Abstract
Tuberculosis (TB) is a recurrent and progressive bacterial disease caused by Mycobacterium tuberculosis (Mtb), posing a significant challenge globally due to its drug resistance. This study focuses on identifying natural phytocompounds from the plant Datura innoxia (leaves), which is well known for its biologically active metabolites. Initially, the current study employed in vitro analysis of 20 phytocompounds, revealing that the natural compound 9, o-vanillin, exhibited the best minimal inhibitory concentration (MIC) which was 12.5 µg/mL, and minimal bactericidal concentration (MBC) was 50 µg/mL, all other phytocompounds showing remarkable antitubercular activity against the Mtb H37Ra strain. The molecular docking and simulation also validated the strong affinity and stable binding interactions between compound 9 and target protein kinase. The pharmacokinetic analysis highlighted the suitable oral bioavailability and no significant CYP450 inhibition for the lead compound 9, reducing the risk for drug-drug interactions. Moreover, the density functional theory analysis of lead compound 9 demonstrated optimal molecular properties, further contributing to the chemical stability and reactivity. Therefore, these results suggest that D. innoxia contains the potent phytocompound o-vanillin, which possesses antitubercular activity and can potentially be used as a drug against TB. However, future studies will focus on in vivo validation and formulation development for clinical applications.
Similar content being viewed by others
Introduction
Tuberculosis (TB) is a lethal, infectious disease. It is caused by the etiological agent Mycobacterium tuberculosis (Mtb)1. It is an aerobic bacillus that can infect any body part, especially the lungs2. According to a recent WHO report on TB, it is estimated that globally, 10.6 million people were infected with TB, with 1.3 million mortalities yearly3, with the highest burden in Asia and Africa4.
Back in ’90 s, there was a significant reduction in TB cases due to the introduction and discovery of antitubercular agents, such as isoniazid5, pyrazinamide, and rifampicin6. Isoniazid is considered the first-line antitubercular defense against TB, but unfortunately, the Mtb strains have shown resistance to isoniazid7. Similarly, rifampicin was also considered an important antitubercular agent, but some unavoidable effects have been reported against this key-player drug in the TB regimen8. Mtb resistance has also caused inefficiency in some of the 2nd line antitubercular drugs such as ethionamide, capreomycin, and kanamycin9. Thereby, with the misuse of these antitubercular drugs, there has been a significant rise in TB cases. Despite the accessibility of WHO-recommended directly observed treatment, short course (DOTS), drug resistance has become a significant barrier to TB eradication globally because of multidrug resistance (MDR) and extensive drug resistance (XDR)10. Although tremendous efforts have been made to discover targeted antitubercular drugs, most have failed to reach the clinical trial phase in the past six decades11. Therefore, the development of novel antitubercular drugs with different inhibitory targets that cause minimal or no side effects is urgently needed.
Notably, plants and humans exhibit significant associations12. Natural products retrieved from plants have always been essential sources of drug interventions13 due to their significant biological activities14. According to floristic studies, there are approximately 500,000 plant species, of which 120,000 contain biologically active metabolites that could serve as therapeutic agents15. Several secondary metabolites are greatly used as antitubercular agents, such as phenols, alkaloids, salicylates, coumarins, flavonoids, indoles, etc. The biomedical sciences domain has now emphasized the identification of therapeutic phytochemicals to overcome the challenges for the treatment of TB16.
The medicinal herb Datura belongs to the Solanaceae family and is commonly referred to as jimsonweed or thornapple17. A variety of Datura species are widely seeded on major continents, including Asia, Africa, Europe, America, and other subtropical and tropical regions18. This plant exhibits both toxic and medicinal properties, and various developing countries use this plant for herbal medicine preparations19. Several species of this plant possess multiple biological properties against various diseases, such as diabetes, cancer, asthma, and TB. Moreover, Datura species also exhibit anti-inflammatory, analgesic, antioxidant, cytotoxic, insecticidal, and neurological activities, as well as wound-healing properties20.
For the effective and economical identification of a potential phytocompound that could be used as an antitubercular drug, an in silico approach has proven to be a reliable method to be employed before actual synthesis21. It should be taken into account that the therapeutic interventions of phytocompounds are evaluated thoroughly, as there could be a chance of discovering more potent modes of action and targets as better candidates for drugs19.
Although natural phytocompounds have been identified for TB treatment, no such studies have systematically evaluated the phytocompounds of D. innoxia against the resistant Mtb H37Ra strain. Therefore, the current study was conducted on the phytocompounds present in the leaves of the D. innoxia plant to identify the antitubercular activity of these phytocompounds under controlled in vitro conditions and followed by an in silico analysis of the phytocompound by molecular docking, pharmacokinetic analysis, DFT calculations, and MD simulation to verify the potential of the compound to be used as a therapeutic drug against TB.
Results
Antitubercular activity
To develop novel antitubercular drugs, we analyzed 20 natural compounds from D. innoxia plant. The results illustrated in Table 1 show the MIC and MBC of these phytocompounds against the Mtb H37Ra strain. The antitubercular activity of the compounds (9), (7), and (12) showed the best inhibitory activity at 12.5 µg/mL, 50 µg/mL, and 50 µg/mL respectively and also have bactericidal activity with MBCs values of 50 µg/mL, 50 µg/mL and 200 µg/mL respectively. On the other hand, the maximum MIC value was observed for compound (11), which was > 200 µg/mL, while all the other remaining compounds (1), (2), (3), (6), (8), (10), (14), (16), (17), (18), (19), and (20) displayed moderately weak activity at approximately 100 µg/mL, and the following compounds (4), (5), (13), and (15) showed extremely weak activity at 200 µg/mL against the Mtb H37Ra strain. The INH positive control, exhibited MIC and MBC values of 0.156 µg/mL. This suggests that out of all 20 phytocompounds, compound (9) showed strong momentous antitubercular activity under controlled in vitro conditions.
