Abstract
Computational methods for generating molecules with specific physiochemical properties or biological activity can greatly assist drug discovery efforts. Deep learning generative models constitute a significant step towards that direction. We introduce a novel approach that utilizes a Reinforcement Learning paradigm, called proximal policy optimization, for optimizing molecules in the latent space of a pretrained generative model. Working in the latent space of a generative model lets us bypass the need for explicitly defining chemical rules when computationally designing molecules. The generation of molecules is achieved through navigating the latent space for identifying regions that correspond to molecules with desired properties. Proximal policy optimization is a state-of-the-art policy gradient algorithm capable of operating in continuous high-dimensional spaces in a sample-efficient manner. We have paired our optimization framework with the latent spaces of two different architectures of autoencoder models showing that the method is agnostic to the underlying architecture. We present results on commonly used benchmarks for molecule optimization that demonstrate that our method has comparable or even superior performance to state-of-the-art approaches. We additionally show how our method can generate molecules that contain a pre-specified substructure while simultaneously optimizing for molecular properties, a task highly relevant to real drug discovery scenarios.
Similar content being viewed by others
Introduction
In drug discovery, a chemical compound should possess multiple properties in addition to biological activity, to be advanced as a clinical candidate. A compound identified to exhibit therapeutic activity, known as a hit, is being structurally modified by medicinal chemists to address liabilities, such as inadequate solubility and activity through a long and iterative process. During that stage, medicinal chemists apply transformations on the starting molecule to design analogues based on the intuition they develop or through reaction-based library enumeration. Given the vast size of chemical space, it becomes significantly challenging to exhaustively evaluate even a small region around a single molecule or scaffold to identify optimal candidates that satisfy a set of objectives. Computational methods for targeted molecule generation can greatly support the exploration of the chemical space in a time-efficient manner and recommend structures previously unexplored by chemists.
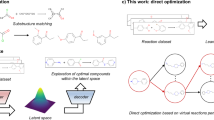
Machine learning and specifically generative deep learning models have been widely explored in chemistry, and in particular drug development, for de novo molecule generation1. Extensions of these methods have also been explored in targeted molecule generation, that is generation of molecules with desired properties, as well as molecule optimization for properties of interest2,3. Current approaches for targeted molecule generation and optimization can be divided into two major categories: those that operate directly on the molecular structure attempting at identifying structural modifications that improve the property of interest, and approaches that operate in the latent space of a generative model indirectly modifying the molecular structure through its latent representation.
The former approaches perform discrete space optimization often utilizing Reinforcement Learning (RL). Such methods can be applied directly on the molecular graph to identify actions, i.e. structural modifications such as inserting or deleting atoms or bonds, that will improve the property of interest4,5. The challenge here is that structural modifications may violate chemical rules resulting in invalid molecular structures, and therefore accounting for chemical knowledge through chemical rules or heuristics is deemed necessary3. An alternative approach is to operate at the substructure level by combining molecular fragments to compose a molecular graph6,7. Finally, sequence-based approaches have also been explored employing RL methods for controlling SMILES generation. Such approaches are built upon language models which offer the advantage of being pre-trained on vast chemical datasets and therefore capturing well the chemical language and validity8,9.
The second approach converts the optimization task into a continuous optimization problem utilizing the latent space of generative models and employing continuous space optimization algorithms such as gradient descent. Under this category, autoencoder architectures such as variational autoencoders are often utilized to form a continuous space, most often operating on the SMILES representation of molecules10. Models that attempt to perform controlled generation of molecular graphs, instead of SMILES, have also been explored11,12. Nevertheless, chemical validity remains a challenge as there is no guarantee that a point in the latent space corresponds to a valid molecule. However, significant advancements have been made in generative models by utilizing novel architectures as well as training modifications to improve validity and continuity of the latent space. Still, the use of continuous space optimization algorithms, such as gradient descent, assumes a smooth space with respect to the property of interest which does not naturally occur. To enforce a structured continuous latent space with respect to a specific property, joint training of generative models with property prediction models has been used10,11.
Other noteworthy approaches that do not strictly fall under the two aforementioned categories include the use of masked language modeling for conditional SMILES generation13, direct translation (graph or sequence based) of molecules into their improved version utilizing (semi) supervised learning, and finally fine-tuning of generative models through RL14.
Herein we introduce MOLRL (Molecule Optimization with Latent Reinforcement Learning), a novel framework for molecule optimization leveraging powerful pre-trained latent space generative models and RL. MOLRL performs optimization in the latent space of a pre-trained generative model using Proximal Policy Optimization (PPO)15, a state-of-the-art policy gradient RL algorithm. PPO can achieve a good trade-off between exploration and exploitation while also maintaining a trust region which is critical for searching challenging environments such as chemical latent spaces. The proposed framework is agnostic to the architecture of the generative model; however, the characteristics of the latent space can have a significant impact on the performance of the optimization method.
In the following, we discuss the properties that the latent space should possess to facilitate optimization, and we evaluate the performance of MOLRL following common literature benchmarks and performing a comparison with state-of-the-art methodologies. We evaluate MOLRL on a single-property constraint optimization task and a multi-objective optimization task for generating molecules predicted to be biologically active while improving molecular properties. Finally, we demonstrate how the introduced method can be effectively used for scaffold-constrained molecule optimization, a high-importance task in drug discovery scenarios.
