Abstract
Cervical cancer is one of the leading causes of death in women, especially in low- and middle-income countries. Early disease detection is crucial for improving survival, but conventional methods are inefficient, costly, and error-prone. In this study, we present a hybrid machine learning framework that diagnoses cervical cancer risk from clinical and behavioral records by analyzing 36 patient attributes such as age, sexual history, smoking habit, hormonal contraceptive use, and sexually transmitted disease history. Missing values were imputed using a Generative Adversarial Imputation Network (GAIN). The Boruta algorithm was then used to identify the most influential diagnostic features, and Random Oversampling (ROS) was applied to correct class imbalance. Dimensional reduction techniques such as Independent Component Analysis (ICA) and Principal Component Analysis (PCA) were used. The diagnostic prediction is generated by a Bayesian Fusion Ensemble (XBFE) that combines outputs from Decision Tree and Random Forest models to estimate each patient’s likelihood of cervical cancer and systematically evaluates contributing risk factors. Principal determinants, including the Schiller test, the Hinselmann test, cytology, age, number of sexual partners, and smoking, were identified using Boruta feature selection. The proposed model achieved an accuracy of 99.88%, a recall of 1.00, and an AUC-ROC score of 1.00, as validated by K-fold cross-validation. To improve interpretability for healthcare clinicians, Explainable AI (XAI) tools such as SHAP and LIME were used. We developed a web-based application for real-time risk estimation. The proposed system provides a reliable and interpretable solution for predicting cervical cancer risk, helping doctors make better decisions, especially in resource-limited settings. human readable.
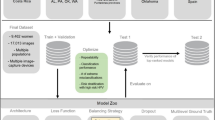
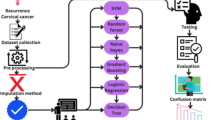
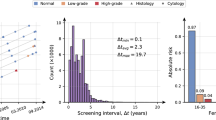
Similar content being viewed by others
Introduction
Cervical cancer is a leading cause of death among women due to premature mortality. According to the World Health Organization (WHO), ovarian, bone, breast, and cervical cancers are among the most significant causes of mortality among women[1]. Human papillomavirus (HPV), a high-risk virus, is the primary perpetrator of cervical cancer, among a myriad of other cancers, including rectal and testicular cancers2. Cervical cancer occurs in four stages. The early phase affects the lymph nodes, and late phases spread beyond the reproductive system, where it can block the renal pelvis and also invade nearby organs3. The disease tends to grow without symptoms in the early phases. In the later stages, abnormal bleeding, pelvic discomfort, and pain during sexual intercourse are experienced by patients4,5. This can lead to infertility, kidney failure, or metastasis of the organs if not treated on time. Risk factors include suppression of the immune system, smoking, and complications during pregnancy.
According to the World Health Organization (WHO), an estimated 604,000 new cases of cervical cancer were reported globally in 2020, making it the fourth most common cancer among women. Due to restricted access to screening and treatment facilities, the impact is disproportionately felt by low- and middle-income countries (LMICs). In 2020, middle- to low-income nations accounted for almost 90% of the 34,200 cervical cancer deaths. HIV causes 5% of cases, and women with HIV are six times more likely to get infected6. Early detection through regular screening techniques, including Pap smears and HPV testing, can dramatically lower death rates; however, conventional diagnostic methods are labor-intensive, need qualified personnel, and can produce false-negative results7. Current research indicates that Machine learning8 in most well-known areas has achieved considerable success across a wide range of fields. Cervical cancer and its early stages can now be predicted by Machine Learning algorithms using various data sources, such as scanning and clinical information. The past few years have witnessed the publishing of different research studies on the use of machine learning and deep learning for the early detection of cervical cancer9,10. Machine learning algorithms are trained on a given dataset to reliably predict cervical cancer symptoms, utilizing past experiences to predict present situations, thereby enhancing the detection of cervical cancer. Risk assessment and early detection of cervical cancer are imperative to improve the treatment outcome and reduce the mortality rate of females. Machine learning methods, when coupled with stringent feature selection, data balancing, and ensemble techniques, offer promising prospects for early, accurate diagnosis.
To address these challenges, this study proposes a hybrid machine-learning model to predict cervical cancer risk. The suggested model combines the latest data imputation, feature selection, sampling, dimensionality reduction, ensemble learning, and explainable AI techniques. The objective is to design a robust, interpretable, and clinically useful system for early diagnosis. The proposed XBFE model is intended as a screening-support tool to identify individuals at elevated risk of cervical cancer based on clinical and behavioral factors. While it does not replace biopsy or cytological confirmation, it serves as an assistive system that helps clinicians early-detect and prioritize high-risk patients for further diagnostic testing.
The key contributions of this work can be summarized as follows:
-
A Generative Adversarial Imputation Network (GAIN) was utilized to handle missing values precisely while ensuring data integrity during model testing and training.
-
The Boruta algorithm was employed to identify the key features from a preliminary feature set of 36 variables. Further feature-space enrichment was achieved through Independent Component Analysis (ICA) and Principal Component Analysis (PCA) to achieve low dimensionality without sacrificing informative signals.
-
Random Oversampling (ROS) was applied to balance highly imbalanced classes in the dataset so that the model could learn from minority and majority class instances equally.
-
A novel Bayesian Fusion ensemble model was presented, which merged Decision Tree and Random Forest classifiers probabilistically with weighting to boost prediction accuracy as well as model robustness.
