Abstract
Dyslipidemia is associated with a low-grade inflammatory process modified by the immune response, mainly monocytes and macrophages. Metabolically activated macrophages (characterized by the expression of CD36, ABCA1 and PLIN2 receptors, and by exhibit trimethylation of the lysine 4 and 27 of the histone 3) have been implicated in obesity and dyslipidemia. However, it has not been reported a population of circulating monocytes with the same phenotype. We hypothesized that continuous exposure to low-density lipoproteins (LDL-c) induces epigenetic changes in monocytes, leading to their polarization into metabolically activated macrophages. We investigated the phenotypic expression of circulating monocyte and macrophage subsets in 23 individuals living with LDL-c dyslipidemia. Using dimensionality reduction and clustering algorithms within a comprehensive analytical model, we analyzed flow cytometry data to evaluate epigenetic changes, specifically H3K4me3 and H3K27me3, in clustered subsets. This model allowed us to identify two statistically significant phenotypes of metabolically activated monocytes that were more prevalent in dyslipidemia patients than in controls, exhibiting distinct expression levels of trimethylation of the lysine 4 and 27 of the histone 3. This approach allowed us to delineate different monocyte and macrophages associated to the metabolically activated phenotype in dyslipidemia.
Similar content being viewed by others
Introduction
Atherosclerosis is characterized by the aggregation of lipids and fibrous material within the arterial intima, leading to plaque formation. Hypercholesterolemia, especially elevated levels of low-density lipoprotein cholesterol (LDL-c) and its oxidized form, has been extensively studied and identified as a major causal factor in atherosclerosis.
The pathogenesis of atherosclerosis is a complex and multifactorial process involving lipid accumulation, endothelial dysfunction, and chronic inflammation1,2,3. Monocytes and Macrophages play essential roles in all stages of atherosclerosis, contributing to lipid uptake, inflammation, and plaque progression4,5. It has been proven that defective monocyte and macrophage function promotes the accumulation of apoptotic cells, which in turn promotes atherosclerosis.
Traditionally, monocyte subsets have been identified by surface marker expression, including the lipopolysaccharide (LPS) receptor CD14 and the Fc gamma receptor III (CD16)3,6,7. It is well established that classical monocytes with CD14 + + CD16 − phenotype, intermediate monocytes (CD14 + CD16+), and nonclassical monocytes (CD14 + CD16++) possess distinct functional properties and contribute differentially to the inflammatory processes in atherosclerotic lesions. For instance, classical monocytes are more prone to differentiate into macrophages that populate atherosclerotic plaques, while intermediate monocytes produce significant quantities of pro-inflammatory molecules. Nonclassical monocytes play a role in patrolling and maintaining endothelial integrity5,6. In addition, macrophages have been classified into two major phenotypes: M1, which are pro-inflammatory and promote plaque growth, and M2, which are anti-inflammatory and involved in tissue repair4,8.
However, emerging evidence suggests a broader spectrum of monocyte and macrophage polarization states, including metabolically activated monocytes (MoMe) and metabolically activated macrophages (MMe)9.
MMe overexpresses ATP-binding cassette transporter (ABCA1), cluster of differentiation 36 (CD36), and perilipin 2 (PLIN2), which are regulated by peroxisome proliferator-activated receptor gamma (PPARγ) and p62 protein. One important feature of MMe is the epigenetic mark characterized by increased trimethylation of lysine 4 on histone 3 (H3K4me3), which is associated with the expression of inflammatory genes (tnf-a, il-6, il-18, mcp-1, cd36, and sr-a), this epigenetic and metabolic reprogramming corresponds to a phenomenon known as “trained immunity.” In this state, cells retain a “memory” of prior inflammatory stimuli, leading to an enhanced inflammatory response upon re-exposure to similar stimuli. These metabolically active cells are believed to be pivotal in chronic inflammation and in the progression of atherosclerosis10,11,12,13,14. Our group has reported one population of monocytes with a metabolically activated profile and the ability to differentiate into MMe in vitro15. Dyslipidemia facilitates this differentiation; however, the metabolic, phenotypic, and epigenetic markers characteristic of MMe in circulating monocytes have not been addressed. In this study, we aimed to identify MoMe not only by surface receptor expression but also by epigenetic mark levels in subjects with dyslipidemia. We hypothesized that continuous exposure to low-density lipoproteins (LDL-c) induces epigenetic changes in monocytes, favoring their polarization into metabolically activated macrophages with atherogenic potential. The use of advanced analysis, such as high-dimensional reduction algorithms and complex clustering analysis, has enhanced the ability to identify cell subsets with greater precision, facilitating the discovery of previously unknown phenotypes16,17,18,19,20.
In this study, we aimed to identify MoMe not only by surface receptor expression but also by epigenetic mark levels in subjects with dyslipidemia.
Materials and methods
Samples
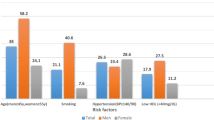
A total of 23 blood samples from patients with a clinical diagnosis of dyslipidemia were analyzed. Dyslipidemia was defined in accordance with the 2022 Mexican Clinical Practice Guidelines for the Diagnosis and Treatment of Dyslipidemia and Atherosclerotic Cardiovascular Disease21. Inclusion criteria included total cholesterol > 200 mg/dL, triglycerides > 150 mg/dL, and low-density lipoprotein (LDL-c) cholesterol > 130 mg/dL, given the elevated cardiovascular risk associated with high LDL-c levels. A control group of 17 healthy individuals with normal lipid profiles (cholesterol < 200 mg/dL, triglycerides < 150 mg/dL, and LDL-c < 100 mg/dL) was included. Clinical data, including age, sex, and relevant medical history (e.g., obesity, hypertension, diabetes mellitus, familial hypercholesterolemia, polycystic ovary syndrome, and overweight), were collected using a standardized clinical assessment tool developed by our laboratory. Lifestyle factors such as dietary habits, alcohol and tobacco use, and physical activity were documented. Anthropometric measurements (weight, height, BMI, waist and hip circumference, body fat percentage, visceral fat, muscle mass percentage, and metabolic age) were obtained using bioelectrical impedance analysis. None of the participants was under pharmacological treatment at the time of blood collection, and all were free from signs of infection. All patients showed secondary dyslipidemia.
