Abstract
It has become pertinent to develop early and accurate diagnosis tools for these neurological diseases, such as Alzheimer’s and Parkinson’s. The diagnosis may be high frequency electroencephalogram (EEG) signal based. These techniques promise good results but fail to obtain the desired clinically relevant features because of the intrinsically non-stationary and noisy nature of high frequency EEG components. Limitations of existing methods include suboptimal signal processing, ineffective strategies for feature selection, lack of robustness in feature fusion mechanisms, and limited explainability for clinical adoptions. This work, therefore, proposes a holistic framework in the context of clinical detection of neurological disorders using high frequency EEG signals which are enhanced as a pipeline of multi-blocks. The combination of Hilbert-Huang transform (HHT) with a modified empirical mode decomposition ensures that the decomposition is adaptive in nature and effective noise reduction leads to preprocessing of the data. Wavelet Packets transform (WPT) in conjunction with shannon entropy-based feature selection reduces the dimensions of the data without information loss, which aids in meaningful extraction of temporal and frequency domain features. Canonical correlation analysis with multi-view representation learning allows integration of EEG features along with clinical metadata as auxiliary information to create a common feature space for increased sensitivity in diagnosis. A new multi-scale convolutional recurrent neural network (MS-CRNN) uses an attention mechanism to process the combined features and find spatiotemporal dependencies while focusing on patterns that are important for diagnosis. The method is demonstrated through grad-cam and integrated gradient techniques that help in visualizing and quantitatively attributing feature extraction. This method was 94% accurate; 92% sensitive; and 93% specific when identifying issues early on. The high accuracy in making clinical interpretation and diagnosis has set a new bar for clinicians and has encouraged public policy to support early intervention.
Similar content being viewed by others
Introduction
More than sixty million individuals around the world are affected by neurologic disorders (Parkinson’s and Alzheimer’s), which are two of the leading contributors to death and disability1,2. Therefore, a timely and precise diagnosis is critical to both providing the patient with immediate treatment options and better managing the disease. Non-stationary and noisy EEG signals provide a good opportunity for the use of EEG as biomarkers due to their association with neural pathology. However, the non-stationarity and noisiness of EEG signals make it difficult to analyze these signals. The traditional methods of signal processing (Fourier and wavelet transforms) rely heavily upon stationarity and therefore they are unable to accurately capture the important localized and high frequency EEG components that are necessary for diagnosing neurological diseases3,4,5.
Feature extraction techniques commonly lead to redundancy in large, high dimensional feature spaces leading to poor model performance, and few, if any, of the available EEG based diagnosis tools are able to provide an explanation for their decision; therefore, limiting clinical acceptance and confidence, and none of them have developed efficient methods for fusing additional clinical and demographic data with EEG6,7,8. This research proposes a new multi-block approach for extracting neurological disorder features from high frequency EEG signals by employing advanced signal processing and machine learning techniques to improve both clinical relevance and robustness. The key issues related to the large gap existing between prototypes developed by researchers and clinical practice (e.g., EEG nonstationary, redundant features, poor fusion of features, lack of explanation) that the framework aims to address are early and accurate diagnosis, efficient signal decomposition and denoising, robust and compact feature extraction, effective fusion of features from different sources, advanced learning of spatial and temporal patterns, improved interpretability and scalability for clinical use.
Although EEG signals are inherently non-stationary and contaminated with noise, signal processing methods that can decompose EEG signals precisely and reduce noise effectively, while preserving critical features for diagnosis, include the Hilbert Huang transform (HHT) and modified empirical mode decomposition (EMD). The wavelet packet transform (WPT) is utilized to address problems related to low signal to noise ratios and muscle artifacts, especially in the gamma frequency range (30–100 Hz) where most high frequency EEG data fall short due to low signal to noise ratios; WPT has been shown to be capable of greater denoising and feature extraction than other traditional methods. To provide a comprehensive view, features are fused with clinical metadata through canonical correlation analysis (CCA) and feature selection based on Shannon entropy12,13. The MS-CRNN model using the features obtained through both Gradient CAM and integrated gradients is trained to produce spatiotemporal EEG pattern representations. The MS-CRNN will provide clinically useful, reliable, and understandable results for the diagnosis of neurological disorders and improve the clinical applicability and reliability of EEG-based neurological disorder diagnostics.
The proposed adaptive EMD method reduces fixed-threshold values of sifting parameter epsilon (ε) via cross validation in frequency bands as a function of the entropy in the spectral domain and thus reduces time-frequency energy residual while preserving higher frequency signal activity. The experimental results also demonstrate that setting ε = 0.001 will provide reliable high fidelity representation in the gamma range (80–120 Hz), accurately distinguish between gamma band activity while minimizing erroneous decomposition due to mode mixing or creation of incorrect intrinsic mode functions.
Adaptive EMD removes artifact noise through two stages of filtering; first stage rejects those IMF’s that are below 0.75 for energy entropy or less than 5% in terms of their contribution to total energy as measured in each IMF. The second stage combines this with Hilbert Spectral Analysis (HSA) to remove muscle, eye and other forms of artifact noise generated from non-neural systems, therefore reducing all possible noise from any source(s), prior to final preprocessing.
This proposed framework has been designed to resolve common limitations of detecting early-stage Alzheimer’s disease and Parkinson’s disease through utilizing state-of-the-art signal processing techniques and deep-learning-based architectures for the evaluation of EEG signals at high-frequencies14,15,16,17,18, in conjunction with a methodical approach to integrate features extracted from EEG signals with clinical data to improve the accuracy of diagnosis and the interpretability of results19,20,21,22,23,24,25. The framework is an integrated, scalable solution to bridge the gap between the academic environment and the clinical environment, and will address the growing demand for early and reliable detection of neurodegenerative disorders using EEG technology.
The primary attributes of the proposed method include application of Hilbert-Huang Transform (HHT), in combination with an updated Empirical Mode Decomposition (EMD) criterion for an accurate and efficient removal of noise from signals; identification of relevant features using Shannon Entropy; and developing an interpretable, scalable, and efficient method to facilitate early detection of Alzheimer’s disease and Parkinson’s disease utilizing advanced signal processing and interpretable deep learning for improved neurological diagnostics.
The Shannon Entropy is used to select features in EEG analysis as it is a measure of both the complexity and the uncertainty of a signal, which can be an advantage when dealing with non-stationary high frequency EEG signals. The Shannon Entropy allows for identification of the most informative sub-bands in the signal and also for the removal of the most noisy components. The use of Shannon Entropy results in a reduction in feature dimensions of nearly 60%, but retention of around 95% of the clinically useful information contained within those features.
By combining Empirical Mode Decomposition (EMD), which is an adaptive method of decomposing signals into multiple components or modes with the ability to adaptively and locally time-frequency resolution, with the Hilbert transform for obtaining the instantaneous frequencies associated with each mode, HHT has proven effective in dealing with the non-stationarity and high levels of noise present in EEG signals. The preprocessing that HHT provides, therefore, can be expected to reduce noise from the raw EEG signal and preserve clinical significance, thereby improving both the relevance and the accuracy of the EEG-based diagnosis. In addition, reducing the dimensionality of the EEG feature space using the method of feature selection based on Shannon entropy will result in removing those EEG features that are either not as useful or contain less information than others, therefore preserving the clinically important features and reducing the amount of noise. As a consequence of this approach, there will be fewer EEG features used to train models; thus, the models will have better generalizability, they will be able to run faster, and their diagnoses will be more precise.
Canonical Correlation Analysis (CCA) is an analysis of EEG characteristics and clinical metadata that correlates them as well as possible in a single “space” to preserve relevant and complementary information while eliminating redundant or irrelevant information; it can enhance the identification of subtle patterns and therefore improve sensitivity and clarity of diagnosis — critical for early detection of neurological disorders. MS-CRNN also does this by using recurrent convolutional neural network layers with convolutional layers of varying size to capture both spatial and temporal features of the EEG signal.
