Abstract
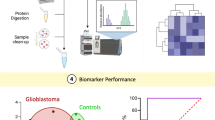
It is difficult to differentiate between glioma recurrence and radiation brain necrosis after radiotherapy in the clinic, but it is very important for patients. The purpose of this study is to observe the clinical significance of cell-free DNA (cfDNA) in the differential diagnosis between them based on liquid biopsy. The SD rats were used to establish a glioma recurrence model and a radiation brain necrosis model. The successful establishment of the two models was verified by MRI, IVFI, HE staining, immunofluorescence and immunohistochemistry. Then, the changes of cfDNA in the two models (B1-SINE in plasma and mtDNA in urine) were analyzed by qPCR. After ionizing particle irradiation, qPCR results showed that the level of mtDNA in urine increased significantly at the 6th week and reached the peak at the 8th week. The B1-SINE of plasma samples increased significantly at the 4th week of irradiation and reached the peak at the 8th week. From week 8 to week 16, when radiation necrosis occurred, the levels of mtDNA in urine and B1-SINE in plasma increased significantly (p < 0.01). In the glioma recurrence model, there was no significant difference in urinary mtDNA level and plasma B1-SINE level between the normal group and the sham surgery group (p > 0.05). These findings demonstrate that cfDNA liquid biopsy, represented by plasma B1-SINE and urinary mtDNA, holds significant clinical translational potential as an early differential diagnostic marker for radiation-induced brain necrosis and glioma recurrence, warranting further validation of its sensitivity and specificity through ROC analysis in clinical samples.
Similar content being viewed by others
Introduction
Glioma is the most common primary malignant brain tumor1. Gliomas are characterized by high incidence, high recurrence rate, high mortality rate, and low cure rate2. Despite the rapid development of medical technology in recent years, glioma remains one of the most lethal tumors in the world3,4. In order to prolong progression-free survival and overall survival after surgery and to improve the therapeutic effect of the disease, current treatment guidelines recommend the combined application of surgery and radiotherapy/chemotherapy5. Radiotherapy is an effective method to treat various intracranial tumors, which can improve the survival time of patients6. Currently, radiotherapy is widely used in conservative treatment, postoperative adjuvant therapy, and Gamma Knife therapy for patients with intracranial tumors7.
Radiation brain necrosis is one of the most serious complications in patients with intracranial tumors following radiation therapy8. Unfortunately, there is no effective way to distinguish radiation brain necrosis from central nervous system tumor recurrence, because both cases show similarly enhanced imaging results and can only be distinguished by clinical symptoms, long-term imaging follow-up, or surgical biopsy9. These methods also provide relatively one-sided information, which often leads to difficulties in differential diagnosis10. The conventional MRI examination revealed enhanced lesions with peripheral edema in both diseases, and the specificity of the enhancement mode and functional imaging for distinguishing between the two diseases was poor. Multi-modal image fusion may solve this problem in the future. At the same time, a small-sample biopsy may only capture necrotic tissue or miss tumor cells, and a biopsy of deep or functional lesions may cause a neurological deficit. The possibility of damage caused by radiotherapy limits the choice of radiotherapy to relatively conservative methods, but the control of glioma recurrence requires greater treatment intensity11. Therefore, it is very important to find a non-invasive and accurate method to distinguish radiation brain necrosis from tumor recurrence at the earliest stage.
Cell-free DNA (cfDNA) refers to DNA that is free of bodily fluids such as blood, urine, and saliva12. It can be derived from many sources, including the natural death of normal cells, tumor cells and necrotic tissues13. At present, cell-free DNA (cfDNA) has been widely used in the study of physiological and pathological diseases, including trauma, inflammatory diseases, radiation injury, malignant tumor detection and prenatal diagnosis, and its research on tumors has received special attention14,15,16. By analyzing the characteristics of cfDNA, we can obtain information for early diagnosis, prognosis evaluation, and monitoring of tumor recurrence17,18. While the progression of other tumor types has outpaced that of CNS malignancies, the level of cfDNA in gliomas has increased significantly19. This method not only provides dynamic information about the progression and response to treatment of intracranial diseases, but also provides information about the genetic characteristics of tumors, enabling the matching and optimization of diagnosis and treatment schemes according to the developing tumors20.
There are many types of cfDNA, including B1 repeat sequence (BRS) and mitochondrial DNA (mtDNA). The B1 family of short repeats is the most abundant DNA sequence in rodents, which corresponds to the ALU family in the human genome21. Hengshan et al. reported that the BRS in mice was significantly increased after exposure to gamma radiation22. Although the changes in cfDNA level of human ALU family in plasma of glioma patients have been reported, the changes in cfDNA level in plasma of radiation brain necrosis patients are still unclear23. Damaged or squeezed cells can release mtDNA, which has antigenicity and can stimulate the cGAS-STING pathway and induce inflammation24. Although mtDNA is not as easy to study as ctDNA, the mutation, deletion or copy number change of mtDNA is considered a potential tumor biomarker that is even more sensitive25. Some specific mtDNA mutations can help distinguish different types of cancer, such as lung cancer and colorectal cancer26,27. More importantly, some mtDNA and nuclear DNA in the blood can tolerate ribozyme degradation and pass into the urine28. This evidence makes urine an important type of liquid biopsy because it is easier to obtain than blood and is not damaged.