The key contributions of this work are:
-
We introduce a novel methodology for targeted molecule generation in the latent space of a pre-trained autoencoder architecture. The method combines powerful generative models pre-trained on massive chemical datasets with state-of-the art RL algorithms for continuous space optimization.
-
We discuss the properties of the latent space that play a critical role in the introduced optimization framework. We additionally discuss the training modifications we applied on a variational autoencoder to improve those properties.
-
We demonstrate the effectiveness of the method for targeted molecule generation by applying it on tasks relevant to drug discovery and utilizing common benchmarks and performing a comparison with state-of-the-art methods.
Results & discussion
Evaluation of pre-trained models & latent space continuity
The effectiveness of the optimization approach will be impacted by the properties of the latent space and the performance of the underlying generative auto-encoder model. To evaluate the quality of the latent space and assess whether the auto-encoder models have been impacted by posterior collapse we investigated the following: i) reconstruction performance of the auto-encoder models, that is the ability of a model to retrieve a molecule from its latent representation, ii) validity rate, which shows how likely it is for a model to generate a valid SMILES, and iii) continuity of the latent space , which reflects the density of the latent space and how the molecules are clustered in the latent space.
Reconstruction and validity
The validity rate will impact training of the RL agent as visiting invalid states will provide no feedback to the agent. The reconstruction rate shows whether the latent vectors have captured sufficient information to reconstruct a molecule which is a necessary requirement for performing optimization in the latent space.
For evaluating reconstruction performance of each model, we sampled 1000 random molecules from the ZINC database not seen during training for both models and encoded each molecule to its respective latent variable \(z_0\) which in turn was decoded back into a molecular structure. As reconstruction performance we report the average Tanimoto similarity between the original molecules and the decoded molecules.
Regarding the validity rate of each model, we sampled 1000 latent vectors from a Gaussian distribution N(0,1) and decoded each vector into a SMILES and subsequently used the RDKit software to parse the SMILES into molecules and assess whether the generated SMILES are syntactically valid. As validity rate, we report the ratio of valid decoded molecules in the 1000 sampled molecules.
Table 1 shows the reconstruction rate and validity of the two models that were paired with the optimization framework. Specifically for the VAE model, we show the model’s performance with two different types of learning rate annealing, logistic and cyclical. In the VAE model trained with logistic annealing we can see the impact of posterior collapse with the model losing its ability to reconstruct molecules from latent representations. The introduction of cyclical annealing though has mitigated this problem bringing a good balance between reconstruction and validity. Similarly, the MolMIM model shows no signs of posterior collapse demonstrating high reconstruction performance and validity rate. In the experiments presented hereafter, we utilize the VAE model trained with cyclical annealing (referred as VAE-CYC) and the MolMIM model.
A more thorough analysis of the VAE performance for different annealing schedules is included in the SI (section S1.3).
Latent space continuity
Next, we evaluate the continuity (or smoothness) of the latent spaces of the two generative models (VAE and MolMIM). The continuity analysis investigates how the molecules are clustered in the latent space. In a continuous space, small perturbations of latent vectors will lead to structurally similar molecules which in turn will enable efficient optimization.
We evaluate continuity as follows: We encode 1000 random molecules from ZINC database to their respective latent variable \(z_0\). Then we perturb the latent variables by adding Gaussian noise with variance \(\sigma\) in a stepwise manner. Finally, we decode the perturbed variables to the respective molecules and measure the Tanimoto similarity between the new molecules and the original ones. The average Tanimoto similarity per step for various levels of noise variance \(\sigma\) is presented in Fig. 1. Since the evaluation presented here only indicates how small perturbations affect structural similarity rather than evaluating generalization, the test molecule overlap with the training datasets minimally affects the continuity.
We observe a sharp decrease in Tanimoto similarity for the VAE models when the added Gaussian noise has variance \(\sigma = 0.25\) and \(\sigma = 0.5\) which means that the perturbed molecules are further structurally from the original ones. However, variance \(\sigma =0.1\) results in a smoother decline of the similarity and therefore improved latent space continuity. Regarding the MolMIM model, Gaussian noise with \(\sigma = 0.25\) and \(\sigma = 0.5\) results in smooth continuity while with \(\sigma =0.1\) the original molecules seem to remain unperturbed.
These findings demonstrate that the latent spaces of the two selected models exhibit continuity and therefore can serve as the space where continuous optimization can be performed.
Average Tanimoto similarity drop after successively applying Gaussian noise to a set of starting molecules with various values of standard deviation \(\sigma\)
Based on the analysis above, the VAE with cyclical annealing schedule (VAE-CYC) and the MolMIM model were selected as our latent models for our downstream tasks.
Single property optimization under constraints
For the evaluation of the optimization performance of MOLRL, we start by adopting a widely used benchmark on constrained optimization introduced by Jin et al.11. The task is to improve the penalized LogP (pLogP) value for a set of 800 molecules while maintaining structural similarity to the original molecules. The pLogP value is defined as the octanol-water partition coefficient, which is a measure of molecular hydrophilicity, penalized by two scores corresponding to synthetic accessibility and presence of long cycles, respectively. The results are reported in Table 2. It should be noted that although this is a popular benchmark in the literature for evaluating molecule optimization approaches, it suffers from a known limitation: an increase in the pLogP value can be achieved by simply increasing the number of carbon atoms (additional information in SI section S4).
