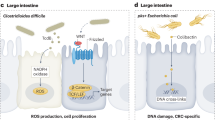
Abstract
c-di-AMP is a bacterial second messenger recognized by host immune sensors such as the STING pathway, linking gut microbiota activity to tumor immunity. This interaction holds significant therapeutic potential particularly for oncologic patients, given the increasingly recognized relationship between gut microbiota and tumor immunity. Recent evidence shows that microbial c-di-AMP can enhance anti-tumor responses and improve the efficacy of PD-1/PD-L1 blockade and radiotherapy. This study identified gut microbial species capable of synthesizing c-di-AMP by mining the Unified Human Gastrointestinal Protein catalogue for diadenylate cyclases (DACs), generating a database of 4,228 DACs across 3,901 species out of 4,744 presents in the Unified Human Gastrointestinal Genome catalogue. Analysis of metagenomic data from 190 healthy subjects and 569 cancer patients (melanoma, NSCLC, renal carcinoma) revealed a significantly higher abundance of DAC-encoding species in healthy microbiota, with no differences between responders and non-responders to immunotherapy. These findings indicate that c-di-AMP-producing bacteria are depleted in cancer-associated microbiota, supporting further studies on their role in modulating anti-tumor immunity.
Similar content being viewed by others
Introduction
Cyclic dimeric adenosine 3’,5’-monophosphate (c-di-AMP) is a ubiquitous and essential second messenger broadly distributed across the world of prokaryotes, being found in nearly all bacterial phyla and numerous archaeal groups1,2,3,4,5,6,7.
This signalling molecule orchestrates a diverse array of physiological processes vital for microbial survival and adaptation. It plays a critical role in maintaining cellular osmotic homeostasis and responding to cell wall stress, and it is involved in monitoring DNA damage and facilitating the recognition of stalled DNA replication forks1,8,9. Beyond structural and genomic integrity, c-di-AMP profoundly influences central metabolism and fatty acid biosynthesis10,11,12,13 and exerts control over fundamental cellular processes such as sporulation, competence development, and cell size regulation8,14,15,16. Its protective roles extend to confer resistance against environmental challenges, such as acid and oxidative stress17,18. Moreover, c-di-AMP is a key regulator of community behaviours, controlling biofilm formation19,20 and cellular differentiation21. For a range of bacterial pathogens, including Borrelia, Listeria, Mycobacterium, Staphylococcus, and Streptococcus, c-di-AMP is an important regulator of virulence22,23,24,25,26.
Beyond its direct impact on bacterial physiology, c-di-AMP can also be recognized by host pattern recognition receptors. Specifically, it is sensed by the endoplasmic reticulum protein STING (STimulator of INterferon Genes), which plays a pivotal role in modulating the host immune response. This interaction holds significant therapeutic potential, particularly in the context of oncologic patients, given the increasingly recognized relationship between the gut microbiota and tumour immunity27,28,29. c-di-AMP released by gut microbiota, particularly in individuals consuming a high-fibre diet, can signal through STING, remodelling macrophages and NK-dendritic cells interactions, ultimately promoting anti-tumour immunity and enhancing the efficacy of PD-1/PD-L1 blockade therapies, utilized to treat melanoma30. Consistently, elevated levels of c-di-AMP were observed in the responder group of hepatocellular carcinoma patients undergoing radiotherapy31. c-di-AMP synergized with radiotherapy, facilitating the maturation and antigen presentation functions of dendritic cells, thereby enhancing the production of interferon-beta (IFN-β) and promoting the activation of cytotoxic CD8 + T cells. This corroborates the hypothesis that c-di-AMP may function as a critical mediator through which the gut microbiota exerts immunomodulatory effects.
The biosynthesis of c-di-AMP is catalysed by the enzyme diadenylate cyclase (DAC), categorized in the five principal classes DisA, CdaA, CdaS, CdaM, and DacZ32. They all share a conserved catalytic domain (DisA_N, DAC) responsible for the cyclization reaction. The varied domain architectures, flanking this catalytic core, led to a more granular functional and phylogenetic distribution, subdividing these five principal classes into 22 distinct families33.
This work aimed to pinpoint the species within the human gut microbiota responsible for c-di-AMP synthesis. This was achieved through the identification of DACs and the subsequent construction of a comprehensive gut DAC database. To accomplish this objective, the Unified Human Gastrointestinal Protein (UHGP) catalogue34 was mined. This catalogue represents an extensive resource, comprising proteins derived from 289,232 genomes clustered into 4,744 distinct species, identified in human gut metagenomes. The DAC database was utilized to quantify the abundance and diversity of intestinal DACs in a curated dataset comprising 190 healthy subjects and 569 oncologic patients affected by metastatic melanoma, non-small-cell lung cancer and renal cell carcinoma. The metagenomic profiles of the patients were obtained from baseline samples collected prior to immunotherapy, with patients later categorized as responders (262) or non-responders (307) according to their clinical response to the treatment. This comparative analysis assessed the presence of differential DACs abundance profiles among healthy, oncologic, responder, and non-responder subjects, to identify other potential indications of the relationship between c-di-AMP and cancer, or c-di-AMP and response to immunotherapy.