With the development of liquid biopsy technology, cfDNA in blood, urine and other body fluids has become a powerful tool for non-invasive tumor monitoring12. Although cerebrospinal fluid contains higher concentrations of biomarkers than plasma, plasma and urine are easier to obtain. Therefore, based on the detection of cfDNA in blood and urine, the value of differential diagnosis between glioma and radiation brain necrosis is worthy of further investigation29. In the previous study, we preliminarily analyzed the plasma cfDNA of glioma patients by high-throughput sequencing30. In this study, we first established a recurrent glioma model and a radiation-induced brain necrosis model. We then used liquid biopsy to monitor changes in cfDNA levels in these two rat models, including urinary mitochondrial DNA (mtDNA) and plasma B1 short interspersed nuclear elements (B1-SINE, BRS). Finally, to explore the application value of these two types of cfDNA liquid biopsy in the early stage of clinical application to distinguish radiation brain necrosis from glioma recurrence.
Methods and materials
Cell culture
The C6-luc glioma cell line was obtained from Nantong University (Nantong, China). The cell line was cultured at 37℃ under 5% CO₂ atmosphere in F12K medium (Gibco BRL; Grand Island, NY, USA) containing 12.5% horse serum and 2.5% fetal bovine serum (Gibco BRL).
Animals
We used 10-week-old Sprague-Dawley rats (weight range: 240–250 g) obtained from the Animal Experimental Center of Anhui Medical University. All animal procedures described herein were performed in accordance with the institutional animal care guidelines approved by the Ethics Committee of the First People’s Hospital of Chuzhou. All studies are reported in accordance with the ARRIVE guidelines. At the end of the experiments, rats were euthanized by an intraperitoneal injection of an overdose of sodium pentobarbital (150 mg/kg).
Glioma model and tumor recurrence
C6-GFP-LUC cells were washed three times with sterile PBS, then resuspended in serum-free F12K medium after centrifugation, and the cell concentration was adjusted to 1 × 105 cells/10µL and stored at 37℃ for later use. The experimental rats were anesthetized with isoflurane and fixed to the brain stereotaxic instrument. A hole was drilled at the skull location coordinates (0.5 mm posterior to the anterior fontanel and 2 mm lateral to the midline), and 10 µL of cell suspension was slowly injected into the right cerebral cortex (depth 3 mm) at a rate of 1 µL/min with a microsyringe, and the needle was left in place for 5 minutes after injection to prevent backflow. Ten days after cell implantation, the formation of intracranial tumor was evaluated by magnetic resonance imaging (MRI) and in vivo fluorescence imaging (IVFI). At this point, the tumors typically reached a diameter of approximately 3–4 mm, confirming successful model establishment. The tumor tissue was then accurately removed according to the imaging localization. After surgery, MRI and IVFI were regularly monitored, and combined with hematoxylin-eosin (HE) staining pathological analysis, tumor recurrence and histological characteristics were systematically evaluated.
Radiotherapy dose planning and irradiation
Prior to radiotherapy, all experimental rats were scanned by computed tomography (CT) (Discovery, GE Medical Systems, USA). The acquired image data were imported into the Eclipse treatment planning system (Varian Medical Systems, USA) for 3D reconstruction and dose calculation. A circular 15 mm diameter irradiation field was constructed using a multi-leaf collimator (MLC) to accurately cover the target area. Six mv high-energy X-ray (dose rate 600 MU/min) from a linear accelerator (LINAC) (Trilogy, Varian Medical Systems, USA) was used for unidirectional irradiation of the right hemisphere of rats. The total irradiation dose was set at 60 Gy (n = 5 in each group), and the machine jump number (MU) was monitored in real time during treatment to ensure dose accuracy (Table 1). The selected radiation dose can reliably induce late-stage brain necrosis with distinct histopathological features in rodents, providing a controlled and standardized platform for investigating necrosis mechanisms and identifying biomarkers31. All irradiation procedures were performed under strict quality control conditions, including daily output dose calibration.
Urine/plasma collection and DNA extraction
All urine samples were collected using a special metabolic cage (manufactured by Shanghai, China Yuyan Instrument Co., Ltd.), which can effectively separate urine from feces. In the process of sample collection, experimental rats were placed in metabolic cages at room temperature (20 1℃) before and after irradiation. To maintain the stability of the sample, the urine collection glass container was placed in an ice bath for about 4 h until a 3 mL urine sample was obtained. At the same time, whole blood samples were collected from each rat (weighing approximately 300 g) by saphenous vein puncture and placed in a polypropylene tube containing EDTA anticoagulant. All biological samples (urine and blood) were centrifuged at 3000 g for 10 min, and the supernatant was separated for subsequent experimental analysis. CfDNA in urine was extracted using the GenElute urine cell-free DNA purification mini kit (Sigma Company, St. Louis, Missouri, USA), and plasma DNA was extracted using the QIAamp MinElute viral DNA extraction kit (Qiagen Company, Dusseldorf, Germany). All procedures strictly followed the standard procedure provided by the manufacturer. Finally, the obtained DNA samples were immediately packaged and stored in the ultra-low temperature refrigerator at −80℃ for a long time to ensure the stability of nucleic acid to meet the needs of subsequent experiments.