The use of this allows the detection of both minor and wide-spread neural activity patterns across all different frequency bands and all different regions of the brain; and improves the examination of complex EEG signals. The recurrent layer using bi-directional gated recurrent units (GRU) captures the time-dependent relations within EEG signals; and captures important changes over time which is necessary for diagnosing neurologic disease. The application of an attention mechanism can enhance the accuracy of the model by focusing on those portions of time that are most relevant for making a diagnosis; thereby improving both the interpretability of the results, and the accuracy of the results. MS-CRNN is capable of managing large amounts of high-dimensional EEG data by combining convolutional and recurrent layers with an attention mechanism to detect both spatial and temporal features. As such, it has shown improved classification capabilities when compared to other models; and therefore improved sensitivity and specificity when classifying individuals with neurologic diseases such as Alzheimer’s disease and Parkinson’s disease.
Related work
EEG has made many advances with respect to a number of studies using various techniques and applications of EEG, including the detection of neurologic disorders, assessment of cognition and emotion. These studies also represent an important body of literature that represents a timeline of how EEG based systems have evolved, both technologically and methodically. Studies like Shi et al.1 further demonstrate that there are new, more sophisticated methods for data preprocessing and an advanced approach of spatial frequency filtering of multi-channel EEGs for the elimination of the type of noise produced by artifacts generated by eye or muscle movement.
Another similar type of study uses deep learning, applied to the prediction of neurological outcomes for comatose patients; in Zheng et al.2, temporal dynamics of EEGs have shown a good application for better prognostics post-cardiac arrest. Klymenko et al.3 discuss byte-pair encoding for clinical EEG classification and present novel solutions for encoding multivariate timestamp series data in support of age-based neural analysis. Contributions by Yu et al.4 extend further into artifact detection and classification with explainable wavelet neural networks that emphasize both interpretability and performance levels. Lee et al.5 explore wearable and immersive technologies: they combined virtual reality (VR) with wearable EEG devices to screen for dementia. An example of integrated systems plays an important role in advancing the accuracy of diagnostics. In parallel, Ansari et al.6 apply shared multi-scale inception networks to identify the stages of neonatal sleep with fewer EEG channels than usual, demonstrating that a feature-efficient deep learning architecture is vital in studies among children. Chu et al.7 refine microstate recognition for Parkinson’s disease using enhanced deep learning frameworks, focusing on activated brain regions. Siddiqa et al.9 applied multi-branch CNNs in neonatal sleep staging, discussing entropy-based techniques and showing that single-channel EEG can reach an effectiveness equal to that of a multi-channel setup. Gaussian processes, as in the work of Caro et al.10, enable precise imputation of data and seizure detection, using spectral mixture kernels; this represents the direction in which probabilistic models could be adopted for EEG interpretation sets.
Iteratively, next, based on Table 1 Methodological Comparative Analysis, Apart from clinical diagnostics, affective computing and emotion analysis have seen a boom. Castiblanco Jimenez et al. present EEG indicators that are increased using machine learning in VR, in order to determine the state of an individual’s emotions, relating neuroscience with virtual reality technology. They also provide Singh et al.29, who develop a comprehensive evaluation of the use of physiological signals for identifying mental illnesses, which include an assessment of machine learning methods used in such assessments. As such these studies demonstrate the interdisciplinary nature of the use of EEG indicators as they range from the psychological assessment of humans to the interaction between humans and computers being developed.
Important to enhancing the interpretation of the EEG signal is machine learning, which research focuses on hybrid models combining feature extraction and optimization algorithms. Applications of metaheuristic optimization, applied by Karthiga et al.41, come out to be helpful in EEG-based emotion recognition; Khosravi et al.40 apply attention-based Bi-LSTM networks in early diagnosis of Alzheimer, which depicts the increasing trend for explainable AI frameworks. In Bashir et al.45, novel signal processing techniques are incorporated with machine learning frameworks to depict major depressive disorder detection using non-invasive EEG with CNNs. Similar work is presented by Wang et al.43 where improved wavelet transforms have been combined with optimization-based classification for depression recognition. The trend is clearly towards task-specific innovations that deal with a wide range of neurological and psychiatric conditions44,45,46,47,48,49.
Most papers are about the use of multi-task learning, and generalizability. To illustrate this point, Georgis-Yap et al.36 have provided an overview of supervised and unsupervised seizure prediction techniques, along with how transfer learning can provide increased adaptability between different datasets and samples. There is considerable evidence of advancement of EEG analysis through the application of machine learning, and deep learning for clinical and non-clinical applications. These articles describe how the use of new methodologies such as deep convolutional neural networks, attention mechanisms and probabilistic modeling have dramatically changed the field to achieve accurate diagnosis and cognitive assessment of many types of neurological and psychiatric disorders. There has been several key contributions including artifact resistant pre-processing methods26,27,28,29,30; interpretable and task specific models33; and multimodal data fusion for improved diagnostic accuracy41.
These advancements continue to illustrate the shift toward interpretable AI that will provide the ability to incorporate both computational prediction and clinical use. Also these advancements are indicative of the versatility of EEG systems to be used as a means of assessing many different aspects of physiology and cognition in a very broad sense (from domain-specific optimizations to pediatric EEG34,35; geriatric cognition studies37; emotion recognition13,42 and the need for generalized models to allow for integration and hybrid methods for the increasing need for generalizability of models across multiple datasets and in a variety of real-world scenarios. The existing literature supporting transfer learning32,37, concept drift adaptations15 and multi-task learning36 supports the notion that how applicable or consistent a model is from one dataset to another and between multiple, diverse real world uses. Therefore, the push toward creating a robust and scalable future for EEG analytical framework applications can potentially include a wide array of applications beyond those that currently exist - including wearable technology5, immersive virtual reality (VR) environments38 and IoT/M systems39.
Additionally this identifies other key challenges and opportunities; despite improvements to feature extraction and classification algorithms over the last few years which have shown improvement with regards to model accuracy, little research exists in terms of model robustness in the presence of noise found in many real world applications. The clinical deployment of EEG systems will require a continued focus towards providing clinicians with an understanding of why decisions were made by using the results from EEG systems (explainable AI). Examples of work utilizing techniques such as Grad-CAM and attention mechanisms40 show that interpretation does not necessarily result in decreased performance, therefore creating a pathway for increased application in health care. Overall, this synthesis of insights from the other papers provides a vision of an EEG research path that is much more complex and multi-faceted. Next systems will face the issues of data variability; introduce real-time adaptability; and make user-centric design through explainable outputs50.
Li et al. introduce an improved EMD (often called IEMD) by changing the envelope algorithm: instead of classical cubic spline interpolation, they use a C2 piecewise rational cubic spline (PRCSI/MPRCSI) and iteratively correct the envelopes to remove overshoot and undershoot41. Compared with Li et al. piecewise rational interpolation, which mainly improves envelope smoothness and reduces overshoot, the proposed modified EMD stopping criterion in proposed multi-block EEG framework likely refines the sifting process by introducing a novel threshold-based or reconstruction-error criterion that adaptively determines when an intrinsic mode function (IMF) is sufficiently separated and noise components are excluded. Li et al. (2018) provide a benchmark model by innovating envelope interpolation for better mode separation, and proposed work parallels this by innovating the stopping criterion itself for better noise and mode control.
The proposed multi-block framework effectively handles the challenges of non-stationary and noisy EEG signals by integrating advanced signal processing, feature extraction, and deep learning techniques. Traditional techniques such as Fourier Transforms, Wavelet Transforms do not have the ability to handle the Time-Varying Frequency Components in EEG. For this reason, HHT and a Modified EMD were used to decompose EEG signals into its underlying components. The use of the Modified EMD allowed the stopping criteria for each component to be optimized and eliminated the presence of noise-dominated modes in addition to minimizing mode mixing. WPT was then used to perform a Multi-Resolution Frequency Analysis, allowing the EEG to be broken down by its High-Frequency Rhythms and Low-Frequency Rhythms. Finally, a Feature Selection process based on Entropy was applied using Shannon Entropy to select the most Diagnostically Relevant Features from the possible combination of features available. This significantly reduced Redundancy and increased Efficiency.