Molecular docking
The lead compound (9) was selected for in silico analysis based on strong evidence of its in vitro antitubercular activity. To further investigate the binding potential of the compound (9), o-vanillin, with the targeted protein kinase (PDB ID: 6B2Q), involved dual inhibition of PknA and PknB, was analyzed as shown in Fig. 1. The binding affinity of compound (9), reference (INH) drug, and CCL (PubChem CID: 50898364) was assessed based on the binding score and interactions with the target protein (PDB ID: 6B2Q). The docking procedure was verified by re-docking the CCL within the allosteric pocket of the protein by following the guidelines mentioned in the methodology section. The superimposed redocked structure shown in Fig. 2 validates the docking procedure as it exhibited an RMSD value of 0.200 Å22. The docking results of compound (9), illustrated in Table 2, displayed a promising docking score of -6.836 kcal/mol, compared to the INH drug.
3-dimensional structure of the protein Pkn (PDB ID = 6B2Q), visualized on Discovery Studio Visualizer, (3D representation: PyMol Molecular Graphics system version 2.4.0).
RMSD value of superimposed original CCL (PubChem CID: 50898364) (blue) and re-docked pose (purple).
The 2D and 3D interactions of compound (9), viewed using Discovery Studio and PyMol (Fig. 3), revealed the formation of a conventional hydrogen bond between VAL-98 amino acid residue and the hydroxy group located at the ortho position, relative to the aldehyde group. The Pi-alkyl interactions were formed between the benzoic moiety and ALA-40, ALA-74, VAL-27, ILE-19, and LEU-148 residues. Additionally, ALA-40 and ALA-74 also formed similar interactions along with MET-95, with the methoxy group present at the meta position to the aldehyde group. Simultaneously, LEU-97 and GLY-100 showed Van Dar Waal interactions, while THR-158 and GLU-96 exhibited a possibility of forming a carbon-hydrogen bond in the compound (9).
In contrast, the CCL showed various conventional hydrogen bonds with the following amino-acid residues of the target protein: LYS-42, ASN-146, GLY-145, GLU-96, and VAL-98. It is noteworthy that the LEU-97 residue showed Van Der Waal’s interaction, and the THR-158 residue showed a carbon-hydrogen bond similar to the compound (9). Moreover, CCL showed Pi-alkyl interactions with similar key amino acid residues (ALA-40, ALA-74, VAL-27, ILE-19, LEU-148, and MET-95) that are involved in the compound (9). These interactions revealed a similar binding pattern and interactions with amino acid residues when compared to the interaction pattern of CCL.
Subsequently, the binding interactions of INH, compared to the compound (9), revealed the formation of a favorable conventional hydrogen bond by the key residues i-e, VAL-98, THR-158, and GLU-96, with the carbonyl and hydroxyl groups of the INH drug. Additionally, non-covalent hydrophobic interactions were also formed by the key residues of the binding pocket, shown in Supplementary Fig. 1. Overall, the interactions highlighted that VAL-98, GLU-98, VAL-27, ALA-40, LEU-148, ILE-19, and THR-158 were considered as the major residues of the binding cavity involved in the inhibitory potential for therapeutic purposes.
3D representation of the complex 6B2Q with the best inhibitory compound (9), shown in (A), and CCL shown in (C) simultaneously, (B) and (D) show the 2D plot of the binding interactions between the compound (9) and CCL against protein Pkn (PDB ID: 6B2Q), respectively, (3D representation: PyMol Molecular Graphics system version 2.4.0; 2D representation: BIOVIA Discovery Studio Client 2021 version 21.1.0.20298).
Pharmacokinetic analysis
The pharmacokinetic analysis of lead compound 9, compared with INH illustrated in Table 3, showed that both compounds fell within Lipinski’s cut-off range, referred to as the Lipinski rule of five (LRF), suggesting a suitable oral bioavailability for these compounds. The lipophilicity was measured as Log Po/w (octanal-water partition coefficient), and solubility was measured as Log S for INH drug and compound 9, revealing that both compounds exhibited a suitable balance between hydrophilicity and hydrophobicity for a drug-like candidate. Subsequently, the drug-likeness of the compound 9 and INH drug indicated their potential use of the compound for therapeutic purposes. However, the medicinal chemistry showed that compound 9 and INH drug showed no pan-assay interference compounds (PAINS) alerts, indicating their reliability as a potent drug candidate. Whereas two Brenk alerts were observed in INH drug i-e., acyle hydrazine and hydrazine, and one Brenk alert was also possessed by the lead compound 9, i-e., aldehyde, indicating few structural liabilities. Moreover, the synthetic accessibility also highlighted the ease of synthesis for both compounds, particularly the lead compound 9. Additionally, the pharmacokinetic descriptors revealed high gastrointestinal (GI) absorption for the compound 9 and INH drug as well as highlighting that both of them were not a substrate p-glycoprotein (PGP), indicating a suitable therapeutic potential of compound (9), compared to the INH drug. Furthermore, compound 9 was also found to be not involved in any type of drug-drug interactions, similar to the INH drug.
Density functional theory (DFT) calculations
The DFT calculations for CCL and compound (9) were performed using Koopman’s theorem to compute the frontier molecular orbitals (FMOs) and global reactivity parameters23, illustrated in Table 4. The FMOs displayed in Fig. 4, highlighted the HOMO and LUMO energy, respectively, depicting electron donating potential and electron accepting ability of the molecules24, illustrated in Table 4. The difference between FMOs, represented as (ΔEGap = ELUMO – EHOMO), is significant in analyzing the electronic transitions of the compounds. The comparative analysis of CCL and compound (9) showed that there is a slight variation in the energy gap (ΔEGap), with compound (9) (0.14975 a.u.) being higher than CCL (0.1331 a.u.), indicating it to be more chemically stable than CCL25.