-
Applied Explainable AI (XAI) methods, such as SHAP and LIME, to interpret the model’s binary diagnosis—predicting whether a patient is likely to have cervical cancer and to ensure transparency in clinical decision-making.
-
Developed a web-based app to enable real-time prediction of cervical cancer risk, with the solution being easily accessible and usable in clinical settings.
All things considered, this study presents a highly accurate, interpretable, and clinically relevant method that closes the gap across machine learning studies and real-world healthcare implementation. The rest of this article is structured as follows. “Related research” section discusses the most relevant literature on predicting cervical cancer and identifies gaps in the methodology. “Methodology” section describes the suggested XBFE framework, such as data preprocessing, feature selection, dimensionality reduction, and model architecture. “Experimental results and analysis” section describes the experimental setup and evaluation metrics, and Sect. 5 provides a detailed analysis of the performance. Explainability results from SHAP and LIME are presented in “Explainable AI” section, whereas the implementation of the real-time web application is addressed in “Deployment in the real world” section. Lastly, “Conclusion and future work” section presents the conclusion and future research directions.
Related research
Numerous studies utilised machine learning (ML), deep learning (DL), and optimization methods to enhance cervical cancer prediction and early diagnosis.
Scientists have developed hybrid ML-based systems with advanced optimization and ensemble techniques to enhance predictive accuracy. Suvanasuthi et al.11 reported a panel of miRNAs coupled with ML models for CC prediction, achieving 90.9% accuracy using POCT (Point-of-Care Testing). Mathivanan et al.12 have employed pre-trained CNN models (AlexNet, InceptionV3, ResNet101, ResNet152), and ResNet152 has achieved 98.08% accuracy. Mohi Uddin et al.13 have employed ensemble models of hard voting of MLP, RF, XGBoost, and PCA with 99.19% accuracy and 100% sensitivity. Shakil et al.14 used SHAP for interpretability and SMOTE/ADASYN, Chi-square, and LASSO with ML algorithms such as DT, LG, NB, RF, KNN, and SVM—where DT achieved the highest performance of 97.60%. Llerberi et al.15 used PSO for feature selection with RF and other models on real hospital data; the RF(PSO) model gave superior results.
Kolasseri et al.16 compared RSF, Cox PH, and Weibull models with SEER data; RSF performed better than the others. Qathrady et al.17 created a web framework (WFC2DS) based on algorithms like ANN, RFC, AdaBoost, KNN, DT, and SVM, and RFC and DT performed with 98.1% accuracy. Chauhan et al.18 presented CHAMP based on ML models, where XGBoost provided maximum accuracy (80.20%). Individual studies also reported significant improvements in model performance and explanation. Glucina et al.19 used ADASYN, SMOTEEN, and oversampling with different ML algorithms; KNN had 95% accuracy. Al.
Mohimeed et al.20 used stacking models (SVM, LR, RF) with SMOTE-Tomek, RFE, and tree-based feature selection, and got 99.44% accuracy. One other study21 used Gradient Boosting on a combined target variable dataset. This study found 98.9% accuracy for cervical cancer and CIN.
A follow-up study22 added SMOTE and RFE for feature selection to the model and developed a stacked ensemble model with high performance. Voting classifiers (LR, DT, RF) with PCA and SMOTE tackled imbalance and increased ROC-AUC and sensitivity in another study23. Parvathi et al.24 developed a web application for cervical cancer and fetal abnormality prediction. Numerous ML models are exclusive to women’s health. Tanimu et al.25 performed a feature selection technique comparison, i.e., RFE, LASSO, and SMOTE-Tomek, wherein Decision Trees obtained an accuracy of 98.72%. Chauhan et al.26 applied non-uniform scaling, SMOTE, and RFE on various machine learning algorithms, wherein Random Forest had the highest result of 98.53%. Al-Batah et al.27 suggested an ensemble method involving LWNB, RF, and logistic regression on hospital datasets, wherein LWNB performed better. Jahan et al.28 evaluated eight ML models with Chi-square, SelectBest, and RF as feature selectors. Among them, MLP performed best (98.10%) with the top 30 features. Curia et al.29 developed a CDSS with ensemble and MLP models and reported 94.5% accuracy and an AUC of 0.98%, aiming to use LIME and SHAP for Explainability.
Previous research has revealed the promise of employing machine learning approaches in cervical cancer prediction. However, there are some critical limitations, and Table 1 summarizes these studies.
-
Most studies employed a single or minimal feature selection technique, without leveraging full hybrid approaches that combine feature selection, extraction, and dimensionality reduction for better model performance.
-
The majority of research utilised inadequate data imputation approaches for dealing with missing values, which has a considerable influence on model dependability and accuracy.
-
Many studies have not employed cross-validation to evaluate model generalizability, potentially overestimating performance and limiting confidence in real-world applicability.
-
There was also minimal emphasis on model interpretability. Few articles employ state-of-the-art explainable AI (XAI) techniques such as SHAP and LIME to provide transparent and clinically interpretable explanations of model predictions.