Ethics
All procedures involving human participants adhered to the World Medical Association’s Declaration of Helsinki. After explaining the project, all participants provided written informed consent, and blood samples were collected. This work was approved by our local research and bioethical committee, Research Ethics Committee of the Faculty of Dentistry UASLP-CONBIOÉTICA-24-CEI-001–20190213, under approval number CEI-FE-014–019.
Cell isolation and culture
Peripheral blood mononuclear cells (PBMCs) were isolated from blood samples of individuals that live with dyslipidemia patients and healthy controls using Ficoll-Paque density gradient centrifugation. Isolated PBMCs were maintained in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS) until use. Cell viability and counting were assessed using the trypan blue exclusion method with a Corning® Automated Cell Counter, following the manufacturer’s protocol. Only samples with a minimum cell viability of 95% were used in subsequent experiments.
CD14 + monocytes were purified with MACS CD14 magnetic beads (Miltenyi Biotec) and cultured in 24-well plates (Costar) at a density of 0.8 million cells per well in RPMI medium supplemented with 10% FBS, 10 nM glutamine, 100 µg/mL streptomycin, and 100 U/mL penicillin. Cell cultures were incubated for six days at 37 °C with 5% CO2. To induce differentiation into metabolically activated macrophages (MMe), monocytes were incubated with 50 ng/mL M-CSF for six days, followed by stimulation with 15 mM glucose, 10 nM insulin, and 0.4 mM palmitate (Sigma-Aldrich) for an additional 24 h. After differentiation, cells were harvested using EDTA and prepared for flow cytometric analysis.
Antibody panels
Antibodies used for flow cytometry included those targeting CD14, CD16, CD36, ABCA1, PLIN2, H3K4me3, H3K9ac, H3K27me3, CD80, CD11b, and CD206. These antibodies were purchased from BD Biosciences (San Jose, CA), Novus Biologicals (Littleton, CO), BioLegend (San Diego, CA), Cell Signaling (San Diego, CA), Invitrogen (San Diego, CA), and BD Pharmingen (San Diego, CA).
Cell staining and flow cytometry acquisition
For flow cytometry analysis, PBMCs and monocyte-derived macrophages were prepared at concentrations of 5 × 10^6 cells/mL and 1 × 10^6 cells/mL, respectively. Cells were placed in 12 × 75 mm tubes for staining. Extracellular antibodies from the designated panel (Supplementary Tables S1 and S2) were added for cell surface staining, and cells were incubated in the dark for 1 h at 4 °C. Following incubation, cells were washed adding 1 mL of PBS and centrifuged at 1500 rpm for 5 min to prepare for intracellular staining.
For intracellular staining, cells were fixed by adding 100 µL of 1% formaldehyde on ice for 10 min, followed by washing with PBS containing 2% FBS and centrifugation at 1500 rpm for 5 min. Cells were permeabilized using 200 µL of 0.1% Triton X-100 in PBS buffer for 10 min at 4 °C, washed with PBS containing 2% FBS, and centrifuged at 1500 rpm for 5 min.
Intracellular and intranuclear staining was performed according to the specifications of the intracellular/intranuclear antibody panel. Cells were incubated with conjugated antibodies for 1 h at 4 °C, washed with 1 mL of PBS containing 2% FBS, centrifuged for 5 min at 1500 rpm, and resuspended in 1 mL of PBS for immediate flow cytometry analysis.
Flow cytometry samples were acquired within three hours of staining using a FACS Canto II cytometer (BD Biosciences, San Jose, CA) at a flow rate of 60 µL/min, and FCS files were exported in version 3.1 format. Flow cytometry data analysis was conducted in FlowJo® version 10.10, supplemented with plugins and R packages (4.1.1) from Bioconductor (3.20)22,23,24.
Enhanced bioinformatic workflow for flow cytometry data analysis
To enhance the clarity and precision of our cellular analysis, particularly for identifying metabolically activated monocyte subpopulations, we employed a comprehensive bioinformatic workflow. This approach integrates advanced computational methods with traditional flow cytometry, minimizing subjective variability and enabling a deeper understanding of complex cellular phenotypes. Our methodology addresses the limitations of manual gating, especially when dealing with high-dimensional datasets and numerous fluorochromes, by automating key analytical steps.
Workflow Overview (Fig. 1):
The analytical pipeline was structured into three primary steps: data pre-processing, dimensionality reduction, and semi-supervised clustering. This systematic approach.
ensures robust data quality and facilitates the identification of distinct cell populations that might otherwise remain undetected.
-
1.
Data Pre-processing and Quality Control: Initial Quality Filtering: Raw FCS (Flow Cytometry Standard) files underwent initial quality control using the flowAI R package. This tool effectively detects anomalies in flow rate, signal acquisition, and dynamic range, ensuring that only high-quality data proceeds to further analysis by removing anomalous patterns and peaks.
-
2.
Automatic Compensation: To correct for spectral overlap between fluorochromes, fluorescence compensation was performed using the Auto-spill algorithm within FlowJo software22,23,24. The automated process accurately identifies positive populations and generates a compensation matrix, which is then applied to all samples, reducing autofluorescence by using unstained controls.
-
3.
Data Normalization and Concatenation: To standardize event numbers, FCS files were downsampled to 25,000 events per sample. These downsampled events were then concatenated into a single, comprehensive FCS file containing 1 × 106 events, with added keywords to maintain sample origin identification. The concatenated data underwent additional refinement using the PeacoQC algorithm, applying specific gain and mean absolute deviation (MAD) limits, to ensure only high-quality events were used for subsequent analyses.
-
4.
Dimensionality Reduction (t-SNE Implementation): We applied t-Distributed Stochastic Neighbor Embedding (t-SNE) to visualize high-dimensional flow cytometry data, representing the multiple measured parameters/markers for each cell, in a more manageable, two-dimensional space. This technique is crucial for discerning complex relationships and structures within the data that are not readily apparent in traditional biaxial plots. Dimensionality reduction simplifies complex datasets by reducing the number of variables while preserving essential information, thereby making patterns easier to visualize. t-SNE analysis was performed using only fluorescence parameters from the processed FCS file, excluding forward and side scatter (FSC and SSC) data to focus purely on marker expression. Six markers for monocyte subsets and seven for macrophage subsets, including epigenetic expression markers (H3K4me3, H3K9ac, H3K27me3), were incorporated.