qPCR for DNA quantification
Urine and plasma DNA samples were analyzed quantitatively by the QCPR method using a 7500 real-time qPCR system (Applied Biosystems, Abilene, TX, USA). Primers and amplification procedures were as described elsewhere22,32. The reaction conditions were 95℃ for 10 min, 95℃ for 10 s, 60℃ for 30 s, and 40 cycles. The upstream primer sequence of plasma-free DNA is 5’CCAGGACACCAGGGCTACAGAG3’, and the downstream primer sequence is 5’CCCGAGTGCTGGGATTAAAG3’. The upstream primer sequence of mtDNA is 5’ AATGGTTGGTTTGTTCAACGATT3’ and the downstream primer sequence is 5’ AGAAACCGACCTGGATTGCTC3’. The upstream primer sequence of GAPDH is 5’ TGGCCTCCAAGGAGTAAGAAAC3’ and the downstream primer sequence is 5’ GGCCTCTCTCTTGCTCTCAGTATC3’. The primers were synthesized by Sangon Biotechnology (Shanghai).
Statistical analysis
Statistical analyses were performed using GraphPad Prism software (version 9.0) and SPSS software (version 27.0). A p-value < 0.05 was considered statistically significant.
Results
Establishment of glioma and recurrent glioma models in rats
As shown in Figs. 1 and 10 days after implantation of C6 cells, a high signal intensity area was detected on T2-weighted MRI (Fig. 1a), and a strong fluorescence signal was obtained by IVFI (Fig. 1d), indicating that the C6 cells were growing well. The postoperative T2-weighted MRI image (Fig. 1b) and IVFI (Fig. 1e) showed that the glioma cells were almost removed. After 4 weeks of lesion resection, the MRI image (Fig. 1c) and IVFI (Fig. 1f) showed that the glioma had grown again.
Establishment of SD rat glioma model and the recurrent glioma model. (a) MRI images at 10 days after C6 cell implantation, (b) postoperative, and (c) 4 weeks after surgery. (d) In vivo fluorescence imaging at 10 days after C6 cell implantation, (e) postoperative, and (f) 4 weeks after surgery.
Simultaneously, the rat brain tissues at each time point were stained with HE for pathological analysis (Fig. 2a). Observation under light microscopy revealed obvious nuclear heteromorphism of the tissues after implantation of C6 cells in the rat cerebral cortex for 10 days, which is the pathological characteristic of a tumor. HE staining after resection of the tumor tissue could still detect residual tumor cells at the edge of the resected tissue. After 4 weeks of tumor resection, vascular proliferation, central necrosis, and nuclear dysplasia were observed in the brain tissues. In addition, immunofluorescence detection of GFAP (Fig. 2b) and immunohistochemical detection of ki67, PCNA, and Bcl-2 (Fig. 2c) further confirmed glioma recurrence.
The detection of SD rat glioma model and the recurrent glioma model. (a) The Hematoxylin-eosin (HE) staining at 10 days after implantation of C6 cells, postoperative, and 4 weeks after surgery. (b) immunofluorescence detection of GFAP, and (c) immunohistochemical detection of ki67, PCNA and Bcl-2.
Establishment of the radiation brain necrosis model
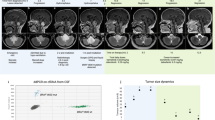
As shown in Fig. 3a, rats were irradiated and MRI scans were performed before irradiation and at 1, 4, 8, 12, and 16 weeks after irradiation. The T2-weighted image of the MRI scan (Fig. 3b) showed circular enhancement of the linear particle irradiation area at week 16, indicating the successful establishment of the radiation brain necrosis model. The pathological HE staining results shown in Fig. 4 showed that the cell edema and degeneration of rat brain tissue appeared in the early stages (1–4 weeks), after which the perivascular space increased, the nucleus condensed, and the local cells underwent gradual punctate necrosis. At the 16th week, extensive necrosis of brain tissue occurred in the targeted irradiation area, demonstrating the occurrence of radiation brain necrosis.
Establishment of SD rat radiation brain necrosis model. (a) Radiotherapy dose planning. (b) MRI scan image of radiation brain necrosis model before irradiation and at 1/4/8/12/16th week of irradiation. The arrow represents the lesion of radiation necrosis.
The HE staining of SD rat radiation brain necrosis model before irradiation and at 1/4/8/12/16th weeks of irradiation.
Changes in urinary MtDNA levels in rats with radiation brain necrosis and glioma recurrence
The level of free mtDNA in urine samples was determined by qPCR. The results (Fig. 5a) showed that the level of mtDNA changed significantly after radiotherapy (n = 5, F = 36.54, p < 0.001). Compared with the pre-treatment level, the urinary mtDNA level increased significantly after 24 h of irradiation, reaching 2.27 times the normal level (p < 0.0001), and then gradually returned to the normal level. At week 6, the urinary mtDNA level again increased significantly, reaching 1.43 times the normal level (p < 0.001), and finally peaked at week 8 (2.21 times the normal level; p < 0.001). From week 8 to the time of radiolabeled necrotic foci on imaging (between weeks 8 and 16), urine mtDNA levels increased with statistical significance at p < 0.01. In the recurrent glioma model, qPCR results (Fig. 5b) showed no significant change in urinary free mtDNA levels compared to the normal and sham surgery groups (n = 5, F = 1.970, p = 0.138).