Despite this limitation, we use this benchmark with caution. In particular, if a large number of carbon atoms is forcefully added to increase the pLogP value, this will be reflected in decreased drug-likeness (QED)16 and synthetic accessibility (SA)17 scores. Therefore, we calculate the average changes in the QED and SA scores (calculated using the RDKit package) in addition to the metrics commonly reported in the literature, i.e. success rate and average property improvement. Success rate is measured as the percentage of molecules for which the pLogP has improved (with respect to the starting molecule \(\Delta\)pLogP>0) while at the same time structural similarity (measured as Tanimoto coefficient on Morgan Fingerprints) is greater or equal than a prespecified cutoff, \(\delta\) Average improvement of the pLogP value is computed among the molecules that have succeeded. We report the average improvement and success rate at two similarity cutoffs (\(\delta\) = 0.4 and 0.6) for MOLRL, when paired with a VAE-CYC model and when paired with a MolMIM model, as well as for a set of state-of-the-art molecule optimization algorithms as reported in the literature. We compute the penalized LogP value using the TDC software package (which provides a normalized pLogP value)18,19. Additionally, we note that the original studies for MolDQN and GCPN measure success rate differently (accounting only for the similarity constraint) while the Back Translation study does not report success rate nor standard deviation of property improvement.
We observe that MOLRL in the MolMIM space results in a higher improvement of the average pLogP value while the MOLRL in the VAE space results in a higher success rate with a starker difference at higher structural similarity cutoff \(\delta\). These differences in the MOLRL performance in the two latent spaces can be explained by the different degree of latent space continuity between the two models (as shown in Figure 1). The MolMIM model has a latent space with a higher degree of continuity, resulting in a denser latent space. This density shapes the environment in which the RL agent operates, indirectly influencing its experiences and thus potentially affecting exploration and exploitation dynamics. On the other hand, less continuity in the VAE space results in a more diverse environment providing the agent with experiences from a broader range of explored states. In summary, a step taken by the agent in a denser space results in more similar molecules. Higher exploitation can result in a higher property improvement while higher exploration can be beneficial for molecules that cannot be improved with small structural changes.
Regarding the method performance in the context of the literature benchmarks, MOLRL on either latent space has achieved success rates on par with the best performing method that is Regression Transformer. Regression Transformer is a language-based model trained on regression tasks using masked language modeling and can additionally be used for conditional molecule generation. This is a stark contrast to the MOLRL method which combines an optimization algorithm with a general-purpose generative model.
Regarding the average property improvement, MOLRL in the MolMIM space has comparable performance with the best performing methods without sacrificing desirable molecular properties as shown in Table 3. Table 3 shows the average changes in the QED and SA scores among the succeeded molecules for MOLRL and MolDQN as a percentage change with respect to the starting molecules scores. As a baseline here we chose an approach that applies RL directly on the molecular structure as opposed to MOLRL which modifies molecules through their latent representation. (MolDQN was chosen here for the additional reason of being the only method we were able to reproduce reported results). We observe that MOLRL compromises QED to improve pLogP however at a significantly smaller extent than MolDQN. On the other side, the SA score is being improved (lower SA score indicates easier synthesis) with a bigger change for the model operating in the MolMIM space. MolDQN though compromises both scores, QED and SA, to achieve higher pLogP values. The smaller effect on QED and the improvement of SA that are observed for MOLRL are a result of working in the latent space of a model that has been trained on a large chemical dataset and therefore has captured the distribution of the properties of the molecules it has been trained on. In the literature, QED and SA changes have not been reported for this benchmark. However, it has been observed that existing approaches tested on the pLogP benchmark often learn a simple policy of just adding carbon chains4.
Multi-objective generation towards biological activity
Next, we evaluate MOLRL on multi-parameter optimization, and we perform a comparison with the current state-of-the-art optimization methods. In particular, we apply MOLRL to generate molecules with biological activity for two targets and at the same time we optimize for drug likeness (QED) and synthetic accessibility (SA), following the benchmark adopted by Jin et al.6. The selected biological targets are two kinases, GSK3\(\beta\) and JNK3, both associated with Alzheimer’s disease. The binding affinity to these targets is estimated using Machine Learning (ML) classification models developed by Li et al.21 that predict the binding probabilities and are available through the PyTDC software package19.
We additionally optimize molecules for binding to each target separately and for dual binding to both targets (without optimizing for QED and SA) and present these experiments and results in the SI (section S5).
We train one RL agent for each latent-space model (VAE-CYC and MolMIM) for 500 epochs and a batch size of 100 with an initial seed of 5 vectors sampled from a Gaussian distribution being the starting states. We additionally apply a restart strategy every 100 epochs to reset the optimization from the top-10 explored states. We additionally run MOLRL for a larger number of epochs (2000) showing that running the optimization longer yields better results.
The reward function for optimizing towards the two targets and the two properties, QED and SA, is defined as:
Where \(w_1 = 0.5\) and \(w_2 = 0.3\) were selected experimentally.
Molecules generated by MOLRL along with the predicted binding probabilities and the closest structurally known inhibitors.