-
5.
Optimization: The t-SNE algorithm was subjected to a rigorous optimization process. Its key parameters—iterations, perplexity, and learning rate—were systematically varied, often guided by Fibonacci numbers. This approach enabled the identification of subtle yet perceptible variations in parameter space, allowing a more thorough exploration than conventional predetermined settings. The aim was to achieve ideal configurations for each t-SNE map that optimally preserved data structure and facilitated refined clustering. Perplexity, a critical parameter that measures effective neighborhood size, was specifically tuned to ensure a balanced representation of both local and global data characteristics in the visualization.
-
6.
Semi-Supervised Clustering (X-shift, FlowSOM, and Cluster Explorer): To objectively delineate specific cell populations and evaluate their concordance with manually gated populations, multiple clustering algorithms were employed.
The X-shift algorithm was initially applied, with K values (nearest neighbors) systematically varied to identify phenotypic relationships and estimate the optimal number of clusters in the dataset. These clusters were mapped onto t-SNE plots for visual validation.
Building upon the cluster counts determined by X-shift, the FlowSOM algorithm was used for further refinement. This plugin-based implementation in FlowJo22,23,24 enabled the creation of a self-organizing map (SOM) grid that consolidates initial clusters into a final set of metaclusters based on identified phenotypic patterns. Only fluorescence parameters relevant to monocyte-macrophage relationships were used, maintaining data integrity by keeping values unscaled.
After the FlowSOM analysis, the Cluster Explorer plugin facilitated interactive visualization of population frequencies and marker profiles. Bar charts and heatmaps were generated to illustrate cluster percentages and marker-intensity distributions, respectively, thereby enhancing clarity of phenotypic differentiation across the identified populations.
Statistical analysis
Final figures derived from FlowJo software and R22,23,24 were exported as BMP or PNG files and minimally edited in a raster graphics editor to maintain graphic integrity. Statistical analyses were performed using GraphPad Prism (v.10). P-values were derived by unpaired Student´s t-tests. Differences were considered statistically significant when p < 0.05, and results were expressed as mean and standard deviation (SD) unless otherwise specified.
Comparative Workflow of Conventional vs. Bioinformatic Analysis of High-Dimensional Flow Cytometry Data in Dyslipidemia.
Results
Circulating levels of classical monocytes are increased in dyslipidemia
We analyzed different cell clusters based on specific receptor expression and the presence of two main epigenetic marks in circulating monocytes from young adults living with dyslipidemia compared to healthy controls. Demographic and clinical data are summarized in Table 1. One important issue we sought to address was the metabolic profile of the subjects in this study; thus, we decided to include young people without prior knowledge of their serum lipid levels, who were not under any treatment and had no other comorbidities. Initially, we compared the metabolic phenotype of MoMe based on the expression of previously described surface markers. To eliminate subjectivity in operator-defined gating thresholds, given the complexity of the datasets, we developed a comprehensive analytical workflow that integrates bioinformatics tools with flow cytometry, facilitating thorough phenotypic analysis of circulating monocytes and monocyte-derived macrophages.
Traditional monocyte subsets were defined by CD14 and CD16 expression in high-quality FCS files using conventional gating strategies. This enabled a comparative analysis of traditional monocyte subsets with lipid-processing markers (ABCA1, PLIN2, CD36) to explore potential distribution patterns within these classifications. We classified monocytes into the classical (CD14 + + CD16−), intermediate (CD14 + + CD16+), and nonclassical (CD14 + CD16++) subsets (Fig. 2A).
Characterization of Monocyte Subsets and Lipid Metabolism Markers in Healthy and Individuals that live with dyslipidemia. (A) Representative dot plots illustrating the traditional gating strategy for CD14 and CD16 to identify classical, intermediate, and non-classical monocytes. Labels differentiate individuals that live with dyslipidemia (n = 23) from healthy controls (n = 17). (B-D) Comparison of the distributions of monocyte subsets between individuals that live with dyslipidemia and healthy groups. (E-G) Analysis of lipid-related markers—ABCA1, PLIN2, and CD36—across monocyte subsets, with marker expression overlaid on CD14/CD16 dot plots to visualize lipid-processing activity within these subsets. Bar graphs display mean ± SD. p-values indicating significance at *p < 0.05.
Comparison of monocyte subsets revealed a statistically significant increase in the percentage of classical monocytes in individuals that live with dyslipidemia compared to controls (mean ± SD: 1.77 ± 0.33% in dyslipidemia versus 0.14 ± 0.07% in controls, P < 0.02 (Fig. 2B-D). To further delineate monocyte subsets, MoMe lipid-processing markers ABCA1, PLIN2, and CD36 were included, based on prior research by our group. These markers were assessed within traditional monocyte categories by overlaying PLIN2, CD36, and ABCA1 positivity onto a CD14/CD16 biaxial dot plot (Fig. 2E-G). This analysis revealed that lipid-processing markers were distributed across all traditional monocyte subsets.
Specific metabolically activated monocyte subpopulations were identified by a novel bioinformatic approach
To refine monocyte subpopulation identification, an unsupervised, nonlinear dimensionality reduction algorithm (t-SNE) was used alongside clustering algorithms. This approach aimed to minimize subjective human intervention, such as manual gating and population categorization. Optimal values for perplexity, learning rate, and iteration were established. Iterations and learning rate were based on Fibonacci values (S1 fig).