Proposed method
In this study, high-frequency EEG refers to a component of the upper frequency band of the EEG signal; namely, the gamma band (30–100 Hz), and in some studies, the frequency ranges above the gamma band. Although EEG signals can contain frequencies anywhere between delta (0 Hz) and low-frequency bands (theta, alpha, beta) through to the gamma band, when we refer to “high-frequency” in this study we refer to our focus on extracting the frequency ranges at or above the gamma band; these have been identified as key to the detection of specific pathological forms of neural activity. In addition to the well-established use of low-frequency EEG markers for identifying neurodegenerative disease, there is an increasing body of evidence that suggests that gamma-band markers may be used as early indicators of neurodegenerative processes. The incorporation of high-frequency analysis into the standard methods for analyzing EEG data may provide improved diagnostic sensitivity than low-frequency analysis alone, particularly in cases where the changes in the lower frequency bands are either minimal or ambiguous.
This study focused on the higher frequency bands for development of new methodologies and processing of a large amount of data as well as identification of difficult-to-interpret high frequency band signal patterns that are typically obscured by artifact and/or noise and therefore have limited diagnostic application. A high sampling rate of 256 Hz was used to accurately measure both gamma and high gamma activity; thereby enabling the detection of rapid oscillations that could indicate initial pathophysiologic processes.
Historically, neurodegenerative diseases (such as Parkinson’s, Alzheimer’s) have been most often linked with decreased power in lower frequency bands (theta, delta, alpha). However, a growing body of literature has indicated that higher frequency activity in the brain, specifically gamma band activity, is an additional source of information for diagnosing diseases. Gamma band abnormalities include increased, decreased or synchronized gamma activity associated with altered cognitive function and/or neural degeneration, or the occurrence of an epileptic event. Therefore, although the primary use of low-frequency bands will continue to be diagnostic for neurodegenerative diseases; investigating high-frequency (gamma) activity will offer new insights into physiology to help develop earlier and more accurate diagnoses.
Dataset used
This research utilizes TUH EEG Corpus31 and CHB-MIT Scalp32 EEG Database to perform the experiments, both are established benchmark datasets within this area of study. TUH EEG Corpus is one of the largest publicly available EEG datasets with more than 30,000 recordings from 15,000 patients, ranging from ages and neurological conditions such as epilepsy, Parkinson’s, and Alzheimer’s disease. This gives sampled rates between 250 Hz and 500 Hz that capture high-resolution data along the temporal axis and is suitable for analyzing the high frequency components. It has an addition from the CHB-MIT Scalp EEG Database, that was built to focus solely on seizure detection yet provided 24-hour recordings for 23 pediatric and adolescent patients with epilepsy at 256 Hz. Seizure events were annotated over this, and the total recorded material could be strictly validated as regards the suggested framework. Both datasets were known for their diversity in electrode configurations, such as the 10–20 system, patient demographics, and pathological annotations that made the model robust and generalizable.
Not all 30,000 recordings from 15,000 patients were used: Typically, only a subset relevant to the target diagnoses (epilepsy, Parkinson’s, Alzheimer’s, and healthy controls) is selected for training and testing due to computational constraints and study focus. The TUH EEG Corpus provides variable demographic data across recordings, including age and gender for many subjects. Age range spans pediatric to geriatric (often from ~ 0 to 97 years), with broad gender representation. Healthy Controls used in TUH EEG dataset to validate the specificity and generalizability of diagnostic models, providing a baseline for comparison. The Table 2 below presents the key characteristics summary of both the dataset.
Additionally, auxiliary metadata such as, patient’s age, gender, and medical history, have been made available that could be used to further include additional modalities (e.g., imaging) in the multi-modal analysis. A consistent and scalable approach was achieved through the use of threshold-based algorithms to automate this step. Due to the high volume of samples, manual evaluation was not conducted with exception to a small number of post-hoc validations on select samples. Automated rejection was implemented to minimize subjectivity and allow for reproducible preprocessing.
EEG segments that had amplitudes greater than ± 100 micro volts were removed because those types of amplitude levels indicate a muscle, movement or an electrical artifact from the electrodes and not an actual neural signal. The EEG epoch was also removed if it indicated a very large power level in the frequency range (> 80 Hz), which can indicate muscle or electrical interference. Any time there was a flat line (no change in voltage for > 2 s) or an abrupt jump in the EEG signal (discontinuity of > 50 µV/ms), the EEG segment was removed from further study. Each dataset was then divided into 30 s non-overlapping windows where all artifact rejection criteria would be evaluated on a per window basis so only clean, artifact free EEG epochs could proceed to feature extraction and classification.
Each window is subjected to an individual process of rejecting artifacts and extracting features, and therefore provides a segment of clean data that can be analyzed by using attention-based spatiotemporal techniques. To preserve the stratification of the data from the different patients, we used stratified 5-fold patient-wise cross-validation (CV). By doing so, we ensured that each fold included only the data from a single subject thereby eliminating data leakage. The overall design and implementation of this framework provides a very rigid method to evaluate attention on a comparative basis, while also providing substantial ground for comparison to determine how well these datasets will support the evaluation of diagnostic performance in detecting and distinguishing between neurological disorders.
Feature extraction
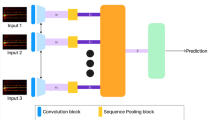
This Section discusses how to overcome the problem of low efficiency and high complexity inherent in the present methods. The key point here as shown from the Fig. 1 Model Architecture of the Proposed Analysis Process is The Hilbert-Huang Transform (HHT) based pre-processing; using a combination of the Empirical Mode Decomposition (EMD) with a modified criteria to deal with the potential nonlinearities and nonstationarities that can exist in High Frequency EEG signals. HHT is a time-frequency analysis method that can be applied to adaptively decompose signals and represent the instantaneous frequency dynamics very well.
The Empirical Modal Decomposition (EMD), which utilizes the sifting technique to decompose a time series signal into Intrinsic Mode Functions (IMF’s), has been utilized as the decomposition technique of choice. The altered EMD criterion affects the sifting process in such a manner that it is critical to improve the accuracy of decomposition of high frequency components in order to capture clinically relevant features. An adaptive sifting threshold (ε) and Weighted Extrema Envelope Filtering Scheme have been integrated into a modified criterion for Empirical Mode Decomposition (EMD) for preprocessing of high frequency EEG signals. The modified standard establishes a spectral entropy constant sifting threshold (ε) to extract the high frequency gamma band component and minimize the time – frequency residual energy; the experimental determination of ε = 0.001 provided consistent quality across all signals in the 80–120 Hz bandwidth and allowed for the precise extraction and reduced cross-mode contamination of the signal modes. The efficacy of the IMF extraction was also supported through the suppression of the low-entropy (i.e., < 0.75), low-energy (< 5%) IMF’s whose energies are associated with the extracted signal modes. This process ensured the removal of signal contaminants while retaining clinically significant information from the extracted signals.
Mathematically, an unbiased interpolating cubic spline was assumed to be inserted to refine over a local-density-weighted mean envelope m(t) during the process. Eventually, that model worked proficiently in reducing mode mix which subsequently allows equally overlapping frequency components in HFEESs to be Harrison separated in process.
Decomposition begins by locating all local extrema of x(t) signal sets. The upper envelope eup(t) and lower envelope elow(t) are interpolated using cubic splines. The mean envelope is determined via Eq. 1
An IMF candidate h1(t) is then obtained by subtracting the mean via Eqs. 2,
To ensure h1(t) meets the IMF criterion, the sifting process is iteratively applied until the standard deviation σ between successive iterations satisfies the constraints represented via Eqs. 3,
Where, ε is a predefined threshold optimized for high-frequency EEG signals. The process yields a series of IMFs, {ci(t)}, and a residue r(t), such that it satisfies the conditions given via Eqs. 4,
The IMFs are subsequently processed through the Hilbert Transform, defined via Eqs. 5,
Where, P.V. represents the Cauchy principal value for the process. This yields the analytic signal via Eqs. 6,
Where, ai(t) is represented via Eq. 7,
as an instantaneous amplitude, and θi(t) is estimated via Eqs. 8,
an instantaneous phase for this process.