The result of global chemical reactivity descriptors illustrated in Table 4, showed that the ionization potential (I) of compound (9) and CCL was 5.84 eV, suggesting higher stability of both compounds26. On the other hand, the electron affinity (A) showed that a higher value of CCL made it more prone to accept the electron while a lower A of compound (9), suggested that it was more likely to be an electron donator27. The electronegativity (χ) outcomes of both compounds were almost similar, while the electrophilicity (ω) revealed that CCL was more reactive than the compound (9). However, the chemical hardness (η) and softness (S) of the compounds revealed that both compounds had comparatively the same level of hardness, whereas compound (9) (0.49 eV) was significantly softer than CCL (0.55 eV). Moreover, the charge distribution in a molecule is visualized as MEP in Fig. 4 to determine the presence of binding sites for receptors or ligands28.
Optimized structural geometries of compound (9) and CCL, depicted in both solvent and gaseous phases (Figures A and B for compound (9), and C and D for CCL), showed MEP. In the illustrations of MEP, the green areas represented neutral electrostatic potential, red indicated negative potential, and blue signified positive potential.
MD simulations
The docking studies showed protein-ligand binding interactions in a static condition while the MD simulation identified the complex’s structural conformation and motion in real time29. To observe the dynamic behavior of compound (9) and CCL with the target protein (PDB ID: 6B2Q), 100ns simulations were run, enabling us to scrutinize the fluctuation and adaptability within the allosteric cavity.
Figure 5(A) shows the RMSD values for the Cα atoms for both compounds. The RMSD plot revealed that compound (9) (represented in orange) and CCL (represented in blue) remained stable and steady throughout the 100 ns trajectory, characterized by minimal scattering in compound (9) for approximately 5 ns at the beginning of the simulations. It is noteworthy that the RMSD value of compound (9) maintains an RMSD below 3Å, indicating its stability within the protein’s binding site.
Figure 5(B), displayed the RMSF analysis for the target protein, providing insights into the real-time behavior of its amino acid residues. The results showed the structural integrity of amino acids. However, a moderate level of fluctuation was observed in compound (9) (represented in orange) from GLY-80 to THR-90 residues of the protein but did not exceed more than 3 Å, indicating structural flexibility and conformational changes within the complex.
The overall compactness of the protein was determined by RGyr, shown in Fig. 5(C). The results showed that the receptor in complex with compound (9) was more compact and consistent during the 100ns trajectory due to a lower level of fluctuations compared to CCL, which showed a slightly higher fluctuation moving towards the N-terminus of the protein, suggesting less compactness of the protein bound to CCL.
Moreover, the total number of hydrogen bond interactions between compound (9) and the target receptor, in comparison to the CCL docked complex, was also analyzed over 100ns simulation trajectory. The plot illustrated in Supplementary Fig. 2 revealed a dynamic binding pattern for CCL with minor fluctuations around 75 ns, indicating transient binding stability. Whereas, the hydrogen bond analysis of the lead compound 9 showed that hydrogen bond donors were more continuous than hydrogen bond acceptors, and mainly fluctuates between 6 and 9 over time. It was noteworthy that no complete loss of hydrogen bond was observed during the 100 ns trajectory with a pronounced fluctuation around 50 ns, indicating flexible and stable conformational changes within the docked complex.
Graphical representation of the simulation. (A) RMSD fluctuation of CCL (blue) and compound (9) (orange), with time on the X-axis and RMSD on the Y-axis. Plot (B) portrayed the RMSF of CCL (blue) and compound (9) (orange), with time on the X-axis and RMSF on the Y-axis. Plot (C) displayed RGyr with time on the X-axis and RGyr on the Y-axis. RMSD = root mean square deviation, RMSF = root mean square fluctuation, RGyr = radius of gyration.
PCA-based FEL analysis
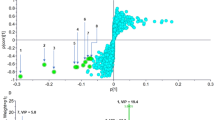
The PCA investigated the conformation dynamics of the complexes during a 100ns simulation trajectory. The plots shown in Fig. 6, revealed that although both complexes showed variation in conformational dynamics. The docked complex with CCL showed a more scattered distribution, highlighting its higher conformational flexibility. Whereas, the lead compound (9) possessed more compact cluster, signifying its greater stability during 100ns simulation trajectory further enhancing its inhibitory efficacy against the target protein.
PCA of the compound (9) in comparison to the CCL. (A) Combined scattered plots for both complexes, (color-coded blue = CCL and orange = P-2). (B&C) individual scattered plots of CCL and compound (9) in complex with target protein 6B2Q, respectively. (D&E) plots colored with Gibb’s energy respectively.
Moreover, the results of FEL (Fig. 7) showed a compact conformation of the ligand binding site in both complexes, indicated as Gibbs free energy across principal components. The plots depicted that the lead compound (9) exhibited a lower energy basin, indicating a more stable binding conformation with minimal energy fluctuation compared to CCL. These results suggested that the compound 9 remains stabilized within the binding pocket of target protein by minimizing the entropic effect, which could ultimately improve its efficacy as a therapeutic agent against drug resistant Mtb.
(A & B) represents the FEL of the CCL and compound (9) across PC1 and PC2. FEL = free energy landscape.
Discussion
Tuberculosis is a lethal communicable disease, specifically caused by Mtb. The reemergence of TB over the past few decades due to drug resistance has posed a significant challenge affecting healthcare. This challenge has necessitated the swift identification and development of new antitubercular drugs30. The field of biological sciences has revolutionized by focusing on the treatment of multiple diseases through natural compounds. Researchers are currently concentrating on utilizing comprehensive in vitro and in silico approaches to explore natural phytocompounds as promising drug candidates. An experimental study conducted by Khan M.U. et al. investigated the efficacy of the natural compound quercetin and armepavine as a potential drug candidate against cystic fibrosis31. Similarly, another study employed in silico studies to search for novel natural compounds against neurological disorders32.
The current study aimed to identify the phytocompounds from the D. innoxia plant which is capable of producing biologically active metabolites against TB20. This research employed in vitro analysis on 20 biologically active phytocompounds of D. innoxia as antitubercular agents, as reported in the literature. The in vitro analysis revealed that compound 9 was capable of maximum inhibitory activity with a minimal inhibitory concentration of 12.5 µg/mL against the Mtb H37Ra strain, suggesting it is a remarkable phytocompound to be used as an antitubercular agent. The potential of compound 9 as an inhibitor was also confirmed by another study that demonstrated the in vitro efficacy of vanillin against Leishmania tropica33. Another experimental research conducted by Ravindran R. et al.34 also aligns with the current study, showing the inhibitory potential of conventional plants against Mycobacterium smegmatis. Similarly, one of the studies showed the MIC value of Datura species against the Mtb H37Rv strain, indicating the potential of biologically active phytocompounds to treat TB35.