We developed an optimized clustering scheme focused on CD14-positive events, which serve as a primary marker for monocyte identification. Initially, CD14-positive events were selected from the total dataset to establish the clustering foundation. CD14 positivity was assessed using the Stain Index equation (S2 fig A), and CD14-positive values were subsequently subjected to dimensionality reduction to organize these events within the t-SNE space. This approach effectively excluded CD14-negative events, centering the analysis on the relevant cell populations defined by our antibody panel. The dimensional reduction was refined by selecting specific parameter values from the original 8 × 8 scheme and implementing a new 4 × 2 t-SNE grid, configured with an iteration value of 1597 and perplexity levels of 89 and 144. Learning rates were systematically set at 13, 21, 34, and 55% to facilitate robust visualization (S2 fig B). This configuration enabled focused clustering of CD14-positive events, thereby enhancing the precision of cell subset delineation. Using Cluster Explorer, we generated heatmaps of expression levels across various fluorescence markers and overlaid them on t-SNE maps to visualize marker distribution patterns. Figure S2C presents the t-SNE heatmaps generated with a perplexity of 89, illustrating marker expression versus the optimized learning rates. Similarly, Figure S2D displays analogous heatmaps for t-SNE maps obtained with a perplexity of 144, further exploring antibody staining and fluorescence marker positivity and their impact on t-SNE clustering within CD14-positive events. Comparative analysis of t-SNE maps at different perplexity levels (144 and 89) revealed distinct spatial separations within the t-SNE islands, particularly for markers CD14, ABCA1, and PLIN2. Each of these markers demonstrated clear, well-separated “islands” in the heatmaps, indicating distinct expression clusters.
Notably, while the heatmap for CD14 may show blue coloration, this does not imply negative CD14 expression. The Cluster Explorer tool transforms fluorescence levels into an intensified color gradient to more precisely delineate differences in expression levels. In the case of CD14, blue shades indicate lower, yet positive, expression, with increasing intensity corresponding to higher expression levels. Although these findings lack formal statistical significance, the visual separations and distinct clustering roles of CD14 and ABCA1 within this analysis are noteworthy. The observed expression patterns suggest a unique role for these markers in cellular clustering, warranting further examination in subsequent analyses (S3 fig).
Patterns of CD16, PLIN2, CD36, ABCA-1, and CD14 expression define four different populations of Circulating monocytes
For automatic subpopulation identification, the X-shift and FlowSOM clustering algorithms were sequentially implemented. We used X-shift to identify preliminary cluster numbers before applying FlowSOM for more stable and precise metaclustering.
Arbitrary parameter selection can introduce potential inconsistencies in clustering and lead to computational inefficiencies. Nevertheless, since X-shift was not intended for final cluster refinement in our study, we utilized it to estimate the minimum number of clusters based on our selected learning rate parameters.
The subsampling limit was set to 13% of CD14-positive events rather than the total number of events in the FCS file. For the nearest neighbors (K) value, we retained the default setting to conduct effective, unsupervised cluster identification, while Euclidean distance was selected as the metric to ensure alignment with t-SNE’s similarity basis.
The results of X-shift clustering revealed seven distinct clusters, with one comprising 50.4% of the analyzed events, followed by clusters containing 19.8%, 13.3%, and 7.79% of the events, respectively. Smaller clusters accounted for between 1.49% and 3.72% of the total events. Using the Cluster Explorer tool, we generated graphs of relative expression levels, revealing phenotypic patterns characteristic of each identified cluster (S4 fig). This unsupervised clustering facilitated the initial identification of phenotypic patterns within the clusters, which were visualized on t-SNE maps (perplexity of 144 and learning rates of 13%, 21%, 34%, and 55%). The cluster-specific heatmap table provided further differentiation by illustrating tonal variations according to maximum fluorescence intensity scales, with relative expression levels indicating phenotypic distinctions among clusters. For instance, clusters 5 and 7 shared notable similarities in their expression patterns. This pattern similarity prompted further analysis to determine whether these phenotypic distinctions were statistically significant and whether merging similar clusters would enhance data clarity. After applying X-shift clustering, FlowSOM was used to validate cluster stability.
We obtained four main FlowSOM metaclusters, which collectively comprise 98% of CD14 + events, specifically: 62.7% for Metacluster 1, 24% for Metacluster 5, 6.89% for Metacluster 4, and 4.71% for Metacluster 2, while remaining clusters each represented less than 1% of total events (Fig. 3A-B). Figure 3C presents a transformed relative expression level graph illustrating the phenotypic and epigenetic patterns of the four main metaclusters.
We assessed expression levels across the four metaclusters using FlowJo’s Spectral Population Viewer22,23,24 to elucidate phenotypic distinctions. Comparisons were made between these levels and traditional monocyte subpopulations. In our two primary metaclusters (1 and 5), we observed comparable expression levels of CD16, PLIN2, and CD36. However, significant differences were observed in CD14 and ABCA1 expression (Fig. 3D).
The metaclusters were defined as follows (Fig. 3E): Metacluster 1: CD14High (100%), CD16Pos (99.4%), ABCA1Pos (71.9%), PLIN2Pos (91.5%), and CD36Pos (71.9%); Metacluster 2: CD14High (100%), CD16Neg (80.2%), ABCA1Pos (99.8%), PLIN2Var (61.4%), and CD36Var (61.1%); Metacluster 4: CD14Low (97.2%), CD16Neg (94.1%), ABCA1Neg (72.3%), PLIN2Pos (66.6%), and CD36Neg (89.1%); Metacluster 5: CD14Low (95.8%), CD16Pos (99.7%), ABCA1Neg (83.7%), PLIN2Pos (90.9%), and CD36Var (57.8%).
With the identification of the four principal metaclusters, we conducted a comparative analysis with traditional monocyte populations identified through manual gating. Figure 3F illustrates this comparison using biaxial dot plots (CD14 on the x-axis and CD16 on the y-axis), showing overlays of our metaclusters and conventionally gated monocyte populations in t-SNE space.
Traditional monocyte subsets were not redefined phenotypically, retaining their classical, intermediate, and non-classical monocyte classifications through CD14 and CD16 gating. However, lipid-processing markers showed variability between the 25th and 75th percentiles, indicating a broader range within traditional subsets than the more consistent metacluster percentiles (Fig. 3G- J). Metacluster 1 (CD14High/CD16Pos/ABCA1Pos/PLIN2Pos/CD36Var) may correspond to the intermediate monocytes, Metacluster 4 (CD14Low/CD16Neg/ABCA1Neg/PLIN2Pos/CD36Neg) corresponds to classical monocytes; Metacluster 5 (CD14Low/CD16Pos/ABCA1Neg/PLIN2Pos/CD36Var) could represent non-classical monocytes, while Metacluster 2 corresponds to MoMe (CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var).