While earlier studies achieved promising results, many were constrained by small or imbalanced datasets, limited preprocessing, and weak validation strategies. Most also lacked interpretability, which limits their clinical application. To address these limitations, this study introduces several targeted methodological improvements. First, rather than relying on a small or single-source dataset, the analysis utilizes the complete UCI cervical cancer risk-factor benchmark dataset and applies 10-fold cross-validation to improve generalizability. Second, a multi-stage preprocessing pipeline is implemented, including GAIN for missing-value imputation, Boruta for feature selection, and a combination of ICA and PCA for dimensionality reduction. This approach addresses deficiencies in prior studies that used minimal or single-step preprocessing. Third, class imbalance, which has previously undermined model reliability, is mitigated through Random Oversampling (ROS) to ensure equal representation of minority cancer cases. Fourth, to improve interpretability, the proposed XBFE model integrates SHAP and LIME, providing transparent explanations for predictions. Collectively, these enhancements address the shortcomings identified in the literature and establish a more robust, interpretable, and clinically relevant framework.
Methodology
In this section, the implementation process of the workflow is shown in Fig. 1 of our study. A description of the dataset is presented and discussed in the section named “Dataset”. After that, data preprocessing begins with several steps. Data is saved after preprocessing and preparation for the “Feature Selection” process. After this step, the saved data is passed to “Dimensionality Reduction Techniques”, which preserves the most important features to enable faster computation in our proposed model. The dataset has been split into a training and test set so that our experiment can be conducted on each set separately. After successfully training our proposed model, we analyzed our model evaluation with other models. Applied model result analysis shows metrics using different graphical representations. With our best model, we apply the “Black Box” technique, such as SHAP and LIME, to show the feature contributions for a specific prediction. It made decision-making easier in model performance. Finally, implementing a web-based application to predict and analyze in real time.
The workflow of the proposed XBFE framework shows data preprocessing, feature selection, model training, and explainability.
Dataset preparation
The dataset Cervical Cancer (Risk Factors) is from Hospital Universitario de Caracas, Venezuela. It is widely adopted as a benchmark in cervical cancer ML research across several peer-reviewed studies. It has 858 instances and 36 patient conditions30. The data includes demographic information, reproductive status, lifestyle information, hormonal contraception, and STDs. The data also provide diagnostic information on cervical cancer, cervical intraepithelial neoplasia, and HPV. The target features include test results from various screening tests, such as Hinselmann, Schiller, Cytology, and Biopsy. The data support the development of models to forecast cervical cancer risk.
Handling missing values
The dataset includes missing data for several clinical features. To preserve data reliability and avoid distortion caused by standard imputation methods such as mean/median, we applied the Generative Adversarial Imputation Network (GAIN). GAIN is based on a generative adversarial framework to capture the data distribution and estimate missing patterns without destroying the statistical relationships among features. This is for accurate reconstruction before modeling.
Dataset balancing
Dataset balancing is an essential process to rectify class imbalances that can negatively impact model performance. We found an imbalance in the dataset by analyzing the distribution of the target variable, ’Biopsy.’ The class distribution showed that the majority class (negative cases) had 803 samples, while the minority class (positive cases) had 55 samples. This significant imbalance may bias a machine learning model toward the majority class, leading it to perform poorly on the minority class. To confirm the imbalance, we calculated the class ratios. This showed that the minority class represented just 6.41% of the data. We used a 10% threshold to assess the size of the imbalance. Since the minority class was below this figure, we determined that the dataset was imbalanced. To address this problem, we used Random OverSampling31(ROS). This technique generates more samples in the smaller group by randomly copying existing samples. In this way, both groups have the same number of samples, which reduces the imbalance. In the dataset, k samples from the minority class with replacement are added:
Making the dataset balanced by ensuring the condition nmajority = nminority. After applying random oversampling, we reassessed the class distribution and confirmed that both classes now had 803 samples. This balanced distribution ensures that the model sees an equal representation of both classes, reducing the possibility of biased predictions.
Feature selection
One crucial aspect of machine learning is feature selection, which aims to improve model performance by removing irrelevant or redundant features. Feature selection reduces overfitting, improves model interpretability, and decreases computational complexity by choosing the most relevant variables. While most of the previous studies rely on a single feature engineering strategy (e.g., only PCA or only filter-based selection), our framework proposes a two-staged hybrid framework (1) Boruta for interpretable, relevance-driven feature selection, followed by (2) ICA and PCA for dimensionality reduction that preserves both statistical independence and maximal variance. The use of this combination improves the model’s performance and clinical interpretability. We used the Boruta algorithm, a powerful wrapper-based feature selection approach32, to select the most significant features for classification. Boruta is a repetitive algorithm that shuffles the original features, produces shadow features, trains a random forest classifier, compares feature importances, retains features that perform well, and either confirms or rejects them. This algorithm identified the most influential features, including the Schiller test, the Hinselmann test, cytology results, age, number of sexual partners, smoking status, hormonal contraceptive use, and STD history. These features collectively improved model accuracy and were further validated through SHAP and LIME analysis, which confirmed that Schiller, Hinselmann, and cytology contributed most to model performance.
The Boruta procedure was applied to a balanced dataset using a random forest classifier as the base estimator. The algorithm ranked features by importance, with rank-1 features being the most relevant. Key variables like age, number of sexual partners, age at first sexual intercourse, smoking status, hormonal contraceptive use, and STD history were selected due to their high correlation with the target variable. For feature ranking, we used Eq. (2):
where: Xj represents the jth feature in the dataset, Ij denotes the importance score of feature Xj, Order(·) is the ranking function that sorts features by their importance scores. Boruta is an iterative algorithm that creates shadow features by shuffling the original features, trains a random forest classifier, compares feature importances, retains the outperforming features, and confirms or rejects them.