Materials and methods
Construction of the DACs database
The Unified Human Gastrointestinal Protein (UHGP) catalogue34, clustered at 100% identity, was downloaded from Unified Human Gastrointestinal Genome collection (UHGG, v2.0.1, https://ftp.ebi.ac.uk/pub/databases/metagenomics/mgnify_genomes/human-gut/v2.0.1/protein_catalogue/). DAC sequences were searched with a two-step approach: first, all UHGP-100 sequences were queried against the HMM profile for the DAC family retrieved from the Pfam database (PF02457) integrated in InterPro (https://www.ebi.ac.uk/interpro/)35 with the hmmsearch function of HMMER v3.4 (hmmer.org). Hits with e-value > 0.01 were discarded. The results were further analysed with InterProScan36 to confirm that the proteins were uniquely identified by the DAC family profile and to assign them to a specific DAC type. The following InterPro family codes were utilized: DisA IPR023763, IPR050338; CdaA IPR034701; CdaS IPR034693, IPR053472; DacZ IPR014499; CdaM NF038327. A non-redundant version of the database with proteins clustered at 95% amino acid identity was built with MMseqs2 release 17-b804f37.
Metagenomes
The dataset herein analysed encompassed 190 metagenomic faecal samples from healthy subjects and 569 oncologic patients affected by metastatic melanoma (MM; 292), non-small-cell lung cancer (NSCLC; 239) and renal cell carcinoma (RCC; 38), classified as 262 responders (R; 158 MM, 90 NSCLC, 14 RCC) and 307 as non-responders (NR; 134 MM, 149 NSCLC, 24 RCC). The reads were collected from NCBI Sequence Read Archive (SRA) and are accessible with the Bioproject accession numbers listed in Suppl. Table 1. FASTQ files were downloaded with the function fasterq-dump of SRA Toolkit v. 3.0.5 (github.com/ncbi/sra-tools) and assessed for quality and primers presence with FastQC v0.11.838 to assure that only high-quality reads (length > 50 bp; quality score > 20) were further analysed. When necessary, the tool Trimmomatic v0.3939 was used for primer removal, employing ILLUMINACLIP setting with default values. Reads originating from human contamination were removed through the mapping of the human genome assembly GRCh38 with the tool bowtie240. Composition profiling was obtained utilising Kraken241 and Bracken42 with UHGG v. 2.0.134 as database. The compositions were imported into QIIME 243 as biom files with kraken-biom44 to conduct diversity analysis. Alpha diversity was investigated with Chao1, Pielou, and Shannon indices, with statistical significance assessed by Kruskal-Wallis test, while beta diversity was explored with Principal Coordinates Analysis (PCoA) based on Aitchison distance and significance among groups was evaluated with PERMANOVA. PCoA results were plotted with R package scatterplot3d v 0.3–4445. LefSe analysis46 was conducted to identify biomarkers characterizing healthy and oncologic patients.
DAC search in genomes and metagenomes
The DACs database was utilized to query the 4,744 reference genomes of UHGG collection with DIAMOND47, considering positive the hits with at least 90% identity and 50% query coverage. This result was linked to the same genomes present in the UHGG database built for Kraken2, in order to identify which species possessed c-di-AMP synthesis capabilities and, in case, which type of DAC. DACs abundance was extrapolated from metagenome’s composition to reflect the relative abundance of the corresponding species. The comparison between DACs abundances of healthy and oncologic subjects was carried out with Kruskal-Wallis test and Wilcoxon pairwise comparison with Benjamini-Hochberg (BH) correction, calculated with R software v. 4.5.1. Spearman correlation tests between DAC abundances and selected microbial taxa were carried out with R software v. 4.5.1.
Results
Screening and clustering of DAC proteins
The UHGP-100 catalogue, encompassing all proteins encoded by gut prokaryotes included in the UHGG, was screened with hmmsearch against the DAC family HMM profile PF02457 (e-value < 0.01). This search retrieved 17,665 proteins, of which 17,586 putative DAC sequences were validated by InterProScan, forming the DAC database-100. These proteins were subsequently clustered at 95% amino acid identity with MMseqs2, yielding the DAC database-95, which comprised 4,141 sequences assigned to the subtypes CdaA (3,310), DisA (788), DacZ (22), and CdaS (21) (Suppl. Spreadsheet 1).