Refined Population Categorization Using FlowSOM Clustering and Metacluster Analysis. FlowSOM clustering, applied post-X-shift, utilized a 15 × 15 Self-Organizing Map (SOM) with seven metaclusters for population categorization. (A-B) t-SNE maps with FlowSOM clustering (perplexity: 144; learning rates: 34% and 55%) show improved metacluster stability and reduced overlap. The major metaclusters, representing 98% of CD14 + events, included Metacluster 2 (54.2%), Metacluster 7 (34.15%), Metacluster 4 (7.05%), and Metacluster 1 (2.95%). (C) The FlowSOM heatmap confirms phenotypic distinctions among metaclusters. (D) Relative expression level transformations reveal phenotypic patterns, highlighting distinctions between metaclusters and traditional X-shift clusters. (E) The Minimal Spanning Tree (MST) illustrates metacluster linkages and depicts phenotypic similarity. (F-G) Comparisons with conventionally gated monocyte subsets reveal both alignment and distinctions. (H-I) FlowJo’s Spectral Population Viewer22,23,24 further characterizes expression levels within metaclusters compared with traditional subsets, showing median intensity lines with percentile boundaries. (J) Metacluster phenotypes were characterized as “Pos,” “Neg,” or “Var” based on marker expression percentages, with “High” descriptors indicating expression intensity.
Dyslipidemia individuals show higher levels of CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var and CD14Low/CD16Neg/ABCA1Neg/PLIN2Pos/CD36Neg Circulating monocytes compared to control group
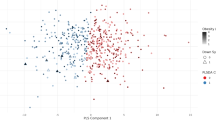
Following the identification and characterization of monocyte metaclusters, we analyzed their distributions across dyslipidemia and control samples to identify group-specific patterns. Using preprocessed keywords dot plots, dyslipidemia and control samples were selected and visualized through t-SNE density maps (Fig. 4A). Notably, dyslipidemia samples displayed elevated event densities within metaclusters 2 and 4, defined as CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var and CD14Low/CD16Neg/ABCA1Neg/PLIN2Pos/CD36Neg, respectively, characterized by distinct marker expression profiles. These findings suggest a potential relevance of these clusters in the context of dyslipidemia, particularly in pathways related to lipid processing and monocyte activation.
To further validate these findings, we generated bar graphs comparing the mean CD14 + monocyte counts and metacluster percentages between dyslipidemia and control samples. Our analysis revealed statistically significant differences, particularly within the Metacluster 2 (CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var), where dyslipidemia samples showed higher proportions (3.05 ± 0.32% vs. 0.70 ± 0.24%, P = 0.0149). Similarly, Metacluster 4 (CD14Low/CD16Neg/ABCA1Neg/PLIN2Pos/CD36Neg) was more prevalent in dyslipidemia individuals than in controls (4.57 ± 0.29% vs. 0.77 ± 0.18%, P = 0.0042) (Fig. 4B-C). Furthermore, we calculated the relative proportions of each metacluster within the total CD14 + monocyte population (Fig. 4D). In dyslipidemia samples, each metacluster showed a significant increase in percentage compared with controls, suggesting a higher overall prevalence of metabolically activated monocytes in subjects that live with dyslipidemia. These results indicate that this condition may drive specific alterations in monocyte subpopulations, contributing to the pathophysiology of metabolic syndrome and atherosclerosis by increasing the prevalence of non-classic pro-inflammatory and lipid-associated macrophage phenotypes.
Comparison of Metacluster Distribution Between Dyslipidemia and Healthy Controls. (A) The t-SNE-generated dot plots, derived from the sample selection plots, illustrate the density variations between dyslipidemia and healthy samples. Individuals that live with dyslipidemia exhibit a higher density of events in the southern region of the t-SNE plot compared to healthy subjects. (B) The application of clustering algorithms to t-SNE maps for dyslipidemia and healthy individuals reveals distinct clustering of monocyte subsets and the phenotypes associated with each identified metacluster. (C) Bar graphs display the percentage of each identified metacluster (1, 2, 4, and 5) within the total monocyte population for healthy and subjects living with dyslipidemia. D) Bar graphs illustrate the percentages of events for each metacluster within the CD14 + monocyte population, showing statistically significant differences across all metaclusters when comparing the percentages between dyslipidemia and healthy samples. Data are presented as mean ± SD. p-values indicating significance at *p < 0.05, **p < 0.01.
Metabolically activated monocytes, MoMe (CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var), from dyslipidemia individuals display higher levels of H3K4me3
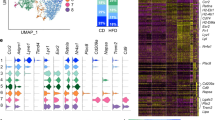
To explore potential epigenetic differences between dyslipidemia and healthy samples, we incorporated H3K4me3 and H3K27me3 markers into our analysis. Figure 5A displays t-SNE maps showing the heatmap of expression levels for these epigenetic markers across the two groups. Each metacluster was analyzed with descriptive statistics (Geometric Mean and Robust SD) for fluorescence intensity, reflecting the distribution and relative abundance of H3K4me3 and H3K27me3. This analysis accounted for the fluorescence intensity and the number of cells represented within each metacluster, providing a detailed perspective on epigenetic marker variability between dyslipidemia and control samples.
These findings disclose statistically significant differences in the expression of H3K4me3 and H3K27me3 across metaclusters (Fig. 5B-E), with notable variability between the two groups. Specifically, within Metaclusters 2 and 4, dyslipidemia samples exhibited significantly higher levels of H3K4me3 than controls, whereas in other metaclusters, both H3K4me3 and H3K27me3 levels were generally elevated in control samples relative to samples from individuals that live with dyslipidemia. This pattern indicates that the physiological condition of individuals that live with dyslipidemia may distinctly modify the epigenetic landscape of specific monocyte subsets, potentially contributing to the inflammatory milieu associated with dyslipidemia and the development of atherosclerotic lesions.