Dimensionality reduction
We used PCA and ICA to reduce dimensionality and uncover relevant patterns33 from high-dimensional data. PCA transforms data into orthogonal components that capture the maximum variance. The key transformation is:
where Y contains principal components, X is standardized data, and V consists of eigenvectors from the covariance matrix.
\({\mathbf{C}} = ~\frac{1}{{n - 1}}~~~{\mathbf{X}}^{T} ~{\mathbf{X}}\) ICA separates mixed signals into statistically independent components.
where its S contains Eq. (4): independent components, X is observed data, and W is the unmixing matrix optimized for non-Gaussianity. The combined approach first applies ICA to separate independent sources, then applies PCA to reduce dimensionality while preserving 95% of the variance, effectively eliminating redundancy and noise. Data with the most significant information and little redundancy and noise.
Applied models
In our research, we used various algorithms to analyze cervical cancer risk factors. We discuss those models below.
Proposed model
In our present study, we introduce a Bayesian Fusion Ensemble (RDBF) framework that fuses Random Forest (RF) and Decision Tree (DT) classifiers via probabilistic weighting to improve the predictive power of cervical cancer risk assessment. Traditional hybrid methods, such as Voting or Stacking ensembles, tend to treat individual models uniformly or use meta-learners, which may lead to spurious complexity or inferior performance when models exhibit different confidence levels. Our approach circumvents these limitations through the dynamic combination of RF and DT predicted probabilities using Bayesian weighting, where each model’s contribution is weighted by its validation accuracy, as in Eqs. (5) and (6).
The proposed model reconciles the virtues of RF and DT, mitigating overfitting via ensemble diversity and learning complex patterns. It classifies voting ensembles by combining probabilities, which lowers computational overhead. The probabilistic approach is consistent with clinical decision-making. Predictions frequently combine confidence scores from a range of diagnostic tests, whereas stacking involves training a meta-model and can suffer from overfitting on small datasets. The implementation of the proposed algorithm, summarized in Figs. 2 and 3, shows the workflow of our proposed model.
Proposed model algorithm.
Architecture of the proposed model.
Random forest
Random Forest is an ensemble learning method that generates numerous decision trees in the training process, ultimately returning the mode of the classes or the mean prediction of individual trees34. Random Forest is renowned for its robustness, accuracy, and efficiency in dealing with large datasets of high dimensionality. For random forest prediction using Eq. (7), where prediction f t in each tree as T and.
In this study, Random Forest was implemented to predict on the given features of the cervical cancer dataset. The model’s ability to handle non-linear relationships and its noise tolerance made it appropriate for this data, which has intricate interactions among variables such as age, sexual partners, and medical history. The ensemble character of Random Forest ensures that it effectively discovers the underlying patterns, thereby making accurate predictions.
Decision tree
Decision Tree is a machine learning technique that can be used for classification and regression. This method creates a tree-like model by splitting a dataset based on key attributes. The internal nodes represent decision points, branches represent choice outcomes, and leaf nodes provide the final prediction35. It operates by recursively partitioning the data until we meet a pre-specified stopping criterion, which may be achieving an optimal depth or a minimum sample size in a leaf node. This research employed the Decision Tree model to analyze the cervical cancer dataset, aiming to predict the probability of stroke occurrence. The model describes interactions among variables such as age, sexual partners, and health history, thereby offering distinct decision paths that are susceptible to clinical interpretation.
Naïve Bayes
Based on Bayes’ Theorem, it’s a probabilistic supervised learning technique primarily used for categorizing high-dimensional data. This classifier assumes conditional independence of features, thus enabling the calculation of probabilities. It calculates the posterior probability for each class and assigns the class with the highest likelihood to the provided input instance36. In this study, Naive Bayes was applied to the cervical cancer dataset for several classification tasks, including predicting the likelihood of stroke. Its effectiveness and ability to handle high-dimensional data make it suitable for medical datasets characterized by numerous features.
Artificial neural network
Artificial Neural Networks (ANNs) are advanced machine learning architectures that excel at recognizing intricate, non-linear patterns in data sets. ANNs are composed of multiple layers of nodes with inter-layer connections, such as input, hidden, and output layers37. Learning occurs by tweaking weights to narrow the gap between predicted outputs and actual outcomes, using algorithms such as backpropagation and gradient descent. In this study, we applied an ANN to the cervical cancer database to model intricate relationships among such features as age, sexual history, and medical history. ANN’s strength and flexibility make it a perfect predictor of outcomes such as the likelihood of stroke, and thus a robust predictive modeling tool.
XGBoost
XGBoost is a gradient-boosting classification and regression technique recognized for its fast execution and good prediction performance. It is an ensemble approach that creates a succession of decision trees, repairing the mistakes of the previous ones and optimizing a differentiable loss function using gradient descent38. The algorithm is highly customizable, with parameters such as tree depth, learning rate, and regularization coefficients being tunable. In this study, XGBoost was applied to the cervical cancer dataset to predict outcomes such as the probability of stroke. Its efficiency, accuracy, and robustness against overfitting make it a promising choice for health care predictive modeling.
K- nearest neighbors
K-Nearest Neighbors (KNN) is a supervised learning method that performs classification and regression. It predicts the output for a new point by locating the nearest neighbors in the training set and utilizing their results. Classification employs majority voting among neighbors, whereas regression uses the mean of neighbor values. KNN was used on the cervical cancer dataset to create predictions, such as stroke probability. The method’s simplicity and efficacy in detecting local patterns make it well-suited to complex feature interactions and data fitting without the need for model training. Following that, we prepared data for model training and performance evaluation. The following section will cover the model’s performance analysis.