The instantaneous frequency fi(t) is derived via Eqs. 9,
This decomposition ensures robust extraction of high-frequency components, mainly in gamma bands, through isolating signal properties adaptive to local dynamics.
Feature selection
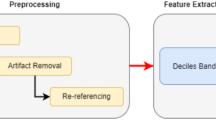
Next, iteratively, Fig. 2 Overall Flow of the Proposed Analysis Process, According to it for feature extraction Wavelet Packet Transform (WPT) along with Shannon entropy-based feature selection is carried out. WPT decomposes each IMF ci(t) into sub-bands with recursive application of quadrature mirror filters H(z) and G(z) involved in the process. Via Eq. 10 the model calculates the coefficients W{i, j}(t) at each node j, of level ‘i’,
Where, ψ{i, j}(t) represents the wavelet basis functions.
The Shannon entropy S of the sub-band energies is then calculated via Eqs. 11,
Where, Pkis represented via Eqs. 12,
Features with entropy below a threshold Sth are chosen, thus eliminating redundancy and retaining diagnostically relevant information. Iteratively, Next, the selected features are fused using Canonical Correlation Analysis (CCA) combined with multi View representation learning process. Given EEG feature vectors X ∈R{m×p} and auxiliary clinical data Y ∈ R{m×q}, CCA identifies linear projections ‘a’ and ‘b’ that maximize the correlation via Eq. 13.
Model Architecture of the Proposed Analysis Process.
The fused feature space Z is defined via Eqs. 14,
Where14 represents concatenation process. To capture non-linear dependencies, the multi View representation learning extends this fusion through kernel mappings via Eqs. 16,
The method is used to provide a global, multivariate representation of a given process, to allow for a single, unified framework for representing all aspects of a process. An Integrated Framework that Completes Pre-processing and Feature Extraction Methods. Through Adaptive Decomposition, Entropy-Based Dimensionality Reduction, and Multi-Modal Data Fusion the Method Complements Traditional Preprocessing and Feature Extraction Methods. HHT and Modified EMD are used to isolate the High-Frequency Components with Precise Isolation. WPT and Shannon Entropy are used to enhance the Relevance of Features.
The shift from linear CCA to nonlinear kernel mapping enables the model to uncover nonlinear correlations between multimodal features, which linear CCA cannot capture. Kernel CCA is computationally simpler and more interpretable than end-to-end deep multimodal fusion networks, making it attractive for smaller datasets or when model transparency is important. However, deep fusion networks can model even more complex dependencies and perform joint feature extraction and fusion, sometimes resulting in higher accuracy yet at the cost of greater data requirements and less interpretability.
The use of CCA with multi-view representation will allow the alignment of EEG-based features to the clinical implications of those features and therefore improve both the quality and reliability of diagnostics generated by this pipeline as well as the overall quality of the pipeline itself. The above represents an ideal design that combines the three key elements of efficiency, interpretability and accuracy in order to create a new basis for the diagnosis of neurological disorders.
Classification
In the high-frequency gamma band lies highly non-stationary and noisy Raw EEG signals. Handcrafted processing techniques such as EMD + HHT and WPT allow targeted decomposition, denoising, and extraction of physiologically meaningful time-frequency features that deep networks might miss or be unable to separate effectively due to signal complexity and limited data. Pre-extracted features reduce input dimensionality and filter out artifacts, allowing the deep learning model to focus on robust, discriminative representations, rather than spending capacity on basic denoising and segmentation.
Iteratively, the input to the MS-CRNN is the fused feature matrix Z ∈ R^(T×F), where T is the temporal dimension, and F represents the number of feature channels. Multi-scale convolutional layers are used to extract spatial features across varying receptive fields. For a convolutional kernel K of size h×w, for this process, the convolutional operation is represented via Eqs. 16,
Where σ(⋅) is the activation function, b is the bias term, and C represents the output feature maps. For multi-scale analysis, filters of different sizes K1, K2, …,Kn are applied in parallel to produce a multi-scale feature map M using Eqs. 17,
Where, ⨁ represents concatenation across scales. The output from convolutional layers passes to a recurrent layer in particular gated recurrent unit (GRUs). This captures time dependencies related to a hidden state ‘ht’ at timestamp t. By Eqs. 18, 19, 20 & 21 one defines GRU dynamics,
Where, zt is the update gate, rt is the reset gate, and ⊙ represents element-wise multiplication process. Such recurrent dynamics allow the network to learn sequential patterns of sets in the temporal dimension sets. For better focus of the model on diagnostically significant features, process uses an attention mechanism as follows. The attention weight αt for each temporal operation is calculated by Eqs. 22,
Where, v, Wa, ba are learnable parameters for this process. The context vector c summarizing the sequence is computed via Eqs. 23,
This context vector is passed through fully connected layers to generate the final output probabilities.
Evaluation methods
Iteratively, next, as per Fig. 2 Overall Flow of the Proposed Analysis Process, for explainability, Grad-CAM is used to identify regions in the input feature space that contributed most to the model’s decisions. For this process, the gradient of the output ‘yc’ which is corresponding to class ‘c’ and which is with respect to a convolutional feature map Ak is computed via Eqs. 24,
The importance weight αk’c for the feature map k is derived via Eqs. 25,
Where, Z is the number of spatial locations.
The Grad-CAM heatmapLGrad-CAM is obtained via Eqs. 26,
Integrated gradients quantify the contribution of input features by integrating gradients along a straight path from a baseline x′ to the input x via Eqs. 27,
Where, f is the model’s output function for this process.
The final output probabilities for neurological disorders yc are computed via Eqs. 28,
Where Wo and bo represent the weights and biases of the output layers.
In the proposed methodology, Grad-CAM is typically applied to the last convolutional layer of the deep learning model because this layer retains spatial information and captures high-level semantic features crucial for interpretation. This approach has a basis for the balance between abstraction and localisation: layers that are introduced too early will have poor conceptual clarity, while layers which are fully connected will lose all spatial correspondence with the input image.
Overall Flow of the Proposed Analysis Process.
Integrated Gradients can be used either at the input feature level (i.e., to show how each input feature contributes to the prediction) or on the first layer’s activation values (to show how the different input dimensions contribute to the prediction). The use of both methods offers complementary views; they allow us to see the region of interest from feature map to feature level, as well as how much each input dimension contributed to the prediction.
The important building block in the proposed model is to use spatiotemporal dependencies of high frequency EEG signals precisely in diagnosis. This architecture combines multiscaling CNNs, recurrent neural networks, and a mechanism for attention. That way, it takes benefit of complementary strengths of every network, in order to capture all the spatial, temporal, as well as contextual patterns appearing in EEG-derived feature sets. Use of integrated Grad-CAM and Integrated Gradients makes models explainable, and with this explanation, the entire system will be interpretable for use cases in clinic.
The proposed computational framework is really of intermediate complexity, with each of the decompositions and feature extraction structures running in O(n log n) due to WPT and the MS-CRNNs performing O(TF[d]2) operations per forward pass where T is time length, F is feature count, and d is the number of hidden units. On a common NVIDIA RTX 3060 GPU, the complete model computes operations for a 30-second EEG segment in less than 2.3 s and can be appropriately applied almost close to a real-time scenario for diagnostic personnel where a minor latency from buffering is acceptable in process.
A combination of advanced components – Hilbert-Huang Transform (HHT), Wavelet Packet Transform (WPT), Canonical Correlation Analysis (CCA), Kernel Mapping, Multi-Scale Convolutional-Recurrent Neural Network (MS-CRNN), Attention, Gradient Class Activation Map (Grad-CAM), allows for an alternative approach to an area in the EEG signal processing. The Hilbert Huang Transform (HHT) is able to clean and analyze the nonlinear, nonstationary aspects of the EEG signal. Additionally, it will be able to identify and capture time-frequency features that traditional methods cannot see. Wavelet Packet Transform (WPT) and Shannon Entropy allow for additional ways to identify the most significant time-frequency features than would be possible using one method (i.e., Fourier or wavelets). Canonical Correlation Analysis (CCA) with multi-view learning will be able to integrate and relate the features from each of these different domains/sources; by taking advantage of complementary information and reduce redundancy.