Taxonomic distribution of DAC-encoding species
The DAC proteins included in database-95 were used to identify the DAC-encoding species. Among the 4,744 UHGG species, 3,901 harbored at least one DAC sequence. The vast majority of DAC-positive organisms were bacteria (3898/3901), with a smaller fraction belonging to archaea (3/3901). Across these 3,901 species, we identified 4,288 DAC sequences, divided into 3,163 CdaA, 1,082 DisA, 22 CdaS, and 21 DacZ. No CdaM sequence was found (Table 1, Suppl. Spreadsheet 1).
DAC-encoding prokaryotic taxa were widely distributed, spanning 19 phyla, 28 classes, 77 orders, 178 families, and 911 genera (Suppl. Spreadsheet 1). The phylum Bacillota A included the largest number of species encoding CdaA (1,549), followed by Bacillota (400) (Table 1). Bacteroidota was also largely characterized by the presence of only CdaA (562/619 species). In contrast, DisA was mainly associated with Actinobacteriota, whereas most species encoding both CdaA and DisA belonged to Bacillota A (176). Verrucomicrobiota were predominantly characterized by the presence of CdaA (39) and, with 19 out of the 21 species identified, represented the main phylum harboring DacZ. Notably, CdaS was exclusively encoded by Bacillota (22 species). With only a few exceptions, Proteobacteria lacked any DAC-encoding genes.
Notable differences in terms of DAC genes distribution emerged at genus level. With the exception of Bifidobacterium adolescentis, all other bifidobacterial species appeared unable to synthesize c-di-AMP. Collinsella spp. typically encoded DisA. Members of Bacteroides, Parabacteroides, Phocaeicola, and Prevotella predominantly carried CdaA, whereas Alistipes spp. encoded either CdaA alone (44) or CdaA together with DisA (10). Among Lactobacillales, the vast majority (160/168) displayed the potential to produce c-di-AMP due to the presence of CdaA. Within the genus Paenibacillus (including clades Paenibacillus A, B, F, and L), multiple gene combinations were identified: 6 strains carried DisA + CdaA + CdaS, 10 carried DisA + CdaA, and one carried CdaA + CdaS.
Most members of Clostridiaceae (44/47) and Peptostreptococcaceae (22/25), as well as Peptococcaceae (class Peptococcia), encoded both DisA and CdaA. By contrast, Lachnospiraceae, Oscillospiraceae, and Ruminococcaceae typically harboured only CdaA, a feature also observed in the genus Akkermansia.
Abundance of DAC-encoding species in healthy versus oncologic patients
Two datasets were generated: one comprising healthy subjects (190, H) and another including cancer patients (569, C) diagnosed with melanoma, non-small-cell lung cancer, or renal cell carcinoma. The latter dataset consisted of baseline faecal samples from patients undergoing immunotherapy, further stratified into responders (262, R) and non-responders (307, NR). To investigate potential differences in DAC abundance, the two metagenome datasets were compared across healthy subjects and oncologic patients, with the latter further subdivided into responders and non-responders to immunotherapy.
Alpha diversity assessed using the Chao1 index was comparable between healthy subjects (H) and cancer patients (C), as well as the subgroups of responders (R) and non-responders (NR), indicating a similar species richness across groups (Fig. 1). In contrast, both Pielou’s evenness and the Shannon index were significantly lower in the C, R, and NR groups compared to H, suggesting that although the total number of species was similar, the distribution of species abundances was less even, and overall diversity was reduced in cancer patients.
Alpha diversity calculated for Chao index, Pielou’s evenness and Shannon index of the healthy, oncologic, NR and R samples. Significance: * p < 0.05. In the boxplots, the upper edge indicates 75th percentile (Q3), the lower edge the 25th percentile (Q1), the inner line the median and the X the mean value. Whiskers extend to 1.5 × the interquartile range (IQR) beyond the first and third quartiles.
Although the beta-diversity plots show considerable overlap among groups (Suppl. Figure 1), PERMANOVA revealed significant differences (p < 0.05) in the overall microbial community composition between healthy subjects and cancer patients, responders, and non-responders, as well as between responders and non-responders to therapy.
From the LEfSe analysis performed to identify taxa enriched in healthy subjects compared with oncological patients (R + NR), one of the most robust biomarkers associated with cancer patients was E. coli, together with all higher taxonomic ranks up to the phylum Proteobacteria (Suppl. Figure 2).
Considering the abundance of DAC-encoding species (Fig. 2a, Suppl. Table 2), healthy subjects harboured, on average, 95.9% of gut bacteria with the potential to synthesize DACs. This proportion was significantly lower in cancer patients, including both R and NR, with mean values of 91.7, 92.2, and 91.2%, respectively. The C, R, and NR groups also exhibited lower medians compared to H (Fig. 2a), accompanied by higher standard deviations (s.d. = .± 6.0%; 11.1%; 12.3%; 9.7% in H, C, NR, and R, respectively), reflecting the presence of several outliers with low and extremely low abundances of c-di-AMP producers in cancer patients. Although not statistically significant, responders showed higher mean and median abundances of DAC-encoding species compared to non-responders, accompanied by a lower standard deviation, suggesting a trend towards a more consistent presence of c-di-AMP producers in R, which could be associated with a healthier or more resilient gut microbiota.