Experimental results and analysis
To address the common limitation of weak validation in prior work, all models were evaluated using 5 and 10-fold cross-validation, and performance was reported per class (precision, recall, F1) in addition to overall accuracy and AUC-ROC. Furthermore, to ensure clinical trust and transparency often missing in black-box models, we integrated dual XAI techniques: SHAP for global feature importance and LIME for local instance-level explanations. Finally, we deployed the model as a real-time web application (Fig. 12) to demonstrate real-world feasibility.
(a–g) Overall model performance across different data-balancing scenarios.
Metrics for evaluation
Performance analysis of the algorithm is essential after completing the model training. These performance metrics are critical. They provide a statistical and graphical presentation of performance39 where include specific task, clustering, regression and other data analysis. In our research performance metrics, we show Accuracy in Eq. (8), Recall in Eq. (10), Precision in Eq. (9), and F1-value in Eq. (11).
Accuracy
The ratio of accurately anticipated observations to total observations.
Precision
The fraction of real positive predictions vs. all anticipated positives.
Recall
The fraction of accurate positive forecasts vs. all actual positives.
F1-value
The harmonic average of accuracy and recall.
Best model selection
The research developed a predictive model to detect cervical cancer risk using advanced machine learning techniques. To manage class imbalance, the model employed the Synthetic Minority Oversampling Technique (SMOTE) or Random Oversampling, followed by a two-stage dimensionality reduction procedure that included Independent Component Analysis and Principal Component Analysis. After that, the Bayesian fusion ensemble approach improved sensitivity to minority class instances and confidence estimates, making it suitable for cervical cancer risk prediction.
Result analysis
Our comprehensive evaluation of machine learning models on the imbalanced cervical cancer dataset yielded critical insights into model performance across different data balancing strategies. Accuracy was still a fundamental performance metric, although precision, recall, and AUC-ROC scores provide a more pertinent assessment of the model’s effectiveness for medical diagnostics.
The use of SMOTE greatly enhanced model stability, with Random Forest achieving high accuracy and XGBoost reaching 95.93% accuracy with enhanced recall. Random Over-Sampling (ROS) was the top performer, with XGBoost achieving 98.34% accuracy, perfect recall, and high precision. The proposed model, which incorporates balancing and ensemble learning techniques, achieved competitive performance, as shown in Table 2.
Now we see the overall model performance for different scenarios at a glance in the bar chart in Fig. 4a–g.
These ROC curves compare the classification performance of all baseline models alongside the proposed model. The Area Under the Curve (AUC) values represent each model’s overall performance, where larger AUCs represent better discrimination between classes. The proposed model demonstrates strong performance across a wide range of scenarios, typically achieving the highest or second-highest AUC values, as shown in Fig. 5.
AUC-ROC curves comparing model performance under various scenarios.
The Bayesian Fusion technique, which combines dimensionality reduction techniques, has significantly improved model performance. The combination of SMOTE, ICA, PCA, and Bayesian Fusion has yielded balanced precision and recall scores above 0.94, with KNN as the top-performing model. The new model maintains an accuracy of 95.93%. Table 3 presents the model performance after applying dimensionality reduction and data-balancing techniques.
The proposed model achieved 99.88% accuracy, perfect recall, and the best AUC-ROC (1.00) due to three innovations: ROS balances class distribution without removing minority patterns, ICA-PCA combination identifies independent risk factors, and Bayesian Fusion ensemble combines Random Forest and Decision Tree predictions. These innovations reduce false negatives and ensure perfect prediction of risk factors in clinical metrics. We now present the overall model performance across different scenarios with dimensionality reduction after data balancing with SMOTE and ROS, as a bar chart. Model stability was assessed by evaluating all configurations using 5-fold and 10-fold cross-validation. The results indicated minimal performance variance (< 0.2%) between the two schemes, confirming robustness to data partitioning. Under the ROS + ICA + PCA setting, the proposed model achieved 99.67% (CV = 5) and 99.50% (CV = 10) accuracy, with a precision of 0.99, a recall of 1.00, and an AUC-ROC of 1.00. This consistency indicates strong generalizability and a low risk of overfitting. Other models also demonstrated stable cross-validation performance, further supporting the reliability of the preprocessing and ensemble design.
We can see that the AUC of all models in Fig. 6a–g is higher than before, as there was no balancing applied. This means that both SMOTE and ROS seem to improve classification in all instances. The proposed model is exceptional, as it achieved a highly enhanced AUC of 0.9776 with SMOTE and a perfect 1.0000 with ROS.
(a–g) Overall model performance on dimensionality reduction after using SMOTE and ROS.
Figure 7 illustrates the AUC-ROC analysis of the evaluated models, including the proposed algorithm.
AUC-ROC curve of different models in different scenarios after data balancing SMOTE and ROS with dimensionality reduction.
Table 4 summarizes the classification results for the binary classes across all models, including the proposed method and the six baseline classifiers.