Changes in the urine and plasma cfDNA levels in SD rat with radiation brain necrosis and glioma recurrence. (a) The changes in the urine mtDNA level in rats with radiation brain necrosis and (b) glioma recurrence. (c) The changes in the plasma B1-SINE level in rats with radiation brain necrosis and (d) glioma recurrence. Y axis in 5c is logarithmic scale (log10. ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Changes in the plasma BRS level in rats with radiation brain necrosis and glioma recurrence
After linear particle irradiation, plasma cell-free BRS levels were measured at each time point (1, 2, 4, 8, 12, and 16 weeks). The results of qPCR (Fig. 5c) showed that the plasma B1-SINE level changed significantly before and after radiotherapy (n = 5, F = 142.6, p < 0.001). The B1-SINE level of plasma free cells increased significantly at week 4 (p < 0.0001) and reached its peak at week 8 (10.61 times the normal level, p < 0.0001). Subsequently, the plasma B1-SINE level increased during 8–16 weeks (p < 0.01), and this increase was significantly higher than that in the control group (p < 0.01). In the recurrent glioma model, qPCR results (Fig. 5d) showed no significant change in plasma BRS levels compared to the normal and sham surgery groups (n = 5, F = 2.650, p > 0.05).
Discussion
Liquid biopsy mainly detects circulating tumor cells, circulating tumor DNA (ctDNA) or exosomes released from the primary tumor site or metastatic site into peripheral blood or other body fluids to achieve the purpose of diagnosing or monitoring tumors33. Liquid biopsy technology has solved the major shortcomings of traditional tissue biopsy technology and attracted much attention for its advantages of non-invasive repeated sampling, early diagnosis, real-time dynamic monitoring and overcoming tumor heterogeneity34,35. Cell-free DNA (cfDNA) is an important research direction and hot spot in liquid biopsy of various malignant tumors, including glioma36. CfDNA is a DNA molecule that can be shed into the blood by cells under both physiological and pathological conditions. The length of cfDNA is between 50 and 300 base pairs, which can be rapidly eliminated by phagocytosis, so it is usually present in low concentrations in the blood of healthy individuals37. A large number of studies have shown that changes in the amount and structure of circulating cfDNA in plasma or serum can be used as biomarkers for the prognosis and diagnosis of various human diseases38. It is also extremely important to study the response of cfDNA to ionizing radiation as a potential biomarker in animal and human body fluids20. Currently, cfNDA is still in the research and exploration stage in clinical practice for distinguishing glioma recurrence from radiation necrosis of the brain. A unified threshold or clinical standard has not yet been established.
Studies have shown that repetitive sequences make up a significant proportion of mammalian genomes (e.g., more than half of human and mouse genomes), suggesting that repetitive sequences have an important influence on the higher structure of chromatin39. BRS is the most abundant short repetitive sequence in rodents, present in all cells21. The activity and expression of BRS are closely related to genomic instability, which plays a key role in the formation of higher chromatin structure and the regulation of gene expression. Tumor cells usually show higher genomic instability, and the mutation, amplification or loss of BRS may be related to the occurrence and development of tumors, which can be used as a potential biomarker for early diagnosis of tumors. Including DNA involved in apoptosis, can enter the plasma, so BRS can represent the total level of free DNA in vivo. In addition, the coagulation process can release DNA from blood cells. To improve the specificity of the test results, we used plasma instead of serum in this study.
Mutation, deletion or copy number change of mtDNA is considered as a potential biomarker of tumor40. Compared with circulating free nuclear DNA, circulating free mtDNA has the characteristics of small genome, many copies and high mutation frequency, and has unique advantages in tumor diagnosis and treatment, showing good application prospects41,42. The study in the mouse glioblastoma model of Cambridge University found that the sensitivity of mtDNA in plasma was better than that of ctDNA in the early stage, and mtDNA could be detected in cerebrospinal fluid and urine25. Urine is commonly used for non-invasive diagnosis and contains a variety of metabolites that are helpful in finding specific biomarkers for various diseases43. Cell-free mtDNA fragments can be degraded by nucleases, but some of these components can be used as transrenal DNA to reach urine44. Therefore, the purpose of this study is to investigate the changes in urinary mtDNA level and plasma BRS level of linear particle irradiated rats.
In the clinical setting, delayed radiation necrosis is slow, occurring several months or even decades after radiation therapy, making it difficult to diagnose45. Because different diagnoses have different treatment schemes, patients with radiation brain necrosis do not need surgery. Therefore, in most cases, the true pathological diagnosis cannot be made46. Radiation-induced brain injury is usually accompanied by an increase in oxidative stress, and mitochondria are an important regulatory center of intracellular oxidative stress. In this study, radiation brain necrosis model and glioma recurrence model were first established. Then, the change of cfDNA level in rats was evaluated by measuring the level of free mtDNA in urine and B1-SINE (BRS) in plasma.