Epigenetic Marker Analysis in Dyslipidemia. (A) t-SNE maps illustrate the heatmap of epigenetic marker expression levels (H3K4me3 and H3K27me3) across samples from individuals that live with dyslipidemia and healthy controls, with fluorescence intensity statistics provided for each marker. (B-E) Statistical comparisons of H3K4me3 and H3K27me3 across metaclusters reveal significant differences between the healthy group and the group of individuals that live with dyslipidemia. Data are presented as mean ± SD. p-values indicating significance at *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Monocyte-derived metabolically activated macrophages show distinctive epigenetic profiles in dyslipidemia compared to healthy subjects
To further analyze the polarization potential of circulating monocytes, the phenotype of monocyte-derived macrophages exposed in a metabolic syndrome-like environment was analyzed. Cultured macrophages were stained for flow cytometry, and t-SNE maps generated across all harvested cells (Fig. 6A) depict the heatmaps of marker expression profiles. The t-SNE visualization indicates a semi-homogeneous distribution of cells, with a prominent group forming a peninsula-like region enriched for CD11b, CD80, and CD14, as well as lipid-processing markers CD36, ABCA1, and PLIN2. Cells expressing CD14 and CD11b were further subjected to secondary dimensional reduction, allowing us to explore phenotypic divergence within these populations. Using x-shift followed by FlowSOM, we identified four phenotypically distinct metaclusters: Metacluster 1: CD80Neg (81.6%), CD11bNeg (71.1%), CD14Pos (96.7%), PLIN2Neg (90.5%), ABCA1Neg (78.2%), CD36Var (59.7%). Metacluster 2: CD80Pos (100%), CD11bNeg (75.8%), CD14Pos (93.1%), PLIN2Pos (98.8%), ABCA1Var (61.2%), CD36Pos (71.2%). Metacluster 3: CD80Neg (93.6%), CD11bPos (99.3%), CD14Neg (95.3%), PLIN2Neg (97.7%), ABCA1Neg (98.6%), CD36Neg (71.3%). Metacluster 4: CD80Pos (100%), CD11bPos (100%), CD14Neg (97.8%), PLIN2Pos (86.4%), ABCA1Neg (98.8%), CD36Pos (99.8%). Metacluster 3, representing CD11bPos macrophages, comprised the majority (60.3%) of observed events, followed by Metacluster 1 (37.8%), which displayed high CD14 expression and variable CD36, while Metaclusters 2 and 4 were less prevalent (0.71% and 1.34%, respectively). Notably, Metacluster 4 exhibited characteristics of the pro-inflammatory M1 macrophage phenotype (CD80Pos, CD11bPos) and metabolic markers PLIN2, CD36 (Fig. 6B). Heatmaps in Fig. 6C illustrate the distribution of CD11bPos and/or CD14Pos cells across the t-SNE landscape, showing distinct localization of both markers. Another notable finding is the extensive expression and spatial distribution of the lipid-processing marker ABCA1, a cell-membrane protein with broad distribution, albeit highly expressed in macrophages. Regarding histone modifications, we observed a higher prevalence of H3K9ac and H3K4me3 in control cells than in dyslipidemia cells. H3K9ac exhibited statistically significant differences in Metaclusters 1, 2, and 4 with lower levels in dyslipidemia samples compared to controls; however, no significant difference was observed in Metacluster 3. For H3K4me3, expression levels were significantly higher in controls across Metaclusters 1, 2, and 3, while no statistically significant difference was found in Metacluster 4 (Fig. 6D). Although descriptive, these findings underscore the importance of epigenetic modifications in dyslipidemia. Future studies should include larger cell populations and an expanded cytometry panel, including additional markers such as CD206, to accurately delineate macrophage subpopulations.
Phenotypic and Epigenetic Profiles of Monocyte-Derived Macrophages. (A) t-SNE maps of total events illustrate the expression of all markers. (B) Selection process for CD14-positive and/or CD11b-positive cells (7.76% of total events) followed by a secondary t-SNE analysis, revealing metacluster populations, along with a heatmap and a transformed expression graph for fluorochrome levels across metaclusters. (C) t-SNE maps of CD11bPos and/or CD14Pos macrophages. (D) Unpaired t-test results for epigenetic comparison across metaclusters indicate significant differences between individuals that live with dyslipidemia and healthy samples. Data are reported as Geometric Mean ± Robust SD. p-values indicating significance at *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Discussion
High-dimensional cytometry in single-cell analysis requires robust clustering methods to accurately capture the complexity of diverse cellular populations, particularly within immune cells26.
Traditional gating methods, while valuable for lower-dimensional analyses, are limited when applied to high-dimensional datasets, as they struggle to simultaneously visualize multiple markers and capture subtle variations within phenotypically similar cell populations19,20,21,22,23,24,25,26,27,28. This study demonstrates that integrating unsupervised and semi-supervised clustering with dimensionality reduction techniques is critical for revealing intricate patterns within monocyte subpopulations that would otherwise remain undetected. Such an approach not only enhances our understanding of cellular heterogeneity but also provides insights into the specific functional roles these subtypes may play in the pathology of dyslipidemia.
To our knowledge, this study represents the first report of circulating MoMe exhibiting a distinctive phenotype that distinguishes them from the three previously reported monocyte populations (classical, intermediate, and non-classical). All four monocyte subpopulations identified displayed variable expression of lipid-processing markers, such as ABCA1 and PLIN2, implicated in the regulation of monocyte inflammatory responses. However, our bioinformatic analysis revealed four distinct monocyte subpopulations, not only by marker positivity but also by receptor expression intensity. Metacluster 1 (CD14High/CD16Pos/ABCA1Pos/PLIN2Pos/CD36Var) may correspond to the intermediate monocytes; Metacluster 4 (CD14Low/CD16Neg/ABCA1Neg/PLIN2Pos/CD36Neg) corresponds to classical monocytes; Metacluster 5 (CD14Low/CD16Pos/ABCA1Neg/PLIN2Pos/CD36Var) could represent non-classical monocytes, while Metacluster 2 corresponds to MoMe (CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var). Traditional gating would likely overlook these subpopulations, underscoring the value of advanced clustering for enhancing biological understanding in inflammatory diseases.