Relative abundance of DAC-encoding species in healthy, oncologic, NR and R subjects (a) and in healthy MM, NSCLC, and RCC (b). Significance: * p < 0.05. In the boxplots, the upper edge indicates 75th percentile (Q3), the lower edge the 25th percentile (Q1), the inner line the median and the X the mean value. Whiskers extend to 1.5 × the interquartile range (IQR) beyond the first and third quartiles.
E. coli, Proteobacteria and Bifidobacteria resulted significantly more abundant in oncologic patients than in healthy subjects (Kruskal-Wallis test, p < 0.05), according to the absence of DAC genes and lower potential to produce c-di-AMP and negatively correlated with DAC abundance (Suppl. Table 3).
When accounting for cancer type, the relative abundance of DAC-encoding species in the healthy cohort was again statistically higher than MM and NSCLC patients, whereas it lacked statistical significance compared with the RCC cohort (Fig. 2b). Comparisons between NR and R patients within the MM, NSCLC, and RCC cohorts presented no significant differences (Suppl. Table 4).
Discussion
This analysis provides a comprehensive overview of DAC-encoding species in the human gut microbiome and their potential relevance in cancer immunotherapy response. The extensive screening of the UHGP-100 catalogue revealed that DAC proteins are widespread across gut prokaryotes, with 3,901 out of 4,744 UHGG species harbouring at least one DAC gene. The predominance of CdaA across most bacterial phyla, and the more limited distribution of DisA, DacZ, and CdaS, highlights the evolutionary and functional diversification of diadenylate cyclases in gut microbes. Bacillota and related clades represented the bulk of DAC-encoding species, whereas Proteobacteria largely lacked these genes.
At finer taxonomic resolution, clear genus-level patterns emerged. While Bifidobacteria, except for B. adolescentis, appeared largely incapable of c-di-AMP synthesis, most of the other dominant taxa such as Bacteroides, Lachnospiraceae, Oscillospiraceae, and Ruminococcaceae exhibited strong potential for DAC production. Notably, several genera of Bacillota/Bacillota A, including Clostridiaceae and Peptostreptococcaceae, encoded multiple DAC classes, suggesting that certain gut bacteria may employ complex c-di-AMP signalling networks.
When comparing healthy subjects and cancer patients, alpha diversity analyses indicated similar species richness across groups, but revealed reduced evenness and Shannon diversity in patients. This pattern suggests that, although the total number of bacterial species is maintained, the gut microbiota of oncologic patients is dominated by fewer taxa, leading to an uneven community structure. Complementing these findings, beta diversity analyses confirmed that the overall microbial composition of patients differed significantly from that of healthy subjects, despite substantial overlap in the ordination plots. This indicates a subtle but consistent shifts in community structure may underlie cancer associated differences.
Importantly, the abundance of DAC-encoding species was significantly lower in cancer patients compared to healthy subjects, reflecting a reduced potential for c-di-AMP production. The fact that different levels of c-di-AMP were found in healthy individuals compared with cancer patients clearly stems from the differing abundances of Proteobacteria and Bifidobacteria. The absence of DAC genes in these two groups indicates a consistent evolutionary trajectory and does not appear to be attributable to the loss of metabolic functions in certain species. These findings indicate that cancer patients generally harbour a gut microbiota with diminished capacity to produce c-di-AMP. This observation raises the intriguing possibility that reduced microbial c-di-AMP synthesis may contribute to tumour development or progression, potentially by impairing host immune homeostasis or altering microbe-mediated signalling pathways. While causality cannot be inferred from these data, the consistent depletion of c-di-AMP-producing bacteria points to a potential mechanistic link between microbial signalling and oncogenesis, as well as a possible role in impairing and diminishing immunotherapy efficacy.
Even though our study also demonstrates that the capacity to produce c-di-AMP is widespread among intestinal bacteria, tumour occurrence seems to be associated with the increased abundance of the two major microbial groups unable to produce c-di-AMP, namely Bifidobacterium and Proteobacteria. Consistently, E. coli, Proteobacteria, and Bifidobacterium negatively correlate with the abundance of DAC-producing species.
The LEfSe analysis comparing healthy individuals with cancer patients highlighted E. coli as one of the most consistent microbial biomarkers associated with cancer, together with higher taxonomic levels up to the phylum Proteobacteria. These observations may suggest that Enterobacteriaceae, through their pro-inflammatory activity, could contribute to tumour development, a process that might not be adequately counteracted by the STING pathway in the context of reduced c-di-AMP levels.