Based on the in vitro inhibitory potential of lead compound 9 against Mtb H37Ra, docking studies were performed along with DFT analysis and MD simulations to gain further knowledge on the potential binding affinity and interactions of the compound against the receptor of this bacterium. A similar approach was also implemented in various studies for the identification of novel compounds against drug-resistant bacteria. This approach is further supported by research conducted on the natural compound propolis, employing both in vitro and in silico analysis, to combat anti-microbial drug resistance19.
The results of the molecular docking showed an acceptable binding affinity with the protein kinase (PDB ID: 6B2Q) in a static condition. The binding interactions of the lead compound 9 exhibited a strong H-bond with the VAL-98 and GLU-96 amino acid residue of the protein, along with Pi-alkyl and van der Waals interactions among key amino acid residues such as ALA-40, ALA-74, VAL-27, ILE-19, and LEU-148 involved in CCL. These interactions showed stable binding conformation of the lead compound within the binding pocket of the target protein, which was further validated by MD simulations. A study also confirmed the inhibitory potential of the target protein kinase B (PknB) as an attractive target of Mtb H37Ra for the identification and discovery of novel drug candidates36. Moreover, experimental research conducted to identify novel inhibitors revealed six compounds as potent drug candidates against a similar target, protein kinase37.
The pharmacokinetic analysis of the lead compound in comparison to the INH drug also provided essential details for the compound to assess its therapeutic potential as an antitubercular agent. The pharmacological results comply with the LRF, underscoring its favorable oral bioavailability. Followed by moderately balanced hydrophilic and hydrophobic profiles highlighting the membrane permeability and absorption of the lead compound 9 within physiological space. Moreover, the lead compound was also observed to be PGP negative, revealing that it is unlikely to be rapidly effluxed from the cells, similar to the INH drug. It is noteworthy that the high Log P value of lead compound 9 suggested its improved permeability, which correlates with its significant binding affinity with the target protein. Likewise, with the lack of CYP450 inhibition in INH drug, the lead compound 9 was also found unlikely to interfere with the metabolic enzymes, thereby reducing the risk for drug-drug interactions as well as altered drug metabolism. However, the aldehyde group of lead compound 9, highlighted as Brenk alert, suggested metabolic liability; simultaneously, this structural feature may also have contributed to its stronger binding interaction with the active site residues of the target protein. Therefore, further optimization of this functional group may enhance the stability of the compound while retaining its bioactivity for therapeutic purposes.
DFT calculations determined the molecular properties of the lead compound 9 in comparison to CCL. The HOMO and LUMO energy levels provided an insight into the donation and acceptance of electrons in the molecule, highlighting its significance for molecular interactions, specifically with protein-ligand binding. The results illustrated in Table 3 show that the significant properties of the compound 9 exhibited strong reactivity and stability. The energy gap is considered a key indicator of chemical stability and reactivity, suggesting greater chemical stability and suitable reactivity for the lead compound 9. This stability and reactivity of compound 9 also aligns with the docking and simulation studies underscoring strong and persistent interactions with key interacting residues (VAL-98, ALA-40, and GLU-96) throughout the simulation trajectory. Additionally, the stable RMSD value and strong hydrogen bonding during simulations also supported the stability and reactivity of the lead compound 9. Moreover, the global reactivity parameters, such as electronegativity and electrophilicity, also showed that CCL is more prone to non-specific interactions due to its higher reactivity, whereas a balanced reactivity of compound 9 contributed to selective binding to the target protein, as visualized in docking studies. Furthermore, higher ionization energy and lower electron affinity of lead compound 9 indicated enhanced stability within the binding pocket of the target protein. Subsequently, the formation of hydrogen bonds with VAL-98 residue and hydrophobic interactions with ALA-40 and LEU-148 also aligns with the molecular electrostatic potential visualized in the MEP analysis. Yernale et al. have also applied this approach to investigate novel compounds as antimicrobial and antioxidant agents. Further, the DFT calculations of natural compounds were also computed by Jayaraman M. et al.38, showing the molecular properties of the natural compound to be used as inhibitors against Mtb H37Ra. Therefore, by integrating DFT findings with molecular docking and simulations, the current study provides an in-depth understanding of the electronic properties of compound 9 contributing to its binding affinity and stability against the target protein, further underscoring its potential as a promising antitubercular agent.
Finally, the compound was subjected to MD simulations to determine the stability of the complex in real time. Other studies have also utilized this approach to observe the stability and conformational changes within the complex39. The results of the current investigation showed that in the phytocompound in complex with the target protein kinase (PDB ID: 6B2Q), the RMSD value remained within an acceptable range of 3 Å, and the RMSF value also remained within the limit of 3 Å, suggesting that fluctuations within the amino acid residues were responsible for the molecular motion, leading to functional modulation. The overall analysis showed that the complex remained steady and stable, showing flexibility during a 100ns simulation trajectory. These results suggested a more specific interaction pattern between ligand and receptor, which correlates to higher binding affinity and sustained inhibition of the target protein. Additionally, the RGyr quantifies the compactness of docked complex. The pronounced compactness in the lead compound 9 correlates with the stable RMSD and RMSF patterns, hence validating its potential as an effective inhibitor for therapeutic purposes. Furthermore, the stability and lower conformational changes in compound 9 docked with target protein underscore its efficacy for a promising drug like candidate for future drug development. Simultaneously, the hydrogen bond analysis also highlighted that compound 9 bind effectively with the receptor due to the formation of stable hydrogen bonds throughout simulation trajectory. This also correlates with the docking results, further supporting the therapeutic efficacy of compound 9 against resistant Mtb H37Ra strain. These results are further supported by a study conducted on the Mtb H37Rv strain on PknG in complex with phytocompounds, showing promising results in the in silico studies40. These findings offered a potent lead compound 9, o-vanillin, as an antitubercular agent for combating TB due to drug resistance.