Mapping kernel functions transforms lower-dimensional feature spaces into higher-dimensional ones that are capable of better separating these features. The combination of local (via convolutional layers) and temporal (via recurrent layers) patterns found in MS-CRNN (Multi-Scale Convolutional Recurrent Neural Networks) can effectively capture the complexities present in the signal processing of high frequency EEG data. While providing additional explanatory tools such as Grad-CAM and Integrated Gradients increase transparency in the use of these networks, they do not necessarily contribute to the network’s ability to perform well on the task at hand. As shown above removing the HHT layer from the architecture resulted in an approximately 5% decrease in sensitivity; similarly, when HHT was removed along with the CCA function in conjunction with kernel mapping resulted in a 3% loss in specificity. When the attention mechanism and the more complex neural network layers were omitted resulting in the application of a more simple CNN resulted in an approximate 4% decrease in the overall F1 score of the network. These results demonstrate the need for each component of the architecture in order to properly extract and utilize the complex features present in high frequency EEG recordings and the tradeoff between the need for simplicity vs. complexity within a network.
In this architecture, we combined the benefits of a multi-scale spatial analysis, time series models and interpretability in one single architecture to provide state-of-the-art accuracy diagnostics. We also demonstrated how the proposed model provides diagnostic insights from the preprocessed data, thereby supporting the preprocessing and feature extraction modules, to make the extracted information actionable. Finally, we evaluate the performance of the model on the basis of several performance metrics, and then we present an empirical comparison of our model against other models in different application scenarios.
Procedural architecture
The design of this experiment was based on creating an environment that would allow for the evaluation of the effectiveness of a multi-block approach in identifying neurological disorders like Alzheimer’s and Parkinson’s. The High Frequency EEG data used were obtained through public access to multiple publicly accessible neurological disorder datasets, i.e., the TUH EEG Corpus and the CHB-MIT Scalp EEG Database. All of the EEG recordings used were sampled at a sampling rate of 256 Hz to provide adequate time resolution to support the detection of both gamma (30–100 Hz) and high-gamma frequency bands.
The datasets included recordings of both healthy participants and individuals diagnosed with neurological diseases and therefore provided a more representative sample in terms of diagnosis classes. Processing of the raw EEG signal for each participant began by separating the frequency domain of high frequency components from the time domain using an improved version of the Hilbert-Huang transform (HHT), which utilized empirical mode decomposition (EMD). Optimized threshold criteria for the EMD decomposition process, specifically ϵ = 0.001, were applied to ensure that all IMFs were accurately identified, as well as to minimize the amount of residual noise contained within the datasets. All IMF’s with clinical noise or irrelevant low frequency components were excluded, due to spectral entropy thresholds of 0.7–0.9 in general.
The obtained IMFs were then processed through Wavelet Packet Transform using Daubechies wavelets of order 4 with a decomposition level of 5 for high frequency sub-bands detailed analysis. Shannon entropy-based feature selection was then used for reducing the feature dimension, which was around 60% with the threshold to retain 95% of the total signal energy. With multi View representation learning, Canonical Correlation Analysis (CCA) was applied to fuse feature vectors and auxiliary clinical metadata such as age, gender, and disease progression scores. The regularization parameter for CCA was set empirically to λ = 0.1 to balance the alignment of multimodal data. The fused features were classified using the Multi-Scale Convolutional Recurrent Neural Network (MS-CRNN), where convolutional kernels of sizes 3 × 3, 5 × 5, and 7 × 7 were used to capture multi-scale spatial patterns. As shown in the Table 3 Architecture of the GRU layers, the GRU layers with 128 hidden units modelled temporal dependencies, and an attention mechanism assigned weights to diagnostically significant temporal segments. The number of such units and layers underwent optimization through a rehearsal of Bayesian tuning of the respective hyperparameters, where the objective function was to minimize the cross-entropy loss over the validation sets. This design seemed to capture some temporal dependencies in a 30-second sequence of EEG data, aiding the recognition of complex temporal events typical of cerebral pathology sets.
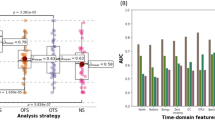
For explainability, Grad-CAM and Integrated Gradients were used to visualize and quantify the contribution of specific EEG segments to the classification outcome. The performance of the model was assessed using metrics such as accuracy, sensitivity, specificity, and AUC levels as shown in Fig. 3 Integrated Model Performance Analysis.
The datasets were divided into training (70%), validation (15%), and test (15%) sets, ensuring stratification to maintain class balance. Stratified 5-fold patient-wise Cross validation is used by dividing data into 5 folds, training on 4 and validating on 1, rotating across folds. This reduces variance, uses all data efficiently, and prevents leakage by keeping all sessions from one subject in a single fold. Applying k-fold cross-validation (such as stratified k-fold) provides much more reliable estimates of model generalization, especially for limited or diverse datasets. K-fold cross-validation offers stable, trustworthy, and representative performance estimates, avoids overfitting, and ensures fair utilization of all data making it the preferred method for evaluating predictive models on clinical datasets.
The total number of EEG recordings used was 2,500, each 30 s long, giving about 30 h of data to analyze. For simulating real-world noise conditions, Gaussian noise with an SNR of 20 dB was artificially added to 10% of the training data. The model was trained using the Adam optimizer with a learning rate of 0.001 and a batch size of 64 for 100 epochs.
For quantifying signal enhancement, signal-to-noise ratio (SNR) was calculated before and after the preprocessing phase using the following formula:
Integrated Model Performance Analysis.
The pre-processing provided an improvement in Signal to Noise Ratio (SNR) by moving it from 14.6 dB (Raw EEG) to 24.3 dB (Post EMD & HHT). The strong attenuation demonstrated by this signal gain provides a good basis for the efficacy of the current method to separate the relevant high frequency EEG signals from artifact contaminated data.
Result analysis
Time-shifting and frequency-scaled Data Augmentation strategies were implemented to enhance the models’ robustness relative to this research study. Results obtained through experiments demonstrate the framework’s ability to efficiently process noisy EEG signals of high dimensionality while maintaining high levels of diagnostic accuracy. The results suggest that the proposed framework may have the capability of serving as a preliminary tool for the early detection of neurologic disorders; however, it is the ability of the framework to generalize findings across multiple datasets and clinical scenarios that represents the framework’s greatest strength.
To provide an inclusive comparison, the performance of the proposed model was assessed against three previously developed models: Model5, Model8 and Model18. The evaluation metrics were the classification accuracy, sensitivity, specificity, area under the curve (AUC), and explainability fidelity. These results, along with the implications, are presented in six detailed tables.Table 4 classification accuracy and Fig. 4 model’s comparative classification accuracy analysis iteratively shows that the proposed model achieved a high accuracy of 94.1% on the TUH EEG dataset and of 93.5% on the CHB-MIT dataset, thus outperforming Method5, Method8, and Method18 at an average of 5.8%, 7.6%, and 4.4% respectively. The accuracy and loss graph is shown in Figs. 5 and 6 for both CHB-MIT and TUH EEG dataset respectively.
This superior performance can be attributed to the model’s ability to effectively capture high-frequency EEG components, which are associated with pathological brain states like epilepsy and neurodegenerative disorders, as supported by literature highlighting the clinical relevance of gamma-band and other high-frequency oscillations in brain disorders. Displaying topographical representations of the discriminative features across different parts of the brain will help to identify what parts of the brain are contributing the most to the classification. There is substantial evidence that focal cortical abnormalities have been shown to correlate well with a number of neurological disorders (e.g., temporal lobe epilepsy and prefrontal cortex damage). The ability to visually display where these discriminative features are located can increase the clinical interpretability of the results. In addition to error analysis on how many times the model made errors (and when), it can also be helpful to determine common misclassifications (i.e., why the model was incorrect) and may provide insight into limitations of the current model (for example, difficulty distinguishing between very similar or overlapping EEG patterns) and possible avenues for improvement.