The cervical cancer risk factor dataset displays classification outputs for different models, and the proposed algorithm performs well, with precision, recall, and F1-scores near 1.00 for both classes. The dataset, containing 858 instances, was divided into 80% training (686 samples) and 20% testing (172 samples) to ensure reliable evaluation. After completing dimensionality reduction on the ROS sample data, we trained our model and fine-tuned the classification report. Figure 8 shows the performance of various classifiers. Among these, the hybrid model achieved the best performance. It correctly classifies 179 of 181 Class-0 and 141 of 143 Class-1 samples with only two false negatives and zero false positives. Random Forest and Decision Tree followed closely, with ANN and XGBoost showing moderate accuracy but minor errors.
(a–g) Confusion matrix of model predictions on the test set.
Explainable AI
Explainable AI (XAI) makes machine learning model decision-making processes more explainable40 and comprehensible to humans. XAI closes the gap between complex models and human understanding, making decision-making and trust in the system feasible. XAI also helps with debugging models, regulatory compliance requirements, and improving model performance. In Explainable AI, we implemented SHAP and LIME to evaluate our model. SHAP is a unified approach to explain the prediction of any machine learning model. The basic idea is to estimate the impact of each feature by considering all subsets of features. For features i SHaply value in Eq. (12) is shown below:
Here, F represents all sets of features and A, a subset of features, while f(s) models the prediction of subset features A. In Fig. 9, we plot a summary plot for the impact of the features in both classes.
SHAP summary plot for comparing feature impact on Class-0 and Class-1.
The proposed model predicts cervical cancer risk using SHAP contribution plots, with top-ranked features such as Schiller, Hinselmann, and cytology indicating higher risk. The model also reveals class-specific information, with age having a marginally more substantial impact on Class-1 classification. The best model predicts the probabilities of Class-0 and Class-1, with probabilities of 0.092 and 0.908, respectively. Age and number of sexual partners reduce the likelihood of Class-1, but a high output of 0.91 indicates the probability of Class-1.
LIME explains each forecast by estimating the model locally using an interpretable model. It perturbs the input data and monitors the resulting changes in predictions, focusing on the model’s local behavior. The explanation was achieved by optimizing the following. Equation 13
Here, the original model is denoted by d, while h is the interpretable model, the weighted kernel x, the loss function as L, and the complexity (g) is penalized. In Fig. 10, we plot the SHAP force plots, which were used to improve the interpretability of the model’s predictions. In Class-0 instances, features such as “Schiller = 1.0” and “Citology = 1.0” had strong negative SHAP values, pushing the prediction toward the negative class. In Class-1 instances, features like “Age = 21.0,” “Number of sexual partners = 4.0,” and “Citology = 1.0” pushed the model toward the high-risk class. The “Hinselmann = 0.0” feature consistently contributed negatively in both cases, suggesting its potential risk. These force plots provide a clear, specific case breakdown of how individual features influenced. The final classification outcome reinforces the value of explainable AI in clinical risk prediction.
SHAP force plots explaining random patient outcome.
The LIME explanation for Fig. 11 was used to improve individual predictions in cancer cases. In an instance of cancer, the model predicted a 91% probability of cancer, influenced mainly by “Schiller = 1.0” and “Citology = 1.0.” However, in a non-cancer instance, all features, including “Schiller,” “Citology,” and “Hinselmann,” had values of 0 and contributed negatively to the cancer class. These LIME visualizations confirm the model’s alignment with clinically relevant factors, enhancing trust and interpretability in high-stakes medical prediction tasks.
LIME explanation for a random patient.
(a, b) Real-time web application for cervical cancer risk analysis.
The LIME explanation provides an exact explanation for a single case, whereas SHAP provides a global explanation of feature importance across the entire dataset. This is how vital clinical tests such as Schiller, Hinselmann, and Citology are to identify the risk of cervical cancer.
Deployment in the real world
The deployment of our cervical cancer risk assessment model focuses on integrating the trained machine learning system into a secure, scalable web application for real-world clinical use. We are putting our model for cervical cancer risk calculation into practice by developing it in a safe and flexible web application for41 use in actual medical practice. To ensure ethical and secure use, the web application is HIPAA- and GDPR-compliant by anonymizing and encrypting all patient information. A drift-monitoring module tracks performance on new data and initiates retraining when distribution shifts occur, maintaining diagnostic reliability and compliance in real-world use.
Figure 12a, b represents the web application UI interface for taking data from the user and showing output.
The backend and frontend were developed using Flask and HTML, CSS, and JavaScript, with machine learning modelling using scikit-learn. The application integrates with hospital databases to automate risk scoring in patient screening, and future enhancements include a mobile app and multilingual capability. This framework enables healthcare practitioners to receive AI-powered risk insights while adhering to medical data compliance requirements.
Table 5 highlights how our proposed study compares to previous studies on cervical cancer risk.
This comparison highlights that most existing models use PCA or similar techniques for feature selection and achieve accuracies between 90.9% and 99.19%. While some research works apply advanced techniques like XGBoost or ensemble learning, hardly any apply XAI (e.g., SHAP). The proposed model outperforms others, achieving 99.88% accuracy via Boruta feature selection, Bayesian fusion, and robust XAI (SHAP & LIME), and real-time deployment via a web app.