Conclusion
The emergence of drug-resistant Mtb poses a significant health challenge. From this perspective, the current study reflected that the phytocompound o-vanillin, retrieved from the plant D. innoxia, exhibited potential biological inhibitory activities, as confirmed by in vitro analysis. Moreover, in silico analysis showed valuable binding affinities and interactions while determining the molecular characteristics of o-vanillin as a suitable antitubercular drug candidate. However, in vivo analyses are yet to be performed to further validate the efficacy of this compound for safe delivery with minimal side effects against TB.
Methodology
The current study employed comprehensive in vitro and in silico methodology, displayed in Fig. 8 for the identification of promising phytocompounds from the D. innoxia plant.
Approach used in the current study for identifying phytocompound against Mtb H37Ra.
In vitro study
Phytocompounds for bioassay
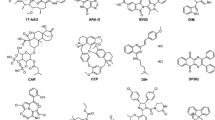
Initially, we explored the anti-tubercular properties of various medicinal plants and found that Datura innoxia was the most potent. Therefore, we selected twenty phytocomponents from Datura innoxia that have biological active in order to investigate its anti-tubercular potential. Twenty natural phytocompounds include trans-ferulic acid, 4-hydroxybenzoic acid, (-)-scopolamine hydrobromide, (-)-scopolamine N-butyl bromide, norharman, p-coumaric acid, anisodamine (7β-hydroxyhyoscyamine), o-vanillin, nicotinic acid, atropine, piperine, scopoletin, Methyl isonicotinate, Methyl isonicotinate N-oxide, d-Damascone, 3-Indoleacetic acid, 3-Methylindole, 2-Aminonicotinic acid, and 2-hydroxy-3-methoxybenzoic acid were purchased in pure form for antitubercular analysis in this study detail. The commercial sources and authentication numbers of the compounds were detailed in Supplementary Table 1.
Compounds dissolution and dilution
Initially, 4 mg of each compound was dissolved per mL of dimethyl sulfoxide (DMSO), followed by two-fold serial dilutions, resulting in a concentration; 200 µg/mL to 0.39 µg/mL for each compound. INH positive control was tested at concentrations range from 0.625 µg/mL to 0.039 µg/mL.
Microorganism strain
The Mtb strain utilized in the current study was the Mycobacterium tuberculosis H37Ra strain (Mtb H37Ra), to evaluate the antitubercular activity of phytocompounds.
Growth conditions of bacteria
The Mtb H37Ra strain was grown in a sterile PETG media bottle containing 10 mL Middlebrook 7H9 broth supplemented with 10% (w/v) OADC (Difco, Becton Dickinson, USA), 0.05% (w/v) tyloxapol, and 0.2% (v/v) glycerol (Sigma Chemical Co.). The growth conditions were maintained at 37 °C with mild shaking at 100 rpm in a shaking incubator (Innova 4900, New Brunswick Scientific, USA) until Mtb H37Ra strain growth reached the log phase with an optical density (OD) of 0.1–0.3 at 600 nm. The culture was diluted to a final OD600 of ~ 0.01 for the antitubercular assay. Simultaneously, Middlebrook 7H10 agar media supplemented with 0.2% glycerol and 10% (w/v) OADC (Difco Becton Dickinson, USA) was prepared to measure the minimum bactericidal concentration (MBC) of Mtb H37Ra.
Determination of minimum inhibitory concentration and minimum bactericidal concentration by microdilution method – MTT assay
3-[4,5-Dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide (MTT) was used to determine the antitubercular activity of the phytocompounds against Mtb H37Ra, with a slight modification of the method according to Martin et al.41. All phytocompounds were diluted two-fold in DMSO and added 5µL in a 96-well microtiter plate and then 95µL of the Mtb H37Ra culture suspension (final OD600 = 0.01) was inoculated into each well, followed by incubation at 37 °C for one week. INH was used as a positive control, and DMSO was used as a negative control because it is not active against this bacterium. Simultaneously, the MBC was recorded as the lowest concentration that killed 99% of the colony-forming units (CFU) in the initial inoculum. For this purpose, 10µL was taken from each well, inoculated onto Middlebrook 7H10 agar plates, incubated at 37 °C, and read after 3 weeks. Then 10µL of MTT solution was poured into each 96-well plate containing the inoculum and incubated at 37 °C overnight. If violet-colored precipitation was observed visually, then 50µL of SDS-DMF formazan solubilization buffer was added, followed by incubation at 37 °C for another 3–4 h. A color change from yellow to violet indicates bacterial growth, and the MICs were interpreted accordingly42. The experiments were performed in triplicate and the MIC was recorded by visual observation.
In silico analysis
Target protein retrieval
Data from the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) (RCSB PDB: Homepage), accessed on September 23, 2024, were used to retrieve the crystal structure of protein kinase (Pkn), PDB ID: 6B2Q, which is majorly involved in signal transduction. The structure retrieved was in PDB format and contained four chains, namely A, B, C, and D, each having a sequence length of 317 amino acids without any mutation at a resolution of 2.88 Å43.
Ligand retrieval
The lead compound was selected as the ligand for the in silico analysis. The phytocompound was retrieved in 2D and 3D structures from PubChem and sketched by using ChemDraw Professional 16.0 software (version: 16.0.1.4.77). The 3D structure was saved in the SDF format for docking purposes and the MOL format for DFT calculations.
Ligand preparation
The Ligprep tool available on Schrödinger’s 2020-3 (Maestro Version 12.5.139) was utilized for ligand preparation of the co-crystal ligand (CCL) and lead compound for obtaining structural optimization. The Epik module was employed to generate 32 poses for each ligand, producing desalts and tautomers at pH 7.0 ± 2. Finally, each ligand was minimized under a force field of OPLS3e44.