In the study, we also focused on possible differences between responders versus not-responder patients, searching for unexplored predictive markers to immunotherapy. Although responders to immunotherapy exhibited slightly higher mean and median abundances of DAC-encoding species than non-responders, the difference was not statistically significant; however, the lower standard deviation among responders suggests a more consistent presence of c-di-AMP producers. When accounting for the type of cancer and the response, the same pattern was confirmed, without reaching statistical significance. However, in MM and NSCLC, the general trend showed higher mean and median and lower standard deviation in R than in NR.
In the gut microbiota-immunotherapy axis, c-di-AMP shapes host immune responses through STING signalling, which triggers type I IFN production in host cells. For example, c-di-AMP produced by A. muciniphila stimulates the activation of intertumoral monocytes, enhances type I IFN secretion, and fosters an immune-active tumour microenvironment30. This, in turn, drives the activation of NK cells and dendritic cells, ultimately supporting the efficacy of immune checkpoint blockade therapy in mice.
These findings underscore the substantial potential of c-di-AMP producers as immune system modulators. In vivo studies have already demonstrated that engineered E. coli strains producing c-di-AMP can effectively reinforce antitumor immune responses and therapeutic potency48.
The utilization of an engineered strain of Salmonella typhimurium to produce and deliver c-di-AMP resulted in an enhanced stimulation of immune system through STING activation in mice49. Furthermore, in animal models, a high-fibre diet has been shown to enrich for the c-di-AMP producer Akkermansia, which in turn triggers type I interferon (IFN-I) production via STING-c-di-AMP interaction30. It is noteworthy that the study of Li et al. (2022)31 correlates the response to anticancer radiotherapy with faecal c-di-AMP levels and identifies the abundance of Bifidobacterium-related taxa across multiple taxonomic levels as a potential biomarker of non-responders. These findings highlight the potential of targeting specific gut bacterial functions, such as c-di-AMP synthesis, to optimize immunotherapy outcomes.
Our results might appear to contrast with limited existing data on c-di-AMP levels in oncologic patients, since we didn’t observe significant differences between R and NR. It is crucial to recognize a key distinction: our study estimated the relative abundance of DAC-encoding species within the gut microbiota, without assessing the level of c-di-AMP in faeces, like observed by Li et al. (2022)31, as well as it did not directly investigate the c-di-AMP effects on immune system stimulation, as seen by Lam et al. (2021)30. However, our findings underscore the potential of c-di-AMP as an immune system modulator and open new and important perspectives that warrant validation through clinical trials to demonstrate that c-di-AMP produced by the gut microbiota can enhance the response to oncologic therapies that benefit from immune stimulation. The scientific literature associating c-di-AMP with STING-driven type I interferon responses in cancer immunity largely relies on review articles27,50,51,52, and our results emphasize the importance of advancing toward clinical validation.
A critical factor that might explain why our results showed no difference between R and NR groups is the unclear mechanism of c-di-AMP release into the extracellular environment and its subsequent uptake by immune cells. In Listeria monocytogenes, c-di-AMP is secreted through the multidrug efflux pumps MdrM, drT, MdrA, and MdrC (MDR), of the MFS superfamily. In the host cytosol, the secreted c-di-AMP induces a host type I interferon transcriptional response53. While it seems counterintuitive for a pathogen to release an inflammatory molecule that could lead to immune detection, one hypothesis suggests that coupling c-di-AMP secretion to MDR activity offers a mechanism independent from phosphodiesterase, the enzyme responsible for c-di-AMP degradation, to reduce intracellular nucleotide levels54. This could be particularly advantageous under various environmental stresses54. Beyond c-di-AMP, MdrT and, to a lesser extent, MdrM also function as efflux pumps for bile acids55. Given that bile-induced transcriptomes significantly overlap with those of major virulence regulators and stress factor, c-di-AMP might also serve as an intracellular stress reporter, with its secretion potentially linked to specific stress responses54. While c-di-AMP secretion has been most thoroughly characterized in L. monocytogenes, MFS transporters are broadly conserved across bacteria, suggesting that the ability to secrete c-di-AMP could extend to many other microorganisms. However, it’s important to note that mutations in all MdrM and MdrT homologs in Staphylococcus aureus resulted in only a negligible change in extracellular c-di-AMP levels56, indicating that c-di-AMP secretion by MDRs may be organism-specific. Therefore, further research is needed to determine if additional c-di-AMP-specific transporters exist beyond MDRs.
Overall, our results suggest that c-di-AMP-producing bacteria are a core component of the healthy gut microbiome, and that reductions in their abundance may characterize the dysbiotic microbiota observed in cancer patients. The trends linking DAC abundance to immunotherapy response, though preliminary, provide a rationale for further mechanistic studies to explore the role of microbial c-di-AMP signaling in modulating anti-tumor immunity. The study provides insights into the human gut microbiota species responsible for c-di-AMP synthesis. These findings can guide the selection of individual species or consortia as potential adjuvants in oncology, enabling in-depth characterization to determine their precise contribution in modulating the immune system.