Following the evaluation strategy by Jin et al.11, we record the top-5000 highest reward molecules generated during the optimization process and compute the following metrics: i) success rate ii) novelty, and iii) diversity. Success rate is computed as the percentage of molecules that have succeeded in the top-5000. A generated molecule is deemed successful if the predicted probabilities for binding towards each target are greater or equal to 0.5, and the following cutoffs for the QED and SA scores are satisfied: \(QED \ge 0.6\) and \(SA \le 4\). Novelty examines the closeness of the generated molecules to the active molecules used for training the predictive classification models for activity to GSK3\(\beta\) and JNK3. In particular, it is computed as the fraction of molecules with nearest neighbor similarity, with respect to the train set, lower than 0.4 among the succeeded ones. Finally, diversity examines how similar or dissimilar structurally the succeeded molecules are. In particular it is computed by subtracting the average pairwise similarities from 1 with higher values indicating lower average similarity and therefore higher diversity. The molecular similarity is calculated as the Tanimoto similarity on Morgan fingerprints.
Table 4 shows the performance of MOLRL trained in the latent space of the VAE-CYC and MOLRL trained in the MolMIM space as well as the performances of state-of-the-art molecule optimization methods as reported in the literature6.
We observe that FaST, which builds molecular graphs by combining fragments using RL, shows a higher success rate across all compared method. FaST and RationaleRL have an advantage in terms of diversity and novelty. Both are graph generation approaches that utilize a set of substructures as building blocks to compose molecular graphs. The substructures are extracted from known active molecules which means that both methods start from a good starting point taking advantage of prior knowledge. Furthermore, it should be noted that the scoring function that guides the optimization across all approaches is based on ML models and therefore the accuracy of the function depends on how far the generated molecules are from the dataset used to develop the ML models22. Given that RationaleRL utilizes substructures extracted from the known inhibitors which have come from the dataset used to train the ML models, the scoring function is expected to be more accurate. On the other side, both REINVENT and MOLRL start from random molecules which could be far from what the ML classifiers have been trained on. Despite that, MOLRL achieves comparable novelty with RationaleRL while FaST has highest overall performance on this task. We additionally note that uniqueness of generated molecules has not been directly accounted for in this benchmark across previous methods; therefore, we report the novelty and diversity metrics on molecules that have satisfied the success criteria and uniqueness is implicitly reflected in the diversity metric.
Utilizing prior knowledge as a starting point can provide an advantage by guiding the optimization process; however, it may also constrain the search space, furthermore this reduces the method’s applicability in scenarios where prior information is not available. In our current benchmarks this effect is not strongly reflected, however we note that the machine learning scoring function itself may inherently encode a bias towards known scaffolds.
In Fig. 2, we visualize two examples from the highest-reward molecules generated by MOLRL (VAE-CYC). The closest structurally known dual inhibitor is visualized next to each generated molecule with the maximum common substructure between the generated and the known inhibitor being highlighted. The common substructures are known hinge motifs in kinase inhibitors. Although MOLRL starts from random molecules, it generates molecules that contain the hinge motifs that are present in known inhibitors guided by the reward function.
Scaffold guided multi-parameter optimization
A common practice in drug discovery is to identify a chemical scaffold that is known to bind to a target or class of targets and use it as a starting point for chemical design and optimization. In the following experiment, we evaluate our method for scaffold-guided molecule optimization, that is, generation of molecules that contain a given substructure and have improved drug-like properties.
In particular, we focus on generating molecules containing the aminopyrimidine substructure (Fig. 3) which is a known hinge binding motif present in certain approved kinase inhibitors and is a commonly used substructure for the design of molecules across numerous therapeutic areas23.
We train the RL agent by rewarding for the presence of the aminopyrimidine substructure and we additionally reward drug-likeness (QED score) and synthetic accessibility (SA).
The reward function for this task is formulated as:
2-Aminopyrimidine substructure
\(w_1\) and \(w_2\) are weights introduced to control the QED and SA scores importance in the reward and are set at 0.5. The QED score range is [0,1] and we normalized the SA score such that it also takes values in the range [0,1] with 0 being difficult to synthesize and 1 being easy to synthesize.
For this experiment, we performed optimization in the latent space of the MolMIM model. The RL agent was trained for 100 epochs with a batch size of 100. We evaluated the performance of the method for this experiment by extracting the following metrics on the top-1000 scored molecules: i) uniqueness, defined as the percent of distinct canonical SMILES, ii) diversity, measured as the average pairwise similarity among the top-1000, and iii) success, measured as the percentage of molecules containing the 2-aminopyrimidine substructure. The results are reported in Table 5. We add the two following baselines: random sampling on the prior, and CMA-ES, a gradient-free greedy search algorithm24. We configured CMA-ES to run with a population size of 20, 5 random initial molecules, 10 iterations, and 10 restarts and \(\sigma = 1\), as implemented in the BioNemo framework.
For all experiments, we fixed the starting initial states to the same 5 randomly selected molecules sampled from ZINC and fixed the total number of reward function calls to 10,000. We observe that CMA-ES has lower success rate than MOLRL but higher diversity while MOLRL outperforms CMA-ES on success and uniqueness.
Additionally, we investigated the effect of the parameter \(\sigma\) for values 0.25, 0.5 and 1.0 on the agent training convergence for 1000 epochs in the VAE-CYC space as shown in Fig. 4. We observe that the agent converges faster with smaller standard deviation values which further reinforces our observation that in this case the exploration is limited, and the agent exploits solutions early on during training. The results are reported in Table 6. Figure 5 shows the evolution of each one of the 3 components of the reward function (QED score, SA score and presence of aminopyrimidine substructure as a binary parameter) during training for \(\sigma =1.0\). We observe that the agent learns to generate molecules with the desired structure while simultaneously improving or maintaining QED and SA.