Protein preparation
The Protein Preparation Wizard tool, easily available on Schrodinger 2020-3 (Maestro Version 12.5.139) was employed to conduct the protein preparation of the target protein, PDB ID: 6B2Q. The missing loops and side chains were filled by the prime job, and the Epik module was employed to generate het states between pH = 7.0 ± 2, and water molecules beyond the diameter of 3 Å were deleted. Moreover, PROPKA was employed at pH = 7.0 to optimize the H-bond scaffolding. The steric hindrance was removed for further refinement and modification, and finally, the protein was minimized under the OPLS3e force field45.
Glide grid generation
Receptor grid generation was done using the receptor grid generation panel in the Maestro glide program available on Schrodinger 2020-3 (Maestro Version 12.5.139) in the centroid of the working space of the CCL for the identification of suitable interactions between the receptor and the ligand46,47.
Molecular docking
The molecular docking of lead compound and INH was done along with CCL on Maestro, Schrodinger 2020-3 (Maestro Version 12.5.139), with the Ligand Docking tool. The receptor grid file and the energy-minimized files of the ligands were imported48 and the docking was set to extra precision, flexible ligand sampling with the per residue scoring of 12Å of grid generation along with the calculation of RMSD to input ligand geometries. Finally, the 2D interactions of the docked complexes were viewed on Discovery Studio Visualizer v21.1.0.20298, and the 3D interactions were viewed on PyMol Molecular Graphic System version 2.4.0.
Pharmacokinetic analysis
The pharmacokinetic analysis of the lead compound and INH drug was performed on SwissADME (SwissADME), an online tool, by applying the molecular fingerprinting technique. The string of canonical SMILES of the query molecule was provided as input data to evaluate the ADME across various descriptors, including physiochemical features, lipophilicity, solubility, pharmacokinetics, drug-likeness, and medicinal chemistry.
DFT calculations
The three-dimensional geometries of the lead compound and CCL were designed and optimized by using GaussView (version 5.0.8). The CPCM model was chosen for the optimization and frequency calculations in the solvent and gaseous phases using the B3LYP method. The frontier molecular orbitals (FMOs) and global reactivity parameters were computed, along with a visualization of the molecular electrostatic potential (MEP) of the molecules.
MD simulations
MD was performed on Desmond to determine the natural setting behavior and stability of the protein-ligand docked complex in real-time29. Initially, Maestro’s Protein Preparation Wizard was used to optimize and minimize the complex. The settings were done by the System Builder Tool, and the complexes were dissolved with the simple point-charge (SPC) solvent model in an orthorhombic box with dimensions of 10Åx10Åx10Å. The physiological conditions were mimicked by adding 0.15 M NaCl. The simulation trajectories were run under the OPLS4 force field at 300 K temperature and 1 atm pressure for NPT production. Nosé–Hoover chain coupling scheme and Martyna–Tuckerman–Klein chain coupling scheme, respectively, were used to control the temperature and pressure49. The trajectories of the docked complexes of o-vanillin and CCL were saved after every 100 ps during a 100 ns simulation. The stability and conformation were analyzed based on the root mean square deviation (RMSD) and root mean square fluctuation (RMSF) during the course of the simulation50. Lastly, Desmond trajectories were converted to XTC format for the computation of the radius of gyration using the gmx gyrate command of GROMACS51.
Principal component analysis (PCA) – based free energy landscape study (FEL)
PCA is a multivariate statistical technique performed by computing the protein’s internal motion denoted by eigenvalues and eigenvectors. The covariance matrix for the C-alpha (Cα) coordinates was generated over a 100 ns simulation trajectory52. PCA was conducted in Python language via the scikit-learn library53. Moreover, FEL analysis was conducted on principal components (PC1 and PC2) to gain insight into conformational states relying on the energy distribution throughout the complex using GROMACS.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Floyd, K., Glaziou, P., Zumla, A. & Raviglione, M. The global tuberculosis epidemic and progress in care, prevention, and research: an overview in year 3 of the end TB era. Lancet Respiratory Med. 6, 299–314. https://doi.org/10.1016/s2213-2600(18)30057-2 (2018).
Li, Y. Y., Cai, R. J., Talbot, E. A. & Wang, Y. T. in Molecular Medical Microbiology 1569–1584Elsevier, (2024).
Geneva Global Tuberculosis Report 2023., (2023).
Organization, W. H. Global Tuberculosis Report 2018WHO., (2018).
Selikoff, I. J., Robitzek, E. H. & Ornstein, G. G. Treatment of pulmonary tuberculosis with Hydrazide derivatives of Isonicotinic acid. J. Am. Med. Assoc. 150, 973–980. https://doi.org/10.1001/jama.1952.03680100015006 (1952).
Daniel, T. M. Rifampin–a major new chemotherapeutic agent for the treatment of tuberculosis. N. Engl. J. Med. 280, 615–616. https://doi.org/10.1056/nejm196903132801112 (1969).
Eldehna, W. M., Fares, M., Abdel-Aziz, M. M. & Abdel-Aziz, H. A. Design, synthesis and antitubercular activity of certain nicotinic acid Hydrazides. Molecules (Basel Switzerland). 20, 8800–8815. https://doi.org/10.3390/molecules20058800 (2015).
Kim, H. J. et al. Real-world experience of adverse reactions-necessitated rifampicin-sparing treatment for drug-susceptible pulmonary tuberculosis. Sci. Rep. 13, 11275. https://doi.org/10.1038/s41598-023-38394-1 (2023).
Bunalema, L., Fotso, G. W., Waako, P., Tabuti, J. & Yeboah, S. O. Potential of Zanthoxylum leprieurii as a source of active compounds against drug resistant Mycobacterium tuberculosis. BMC Complement. Altern. Med. 17 https://doi.org/10.1186/s12906-017-1602-x (2017).
Sharma, D. & Yadav, J. P. An overview of phytotherapeutic approaches for the treatment of tuberculosis. Mini Rev. Med. Chem. 17, 167–183. https://doi.org/10.2174/1389557516666160505114603 (2017).