Moreover, our exploration of epigenetic markers, specifically H3K4me3 and H3K27me3, in healthy subjects and individuals living with dyslipidemia offers preliminary insights into the potential role of histone modifications in monocyte heterogeneity. These modifications are linked to trained immunity. Immunological training is linked to epigenetic reprogramming of cells, including histone acetylation and methylation at gene promoters. This innate immune memory can be induced by infectious or non-infectious agents, such as endogenous molecules, including oxidized LDL-cor environmental pollutants. In this regard, we found that MoMe from subjects that live with dyslipidemia displayed elevated levels of H3K4me3, a mark associated with increased expression of typically inflammatory genes, whereas in control subjects, as expected, this mark was predominantly observed in the expected pro-inflammatory populations (classical and intermediate monocytes). This specific epigenetic modification in MoMe from individuals that live with dyslipidemia aligns with the hypothesis that exposure to elevated levels of serum lipids (ox-LDL-c or others) can induce epigenetic modifications and immune training in specific macrophage populations29,30,31,32.
Our findings concerning monocyte-derived MMe are significant, albeit challenging to fully explain. Nearly all differentiated macrophage populations from control subjects exhibited increased H3K4me3 and H3K9ac, suggesting higher transcriptional activity of several potentially pro-inflammatory genes. This observation was contrary to our initial hypothesis, which predicted that trained monocytes from individuals that live with dyslipidemia would maintain their epigenetic marks when differentiated into macrophages. We cannot discard the possibility that in vitro differentiation under highly controlled metabolic stimuli may account for our findings. However, it would be relevant to elucidate whether the observed decrease in H3K9ac in MMe from individuals that live with dyslipidemia could translate into a functional defect beyond cytokine production, such as impaired phagocytic capacity, or, conversely, represent a compensatory response.
Most differentiated macrophage subpopulations from controls showed high levels of H3K4m3 and H3K9ac. In this regard, it has been reported that not only M1 but also M2 macrophages rely on histone methylation and acetylation processes to execute their reparative functions33.
It is important to note that our study aimed to evaluate the effect of dyslipidemia on monocyte immune training in the absence of other metabolic abnormalities. Therefore, we included young people who had not received a previous diagnosis of dyslipidemia and were not under any statin or lipid-lowering drugs. However, we cannot dismiss the fact that our control group had a family history of hypercholesterolemia, and the subjects included in the dyslipidemia group exhibited mild disease. Another limitation in our work was the small sample size. It is important to consider that these findings may represent only a stage in the process, as part of a continuum illustrating the natural progression of atherogenesis. It is necessary to evaluate cytokine and functional activity of the described populations to fully comprehend the physiological implications of our findings.
In summary, this study demonstrates that combining and optimizing clustering algorithms offers a robust tool for unraveling cellular heterogeneity within complex immune cell populations. This approach facilitates the discovery of cell subtypes, specifically the CD14High/CD16Neg/ABCA1Pos/PLIN2Var/CD36Var MoMe and underscores the immune training of circulating monocytes fromindividuals that live with dyslipidemia.
By enhancing our ability to identify and characterize disease-relevant cell subsets, advanced clustering methodologies hold promise for advancing diagnostics, prognostics, and therapeutics in dyslipidemia and other chronic inflammatory diseases. This work lays a foundation for ongoing refinement in single-cell analytics, with the goal of leveraging cellular diversity for clinical insights and therapeutic innovations.
An important strength of this study is the bioinformatic approach applied. The implementation of Fibonacci-derived parameters enhanced cluster specificity by focusing on key markers, such as CD14, which plays a significant role in monocyte classification. This approach reduced noise and facilitated the resolution of unique phenotypic patterns in markers relevant to dyslipidemia, such as ABCA1 and PLIN2.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
References
Wolf, D. & Ley, K. Immunity and inflammation in atherosclerosis. Circ. Res. 124, 315–327. https://doi.org/10.1161/CIRCRESAHA.118.313591 (2019).
Fernandez, D. M. & Giannarelli, C. Immune cell profiling in atherosclerosis: role in research and precision medicine. Nat. Rev. Cardiol. 19, 43–58. https://doi.org/10.1038/s41569-021-00589-2 (2022).
Honold, L. & Nahrendorf, M. Resident and monocyte-derived macrophages in cardiovascular disease. Circ. Res. 122, 113–127. https://doi.org/10.1161/CIRCRESAHA.117.311071 (2018).
Bobryshev, Y. V. et al. Macrophages and their role in atherosclerosis: pathophysiology and transcriptome analysis. BioMed Res. Int. 9582430 (2016). (2016). https://doi.org/10.1155/2016/9582430
Hilgendorf, I., Swirski, F. K. & Robbins, C. S. Monocyte fate in atherosclerosis. Arterioscler. Thromb. Vasc Biol. 35, 272–279. https://doi.org/10.1161/ATVBAHA.114.303565 (2015).
Blagov, A. V. et al. The role of macrophages in the pathogenesis of atherosclerosis. Cells 12, 522. https://doi.org/10.3390/cells12040522 (2023).
Liu, X. et al. Targeting foam cell formation and macrophage polarization in atherosclerosis: the therapeutic potential of rhubarb. Biomed. Pharmacother. 129, 110433. https://doi.org/10.1016/j.biopha.2020.110433 (2020).
Amengual, J. & Barrett, T. J. Monocytes and macrophages in atherogenesis. Curr. Opin. Lipidol. 30, 401–408. https://doi.org/10.1097/MOL.0000000000000634 (2019).
Kratz, M. et al. Metabolic dysfunction drives a mechanistically distinct Proinflammatory phenotype in adipose tissue macrophages. Cell. Metab. 20, 614–625. https://doi.org/10.1016/j.cmet.2014.08.010 (2014).
Chen, L. et al. Molecular mechanisms for ABCA1-mediated cholesterol efflux. Cell. Cycle. 21, 1121–1139. https://doi.org/10.1080/15384101.2022.2042777 (2022).
Jacobo-Albavera, L. et al. The role of the ATP-binding cassette A1 (ABCA1) in human disease. Int. J. Mol. Sci. 22, 1593. https://doi.org/10.3390/ijms22041593 (2021).
Loix, M. et al. Perilipin-2 limits remyelination by preventing lipid droplet degradation. Cell. Mol. Life Sci. 79, 515. https://doi.org/10.1007/s00018-022-04547-0 (2022).