Data availability
UHGP fasta file, UHGG Kraken2 database and UHGG reference genomes are available at UHGG repository (https://ftp.ebi.ac.uk/pub/databases/metagenomics/mgnify_genomes/human-gut/v2.0.1/). Accession numbers of metagenomes utilized in this work are reported in Suppl. Table 1.
References
Witte, G., Hartung, S., Buttner, K. & Hopfner, K. P. Structural biochemistry of a bacterial checkpoint protein reveals diadenylate cyclase activity regulated by DNA recombination intermediates. Mol. Cell. 30, 167–178. https://doi.org/10.1016/j.molcel.2008.02.020 (2008).
Corrigan, R. M. & Gründling, A. Cyclic di-AMP: another second messenger enters the fray. Nat. Rev. Microbiol. 11, 513–524. https://doi.org/10.1038/nrmicro3069 (2013).
Römling, U. Great times for small molecules: c-di-AMP, a second messenger candidate in bacteria and archaea. Sci. Signal. 1, pe39. https://doi.org/10.1126/scisignal.133pe39 (2008).
Braun, F. et al. Cyclic nucleotides in archaea: Cyclic di-AMP in the archaeon haloferax volcanii and its putative role. Microbiologyopen 8(9), e00829. https://doi.org/10.1002/mbo3.829 (2019).
He, J., Yin, W., Galperin, M. Y. & Chou, S. H. Cyclic di-AMP, a second messenger of primary importance: tertiary structures and binding mechanisms. Nucleic Acids Res. 48, 2807–2829. https://doi.org/10.1093/nar/gkaa112 (2020).
Yin, W. et al. A decade of research on the second messenger c-di-AMP. FEMS Microbiol. Rev. 44, 701–724. https://doi.org/10.1093/femsre/fuaa019 (2020).
Stülke, J. & Krüger, L. Cyclic di-AMP signaling in bacteria. Annu. Rev. Microbiol. 74, 159–179. https://doi.org/10.1146/annurev-micro-020518-115943 (2020).
Oppenheimer-Shaanan, Y., Wexselblatt, E., Katzhendler, J., Yavin, E. & Ben-Yehuda, S. c-di-AMP reports DNA integrity during sporulation in Bacillus subtilis. EMBO Rep. 12, 594–601. https://doi.org/10.1038/embor.2011.77 (2011).
Zhang, L. & He, Z. G. Radiation-sensitive gene A (RadA) targets DisA, DNA integrity scanning protein A, to negatively affect cyclic di-AMP synthesis activity in Mycobacterium smegmatis. J. Biol. Chem. 288, 22426–22436. https://doi.org/10.1074/jbc.M113.464883 (2013).
Sureka, K. et al. The cyclic dinucleotide c-di-AMP is an allosteric regulator of metabolic enzyme function. Cell 158, 1389–1401. https://doi.org/10.1016/j.cell.2014.07.046 (2014).
Whiteley, A. T., Garelis, N. E., Peterson, B. N., Choi, P. H. & Tong, L. c-di-AMP modulates Listeria monocytogenes central metabolism to regulate growth, antibiotic resistance and osmoregulation. Mol. Microbiol. 104, 212–233. https://doi.org/10.1111/mmi.13622 (2017).
Tang, Q. et al. Functional analysis of a c-di-AMP-specific phosphodiesterase MsPDE from Mycobacterium smegmatis. Int. J. Biol. Sci. 11, 813–824. https://doi.org/10.7150/ijbs.11797 (2015).
Krüger, L. et al. Sustained control of pyruvate carboxylase by the essential second messenger cyclic di-AMP in Bacillus subtilis. mBio. 13, e0360221. https://doi.org/10.1128/mbio.03602-21 (2022).
Mehne, F. M. et al. Control of the diadenylate cyclase CdaS in Bacillus subtilis: an autoinhibitory domain limits cyclic di-AMP production. J. Biol. Chem. 289, 21098–21107. https://doi.org/10.1074/jbc.M114.562066 (2014).
Zheng, C. et al. Functional analysis of the sporulation-specific diadenylate cyclase CdaS in Bacillus thuringiensis. Front. Microbiol. 6, 908. https://doi.org/10.3389/fmicb.2015.00908 (2015).
Zarrella, T. M., Yang, J., Metzger, D. W. & Bai, G. The bacterial second messenger cyclic di-AMP modulates the competence state in Streptococcus pneumoniae. J. Bacteriol. 202, e00691–e00619. https://doi.org/10.1128/JB.00691-19 (2020).
Bowman, L., Zeden, M. S., Schuster, C. F., Kaever, V. & Gründling, A. New insights into the cyclic di-adenosine monophosphate (c-di-AMP) degradation pathway and the requirement of the cyclic dinucleotide for acid stress resistance in Staphylococcus aureus. J. Biol. Chem. 291, 26970–26986 (2016).