Bahuguna, A. & Rawat, D. S. An overview of new antitubercular drugs, drug candidates, and their targets. 40, 263–292, (2020). https://doi.org/10.1002/med.21602
Khadka, D. et al. The use of medicinal plants to prevent COVID-19 in Nepal. 17, 26, (2021). https://doi.org/10.1186/s13002-021-00449-w
Fabricant, D. S. & Farnsworth, N. R. The value of plants used in traditional medicine for drug discovery. Environ. Health Perspect. 109 (Suppl 1), 69–75. https://doi.org/10.1289/ehp.01109s169 (2001).
Atanasov, A. G., Zotchev, S. B. & Dirsch, V. M. Natural products in drug discovery: advances and opportunities. 20, 200–216, (2021). https://doi.org/10.1038/s41573-020-00114-z
KARDI, C. & SEKIOU, S. Valorisation D’une Plante Poussant Spontanément Dans La Région De Tipaza (Algérie), Par L’analyse Phytochimique Et L’étude Des Activités Biologiques De Ses Extraits (Universite laarbi tebessi tebessa, 2020).
Chopra, B. & Dhingra, A. K. Natural products: A lead for drug discovery and development. 35, 4660–4702, (2021). https://doi.org/10.1002/ptr.7099
Sharma, M., Dhaliwal, I., Rana, K. & Phytochemistry Pharmacology, and toxicology of Datura Species-A review. 10, (2021). https://doi.org/10.3390/antiox10081291
Gaire, B. P. & Subedi, L. A review on the Pharmacological and toxicological aspects of Datura stramonium L. J. Integr. Med. 11, 73–79. https://doi.org/10.3736/jintegrmed2013016 (2013).
Islam, S. et al. Antibacterial potential of propolis: molecular docking, simulation and toxicity analysis. AMB Express. 14, 81. https://doi.org/10.1186/s13568-024-01741-0 (2024).
Alam, W., Khan, H., Khan, S. A., Nazir, S. & Akkol, E. K. Datura Metel: A review on chemical constituents, traditional uses and Pharmacological activities. Curr. Pharm. Design. 27, 2545–2557. https://doi.org/10.2174/1381612826666200519113752 (2021).
Maddeboina, K. et al. Recent advances in multitarget-directed ligands via in Silico drug discovery. Drug Discovery Today. 29 https://doi.org/10.1016/j.drudis.2024.103904 (2024).
da Fonseca, A. M. & Caluaco, B. J. Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking, and approach with molecular dynamics, RMSD, RMSF, H-Bond, SASA and MMGBSA. 66, 1919–1933, (2024). https://doi.org/10.1007/s12033-023-00831-x
Elkaeed, E. B., Yousef, R. G. & Design, Synthesis, Docking, D. F. T. MD Simulation Studies of a New Nicotinamide-Based Derivative: In Vitro Anticancer and VEGFR-2 Inhibitory Effects. 27, (2022). https://doi.org/10.3390/molecules27144606
Ahamed, F. M. et al. Molecular dynamics simulation, QSAR, DFT, molecular docking, ADMET, and synthesis of Ethyl 3-((5-Bromopyridin-2-yl) Imino) butanoate analogues as potential inhibitors of SARS-CoV-2. Polycycl. Aromat. Compd. 44, 294–312 (2024).
Mallikarjuna Reddy, G., Camilo, A. Jr. & Raul Garcia, J. Pyrrole-2,5-dione analogs as a promising antioxidant agents: microwave-assisted synthesis, bio-evaluation, SAR analysis and DFT studies/interpretation. Bioorg. Chem. 106, 104465. https://doi.org/10.1016/j.bioorg.2020.104465 (2021).
Ejaz, S. A., Aziz, M., Zafar, Z., Akhtar, N. & Ogaly, H. A. Revisiting the inhibitory potential of protein kinase inhibitors against NEK7 protein via comprehensive computational investigations. Sci. Rep. 13, 4304. https://doi.org/10.1038/s41598-023-31499-7 (2023).
Rahman, J., Tareq, A. M. & Biological Evaluation, D. F. T. Calculations and molecular Docking studies on the antidepressant and cytotoxicity activities of Cycas pectinata Buch.-Ham. Compounds 13 https://doi.org/10.3390/ph13090232 (2020).
Kawsar, S. M. et al. Chemical descriptors, PASS, molecular docking, molecular dynamics and ADMET predictions of glucopyranoside derivatives as inhibitors to bacteria and fungi growth. Org. Commun. 15, 203 (2022).
De Vivo, M., Masetti, M., Bottegoni, G. & Cavalli, A. Role of molecular dynamics and related methods in drug discovery. J. Med. Chem. 59, 4035–4061. https://doi.org/10.1021/acs.jmedchem.5b01684 (2016).
Yadav, P. & Challenges Solutions for recent advancements in Multi-Drugs resistance tuberculosis: A review. Microbiol. Insights. 16, 11786361231152438. https://doi.org/10.1177/11786361231152438 (2023).
Khan, M. U. et al. Identification of novel natural compounds against CFTR p.Gly628Arg pathogenic variant. 14, 99, (2024). https://doi.org/10.1186/s13568-024-01762-9
Sakhawat, A. et al. Natural compound targeting BDNF V66M variant: insights from in Silico Docking and molecular analysis. 13, 134, (2023). https://doi.org/10.1186/s13568-023-01640-w
Ur Rahman, M., Khan, M., Khan, S. W. & Khan, R. U. Novel schiff bases of Vanillin: potent inhibitors of macrophage harbored leishmania tropica. 47, 619–629, (2023). https://doi.org/10.1007/s12639-023-01594-7
Ravindran, R., Chakrapani, G., Mitra, K. & Doble, M. Inhibitory activity of traditional plants against Mycobacterium smegmatis and their action on filamenting temperature sensitive mutant Z (FtsZ)-A cell division protein. PloS One. 15, e0232482. https://doi.org/10.1371/journal.pone.0232482 (2020).