Conclusion and future work
This study presents a hybrid ensemble learning strategy for early cervical cancer detection to enhance diagnostic accuracy and support clinical decision-making. The proposed framework embraces cutting-edge preprocessing techniques coupled with Generative Adversarial Imputation Networks (GAIN) to address issues of missing data, and employs Boruta to extract pertinent features from an 858-patient dataset with 36 attributes. To address data imbalance and dimensionality, Random Oversampling (ROS), Independent Component Analysis (ICA), and Principal Component Analysis (PCA) were employed. A Random Forest and Decision Tree classifier Bayesian fusion ensemble model was built. The ensemble yielded improved 99.88% accuracy, 0.99 precision, 1.00 recall, 0.99 F1-score, and 1.00 AUC-ROC and was confirmed using K-fold cross-validation. The interpretability of the model was facilitated by the use of SHAP and LIME, making transparency in a clinical environment possible. A real-time web-based prediction tool has also been developed, promoting research practicality. The current study is limited to a single data set. Future research will explore the application of large and heterogeneous datasets, the incorporation of multimodal clinical data, and the deployment of deep and federated learning techniques to enhance generalization and clinical significance.
Data availability
The data set used in this study is accessible from UCI https://archive.ics.uci.edu/dataset/383/cervical+cancer+risk+factors. The dataset includes 858 patients’ historical medical records, habits, and demographic details.
References
Johnson, C. A. et al. Cervical cancer: An overview of pathophysiology and management. Semin Oncol. Nurs. 35, 166–174 (2019).
Stelzle, D. et al. Estimates of the global burden of cervical cancer associated with Hiv. Lancet Glob. Health. 9, e161–e169 (2021).
WebMD. Cervical cancer. (2025). https://www.webmd.com/cancer/cervical-cancer/default.htm. Accessed 24 Apr 2025.
Nithya, B. & Ilango, V. Evaluation of machine learning based optimized feature selection approaches and classification methods for cervical cancer prediction. SN Appl. Sci. 1, 1–16 (2019).
Issah, F., Maree, J. E. & Mwinituo, P. P. Expressions of cervical cancer-related signs and symptoms. Eur. J. Oncol. Nurs. 15, 67–72 (2011).
Sung, H. et al. Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Cancer J. Clin. 71, 209–249 (2021).
Ali, M. M. et al. Machine learning-based statistical analysis for early-stage detection of cervical cancer. Comput. Biol. Med. 139, 104985 (2021).
Edafetanure-Ibeh, F. T. Evaluating machine learning algorithms for cervical cancer prediction: A comparative analysis. OSF Prepr (2024).
Khamparia, A. et al. Dcavn: cervical cancer prediction and classification using a deep convolutional and variational autoencoder network. Multimed Tools Appl. 80, 30399–30415 (2021).
Alam, T. M. et al. Cervical cancer prediction through different screening methods using data mining. Int. J. Adv. Comput. Sci. Appl. 10, 9 (2019).
Suvanasuthi, R. et al. Analysis of precancerous lesion-related MicroRNAs for early diagnosis of cervical cancer in the Thai population. Sci. Rep. 15, 142. https://doi.org/10.1038/s41598-024-84080-1 (2025).
Mathivanan, S. et al. Enhancing cervical cancer detection and robust classification through a fusion of deep learning models. Sci. Rep. 14, 10812. https://doi.org/10.1038/s41598-024-61063-w (2024).
Uddin, K. M. M., Mamun, A., Chakrabarti, A., Mostafiz, A., Dey, S. K. & R. & An ensemble machine learning-based approach to predict cervical cancer using hybrid feature selection. Neurosci. Inf. 4, 100169 (2024).
Shakil, R., Islam, S. & Akter, B. A precise machine learning model: detecting cervical cancer using feature selection and explainable Ai. J. Pathol. Inf. 15, 100398. https://doi.org/10.1016/j.jpi.2024.100398 (2024).
Ileberi, E. & Sun, Y. Machine learning-assisted cervical cancer prediction using particle swarm optimization for improved feature selection and prediction. IEEE Access. 12, 152684–152695. https://doi.org/10.1109/ACCESS.2024.3469869 (2024).
Kolasseri, A. E. & B, V. Comparative study of machine learning and statistical survival models for enhancing cervical cancer prognosis and risk factor assessment using seer data. Sci. Rep. 14, 22203. https://doi.org/10.1038/s41598-024-72790-5 (2024).
Qathrady, M. A. et al. A novel web framework for cervical cancer detection system: A machine learning breakthrough. IEEE Access. 12, 41542–41556. https://doi.org/10.1109/ACCESS.2024.3377124 (2024).
Chauhan, R., Goel, A., Alankar, B. & Kaur, H. Predictive modeling and web-based tool for cervical cancer risk assessment: A comparative study of machine learning models. MethodsX 12, 102653 (2024).
Glucˇina, M., Lorencin, A., And¯elic´, N. & Lorencin, I. Cervical cancer diagnostics using machine learning algorithms and class balancing techniques. Appl. Sci. 13, 1061 (2023).
AlMohimeed, A., Saleh, H., Mostafa, S., Saad, R. M. & Talaat, A. S. Cervical cancer diagnosis using stacked ensemble model and optimized feature selection: an explainable artificial intelligence approach. Computers 12, 200 (2023).
Gopinath, B. & Santhi, R. & Dhivya Praba, R. A machine learning based decision support system to predict the presence of cervical cancer. in 2nd International Conference on Advancements in Electrical, Electronics, Communication, Computing and Automation (ICAECA) (IEEE, 2023).
Bhavani, C. & Govardhan, A. Cervical cancer prediction using stacked ensemble algorithm with Smote and Rferf. Mater. Today: Proc. https://doi.org/10.1016/j.matpr.2021.07.269 (2021).
