk factors of Anaplasma spp. in German small ruminant flocks. Animals 11, 2793. https://doi.org/10.3390/ani11102793 (2021).
Bowman, L. & Palmer, T. The type VII secretion system of Staphylococcus. Annu. Rev. Microbiol. 75, 471–494. https://doi.org/10.1146/annurev-micro-012721-123600 (2021).
Cheng, A. G. et al. Contribution of coagulases towards Staphylococcus aureus disease and protective immunity. PLoS Pathog. 6, e1001036. https://doi.org/10.1371/journal.ppat.1001036 (2010).
Pickering, A. C. et al. Evolutionary and functional analysis of coagulase positivity among the Staphylococci. mSphere 6, e0038121. https://doi.org/10.1128/mSphere.00381-21 (2021).
Viana, D. et al. Adaptation of Staphylococcus aureus to ruminant and equine hosts involves SaPI-carried variants of von Willebrand factor-binding protein. Mol. Microbiol. 77, 1583–1594. https://doi.org/10.1111/j.1365-2958.2010.07312.x (2010).
Madani, A., Garakani, K. & Mofrad, M. R. K. Molecular mechanics of Staphylococcus aureus adhesin, CNA, and the Inhibition of bacterial adhesion by stretching collagen. PLoS ONE. 12, e0179601. https://doi.org/10.1371/journal.pone.0179601 (2017).
Deurenberg, R. H. et al. The Staphylococcus aureus lineage-specific markers collagen adhesin and toxic shock syndrome toxin 1 distinguish multilocus sequence typing clonal complexes within spa clonal complexes. Diagn. Microbiol. Infect. Dis. 65, 116–122. https://doi.org/10.1016/j.diagmicrobio.2009.07.007 (2009).
Monecke, S. et al. Molecular characterization of chimeric Staphylococcus aureus strains from waterfowl. Microorganisms 12, 39. https://doi.org/10.3390/microorganisms12010096 (2024).
Bruce, S. A. et al. Shared antibiotic resistance and virulence genes in Staphylococcus aureus from diverse animal hosts. Sci. Rep. 12, 4413. https://doi.org/10.1038/s41598-022-08230-z (2022).
Conceição, T., Coelho, C., de Lencastre, H. & Aires-de-Sousa, M. High prevalence of biocide resistance determinants in Staphylococcus aureus isolates from three African countries. Antimicrob. Agents Chemother. 60, 678–681. https://doi.org/10.1128/aac.02140-15 (2016).
Ogundipe, F. O. et al. Antimicrobial resistance and virulence of Methicillin-Resistant Staphylococcus aureus from human, chicken and environmental samples within live bird markets in three Nigerian cities. Antibiot. (Basel). 9, 588. https://doi.org/10.3390/antibiotics9090588 (2020).
Lade, H., Joo, H. S. & Kim, J. S. Molecular basis of non-β-Lactam antibiotics resistance in Staphylococcus aureus. Antibiotics (Basel) 11, 1378. https://doi.org/10.3390/antibiotics11101378 (2022).
Costa, S. S., Viveiros, M., Amaral, L. & Couto, I. Multidrug efflux pumps in Staphylococcus aureus: An update. Open. Microbiol. J. 7, 59–71. https://doi.org/10.2174/1874285801307010059 (2013).
Navarro, T. et al. Farm Trial: Use of an autogenous vaccine against Staphylococcus aureus subsp. anaerobius. In Proceedings of 9th International Sheep Veterinary Congress, Harrogate, UK, 22.-26.05.2017.
Ambros, C. et al. Morel’s Disease bei kleinen Wiederkäuern (Morel’s Disease in Small Ruminants). In Proceedings of Tagung der DVG-Fachgruppe Kleine Wiederkäuer & Neuweltkamele, Giessen, 11.-13.10.2023; pp. 1–2.
Park, S. & Ronholm, J. Staphylococcus aureus in agriculture: Lessons in evolution from a multispecies pathogen. Clin. Microbiol. Rev. 34, 27. https://doi.org/10.1128/cmr.00182-20 (2021).
Turner, D. P. J., Pye, S. M. & Taylor, R. E. Catalase-negative Staphylococcus aureus septicaemia. J. Infect. 38, 132–133. https://doi.org/10.1016/S0163-4453(99)90085-3 (1999).
Over, U., Tüç, Y. & Söyletir, G. Catalase-negative Staphylococcus aureus: A rare isolate of human infection. Clin. Microbiol. Infect. 6, 681–682. https://doi.org/10.1046/j.1469-0691.2000.00153.x (2000).
Friedberg, B., Hauer, E., Belkhirat, M., Watine, J. & Le Coustumier, A. Catalase-negative Staphylococcus aureus: A rare cause of catheter-related bacteremia. Clin. Microbiol. Infect. 9, 1253–1255. https://doi.org/10.1111/j.1469-0691.2003.00760.x (2003).
Acknowledgements
Special thanks go to Patrick Wysocki (Institute of Epidemiology, Friedrich-Loeffler-Institut, Greifswald - Isle of Riems, Germany) for his kind support in providing Figure 1. We would also like to thank Frances Sherwood-Brock for proofreading the manuscript to ensure correct English.
Funding
Open Access funding enabled and organized by Projekt DEAL.
This research received no external funding. We acknowledge financial support by the Open Access Publication Fund of the University of Veterinary Medicine Hannover, Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization: BUB, SM; Methodology: BUB, SM; Formal analysis: SM; Validation: BUB, CA, KB, JH, JMR, SM; Investigation: BUB, CA, KB, JH, JMR, OS-J, CS, ES, TB, SM; Data curation: BUB, KB, SM; Writing – original draft: BUB, JMR, SM; Writing – Review & Editing: BUB, CA, KB, JH, JMR, OS-J, CS, ES, TB, MG, SM; Visualization: BUB, SM; Resources: KB, JMR, MG, SM; Supervision: BUB, SM.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
The study was approved as it complied with the animal welfare guidelines of the University of Veterinary Medicine Hannover, the German Animal Welfare Act, and Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the protection of animals used for scientific purposes. All animal procedures were reviewed and approved by the Ethics Committee of the University of Veterinary Medicine Hannover (approval code: TiHo_EA_14_24–24) and were conducted in accordance with the ARRIVE guidelines. Animal-derived data were obtained exclusively for diagnostic purposes. Consequently, the use of these data did not require additional authorization from the IACUC or the competent authority under Directive 2010/63/EU.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bauer, B.U., Ambros, C., Boll, K. et al. Presence and genetic variability of Staphylococcus aureus subsp. anaerobius isolated from small ruminants in Central Europe. Sci Rep 15, 13493 (2025). https://doi.org/10.1038/s41598-025-97300-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-97300-z