The application of BRS in tumor is mainly reflected in the aspects of early diagnosis marker, monitoring tumor progression and evaluation of treatment effect. The study of mtDNA in tumor reveals its important role in metabolism, genetic stability and cell function. This study found that cfDNA (mtDNA and B1-SINE), which changed significantly with time, was detected in both plasma and urine in the radiation brain necrosis model. However, in the glioma recurrence model, no obvious changes in the content of two types of cfDNA were observed. The reason may be that the spread of cfDNA is limited by the existence of the blood-brain barrier, and the amount of cfDNA released by inflammation caused by surgical injury and tumor recurrence is relatively small47. Studies have shown that the level of cfDNA in cerebrospinal fluid (CSF) is much higher than that in plasma or urine, and the difference can reach several orders of magnitude48,49. However, compared to blood and urine, CSF collection is more harmful and riskier to the patient, so it is not the first choice.
In the radiation brain necrosis model, the acute peak and delayed onset of urinary mtDNA levels may indicate that the integrity of the mitochondrial membrane is compromised shortly after irradiation, resulting in release of mtDNA into the cytoplasm or body fluids. When delayed radiation necrosis occurs, a large number of cells die, resulting in the second peak of mtDNA. Urine free DNA can be affected by nuclease and the renal barrier, making it less sensitive to plasma cfDNA levels. At the same time, it is also possible that the B1-SINE level in blood is rich, which leads to the release of a large number of B1-SINE when cells rupture, thus affecting the increase in cell-free B1-SINE level. Consistent with the results of this study, previous studies have found that free mitochondrial DNA in urine significantly increases within three days after x-ray irradiation in rats, and the concentration increases with increasing radiation intensity32.
In human samples, cfDNA in the serum of glioma patients was sequenced in the previous research, and the results showed that it was not a perfect marker for diagnosing glioma and healthy individuals30. However, some researchers have found that cfDNA in the urine of glioma patients is apparently more fragmented than that of healthy individuals, suggesting that it has diagnostic potential50. Studies of radiotherapy for glioma have shown that the level of cfDNA before radiotherapy is an independent risk factor for prognosis20. Although the results of many studies are not completely consistent and the types of cfDNA involved are different, liquid biopsy based on cfDNA has great potential to distinguish radiation brain necrosis from glioma recurrence. Current research also shows that cfDNA from radiation brain necrosis and glioma recurrence apparently have different characteristics and mechanisms. Recurrent glioma cells destroy the blood-brain barrier through invasive growth and release a variety of cfDNA with cell apoptosis, necrosis and active secretion. These cfDNAs not only contain specific mutations of primary tumors (such as IDH, TERT, EGFR, etc.), but also have some special epigenetic features, such as methylation29,51,52. CfDNA associated with radiation brain necrosis lacks specific tumor gene variation and may carry radiation-induced genomic damage markers, such as increased microsatellite instability and deletion/amplification patterns of specific chromosomal regions. Due to the small sample size and the variety of mutation detection methods used in most studies of cfDNA in glioma, the results have been inconsistent, which also presents a challenge for follow-up research.
In this study, at the designated endpoint (4 weeks post-surgery), the recurrent glioma model showed no significant elevation in plasma BRS or urinary mtDNA. This finding cannot be attributed to insufficient tumor burden but is more likely related to limitations in the detection time window and differences in cfDNA release dynamics. First, in compliance with animal ethics guidelines, the experiment was terminated once the tumor reached a predetermined volume. At this time point, the process of cfDNA release from the tumor, passage across a potentially incompletely permeable blood–brain barrier (BBB), and accumulation in the peripheral circulation to detectable levels may not yet have reached a plateau. Second, unlike the acute, fulminant cell death and DNA release triggered by radiation necrosis, tumor-derived cfDNA originates predominantly from the relatively slow process of apoptosis, resulting in a lower release rate and a more diffuse signal that can be easily masked by background fluctuations in single-time-point measurements. Finally, differences in BBB permeability to DNA fragments of varying sizes may explain the modest upward trend observed for small-fragment mtDNA in the tumor group. Together, these factors suggest that a detection strategy based on total cfDNA concentration at a single time point may have limited sensitivity for monitoring early or progressing brain tumor recurrence. Future studies should be designed to incorporate synchronous multi-point sampling (liquid biopsy combined with tissue biopsy) to establish direct correlations between cfDNA levels and quantitative histopathological indicators, and to integrate tumor-specific markers such as mutant ctDNA or methylation signatures, in order to capture recurrence signals with higher specificity and sensitivity.
The detection of molecular biomarkers in body fluids is of great importance for the non-invasive diagnosis of various diseases, which requires active exploration and in-depth research53. As a non-invasive biomarker, cfDNA has made remarkable progress in recent years in early diagnosis, monitoring recurrence and predicting curative effect of glioma36,54. The application of B1-SINE in tumor is mainly reflected in the aspects of early diagnosis marker, monitoring tumor progression and evaluating treatment effect. The study of mtDNA in tumor reveals its important role in metabolism, genetic stability and cell function. The focus of this study is to observe whether cfDNA, as a potential adjunctive diagnosis, can guide long-term monitoring of blood and urine samples after radiotherapy to evaluate the toxicity of radiotherapy. Although the results of this study suggest that B1-SINE and mtDNA can be used as differential diagnostic markers for radiation-induced brain necrosis and glioma recurrence, there are still some shortcomings in this method.