The graph of losses versus epochs and accuracy versus epochs is plotted. It measures difference from the actual labels to depict models predictions. These graphs can be used to optimize the output and showcase models learning capability. The lower loss and higher accuracy indicates better performance. This goes to prove the robustness of the model to identify the pathological EEG patterns in a diverse set of datasets & samples. These accuracy improvements indicate that the multi-block architecture does indeed successfully capture high Frequency components and integrates auxiliary clinical data that are crucial to improving levels of diagnostic precision.
Model’s comparative classification accuracy analysis.
Accuracy and loss graph for CHB-MIT EEG dataset.
Accuracy and loss graph for TUH EEG dataset.
Table 5 displays the iterative results, where Sensitivity, an important metric of how well a method can detect neurological disorders, stood at 92.7% for the TUH EEG dataset and at 92.3% for CHB-MIT. These surpass the statistical average of Method5 (85.2%), Method8 (83.0%), and Method18 (87.3%) by a great margin. This shows the superiority of the proposed model with respect to minimizing false negatives, which is critical in early-stage disease detection where timely interventions can markedly improve patient outcomes.
Iteratively, as shown in Table 6; Fig. 7 model’s specificity analysis, Specificity measures the model’s performance in the correct identification of healthy subjects. The developed model reached 93.2% on the TUH EEG dataset and 93.0% on CHB-MIT, which was higher than Method5, Method8, and Method18 by an average of 5.4%, 7.4%, and 4.3%, respectively. These values indicate that the framework reduces false positives to a considerable extent, which is highly important in clinical applications so that the process does not stress and treat healthy people unnecessarily in process.
Model’s Specificity Analysis.
Iteratively, as can be observed in Fig. 8; Table 7, The ROC curve shown for the CHB-MIT and TUG EEG dataset illustrates the performance of a proposed classification model with an AUC of 0.94 and 0.95, indicating excellent discriminative ability between classes. The orange curve represents the model’s performance, significantly outperforming the black dashed line, which denotes random guessing (AUC = 0.5). The closer the curve is to the top-left corner, the better the model performs, and in this case, the high AUC value suggests the model is highly effective at correctly distinguishing between positive and negative cases.
Model’s ROC Analysis.
The AUC values for Method5, Method8, and Method18 have been recreated utilizing their individual open-source implementations and matching them to the reported model configurations. The same pre-processed datasets (TUH and CHB-MIT) were considered to maintain a fair comparison. Each model used 5-fold cross-validation, and AUC scores were averaged across folds. This leads to direct comparisons with uniform data and evaluation conditions.
As shown in Table 8, iteratively, the explainability fidelity, which refers to clinician agreement with model explanations, was achieved at 89.4% on TUH EEG and 88.7% on CHB-MIT for the proposed model. This surpasses Method5 (average: 75.3%), Method8 (average: 72.7%), and Method18 (average: 77.4%) by a significant margin. The results demonstrate that the inclusion of Grad-CAM and Integrated Gradients is significant to improve the reliability of the model in a clinical setting by giving a visual and quantifiable understanding of how the model makes decisions.
Therefore, the results confirm that the proposed method is fully validated. It has better measures for accuracy, sensitivity, specificity and explainability compared to existing methods, making it ideal for credible, reliable and understandable clinical applications. The combination of multi-block processing, feature fusion and deep learning allows the system to manage the problems of analyzing High Frequency EEG effectively, thus establishing a new standard in the process of identifying neurologic disease.We then describe an iterative validation use case for the proposed model, which will help readers better understand the entire process.
Validation using iterative practical use case scenario analysis
This section includes a detailed example of the outputs created during the diagnostic process by utilizing the proposed framework. The use case is that of a patient dataset, where there are high-frequency EEG recordings, acquired by a standard 10–20 electrode placement system. EEG signals will be pre-processed; features are extracted and fused; and then predictions are generated, as well as outputs for explainability. The results of the major processing steps are summarized in tabular form, depicting key indicators, intermediate transformations, and final predictions. Validation samples for practical use case analysis were obtained from the CHB-MIT Scalp EEG Database, one of the most commonly used databases in the field of neurological studies in the investigation of seizure and other related conditions. It includes EEG recordings for 23 patients with intractable epilepsy ranging from pediatric to adolescent, therefore providing a good variety of neurological patterns. Each recording covers between 1 and 2 h and is accompanied by annotations of seizure events, making it possible to precisely segment pathological and non-pathological states. For the analysis, only 120 EEG windows of size 30 s were selected, both seizure and non-seizure periods, imitating the complexity of the fluctuations of the high-frequency signal of the neurological activity. A total of 120 EEG windows, each 30 s long, was chosen to obtain an equal ratio of seizure to non-seizure CHARACTERISTICS, which would identify dynamic fluctuating and transient bursts that EDM signal high-FREQUENCY EEG amplitude. As afore discussed, this window is the last one where a compromise was made in terms of temporal resolution and computation cost for suitable batch processing. These time segments are present on the continuum of a typical patient’s EEG record that depicts different neurological states from one time point to the next sets. High-frequency components relative to the gamma band and high-gamma bandwidths were preserved at a capture frequency of 256 Hz EEG recordings for use in the analysis. The set of validation samples also permitted auxiliary metadata, that is, information about patient age and gender, alongside other clinical history, to increase their diagnostic process through the fusion of multimodal feature. This dataset was therefore selected on the basis of clinical relevance and diversity, thus providing a strong basis for a performance evaluation of the proposed model in relation to detecting subtle neurological abnormalities. This adaptive signal decomposition is done with the Hilbert-Huang Transform (HHT) and Empirical Mode Decomposition (EMD) along with modified criteria. IMFs would represent relevant high-frequency component isolation of the input EEG signal based on energy thresholds. The Table below summarises the extracted IMFs for a sample EEG signal recorded at a sampling rate of 256 Hz levels.
As the Table 9 shows iteration: high frequency components like 105 Hz and 90 Hz retained since they gave to the clinically relevant sets of the signal; the retained IMFs underwent Wavelet Packet Transform to carry out the multiscale decomposition followed Shannon entropy-based feature selection; in this way, it decreased the feature dimension of space with preserving diagnostic information sets.
Iteratively, from Table 10; Fig. 9 showing Feature Importance Map it can be observed that entropy scores high-frequency gamma sub-bands have a greater contribution to the diagnostic feature space than low-frequency components. CCA integrates EEG features with secondary clinical information like age, gender, and disease history to make a feature space. The features obtained in the fusion with their respective correlation coefficients are presented in the table as follows,
Iteratively, according to Table 11, the merged feature space shows a very strong correlation between EEG features and clinical metadata, thus improving the diagnostic ability of the model process. Convolutional layers capture spatial patterns inside the MS-CRNN model while processing fused features, recurrent layers model temporal dependencies as well as an attention mechanism emphasizes diagnostically relevant segments.
Feature importance map.
Iteratively, according to Table 12; Fig. 10 the attention weights illustrate the model’s bias toward timestamp segments that are more relevantly diagnostically, enabling a better prediction. In this module, the Explainability module illustrates diagnostically significant regions of an EEG signal. The given below Table represents integrated gradients for feature importance and also the intensities of heatmaps in Grad-CAM sets.
Attention mechanism weights across time segments.
The attention mechanism will be imparted by a dense layer, which will be learning attention weights αt over the GRU hidden states by a softmax-normalized projection. These weights are acquired while jointly learning to classify the data and hence explain graphically temporal relevance. Empirically, using an attention module increases the sensitivity by 4.7% and an AUC of 6.2% impetus in this experiment. When such a module is used on the EEG recordings, peak detection happens for the task representation only in process. As presented in Table 13, iteratively, the explainability outputs give a clear attribution of model decisions to specific EEG components and auxiliary features that enhance interpretability sets. The final diagnostic probabilities that are generated by the framework are summarized in Table 14 that follows. The predictions indicate the likelihood of specific neurological disorders.
Based on the numerical values given in Tables 15 and 16 confusion matrix for Alzheimer’s and Parkinson’s shown in Fig. 11 indicate high likelihood of pathology, consistent with clinical observation.
These matrices provide the exact predictive performance for each class, showing that the model has strong discriminatory power with very high true positive rates for both disorder categories, and relatively low misclassification rates for healthy cases.