Kumawat, G. et al. Prognosis of cervical cancer disease by applying machine learning techniques. J. Circuits Syst. Comput. 32, 2350019 (2023).
Parvathi, A. J., Gopika, H., Suresh, J., Sree, S. L. & Harikumar, S. Machine learning based approximate query processing for women health analytics. Procedia Comput. Sci. 218, 174–188. https://doi.org/10.1016/j.procs.2022.12.413 (2023).
Tanimu, J. J., Hamada, M., Hassan, M., Kakudi, H. & Abiodun, J. O. A machine learning method for classification of cervical cancer. Electronics 11, 463 (2022).
Chauhan, N. K. & Singh, K. Performance assessment of machine learning classifiers using selective feature approaches for cervical cancer detection. Wirel. Pers. Commun. https://doi.org/10.1007/s11277-022-09467-7 (2022).
Al-Batah, M. S. et al. Early prediction of cervical cancer using machine learning techniques. Jordanian J. Comput. Inf. Technol 8 (2022).
Jahan, S. et al. Automated invasive cervical cancer disease detection at early stage through suitable machine learning model. SN Appl. Sci. 3, 1–17 (2021).
Curia, F. Cervical cancer risk prediction with robust ensemble and explainable black boxes method. Heal Technol. 11, 875–885. https://doi.org/10.1007/s12553-021-00554-6 (2021).
UCI Machine Learning Repository. (2025). https://archive.ics.uci.edu/dataset/383/cervical+cancer+risk+factors Accessed 5 Nov 2025.
Yang, C., Fridgeirsson, E. A., Kors, J. A., Reps, J. M. & Rijnbeek, P. R. Impact of random oversampling and random undersampling on the performance of prediction models developed using observational health data. J. Big Data 11, 7. https://doi.org/10.1186/s40537-023-00857-7 (2024).
Zhou, H., Xin, Y. & Li, S. A diabetes prediction model based on Boruta feature selection and ensemble learning. BMC Bioinform. 24, 224. https://doi.org/10.1186/s12859-023-05300-5 (2023).
Artoni, F., Delorme, A. & Makeig, S. Applying dimension reduction to Eeg data by principal component analysis reduces the quality of its subsequent independent component decomposition. NeuroImage 175, 176–187. https://doi.org/10.1016/j.neuroimage.2018.03.016 (2018).
Breiman, L. Random forests. Mach. Learn. 45, 5–32. https://doi.org/10.1023/a:1010933404324 (2001).
Mienye, I. D., Sun, Y. & Wang, Z. Prediction performance of improved decision tree-based algorithms: a review. Procedia Manuf. 35, 698–703. https://doi.org/10.1016/j.promfg.2019.06.011 (2019).
Maheswari, S. & Pitchai, R. Heart disease prediction system using decision tree and Naive Bayes algorithm. Curr. Med. Imaging Rev. 14 https://doi.org/10.2174/1573405614666180322141259 (2018).
Abut, S., Okut, H. & Kallail, K. J. Paradigm shift from artificial neural networks (anns) to deep convolutional neural networks (dcnns) in the field of medical image processing. Expert Syst. Appl. 244, 122983. https://doi.org/10.1016/j.eswa (2024).
Zhang, J., Wang, R., Lu, Y. & Huang, J. Prediction of compressive strength of geopolymer concrete landscape design: application of the novel hybrid rf–gwo–xgboost algorithm. Buildings 14, 591. https://doi.org/10.3390/buildings14030591 (2024).
Rainio, O., Teuho, J. & Klén, R. Evaluation metrics and statistical tests for machine learning. Sci. Rep. 14, 1–14. https://doi.org/10.1038/s41598-024-56706-x (2024).
Islam, O., Assaduzzaman, M. & Hasan, M. Z. An explainable AI-based blood cell classification using optimized convolu- Tional neural network. J. Pathol. Inf. 15, 100389. https://doi.org/10.1016/j.jpi.2024.100389 (2024).
wahidpanda. Github - wahidpanda/cervical-cancer-risk-prediction-web-application: Predicting cervical cancer risk factors in real time. (2025). https://github.com/wahidpanda/Cervical-Cancer-Risk-Prediction-Web-Application Accessed 31 May 2025.
Acknowledgements
The authors want to thank Multimedia University, Malaysia.
Author information
Authors and Affiliations
Contributions
O.I. and M.A. contributed equally to the conceptualization, methodology design, and overall implementation of the study. S.A. was responsible for data preprocessing, imputation using GAIN, and feature engineering using Boruta, PCA, and ICA. Nafiz Fahad developed and deployed the real-time web-based application for clinical usability and integrated the machine learning pipeline using Flask. M.J.H. supervised the entire research, validated the results, performed critical revisions, and contributed to the final manuscript editing and review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study utilized. A Publicly available, Fully Anonymized Dataset (‘Cervical Cancer (Risk Factors)’) from the UCI Machine Learning Repository. According To the Repository’s documentation, the datA Were Originally Collected with Institutional Ethical Approval and Informed Consent at the Hospital Universitario De Caracas, Venezuela. No additional ethical clearance was required for this secondary analysis.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Islam, O., Assaduzzaman, M., Akter, S. et al. Enhanced cervical cancer diagnosis using a novel Bayesian fusion ensemble method with explainable AI. Sci Rep 16, 12306 (2026). https://doi.org/10.1038/s41598-026-35334-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35334-7