Grimaldi, V. et al. Epigenetic reprogramming in atherosclerosis. Curr. Atheroscler Rep. 17, 476. https://doi.org/10.1007/s11883-014-0476-3 (2015).
Bekkering, S. et al. Innate immune cell activation and epigenetic remodeling in symptomatic and asymptomatic atherosclerosis in humans in vivo. Atherosclerosis 254, 228–236. https://doi.org/10.1016/j.atherosclerosis.2016.10.019 (2016).
Martínez-Shio, E. B. et al. Differentiation of Circulating monocytes into macrophages with metabolically activated phenotype regulates inflammation in dyslipidemia patients. Clin. Exp. Immunol. 208, 83–94. https://doi.org/10.1093/cei/uxac013 (2022).
Ujas, T. A., Obregon-Perko, V. & Stowe, A. M. A guide on analyzing flow cytometry data using clustering methods and nonlinear dimensionality reduction (tSNE or UMAP). Methods Mol. Biol. 2616, 231–249 (2023).
Siddiqui, S. & Livák, F. Principles of advanced flow cytometry: A practical guide. Methods Mol. Biol. 2580, 89–114. https://doi.org/10.1007/978-1-0716-2740-2-5 (2023).
Brestoff, J. R. & Frater, J. L. Contemporary challenges in clinical flow cytometry: small samples, big data, little time. J. Appl. Lab. Med. 7, 931–944. https://doi.org/10.1093/jalm/jfab176 (2022).
Ask, E. H. et al. MetaGate: interactive analysis of high-dimensional cytometry data with metadata integration. Patterns 5, 100989. https://doi.org/10.1016/j.patter.2024.100989 (2024).
Roca, C. P. et al. AutoSpill is a principled framework that simplifies the analysis of multichromatic flow cytometry data. Nat. Commun. 12, 2890. https://doi.org/10.1038/s41467-021-23126-8 (2021).
Pavia-López, Abel, A. et al. Mexican clinical practice guideline for the diagnosis and treatment of dyslipidemias and atherosclerotic cardiovascular disease. Arch. Cardiol. Méx. 92, 1–62. https://doi.org/10.24875/acm.m22000081 (2022).
BD Life Sciences. FlowJo™ Software, version 10.10. (2023). https://www.flowjo.com/
R Core Team. R: A language and environment for statistical computing, version 4.1.1. R Foundation for Statistical Computing, Vienna, Austria. (2021). https://www.R-project.org/
Bioconductor Bioconductor: Open software for bioinformatics, release 3.20. (2023). https://bioconductor.org/
Van der Maaten, L. J. P. & Hinton, G. E. Visualizing high-dimensional data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Liu, J. & Vinck, M. Improved visualization of high-dimensional data using the distance-of-distance transformation. PLoS Comput. Biol. 18, e1010764. https://doi.org/10.1371/journal.pcbi.1010764 (2022).
Robinson, J. P. et al. Flow cytometry: the next revolution. Cells 12, 1875. https://doi.org/10.3390/cells12141875 (2023).
McKinnon, K. M. Flow cytometry: an overview. Curr. Protoc. Immunol. 120, 5.1.1–5.1.11 (2018). https://doi.org/10.1002/cpim.40
Riksen, N. P. & Netea, M. G. Immunometabolic control of trained immunity. Mol. Aspects Med. 77, 100897. https://doi.org/10.1016/j.mam.2020.100897 (2021).
Flores-Gomez, D., Bekkering, S., Netea, M. G. & Riksen, N. P. Trained immunity in atherosclerotic cardiovascular disease. Arterioscler. Thromb. Vasc Biol. 41, 62–69. https://doi.org/10.1161/ATVBAHA.120.314216 (2021).
Kuznetsova, T., Prange, K. H. M., Glass, C. K. & de Winther, M. P. J. Transcriptional and epigenetic regulation of macrophages in atherosclerosis. Nat. Rev. Cardiol. 17, 216–228. https://doi.org/10.1038/s41569-019-0265-3 (2020).
Shi, Y. et al. Epigenetic regulation in cardiovascular disease: mechanisms and advances in clinical trials. Signal. Transduct. Target. Ther. 7, 200. https://doi.org/10.1038/s41392-022-01055-2 (2022).
Yan, L. et al. Macrophage plasticity: signaling pathways, tissue repair, and regeneration. MedComm 8, e658. https://doi.org/10.1002/mco2.658 (2020).
Acknowledgements
Jorge Flores Ramírez from Clinical Specialties Laboratory for performing lipid measurements and Centro de Salud Universitario, Universidad Autónoma de San Luis Potosí, for its invaluable support in recruiting patients.
Funding
Funding information: This work was supported by the grant “FONDO SECTORIAL DE INVESTIGACIÓN PARA LA EDUCACIÓN” by Consejo Nacional de Humanidades, Ciencia y Tecnología, CONAHCYT, CB2017-2018 No. A1-S-43425, México. https://conahcyt.mx/ (to AM-U). And, CONSEJO POTOSINO DE CIENCIA Y TECNOLOGÍA, “COPOCYT”, Fid. 23871-Multas Electorales de Proyectos para fortalecer la formación de recursos humanos en investigación, desarrollo tecnológico e innovación (to AM-U).
Author information
Authors and Affiliations
Contributions
All authors discussed the results and contributed to the final manuscript. R-T.R. processed the experimental data, performed the analysis, made all the computational work, drafted the manuscript and designed the figures. R-S.S contributed to sample preparation, experiments and contributed to the interpretation of the results.G-H.M.J contributed to sample preparation, experiments and contributed to the interpretation of the results.M-S.E.B contributed to sample preparation, experiments and contributed to the interpretation of the results.E-U.C.D Contributed to conceive the study, was in charge of evaluation of all clinical aspects, recruitment of patients and analysis of the results. M-U.A.E. Conceived and supervised the project. Obtained the financial support. Evaluated the analyzed results. Elaborated the final version of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ramírez-Torres, R., Ramírez-Segovia, S.G., González-Huerta, M.J. et al. Bioinformatic characterization and automated detection of metabolically activated monocyte subpopulations in dyslipidemia. Sci Rep 16, 6170 (2026). https://doi.org/10.1038/s41598-026-36678-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-36678-w