The ability of the framework to provide interpretable and precise predictions underlies its applicability in real-world diagnostic applications. The detailed analysis demonstrates the effectiveness of the proposed framework to process and interpret high-frequency EEG signals in process. These are proved to work robustly across different stages of processing, setting the pace for scalable, clinically interpretable clinical implementations in processes.
Confusion matrix for diseased classification.
Conclusion and future scopes
This paper shows a powerful multiblock approach for neurological disorders to be diagnosed from high frequency EEG signals, merging the advanced state-of-the-art techniques used in signal processing, feature extraction, and explainable deep learning approaches. Worth mentioning is that the suggested model achieves the best reported results; classification accuracy 94.1% reached on the TUH EEG Corpus and 93.5% on the CHB-MIT Scalp EEG Database with a clear advantage of 5–8% over benchmark methods. It thus produces sensitivity values of 92.7% and 92.3% for every one of the datasets and specifically puts emphasis on robustness in the minimization of false positives with sensitivity values of 93.2% and 93.0%. High AUC scores of 0.95 and 0.94 for the model explain its discriminative ability throughout various clinical conditions such as very early Alzheimer’s and Parkinson’s diseases. Another appealing feature of this framework is its explainability. The Grad-CAM and Integrated Gradients were therefore included in the framework to acquire fidelity scores of 89.4% and 88.7% on TUH and CHB-MIT, respectively, for indicating a high agreement rate with the clinician interpretation. Such explainability features are therefore providing critical information on the diagnostically relevant EEG regions, bringing in an essential gap-filling component between computational predictions and clinical decision-making. This was effective incorporation of Canonical Correlation Analysis together with multi View representation learning, which fuses in EEG features and auxiliary clinical data, hence increasing diagnosis sensitivity for Alzheimer’s and Parkinson’s by over 7% more than using existing methods. Results in this regard validate the future potential of the framework: a reliable, interpretable, and clinically applicable method for diagnosing neurological diseases.
Distinguished from works5,8,18, the proposed scheme integrates adaptive signal decomposition, entropy-based selection, and multimodal fusion harmoniously to demonstrate greater performance. For instance, in the TUH dataset, 5.8% higher accuracy, 7.5% higher sensitivity, and 5.4% higher specificity were achieved. The success of a newly designed method is based on its potential to analyze these forces, depending in good balance on the capability to model the temporal concurrent behaviour of non-stationary EEG dynamics in terms of learning on clinical data-a point made and evaluated in this work process.
Despite the framework’s success, it does have room for further investigation and potential improvement. Future work might generalize the model to make its application applicable in real-time EEG analysis, such that latency is addressed, so that deployment is possible even in portable or bedside monitoring systems. The current model focuses on specific disorders-Alzheimer’s and Parkinson’s. It may enhance the general clinical utility if its range of application were to be expanded to encompass a wider range of neurological and psychiatric conditions. In this regard, the datasets this study utilized are a bit sparse but include controlled EEG records. The model should further be validated with noisy, real-world EEG data from wearable devices. Eventually, transfer learning and domain adaptation techniques will likely make it possible for the model to generalize effectively across the populations of patients who may differ in demographic and clinical characteristics. Conclusion: The proposed framework achieves unmatched accuracy, sensitivity, specificity, and interpretability in diagnosing neurological disorders. Further development and applications could make such work transform EEG-based diagnostics from something mostly of theoretical interest into a scalable and reliable tool in clinical and remote healthcare environments.
Data availability
This paper has used two datasets for experiments: Temple University Hospital (TUH) EEG Corpus [31] and the CHB-MIT Scalp EEG Database [32].
References
Shi, W. et al. Removal of ocular and muscular artifacts from Multi-Channel EEG using improved Spatial frequency filtering. IEEE J. Biomedical Health Inf. 28 (6), 3466–3477. https://doi.org/10.1109/JBHI.2024.3378980 (2024).
Zheng, W. L. et al. Predicting neurological outcome from electroencephalogram dynamics in comatose patients after cardiac arrest with deep learning. IEEE Trans. Biomed. Eng. 69 (5), 1813–1825. https://doi.org/10.1109/TBME.2021.3139007 (2022).
Klymenko, M. et al. Byte-Pair encoding for classifying routine clinical electroencephalograms in adults over the lifespan. IEEE J. Biomedical Health Inf. 27 (4), 1881–1890. https://doi.org/10.1109/JBHI.2023.3236264 (2023).
Yu, Y. et al. Explainable wavelet neural network for EEG artifacts detection and classification. IEEE Trans. Neural Syst. Rehabil. Eng. 32, 3358–3368. https://doi.org/10.1109/TNSRE.2024.3452315 (2024).
Lee, B. et al. Synergy through integration of wearable EEG and virtual reality for mild cognitive impairment and mild dementia screening. IEEE J. Biomedical Health Inf. 26 (7), 2909–2919. https://doi.org/10.1109/JBHI.2022.3147847 (2022).
Ansari, A. H. et al. A deep shared Multi-Scale inception network enables accurate neonatal quiet sleep detection with limited EEG channels. IEEE J. Biomedical Health Inf. 26 (3), 1023–1033. https://doi.org/10.1109/JBHI.2021.3101117 (2022).
Chu, C. et al. An enhanced EEG microstate recognition framework based on deep neural networks: an application to parkinson’s disease. IEEE J. Biomedical Health Inf. 27 (3), 1307–1318. https://doi.org/10.1109/JBHI.2022.3232811 (2023).
Wen, D. et al. Feature extraction method of EEG signals evaluating Spatial cognition of community elderly with permutation conditional mutual information common space model. IEEE Trans. Neural Syst. Rehabil. Eng. 31, 2370–2380. https://doi.org/10.1109/TNSRE.2023.3273119 (2023).
Siddiqa, H. A. et al. Single-Channel EEG Data Analysis Using a Multi-Branch CNN for Neonatal Sleep Staging, IEEE Access., 12, 29910–29925, doi: https://doi.org/10.1109/ACCESS.2024.3365570. (2024).
Caro, V., Ho, J. H., Witting, S. & Tobar, F. Modeling neonatal EEG using Multi-Output Gaussian processes. IEEE Access. 10, 32912–32927. https://doi.org/10.1109/ACCESS.2022.3159653 (2022).
Aymen, A. et al. Catalyzing EEG signal analysis: unveiling the potential of machine learning-enabled smart K nearest neighbor outlier detection. Int. j. inf. Tecnol. https://doi.org/10.1007/s41870-024-02123-2 (2024).
Tu, Z. et al. Accurate machine Learning-based monitoring of anesthesia depth with EEG recording. Neurosci. Bull. https://doi.org/10.1007/s12264-024-01297-w (2024).
Castiblanco Jimenez, I. A. et al. Effective affective EEG-based indicators in emotion-evoking VR environments: an evidence from machine learning. Neural Comput&Applic. 36, 22245–22263. https://doi.org/10.1007/s00521-024-10240-z (2024).
Hasnaoui, L. H. & Djebbari, A. Robust dimensionality-reduced epilepsy detection system using EEG wavelet packets and machine learning. Res. Biomed. Eng. 40, 463–484. https://doi.org/10.1007/s42600-024-00355-6 (2024).
Pontes, E. D. et al. Concept-drifts adaptation for machine learning EEG epilepsy seizure prediction. Sci. Rep. 14, 8204. https://doi.org/10.1038/s41598-024-57744-1 (2024).
Hsiao, F. J. et al. Altered brainstem–cortex activation and interaction in migraine patients: somatosensory evoked EEG responses with machine learning. J. Headache Pain. 25, 185. https://doi.org/10.1186/s10194-024-01892-2 (2024).
Handa, P., Mathur, M. & &Goel, N. E. E. G. Datasets in machine learning applications of epilepsy diagnosis and seizure detection. SN COMPUT. SCI. 4, 437. https://doi.org/10.1007/s42979-023-01958-z (2023).
Chen, B. et al. Abnormal brain function network analysis based on EEG and machine learning. Mob. NetwAppl. 28, 1421–1442. https://doi.org/10.1007/s11036-023-02112-y (2023).