First, the acute necrosis induced by the single high‑dose model may differ pathophysiologically from the delayed necrosis observed after clinical fractionated radiotherapy. Future studies employing fractionated irradiation regimens would better simulate the clinical scenario. Second, urinary mtDNA is also a sensitive biomarker of acute kidney injury. Systemic responses in irradiated rats, such as renal function, require systematic assessment to more accurately confirm the brain‑specific origin of urinary mtDNA. Third, the sample size in this exploratory study was relatively small for each group. Although post‑hoc power analysis indicated sufficient power to detect large effect sizes, the limited sample size may reduce our ability to identify smaller cfDNA changes in the glioma recurrence group, increasing the risk of a Type II error. Validation in larger animal cohorts is therefore needed. Fourth, the clinical application of cfDNA requires further exploration. Beyond absolute concentration, characteristics such as cfDNA methylation status and fragment size distribution (fragmentomics) have also been shown to hold important value in tumor differential diagnosis. Fifth, although mouse‑related BRS (B1‑SINE) was successfully amplified in rat samples, rat‑specific sequences warrant further investigation. Future studies integrating multi‑dimensional cfDNA analyses are expected to further improve the accuracy of distinguishing radiation necrosis from tumor recurrence50. Furthermore, the C6 glioma model used here is an allogeneic transplant (C6 cells derived from Wistar rats into SD rat hosts), resulting in major histocompatibility complex (MHC) mismatch. This may elicit an immune rejection response against the implanted tumor in the host. Immune‑mediated tumor cell killing is a significant source of circulating cfDNA. Therefore, the cfDNA detected in this model may partly reflect contributions from the immune response, adding an additional layer of complexity to the interpretation of cfDNA signal differences between the tumor and necrosis groups. Future studies using immunodeficient mice or syngeneic tumor models would help to more purely dissect tumor‑derived cfDNA signatures.
It is very necessary and useful to test the blood and urine samples of patients with radiation necrosis and glioma recurrence, but it is necessary to establish standardized procedures for sample collection, separation, and analysis to ensure the accuracy of the results47. In general, cfDNA analysis provides a new molecular dimension for differentiating radiation brain necrosis from glioma recurrence, and its multiparameter characteristics can compensate for the shortcomings of traditional diagnostic methods. The accuracy of distinguishing radiation brain necrosis from glioma recurrence can be improved based on different cfDNA, combined with expression level, mutation spectrum, and inflammatory factor analysis. Although there are still technical challenges, with the improvement of detection sensitivity and the progress of bioinformatics analysis, cfDNA is expected to become an important part of accurate diagnosis and treatment of neural tumors. Therefore, in the future research, how to use the existing science and technology to achieve efficient and accurate detection and simultaneously detect a variety of cfDNA to improve the identification efficiency is an important research direction12,55.
Conclusions
In short, we successfully established two rat models (recurrent glioma and radiation brain necrosis) and found that two types of cfDNA (B1-SINE in plasma and mtDNA in urine) showed no significant changes in the glioma recurrence model but exhibited substantial changes in the radiation brain necrosis model. These findings provide new insights and evidence to differentiate radiation brain necrosis from glioma recurrence and develop individualized treatment plans. They also reveal that liquid biopsies based on cfDNA have great potential as markers for the differential diagnosis of these two diseases and warrant further study in clinical samples.
Data availability
All data supporting the findings of this study are available within the paper.
Abbreviations
- MRI:
-
magnetic resonance imaging
- IVFI:
-
in vivo fluorescence imaging
- HE:
-
hematoxylin and eosin
- BRS:
-
B1 repeat sequence
- qPCR:
-
quantitative real-time polymerase chain reaction
- cfDNA:
-
cell-free DNA
- mtDNA:
-
mitochondrial DNA
References
Wang, L. M., Englander, Z. K., Miller, M. L. & Bruce, J. N. Malignant glioma. Adv. Exp. Med. Biol. 1405, 1–30 (2023).
Hou, S., Chen, Y., Jin, C. & Lin, N. Integrative analysis of bulk RNA-seq and scRNA-seq data indicates the prognostic and Immunologic values of SERPINH1 in glioma. Environ. Toxicol. 39 (6), 3654–3665 (2024).
Hou, S. et al. Prognostic value of hematologic Prealbumin/Fibrinogen ratio in patients with glioma. World Neurosurg. 160, e442–e453 (2022).
Weller, M. et al. Glioma. Nat. Rev. Dis. Primers. 10 (1), 33 (2024).
Berger, T. R., Wen, P. Y., Lang-Orsini, M. & Chukwueke, U. N. World health organization 2021 classification of central nervous system tumors and implications for therapy for Adult-Type gliomas: A review. JAMA Oncol. 8 (10), 1493–1501 (2022).
Gritsch, S., Batchelor, T. T. & Gonzalez Castro, L. N. Diagnostic, therapeutic, and prognostic implications of the 2021 world health organization classification of tumors of the central nervous system. Cancer 128 (1), 47–58 (2022).
Zhao, M. et al. PGC1alpha degradation suppresses mitochondrial biogenesis to confer radiation resistance in glioma. Cancer Res. 83 (7), 1094–1110 (2023).