Mean reward improvement over optimization epochs for different values of \(\sigma\)
Evolution of each parameter in multi-parameter optimization over the optimization epochs
Top molecules generated at the last optimization epoch with the aminopyrimidine structure highlighted.
Finally, in Fig. 6, we show the top-scored molecules generated at the last optimization epoch along with the QED and SA score. The molecules show an overall good range of QED and SA scores while retaining the 2-aminopyrimidine substructure highlighted in orange.
The RL agent can effectively exploit regions of the latent space that satisfy the reward function while satisfying multiple criteria simultaneously. The diversity of the generated molecules demonstrates the ability of the RL agent to explore several solutions in the latent space. Since PPO maintains benefits of trust region policy optimization (TRPO) we can efficiently control the generation process in regions of latent space of interest. We note that we do not need to pre-define chemical modification rules and the agent reduces the complexity of explicit chemical modification to vector modification in the latent space.
Conclusions
We have introduced MOLRL, a novel approach for targeted molecule generation combining powerful generative models and Reinforcement Learning. Molecular optimization takes place in the latent space of an autoencoder generative model trained on molecules. The latent space provides a continuous representation of molecules suitable for facilitating optimization tasks and bypassing the need for explicitly accounting for chemical validity through pre-defined chemical rules or heuristics. MOLRL shows superior or competitive performance across a variety of tasks for targeted molecular generation and multi-parameter optimization when compared with established methods.
With the continuous advances in generative models, the latent spaces are becoming denser covering a larger chemical space. At the same time, we are witnessing an increasing adoption of these models in drug discovery. Being able to efficiently navigate these spaces to identify novel molecules with desirable properties can greatly assist the drug discovery process.
Nevertheless, a significant roadblock for adopting methodologies based on generative ML models in practical applications in small molecule drug discovery is synthesizability of generated molecules.
As we are moving beyond the known chemical space generating novel structures, synthesizability is not guaranteed. This could be addressed either at the pretraining stage, when the latent space is being formed, or during the optimization phase as part of the reward function. However, the effectiveness of the tools for estimating synthesizability of generated molecules remains inadequate. Optimization efficiency - that is, the degree of exploration needed to reach an optimal solution - is an additional challenge for practical applications, especially when accommodating computationally demanding reward functions such as molecular coupling.
Finally, to assess diversity during optimization, we employed internal diversity for consistency with prior benchmarks; however, we acknowledge its limitations in capturing scaffold-level diversity and highlight newer metrics like #Circles25 as important directions for future work.
Methods
MOLRL consists of two major components: a latent space generative model, and a RL agent. The generative model is a pre-trained encoder-decoder model whose latent space encodes the chemical space where the RL agent operates in. The RL agent is being trained to navigate the latent space using PPO, a policy gradient RL algorithm. The reward function provides feedback to the agent for learning how to navigate the space to identify molecules with desired properties. During training, only the RL agent is being trained while the generative model has been pre-trained and its weights are fixed. In the following, we discuss each component in more detail.
Pre-trained generative model
The generative model facilitates the connection between the chemical space and the latent space and provides a continuous representation for a molecule. It is an encoder-decoder architecture, with the encoder translating each chemical molecule into a latent vector z in a multi-dimensional continuous space, and the decoder retrieving the molecular structure from a latent vector \(z^{\prime }\). The optimization takes place in the latent space and the optimized structure is being retrieved using the decoder.
The optimization framework is agnostic to the architecture of the encoder and the decoder. Nevertheless, the properties of the latent space will greatly impact the optimization performance. In particular, the existence of “holes” in latent space or regions of the space with invalid molecules, will impact the ability of the agent to interpolate into the space. An additional parameter is the continuity or smoothness of the latent space which will facilitate optimization of molecules through a local search. Finally, the “coherence” between the encoder and decoder, that is the generation of molecules conditionally on the latent representation as opposed to random sampling, can greatly impact the ability to control the generation of molecules with desired properties. Certain encoder-decoder architectures are known to suffer from a phenomenon known as posterior collapse which results in a decoder that generates molecules independently of the latent variable26.
In this work, we have evaluated the performance of MOLRL on two different encoder-decoder architectures, a variational autoencoder (VAE)27 and an auto-encoder trained with Mutual Information Machine (MIM) learning28. The two models utilize a different loss function during training and therefore they result in differently structured latent spaces. In the following, we discuss the two architectures and provide remarks on the efforts to improve the properties of the latent space.
Variational autoencoder
The variational autoencoder (VAE)27 gives a probabilistic representation of a molecule in the latent space with each attribute of a latent vector corresponding to a distribution. We have implemented both the encoder and decoder as gated recurrent units (GRU) and we make use of the SMILES notation29 for representing chemical molecules. We have adapted an architecture similar to what is reported by Bowman et al.30.
The loss function of a VAE model has two terms: the reconstruction loss \(L_E\) and the Kullback-Leibler (KL) divergence \(L_R\) between the learned distribution and the true distribution which we assume to be a Gaussian distribution N(0,1). We adopt the \(\beta\)-VAE loss31 which introduces a \(\beta\) parameter in the KL term:
The training objective is to maximize the lower bound on the marginal likelihood of a data point x:
The encoder \(q_{\phi } (z|x)\) transforms input data x into a distribution over latent variables z and the decoder \(p_{\theta } (x|z)\) reconstructs data x from the latent variable z, with \(\phi\) and \(\theta\) being the encoder and decoder parameters, respectively.