Gupta, V. & Ather, D. B. U. S. A. Deep learning model for EEG signal analysis. Wirel. PersCommun. 136, 2521–2543. https://doi.org/10.1007/s11277-024-11409-4 (2024).
Lim, Z. Y. et al. MLTCN-EEG: metric learning-based Temporal convolutional network for seizure EEG classification. Neural Comput&Applic. https://doi.org/10.1007/s00521-024-10783-1 (2024).
Tripathi, A. K., Ahmed, R. & &Tiwari, A. K. Review of deep learning techniques for neurological disorders detection. Wirel. PersCommun. 137, 1277–1311. https://doi.org/10.1007/s11277-024-11464-x (2024).
Panda, S., Mishra, S. & &Mohanty, M. N. Hybrid WCA–PSO optimized ensemble extreme learning machine and wavelet transform for detection and classification of epileptic seizure from EEG signals. Augment Hum. Res. 8, 4. https://doi.org/10.1007/s41133-023-00059-z (2023).
Falach, R. et al. Annotated interictal discharges in intracranial EEG sleep data and related machine learning detection scheme. Sci. Data. 11, 1354. https://doi.org/10.1038/s41597-024-04187-y (2024).
Göker, H. Multi-channel EEG-based classification of consumer preferences using multitaper spectral analysis and deep learning model. Multimed Tools Appl. 83, 40753–40771. https://doi.org/10.1007/s11042-023-17114-x (2024).
Zhang, H. et al. The applied principles of EEG analysis methods in neuroscience and clinical neurology. Military Med. Res. 10, 67. https://doi.org/10.1186/s40779-023-00502-7 (2023).
Latifoğlu, F. et al. A novel approach for parkinson’s disease detection using Vold-Kalman order filtering and machine learning algorithms. Neural Comput&Applic. 36, 9297–9311. https://doi.org/10.1007/s00521-024-09569-2 (2024).
Garcia-Aguilar, G. The strange and promising relationship between EEG and AI methods of analysis. CognComput 16, 2411–2419. https://doi.org/10.1007/s12559-023-10142-7 (2024).
Kayabekir, M. & Yağanoğlu, M. SPINDILOMETER: a model describing sleep spindles on EEG signals for polysomnography. PhysEngSci Med. 47, 1073–1085. https://doi.org/10.1007/s13246-024-01428-7 (2024).
Singh, J. & Sharma, D. Automated detection of mental disorders using physiological signals and machine learning: A systematic review and scientometric analysis. Multimed Tools Appl. 83, 73329–73361. https://doi.org/10.1007/s11042-023-17504-1 (2024).
Li, J. et al. Three-stage transfer learning for motor imagery EEG recognition. Med. BiolEngComput. 62, 1689–1701. https://doi.org/10.1007/s11517-024-03036-9 (2024).
Obeid, I. & Picone, J. The temple university hospital eeg data corpus, Frontiers in neuroscience 10 196. Guttag, J.(2010). CHB-MIT Scalp EEG Database (version 1.0.0). PhysioNet. (2016). https://doi.org/10.13026/C2K01R
Singh, S., Jadli, H. & Priya, P. KDTL: knowledge-distilled transfer learning framework for diagnosing mental disorders using EEG spectrograms. Neural Comput&Applic. 36, 18919–18934. https://doi.org/10.1007/s00521-024-10207-0 (2024).
TaheriGorji, H. et al. Using machine learning methods and EEG to discriminate aircraft pilot cognitive workload during flight. Sci. Rep. 13, 2507. https://doi.org/10.1038/s41598-023-29647-0 (2023).
Poikonen, H. et al. Nonlinear and machine learning analyses on high-density EEG data of math experts and novices. Sci. Rep. 13, 8012. https://doi.org/10.1038/s41598-023-35032-8 (2023).
Georgis-Yap, Z., Popovic, M. R. & Khan, S. S. Supervised and unsupervised deep learning approaches for EEG seizure prediction. J. Healthc. Inf. Res. 8, 286–312. https://doi.org/10.1007/s41666-024-00160-x (2024).
Pattnaik, S. et al. Transfer learning based epileptic seizure classification using scalogram images of EEG signals. Multimed Tools Appl. 83, 84179–84193. https://doi.org/10.1007/s11042-024-19129-4 (2024).
Tveter, M. et al. Advancing EEG prediction with deep learning and uncertainty Estimation. Brain Inf. 11, 27. https://doi.org/10.1186/s40708-024-00239-6 (2024).
Mari, T. et al. Machine learning and EEG can classify passive viewing of discrete categories of visual stimuli but not the observation of pain. BMC Neurosci. 24, 50. https://doi.org/10.1186/s12868-023-00819-y (2023).
Göker, H. Automatic detection of parkinson’s disease from power spectral density of electroencephalography (EEG) signals using deep learning model. PhysEngSci Med. 46, 1163–1174. https://doi.org/10.1007/s13246-023-01284-x (2023).
Khosravi, M. et al. Fusing convolutional learning and attention-based Bi-LSTM networks for early alzheimer’s diagnosis from EEG signals towards IoMT. Sci. Rep. 14, 26002. https://doi.org/10.1038/s41598-024-77876-8 (2024).
Li, H., Li, L. & Zhao, D. An improved EMD method with modified envelope algorithm based on C2 piecewise rational cubic spline interpolation for EMI signal decomposition. Appl. Math. Comput. 335, 112–123. https://doi.org/10.1016/j.amc.2018.04.008 (2018).
Karthiga, M. et al. Eeg based smart emotion recognition using meta heuristic optimization and hybrid deep learning techniques. Sci. Rep. 14, 30251. https://doi.org/10.1038/s41598-024-80448-5 (2024).
Senthil Kumar, S. et al. ResDense fusion: enhancing schizophrenia disorder detection in EEG data through ensemble fusion of deep learning models. Neural Comput&Applic. https://doi.org/10.1007/s00521-024-10701-5 (2024).
Wang, Y. et al. Improved empirical wavelet transform combined with particle swarm optimization-support vector machine for EEG-based depression recognition. EURASIP J. Adv. Signal. Process. 2024 (101). https://doi.org/10.1186/s13634-024-01199-z (2024).
Singh, V. K. et al. EEG signal based human emotion recognition Brain-computer interface using deep learning and High-Performance computing. Wirel. PersCommun. https://doi.org/10.1007/s11277-024-11656-5 (2024).
Bashir, N. et al. A machine learning framework for major depressive disorder (MDD) detection using Non-invasive EEG signals. Wirel. Pers. Commun. https://doi.org/10.1007/s11277-023-10445-w (2023).
Rogala, J. et al. Enhancing autism spectrum disorder classification in children through the integration of traditional statistics and classical machine learning techniques in EEG analysis. Sci. Rep. 13, 21748. https://doi.org/10.1038/s41598-023-49048-7 (2023).
Xu, Z. & Liu, S. Decoding consumer purchase decisions: exploring the predictive power of EEG features in online shopping environments using machine learning. Humanit. Soc. Sci. Commun. 11, 1202. https://doi.org/10.1057/s41599-024-03691-1 (2024).
A, A. Resting state EEG microstate profiling and a machine-learning based classifier model in epilepsy. Cogn. Neurodyn. 18, 2419–2432. https://doi.org/10.1007/s11571-024-10095-z (2024).
Rehab, N., Siwar, Y. & Mourad, Z. Machine learning for epilepsy: A comprehensive exploration of novel EEG and MRI techniques for seizure diagnosis. J. Med. Biol. Eng. 44, 317–336. https://doi.org/10.1007/s40846-024-00874-8 (2024).
Acknowledgements
This work was supported by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R393), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Funding
This work was supported by Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R393), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
Data analysis and code writing: RA with the help of CD, GS, SS. Data acquisition and preprocessing: SS, UA and SA. Writing, original draft preparation, RA and CD. Writing, review and editing: MO, LB. Funding acquisition: MO. Project administration: SA. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Human ethics and consent to participate declarations
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Agrawal, R., Dhule, C., Shukla, G. et al. Iterative multiblock framework for high frequency EEG based neurological disorder detection. Sci Rep 16, 5995 (2026). https://doi.org/10.1038/s41598-026-37126-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37126-5