Mayo, Z. S., Billena, C., Suh, J. H., Lo, S. S. & Chao, S. T. The dilemma of radiation necrosis from diagnosis to treatment in the management of brain metastases. Neuro Oncol. 26 (12 Suppl 2), S56–S65 (2024).
Devan, S. P. et al. Towards differentiation of brain tumor from radiation necrosis using multi-parametric MRI: preliminary results at 4.7 T using rodent models. Magn. Reson. Imaging. 94, 144–150 (2022).
Aseel, A., McCarthy, P. & Mohammed, A. Brain magnetic resonance spectroscopy to differentiate recurrent neoplasm from radiation necrosis: A systematic review and meta-analysis. J. Neuroimaging. 33 (2), 189–201 (2023).
Bourbonne, V. et al. Diagnosis and management of brain radiation necrosis. Cancer Radiother. 28 (6–7), 547–552 (2024).
Zhang, K., Fu, R., Liu, R. & Su, Z. Circulating cell-free DNA-based multi-cancer early detection. Trends Cancer. 10 (2), 161–174 (2024).
De Borre, M. et al. Cell-free DNA methylome analysis for early preeclampsia prediction. Nat. Med. 29 (9), 2206–2215 (2023).
Mathios, D. et al. Detection and characterization of lung cancer using cell-free DNA fragmentomes. Nat. Commun. 12 (1), 5060 (2021).
Martin-Alonso, C. et al. Priming agents transiently reduce the clearance of cell-free DNA to improve liquid biopsies. Science 383 (6680), eadf2341 (2024).
Gaitsch, H., Franklin, R. J. M. & Reich, D. S. Cell-free DNA-based liquid biopsies in neurology. Brain 146 (5), 1758–1774 (2023).
Bao, H. et al. Letter to the editor: an ultra-sensitive assay using cell-free DNA fragmentomics for multi-cancer early detection. Mol. Cancer. 21 (1), 129 (2022).
Wadden, J., Ravi, K., John, V., Babila, C. M. & Koschmann, C. Cell-Free tumor DNA (cf-tDNA) liquid biopsy: current methods and use in brain tumor immunotherapy. Front. Immunol. 13, 882452 (2022).
Johnson, K. C. & Verhaak, R. G. W. Serum cell-free DNA epigenetic biomarkers aid glioma diagnostics and monitoring. Neuro Oncol. 23 (9), 1423–1424 (2021).
Husain, A. et al. Dynamics of cell-free DNA in predicting response in adult diffuse glioma on chemoradiotherapy. Cancer Genet. 268–269, 55–63 (2022).
Krayev, A. S. et al. The nucleotide sequence of the ubiquitous repetitive DNA sequence B1 complementary to the most abundant class of mouse fold-back RNA. Nucleic Acids Res. 8 (6), 1201–1215 (1980).
Zhang, H. et al. B1 sequence-based real-time quantitative PCR: a sensitive method for direct measurement of mouse plasma DNA levels after gamma irradiation. Int. J. Radiat. Oncol. Biol. Phys. 74 (5), 1592–1599 (2009).
Chen, J. et al. Detection of serum Alu element hypomethylation for the diagnosis and prognosis of glioma. J. Mol. Neurosci. 50 (2), 368–375 (2013).
Hu, M. et al. ATM Inhibition enhances cancer immunotherapy by promoting MtDNA leakage and cGAS/STING activation. J Clin. Invest 131 (3), e139333 (2021).
Mair, R. et al. Measurement of plasma Cell-Free mitochondrial tumor DNA improves detection of glioblastoma in Patient-Derived orthotopic xenograft models. Cancer Res. 79 (1), 220–230 (2019).
Bulgakova, O. et al. The level of free-circulating MtDNA in patients with radon-induced lung cancer. Environ. Res. 207, 112215 (2022).
Xu, Y. et al. Quantitative detection of Circulating MT-ND1 as a potential biomarker for colorectal cancer. Bosn J. Basic. Med. Sci. 21 (5), 577–586 (2021).
Umansky, S. R. & Tomei, L. D. Transrenal DNA testing: progress and perspectives. Expert Rev. Mol. Diagn. 6 (2), 153–163 (2006).
Miller, A. M. et al. Tracking tumour evolution in glioma through liquid biopsies of cerebrospinal fluid. Nature 565 (7741), 654–658 (2019).
Sun, J. et al. Examination of plasma Cell-Free DNA of glioma patients by whole exome sequencing. World Neurosurg. 125, e424–e428 (2019).
Zhao, Z. F., Yang, L. Z., Jiang, C. L., Zheng, Y. R. & Zhang, J. W. Gamma knife irradiation-induced histopathological changes in the trigeminal nerves of rhesus monkeys. J. Neurosurg. 113 (1), 39–44 (2010).
Abdullaev, S. A., Minkabirova, G. M., Bezlepkin, V. G. & Gaziev, A. I. Cell-free DNA in the urine of rats exposed to ionizing radiation. Radiat. Environ. Biophys. 54 (3), 297–304 (2015).
Ho, H. Y., Chung, K. K., Kan, C. M. & Wong, S. C. Liquid biopsy in the clinical management of cancers. Int J. Mol. Sci 25 (16), 8594 (2024).