One difficulty when training a VAE is the vanishing of the KL term where \(q_{\phi } (z|x)\) essentially becomes equal to p(z) reducing \(q_{\phi } (z|x)\) to a point estimate. This is known as posterior collapse and when it occurs the latent space becomes uninformative32 and therefore not amenable for molecular optimization.
To mitigate posterior collapse, we applied a cyclical annealing training schedule32 on the \(\beta\) term of the KL term \(L_R\). Cyclical annealing schedule is formulated as:
Where M, is the number of cycles, R is the proportion used to increase \(\beta\) within a cycle, T is the total number of training steps, t is the current step in training and f is a monotonically increasing function. For our training we used R = 0.7, M = 20. These parameters were selected by hyperparameter tuning using the reconstruction and validity of the trained models as criteria (as defined in section 4).
The VAE model was trained on 10 million molecules from the PubChem database33 made available by Chithrananda et al.34. The molecules were standardized to canonical SMILES without stereochemistry using the RDKit package35.
Additional information on the architecture of the VAE and the training scheme used, is provided in the SI (section S1).
MolMIM
MolMIM is a probabilistic transformer-based encoder-decoder model trained with a Mutual Information Machine (MIM) loss. Contrary to the VAE loss which minimizes the divergence between the latent space distribution and the prior, the MIM loss maximizes the mutual information between an observation and its latent representation. This loss prevents the decoder from ignoring the latent variable in decoding which in turn naturally prevents posterior collapse. Additionally, the loss function in the MolMIM model imposes a low marginal entropy in the latent representation which encourages the formation of a well clustered latent space.
We have adopted the publicly available MolMIM model introduced by Reidenbach et al.28 and made available through NVIDIA BioNemo an open-source framework for model development in drug discovery36. This model has been trained on 1B chemical molecules from the ZINC database using the following loss function for a given data point x:
With the model encoder being formulated as \(q_{\theta }(z|x)\) and the model decoder being \(p_{\theta } (x|z)\) and P(x) being the distribution of the training dataset and P(z) the latent space prior.
Reinforcement learning
MOLRL methodology: the latent representation z of an input molecule is being perturbed through an action a sampled from the policy network output. The perturbed latent vector \(z^{\prime }\) is being decoded into molecules which are scored using the reward function. The states z, actions a and rewards R are being collected to update the policy network
Optimization of molecules takes place in the latent space of an encoder-decoder model through a RL paradigm. An overview of the MOLRL methodology is presented in Figure 7. An RL agent interacts with the environment (the latent space) through a Markov decision process (MDP) defined by a set of states (\(s_t \in S\)), a set of actions (\(a_t \in A\)), rewards (\(R_t \in R\)) and transition probabilities \(P({s_{t+1} | s_t, a_t})\). The states correspond to points in the latent space, the actions determine how a latent vector will be modified and finally the rewards are determined by a (task-specific) predefined reward function which evaluates the outcome of each action.
The initial state \(s_0\) is selected either as a fixed starting molecule encoded by the latent model’s encoder \(s_0=z_0\) or seed vectors sampled from a Gaussian distribution representing random points in the latent space. For optimization applications where a starting set of molecules is available and the task is to further improve properties or activity, the first option is chosen. However, when designing molecules with specific activity or properties and no starting point is available, then the initial states are random points in the latent space.
Since we are not concerned with stepwise transitions of the agent, we treat transitions as single final actions, where the maximum number of steps we consider is \(t=1\) per state. Accordingly, rewards \(R_t\) are computed as single step rewards ignoring discounted rewards calculation, a practice that is common in policy gradient optimization.
The policy network is composed of two modules, (1) the actor module denoted as \(\pi _{\theta }\) which outputs a mean vector \(\mu _{\theta }\) given a state and (2) the critic network denoted as \(Q_{\theta }\), which outputs an estimate of the reward at a given state, since the two modules share a hidden layer the actor-critic network parameters are denoted by \(\theta\). Details about the actor-critic network architecture are included in supplementary information (S2).
At each epoch, a batch of actions is sampled from a multivariate Gaussian distribution \(N(\mu _{\theta },\Sigma )\) parameterized by the policy network \(\pi _{\theta }\). The diagonal covariance matrix \(\Sigma\) is non-parameterized and is specified as \(\Sigma = \sigma I^2\) with \(\sigma\) being the standard deviation parameter. The transition dynamics is defined as follows, where \(\mu _{\theta }\) is the mean vector output of the actor network \(\pi _{\theta }\)
The RL agent traverses the latent space by adding Gaussian noise to its current position with a mean value parameterized by the policy network.
The parameter \(\sigma\) in the covariance matrix controls the agent’s exploration-exploitation trade-off with larger values allowing more exploration of the space while lower values are resulting in a more local search. We reduce exploration over time to allow convergence of the agent’s loss function by introducing a decaying factor \(\gamma\) for the standard deviation \(\sigma\) with being set to \(\gamma =0.9995\) for all experiments.
The agent is being trained by collecting a batch of experiences from the environment in the form of \((a_t, s_t, R_t)\) at time t and the policy parameters are updated by computing the proximal policy optimization (PPO) loss15 for N off-line update steps. Advantages \(A_t\) are estimated by the critic \(Q_\theta\) such that \(A_t = Q_\theta (s_t, a_t) - R_t\) with \(R_t\) having been normalized to 0 mean and standard deviation of 1 for each batch.