Rahgozar, N., Khaniki, B., Sardari, S. & G. & Evaluation of antimycobacterial and synergistic activity of plants selected based on cheminformatic parameters. Iran. Biomed. J. 22, 401–407. https://doi.org/10.29252/.22.6.401 (2018).
Thongdee, P. & Hanwarinroj, C. Virtual screening identifies novel and potent inhibitors of Mycobacterium tuberculosis PknB with antibacterial activity. 62, 6508–6518, (2022). https://doi.org/10.1021/acs.jcim.2c00531
Vieira, T. F. & Martins, F. G. In Silico Identification of Possible Inhibitors for Protein Kinase B (PknB) of Mycobacterium tuberculosis. 26, (2021). https://doi.org/10.3390/molecules26206162
Jayaraman, M., Gosu, V., Kumar, R. & Jeyaraman, J. Computational insights into potential marine natural products as selective inhibitors of Mycobacterium tuberculosis InhA: A structure-based virtual screening study. Comput. Biol. Chem. 108, 107991. https://doi.org/10.1016/j.compbiolchem.2023.107991 (2024).
Sakhawat, A. et al. In Silico and in vitro analyses to investigate the effects of vitamin C on VEGF protein. J. Taibah Univ. Med. Sci. 19, 775–789. https://doi.org/10.1016/j.jtumed.2024.06.008 (2024).
Nyambo, K. et al. Molecular docking, molecular dynamics simulations and binding free energy studies of interactions between Mycobacterium tuberculosis Pks13, PknG and bioactive constituents of extremophilic bacteria. Sci. Rep. 14, 6794. https://doi.org/10.1038/s41598-024-57124-9 (2024).
Martin, A. et al. Multicenter study of MTT and resazurin assays for testing susceptibility to first-line anti-tuberculosis drugs. Int. J. Tuberculosis Lung Disease: Official J. Int. Union against Tuberculosis Lung Disease. 9, 901–906 (2005).
Vilchèze, C. et al. Novel inhibitors of InhA efficiently kill Mycobacterium tuberculosis under aerobic and anaerobic conditions. Antimicrob. Agents Chemother. 55, 3889–3898. https://doi.org/10.1128/aac.00266-11 (2011).
Wang, T. et al. Mtb PKNA/PKNB dual Inhibition provides selectivity advantages for inhibitor design to minimize host kinase interactions. ACS Med. Chem. Lett. 8, 1224–1229. https://doi.org/10.1021/acsmedchemlett.7b00239 (2017).
Mishra, A., Mulpuru, V. & Mishra, N. An interaction network driven approach for identifying cervical, endometrial, vulvar carcinomic biomarkers and their Multi-targeted inhibitory agents from few widely available medicinal plants. 195, 6893–6912, (2023). https://doi.org/10.1007/s12010-023-04441-w
Modanwal, S., Mishra, A. & Mishra, N. An integrative analysis of GEO data to identify possible therapeutic biomarkers of prostate cancer and targeting potential protein through Zea mays phytochemicals by virtual screening approaches. J. Biomol. Struct. Dyn. 1–21. https://doi.org/10.1080/07391102.2023.2283163 (2023).
Rajput, D., Jain, D., Kashaw, S. K. & Patil, U. K. Molecular Docking studies on phytoconstituent isolated from Nyctanthes arbortristis Linn. Int. J. Pharm. Invest. 14, 399–408. https://doi.org/10.5530/ijpi.14.2.50 (2024).
Seeliger, D. & de Groot, B. L. Conformational transitions upon ligand binding: holo-structure prediction from apo conformations. PLoS computational biology 6, e1000634, (2010). https://doi.org/10.1371/journal.pcbi.1000634
Ikram, S., Ahmad, F., Ahmad, J. & Durdagi, S. Screening of small molecule libraries using combined text mining, ligand- and target-driven based approaches for identification of novel granzyme H inhibitors. J. Mol. Graph. Model. 105, 107876. https://doi.org/10.1016/j.jmgm.2021.107876 (2021).
Khan, M. F. et al. Exploring optimal drug targets through subtractive proteomics analysis and pangenomic insights for tailored drug design in tuberculosis. Sci. Rep. 14, 10904. https://doi.org/10.1038/s41598-024-61752-6 (2024).
Rehman, H. M. & Sajjad, M. Identification of NS2B-NS3 protease inhibitors for therapeutic application in ZIKV infection: A Pharmacophore-Based High-Throughput virtual screening and MD simulations approaches. 11, (2023). https://doi.org/10.3390/vaccines11010131
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinf. (Oxford England). 29, 845–854. https://doi.org/10.1093/bioinformatics/btt055 (2013).
Gosu, V. et al. Deciphering the intrinsic dynamics of unphosphorylated IRAK4 kinase bound to type I and type II inhibitors. Comput. Biol. Med. 160, 106978. https://doi.org/10.1016/j.compbiomed.2023.106978 (2023).
Morita, S. Chemometrics and related fields in Python. Anal. Sciences: Int. J. Japan Soc. Anal. Chem. 36, 107–111. https://doi.org/10.2116/analsci.19R006 (2020).
Acknowledgements
The authors gratefully acknowledge the financial support from the Higher Education Commission of Pakistan under the International Research Support Initiative Program (IRSIP).
Author information
Authors and Affiliations
Contributions
Study design and conceptualization, M.S.A., A.D.B., W.T.S.; Provision of materials and Experimental procedures, S.A.K., M.A.R., and Z.J.; Acquisition of data and formal analyses, H.E., M.A., and S.M.Q.; Writing—original draft, S.A.K., and M.U.K.; Visualization and BioRender figures, H.E., M.U.K., and S.M.Q.; Writing—review and editing, A.D.B, M.S.A., W.T.S., U.M.K., M.A.R., and S.A.K. All authors contributed to manuscript revision, read, and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khan, S.A., Rather, M.A., Jia, Z. et al. In vitro investigation of Datura innoxia phytocompounds against Mycobacterium tuberculosis H37Ra strain in association with in silico studies. Sci Rep 15, 33454 (2025). https://doi.org/10.1038/s41598-025-99053-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99053-1