Pessoa, L. S., Heringer, M. & Ferrer, V. P. CtDNA as a cancer biomarker: A broad overview. Crit. Rev. Oncol. Hematol. 155, 103109 (2020).
Nikanjam, M., Kato, S. & Kurzrock, R. Liquid biopsy: current technology and clinical applications. J. Hematol. Oncol. 15 (1), 131 (2022).
Birko, Z., Nagy, B., Klekner, A. & Virga, J. Novel molecular markers in Glioblastoma-Benefits of liquid biopsy. Int J. Mol. Sci 21 (20), 7522 (2020).
Qi, T. et al. Cell-Free DNA fragmentomics: the novel promising biomarker. Int J. Mol. Sci 24 (2), 1503 (2023).
Hackshaw, A., Clarke, C. A. & Hartman, A. R. New genomic technologies for multi-cancer early detection: rethinking the scope of cancer screening. Cancer Cell. 40 (2), 109–113 (2022).
Li, S. & Shen, X. Long interspersed nuclear element 1 and B1/Alu repeats blueprint genome compartmentalization. Curr. Opin. Genet. Dev. 80, 102049 (2023).
Kopinski, P. K., Singh, L. N., Zhang, S., Lott, M. T. & Wallace, D. C. Mitochondrial DNA variation and cancer. Nat. Rev. Cancer. 21 (7), 431–445 (2021).
Peng, F. et al. Circulating cell-free MtDNA as a new biomarker for cancer detection and management. Cancer Biol. Med. 21 (2), 105–110 (2023).
Liu, Y. et al. Aberrant fragmentomic features of Circulating cell-free mitochondrial DNA as novel biomarkers for multi-cancer detection. EMBO Mol. Med. 16 (12), 3169–3183 (2024).
Chen, C. K., Liao, J., Li, M. S. & Khoo, B. L. Urine biopsy technologies: cancer and beyond. Theranostics 10 (17), 7872–7888 (2020).
Husain, H. et al. Monitoring daily dynamics of early tumor response to targeted therapy by detecting Circulating tumor DNA in urine. Clin. Cancer Res. 23 (16), 4716–4723 (2017).
Ninatti, G., Pini, C., Gelardi, F., Sollini, M. & Chiti, A. The role of PET imaging in the differential diagnosis between radiation necrosis and recurrent disease in irradiated Adult-Type diffuse gliomas: A systematic review. Cancers (Basel) 15 (2), 364 (2023).
Ohno, M. et al. Assessment of therapeutic outcome and role of reirradiation in patients with radiation-induced glioma. Radiat. Oncol. 17 (1), 85 (2022).
Yekula, A. et al. Liquid biopsy strategies to distinguish progression from pseudoprogression and radiation necrosis in glioblastomas. Adv. Biosyst. 4 (12), e2000029 (2020).
Pages, M. et al. Liquid biopsy detection of genomic alterations in pediatric brain tumors from cell-free DNA in peripheral blood, CSF, and urine. Neuro Oncol. 24 (8), 1352–1363 (2022).
Izquierdo, E. et al. Droplet digital PCR-based detection of Circulating tumor DNA from pediatric high grade and diffuse midline glioma patients. Neurooncol Adv. 3 (1), vdab013 (2021).
Mouliere, F. et al. Fragmentation patterns and personalized sequencing of cell-free DNA in urine and plasma of glioma patients. EMBO Mol. Med. 13 (8), e12881 (2021).
Piccioni, D. E. et al. Analysis of cell-free Circulating tumor DNA in 419 patients with glioblastoma and other primary brain tumors. CNS Oncol. 8 (2), CNS34 (2019).
Azad, T. D., Jin, M. C., Bernhardt, L. J. & Bettegowda, C. Liquid biopsy for pediatric diffuse midline glioma: a review of Circulating tumor DNA and cerebrospinal fluid tumor DNA. Neurosurg. Focus. 48 (1), E9 (2020).
Cescon, D. W., Bratman, S. V., Chan, S. M. & Siu, L. L. Circulating tumor DNA and liquid biopsy in oncology. Nat. Cancer. 1 (3), 276–290 (2020).
Patel, J. et al. Liquid biopsy in H3K27M diffuse midline glioma. Neuro Oncol. 26 (Supplement_2), S101–S109 (2024).
Liu, Y. et al. Optimized PCR-Based enrichment improves coverage uniformity and mutation detection in mitochondrial DNA Next-Generation sequencing. J. Mol. Diagn. 22 (4), 503–512 (2020).
Funding
This work was supported by Health Research Program of Anhui (AHWJ2023A30052), Chuzhou Science and Technology Program (2023ZD032).
Author information
Authors and Affiliations
Contributions
Conception: Shiqiang Hou and Yinan Chen; Experiments and analysis of data: Junlong Sun and Chunjing Jin; Preparation of the manuscript: Junlong Sun and Chunjing Jin; Revision for important intellectual content: Yinan Chen; Supervision: Shiqiang Hou.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
This study was approved by the Ethics Committee of the First People’s Hospital of Chuzhou.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sun, J., Jin, C., Chen, Y. et al. Liquid biopsy of plasma and urinary CfDNA differentiates glioma recurrence from radiation brain necrosis in preclinical models. Sci Rep 16, 7123 (2026). https://doi.org/10.1038/s41598-026-37493-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-37493-z