Rubin, B. E. et al. High-throughput interaction screens illuminate the role of c-di-AMP in cyanobacterial nighttime survival. PLoS Genet. 14, e1007301. https://doi.org/10.1371/journal.pgen.1007301 (2018).
Gundlach, J., Rath, H., Herzberg, C., Mäder, U. & Stülke, J. Second messenger signaling in Bacillus subtilis: accumulation of cyclic di-AMP inhibits biofilm formation. Front. Microbiol. 7, 804. https://doi.org/10.3389/fmicb.2016.00804 (2016).
Peng, X., Zhang, Y., Bai, G., Zhou, X. & Wu, H. Cyclic di-AMP mediates biofilm formation. Mol. Microbiol. 99, 945–959. https://doi.org/10.1111/mmi.13277 (2016).
Latoscha, A. et al. c-di-AMP hydrolysis by the phosphodiesterase AtaC promotes differentiation of multicellular bacteria. Proc. Natl. Acad. Sci. U S A. 117, 7392–7400. https://doi.org/10.1073/pnas.1917080117 (2020).
Bai, Y. et al. Two DHH subfamily 1 proteins in Streptococcus pneumoniae possess cyclic di-AMP phosphodiesterase activity and affect bacterial growth and virulence. J. Bacteriol. 195, 5123–5132. https://doi.org/10.1128/JB.00769-13 (2013).
Witte, C. E. et al. Cyclic di-AMP is critical for Listeria monocytogenes growth, cell wall homeostasis, and establishment of infection. mBio 4, e00282–e00213. https://doi.org/10.1128/mBio.00282-13 (2013).
Yang, J. et al. Deletion of the cyclic di-AMP phosphodiesterase gene (cnpB) in Mycobacterium tuberculosis leads to reduced virulence in a mouse model of infection. Mol. Microbiol. 93, 65–79. https://doi.org/10.1111/mmi.12641 (2014).
Ye, M. et al. DhhP, a cyclic di-AMP phosphodiesterase of Borrelia burgdorferi, is essential for cell growth and virulence. Infect. Immun. 82, 1840–1849. https://doi.org/10.1128/IAI.00030-14 (2014).
Moradali, M. F. et al. Atypical cyclic di-AMP signaling is essential for Porphyromonas gingivalis growth and regulation of cell envelope homeostasis and virulence. NPJ Biofilms Microbiomes. 8, 53. https://doi.org/10.1038/s41522-022-00316-w (2022).
Lu, Y. et al. Gut microbiota influence immunotherapy responses: mechanisms and therapeutic strategies. J. Hematol. Oncol. 15(1), 47. https://doi.org/10.1186/s13045-022-01273-9 (2022).
Wang, M., Zhang, L., Chang, W. & Zhang, Y. The crosstalk between the gut microbiota and tumor immunity: implications for cancer progression and treatment outcomes. Front. Immunol. 13, 1096551. https://doi.org/10.3389/fimmu.2022.1096551 (2023).
Gazzaniga, F. S. & Kasper, D. L. The gut microbiome and cancer response to immune checkpoint inhibitors. J. Clin. Invest. 135(3), e184321. https://doi.org/10.1172/JCI184321 (2025).
Lam, K. C. et al. Microbiota triggers STING-type I IFN-dependent monocyte reprogramming of the tumor microenvironment. Cell 184(21), 5338–5356e21. https://doi.org/10.1016/j.cell.2021.09.019 (2021).
Li, Z. et al. Gut microbiota modulate radiotherapy-associated antitumor immune responses against hepatocellular carcinoma via STING signaling. Gut Microbes. 14(1), 2119055. https://doi.org/10.1080/19490976.2022.2119055 (2022).
Commichau, F. M., Heidemann, J. L., Ficner, R. & Stülke, J. Making and breaking of an essential poison: the cyclases and phosphodiesterases that produce and degrade the essential second messenger cyclic di-AMP in bacteria. J. Bacteriol. 201(1), e00462–e00418. https://doi.org/10.1128/JB.00462-18 (2018).
Galperin, M. Y. All DACs in a row: domain architectures of bacterial and archaeal diadenylate cyclases. J. Bacteriol. 205(4), e0002323. https://doi.org/10.1128/jb.00023-23 (2023).
Almeida, A. et al. A unified catalog of 204,938 reference genomes from the human gut microbiome. Nat. Biotechnol. 39(1), 105–114. https://doi.org/10.1038/s41587-020-0603-3 (2021).
Blum, M. et al. The interpro protein families and domains database: 20 years on. Nucleic Acids Res. 49(D1), D344–D354. https://doi.org/10.1093/nar/gkaa977 (2021).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30(9), 1236–1240. https://doi.org/10.1093/bioinformatics/btu031 (2014).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35(11), 1026–1028. https://doi.org/10.1038/nbt.3988 (2017).