The PPO loss \(L^{PPO}\) contains three terms and is calculated as:
With \(L^{clip}\) being the clipped surrogate loss (actor loss) \(c_1\) was set to 0.5 and \(c_2\) was set to 0.01, \(L^{VF}\) being the value function loss (critic loss), and \(L^{Entropy}\) the entropy regularization term, defined as follows:
Where \(r_t(\theta )\) is the ratio of the parameters between the updated policy parameters and old parameters, where old parameters are considered the parameters for the last update step and calculated as:
and \(\epsilon\) being the clipping ratio threshold and set to 0.2. The clipped loss prevents overly large updates to the network parameters and mimics a trust region policy15. Additional benchmarks on the effects of clipping on training are included in the Supplementary information (S6).
\(L^{VF}\) is computed as the mean squared error between the estimated reward by the critic and the actual reward:
and finally, \(L^{Entropy}\) is the entropy regularization term to encourage exploration and is weighted by a parameter c:
with \(\sigma\) being the standard deviation parameter.
Noting that \(\sigma\) has a major impact on the exploration-exploitation trade-off and can be either set to a prespecified value or parameterized by the policy network. When \(\sigma\) is pre-specified, the entropy regularization term in the PPO loss function becomes ineffective since \(\sigma\) is fixed and independently controlled. In the latter case where \(\sigma\) is being parametrized by the policy network, the entropy regularization term has a larger effect on exploration-exploitation.
In the case where \(\sigma\) is being set to a prespecified value, we introduce a decay factor \(\gamma\) (with a prespecified value of 0.9995) for reducing \(\sigma\) over time to control exploration and allow convergence of the agent’s loss function making the agent’s action become more deterministic as training proceeds.
All states visited throughout the optimization procedure and respective rewards are being recorded and finally the top-k generated molecules with the highest rewards are selected.
As an additional way to control the exploration-exploitation trade-off, we introduce a restart strategy according to which the optimization process restarts every N epoch from the best explored states, that is the starting states at every restart are the highest reward states visited up to that point.
An ablation study with the PPO algorithm being replaced by the REINFORCE37 algorithm, a standard policy gradient RL algorithm, is presented in the supplementary material (section S3), showing that PPO is more efficient in the latent space of both pre-trained models.
Data availability
The VAE and MolMIM models have been trained and evaluated on public datasets as referenced in the manuscript. The data used in the single property optimization benchmark and in the multi-object molecule generation benchmark are also publicly available and referenced in the manuscript. The MOLRL code has been submitted as supplementary material for the review process and will be made available through a code repository upon publication.
References
Anstine, D.M. & Isayev, O. Generative models as an emerging paradigm in the chemical sciences. J. Am. Chem. Soc. 145(16), 8736–8750 (2023) (Publisher: American Chemical Society).
Xia, Y. A comprehensive review of molecular optimization in artificial intelligence-based drug discovery - Xia - 2024. In Quantitative Biology (Wiley Online Library, 2024).
Zhang, O. et al. Deep lead optimization: Leveraging generative AI for structural modification. J. Am. Chem. Soc. (2024). (Publisher: American Chemical Society).
Zhou, Z., Kearnes, S., Li, L., Zare, R.N. & Riley, P. Optimization of molecules via deep reinforcement learning. Sci. Rep. 9(1), 10752 (2019) (Publisher: Nature Publishing Group).
You, J., Liu, B., Ying, R., Pande, V., & Leskovec, J. Graph convolutional policy network for goal-directed molecular graph generation. In 32nd Conference on Neural Information Processing Systems (NeurIPS 2018) (2018).
Jin, W., Barzilay, R. & Jaakkola, T. Multi-Objective Molecule Generation using Interpretable Substructures. arXiv:2002.03244 [cs, stat] (2020).
Chen, B., Fu, X., Barzilay, R., & Jaakkola, T. Fragment-Based Sequential Translation for Molecular Optimization. arXiv:2111.01009 (2021).
Mazuz, E., Shtar, G., Shapira, B. & Rokach, L. Molecule generation using transformers and policy gradient reinforcement learning. Sci. Rep. 13(1), 8799 (2023) (Publisher: Nature Publishing Group).
Olivecrona, M., Blaschke, T., Engkvist, O. & Chen, H. Molecular de-novo design through deep reinforcement learning. J. Cheminform. 9(1), 48 (2017).
Gómez-Bombarelli, R. et al. Automatic chemical design using a data-driven continuous representation of molecules. ACS Central Sci. 4(2), 268–276 (2018) (Publisher: American Chemical Society).
Jin, W., Barzilay, R., & Jaakkola, T. Junction tree variational autoencoder for molecular graph generation. In Proceedings of the 35th International Conference on Machine Learning. 2323–2332. (PMLR, 2018). (ISSN: 2640-3498).
Zang, C., & Wang, F. MoFlow: An invertible flow model for generating molecular graphs. In Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, KDD ’20, New York, NY, USA. 617–626 (Association for Computing Machinery, 2020).
Born, J. & Manica, M. Regression transformer enables concurrent sequence regression and generation for molecular language modelling. Nat. Mach. Intell. 5(4), 432–444 (2023) (Publisher: Nature Publishing Group).
Sob, U.A., Mbou, L., Qiulin, A., Miguel, B., Oliver, S., Andries P., & Pretorius, A. Generative Model for Small Molecules with Latent Space RL Fine-Tuning to Protein Targets (2024). arXiv:2407.13780.