Andrews, S. FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30(15), 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with bowtie 2. Nat. Methods. 9(4), 357–359. https://doi.org/10.1038/nmeth.1923 (2012).
Wood, D. E., Lu, J. & Langmead, B. Improved metagenomic analysis with Kraken 2. Genome Biol. 20(1), 257. https://doi.org/10.1186/s13059-019-1891-0 (2019).
Lu, J., Breitwieser, F. P., Thielen, P. & Salzberg, S. L. Bracken: estimating species abundance in metagenomics data. PeerJ Comput. Sci. 3, e104. https://doi.org/10.7717/peerj-cs.104 (2017).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37(8), 852–857. https://doi.org/10.1038/s41587-019-0209-9 (2019).
Dabdoub, S. M. kraken-biom: Enabling interoperative format conversion for Kraken results (Version 1.2) [Software]. https://github.com/smdabdoub/kraken-biom (2016).
Ligges, U. & Mächler, M. Scatterplot3d - an R package for visualizing multivariate data. J. Stat. Softw. 8(11), 1–20. https://doi.org/10.18637/jss.v008.i11 (2003).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12(6), R60. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Buchfink, B., Reuter, K. & Drost, H. G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods. 18, 366–368. https://doi.org/10.1038/s41592-021-01101-x (2021).
Jiang, Y., Li, X., Qian, F., Sun, B., Wang, X. & et Fine-tuning bacterial cyclic di-AMP production for durable antitumor effects through the activation of the STING pathway. Research 6, Article 0102. https://doi.org/10.34133/research.0102 (2023).
Huang, Y., Piao, L. & Liu, X. Enhancing tumor-specific immunity with SLdacA: A attenuated Salmonella-mediated c-di-AMP delivery system targeting the STING pathway. Int. J. Pharm. 666, 124759. https://doi.org/10.1016/j.ijpharm.2024.124759 (2024).
Wang, B. et al. Clinical applications of STING agonists in cancer immunotherapy: current progress and future prospects. Front. Immunol. 15, 1485546. https://doi.org/10.3389/fimmu.2024.1485546 (2024).
Cheng, X., Ning, J., Xu, X. & Zhou, X. The role of bacterial cyclic di-adenosine monophosphate in the host immune response. Front. Microbiol. 13, 958133. https://doi.org/10.3389/fmicb.2022.958133 (2022).
Zarrella, T. M. & Bai, G. The bacterial second messenger cyclic di-AMP and inflammation. In: (ed Wang, Y. X.) Inflammation. Springer, Cham. https://doi.org/10.1007/978-3-031-55254-0_1-1 (2025).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. C-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705. https://doi.org/10.1126/science.1189801 (2010).
Huynh, T. N. & Woodward, J. J. Too much of a good thing: regulated depletion of c-di-AMP in the bacterial cytoplasm. Curr. Opin. Microbiol. 30, 22–29. https://doi.org/10.1016/j.mib.2015.12.007 (2016).
Quillin, S. J., Schwartz, K. T. & Leber, J. H. The novel Listeria monocytogenes bile sensor BrtA controls expression of the cholic acid efflux pump MdrT. Mol Microbiol. 81(1), 129–142. https://doi.org/10.1111/j.1365-2958.2011.07683.x (2011).
Gries, C. M. et al. Cyclic di-AMP released from Staphylococcus aureus biofilm induces a macrophage type I interferon response. Infect. Immun. 84(12), 3564–3574. https://doi.org/10.1128/IAI.00447-16 (2016).
Funding
The research activities have been funded by PIANO NAZIONALE DI RIPRESA E RESILIENZA(PNRR) - MISSIONE 4 COMPONENTE 2, “Dalla ricerca all’impresa” INVESTIMENTO 1.3, Creazione di “Partenariati estesi alle università, ai centri di ricerca, alle aziende per il finanziamento di progetti di ricerca di base”, finanziato dall’Unione europea - NextGenerationEU” - Progetto identificato con codice PE00000019, Titolo “HEAL ITALIA” - Spoke 5 - CUP E93C22001860006 Avviso MUR DD. 341 del 15.03.2022.
Author information
Authors and Affiliations
Contributions
Conceptualization: Rossi M, Amaretti A, Raimondi SInvestigation: Candeliere F, Sola LFormal analysis: Candeliere F, Busi ESoftware: Candeliere FVisualization: Candeliere F, Amaretti A, Pedroni SWriting - original draft: Candeliere F, Rossi M, Pedroni S, Sola L, Amaretti A, Raimondi S, Greco S, Dominici MSupervision: Rossi MAll authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Candeliere, F., Sola, L., Busi, E. et al. Altered abundance in cancer patients gut of diadenylate cyclase-encoding bacteria. Sci Rep 16, 6070 (2026). https://doi.org/10.1038/s41598-026-35425-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-35425-5