Schulman, J., Wolski, F., Dhariwal, P., Radford, A. & Klimov, O. Proximal Policy Optimization Algorithms. arXiv:1707.06347 [cs] (2017).
Bickerton, G. Richard., P., Gaia V., Besnard, J., Muresan, S. & Hopkins, A.L. Quantifying the chemical beauty of drugs. Nat. Chem. 4(2), 90–98 (2012) (Publisher: Nature Publishing Group).
Ertl, P. & Schuffenhauer, A. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. J. Cheminform. 1(1), 8 (2009).
Therapeutics Data Commons. https://tdcommons.ai/.
Huang, K., Fu, T., Gao, W., Zhao, Y., Roohani, Y., Leskovec, J., Coley, C.W., Xiao, C., Sun, J., & Zitnik, M. Therapeutics data commons: Machine learning datasets and tasks for drug discovery and development. In 35th Conference on Neural Information Processing Systems (NeurIPS 2021) (Track on Datasets and Benchmarks, 2021).
Fan, Y. et al. Back translation for molecule generation. Bioinformatics 38(5), 1244–1251 (2021) (12).
Li, Y., Zhang, L. & Liu, Z. Multi-objective de novo drug design with conditional graph generative model. J. Cheminform. 10(1), 33 (2018).
Weaver, S. & Gleeson, M.P. The importance of the domain of applicability in QSAR modeling. J. Mol. Graph. Model. 26(8) (2008). (Publisher: J Mol Graph Model).
Filho, E.V., Pinheiro, E.M.C., Pinheiro, S. & Greco, S.J. Aminopyrimidines: Recent synthetic procedures and anticancer activities. Tetrahedron 92, 132256 (2021).
Hansen, N. The CMA Evolution Strategy: A Tutorial. arXiv:1604.00772 (2023).
Renz, P., Luukkonen, S. & Klambauer, G. Diverse hits in de novo molecule design: Diversity-based comparison of goal-directed generators. J. Chem. Inf. Model. 64(15), 5756–5761 (2024).
Yan, C., Yang, J., Ma, H., Wang, S. & Huang, J. Molecule sequence generation with rebalanced variational autoencoder loss. J. Comput. Biol. 30(1), 82–94 (2023) (Publisher: Mary Ann Liebert, Inc., publishers).
Kingma, D.P., & Welling, M. Auto-encoding variational Bayes. In 2nd International Conference on Learning Representations, ICLR 2014, Banff, AB, Canada, April 14-16, 2014, Conference Track Proceedings (2014).
Reidenbach, D., Livne, M., Ilango, R.K., Gill, M. & Israeli, J. Improving Small Molecule Generation using Mutual Information Machine (2023). arXiv:2208.09016 (Published at the MLDD workshop, ICLR 2023).
Weininger, D. Smiles, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28(1), 31–36 (1988).
Bowman, S.R., Vilnis, L., Vinyals, O., Dai, A.M., Jozefowicz, R. & Bengio, S. Generating Sentences from a Continuous Space (2015).
Higgins, I. et al. Beta-VAE: Learning basic visual concepts with a constrained variational framework. In International Conference on Learning Representations (2016).
Fu, H., Li, C., Liu, X., Gao, J., Celikyilmaz, A. & Carin, L. Cyclical annealing schedule: A simple approach to mitigating KL vanishing. In (Burstein, J., Doran, C. & Solorio, T. Ed.), Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers), Minneapolis, Minnesota. 240–250. (Association for Computational Linguistics, 2019).
Kim, S. et al. Pubchem in 2021: New data content and improved web interfaces. Nucleic Acids Res. 49(D1), D1388–D1395 (2020) (11).
Chithrananda, S., Grand, G., & Ramsundar, B. ChemBERTa: Large-Scale Self-Supervised Pretraining for Molecular Property Prediction. arXiv (2020).
RDKit: Open-Source Cheminformatics Software. https://www.rdkit.org/.
John, P.S. et al. Bionemo Framework: A Modular, High-Performance Library for AI Model Development in Drug Discovery (2024).
Sutton, R.S., McAllester, D., Singh, S. & Mansour, Y. Policy gradient methods for reinforcement learning with function approximation. In Advances in Neural Information Processing Systems. Vol. 12. (MIT Press, 1999).
Acknowledgements
We thank Parul Doshi and Atli Thorarensen from Cellarity for supporting this work. We also extended our gratitude to Vega Shah and the NVIDIA team for reviewing the manuscript and their insightful feedback on key experiments and benchmarks.
Author information
Authors and Affiliations
Contributions
R.H, conceived the initial idea of applying proximal policy optimization on latent representations for molecular optimization and authored the manuscript. E.L, co-led the vision for the experiments and the method application, co-authored the manuscript and applied the single-property optimization experiments. Z.L, conceived the initial idea of reducing molecular modifications to vector modifications and showed the initial proof of concept. X.Y, trained multiple language models for experimentation and handled cross collaborative communication. D.B, co-designed multiple code components for the method. G.B, supervised the work and the preparation of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
Authors are current or previous employees of Cellarity Inc. or NVIDIA and may hold equity.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Haddad, R., Litsa, E.E., Liu, Z. et al. Targeted molecular generation with latent reinforcement learning. Sci Rep 15, 15202 (2025). https://doi.org/10.1038/s41598-025-99785-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-99785-0