Abstract
Chromosome image classification typically relies on ImageNet pre-training, yet the potential of leveraging intermediate domains from related staining techniques remains largely underexplored. Here, we evaluate two-step transfer learning–where classifiers are first fine-tuned on an intermediate domain before targeting the final classification task–across Q-band (BioImLab dataset) and G-band (CIR dataset) chromosome classification. Each dataset serves as intermediate domain for the other. Across 11 architecture families and three training approaches, models achieve improvements when domain similarity is high and data quality is limited: modern architectures (ConvNeXt, Swin Transformer, ViT, MobileNetV3) show + 0.8 to + 3.3 percentage point gains in Macro-F1 on Q-band classification, while traditional CNNs benefit less or show no improvement. On the higher-quality G-band dataset, all architectures approach performance saturation, with minimal gains from two-step transfer (+ 0.1 to + 0.7 percentage points). Consistent results across both transfer directions demonstrate that, with appropriate architecture selection and intermediate domain similarity, two-step transfer learning can boost performance when target datasets are challenging, while ImageNet pre-training alone suffices for high-quality data. The code is publicly available at https://github.com/MuscleOne/chromosome_TL.
Similar content being viewed by others
Introduction
Automated chromosome image classification plays a critical role in cytogenetic diagnostics, enabling the identification of chromosomal abnormalities associated with genetic disorders and cancer1,2,3. Human karyotyping involves classifying 46 chromosomes (22 pairs of autosomes numbered 1-22 by decreasing size, plus sex chromosomes X and Y) into 24 classes based on size, morphological features, and banding patterns obtained via staining techniques such as Q-banding and G-banding (Fig. 1)1. Automated karyotyping systems have become promising tools for replacing time-consuming manual operations4, yet training deep learning classifiers for these specialized medical imaging tasks faces inherent data scarcity.
A G-banding chromosomes karyotype from a human male. Banding patterns refer to the dark and light patterns along the medial axis of the individual chromosome.
Transfer learning with ImageNet pre-training has become standard practice in medical imaging analysis tasks5,6,7, including chromosome image classification8,9,10,11,12. ImageNet-pretrained architectures have been observed to bring benefits to downstream tasks with limited labeled data by transferring knowledge learned from large-scale datasets13,14,15. However, the significant domain distance between general natural images and specialized chromosome images raises questions about transfer effectiveness, and the private datasets used in many studies make it difficult to reproduce results and systematically compare training approaches.
The use of intermediate domains in transfer learning has been explored in various medical imaging contexts7,16,17,18, yet systematic investigation in chromosome classification remains lacking. This approach, known as two-step transfer learning, involves sequential fine-tuning on two datasets: first on an intermediate domain closer to the target domain, then on the target domain itself. Ray et al.7 demonstrated that domain-specific pre-trained weights from related histopathology staining techniques (H&E and IHC) provided performance boosts over ImageNet weights. Alzubaidi et al.18 used general skin images as an intermediate domain for diabetic foot ulcer classification. These studies suggest that intermediate domains, which act as additional information sources beyond ImageNet, can refine decision boundaries when appropriately selected. In chromosome karyotyping, multiple staining techniques (e.g., Q-banding and G-banding) produce morphologically similar but visually distinct images, presenting a unique opportunity: when and why do intermediate domains from related staining techniques improve chromosome classification beyond direct ImageNet transfer?
In this work, we evaluate the utility of intermediate domains in chromosome classification by comparing three training approaches: training from scratch, ImageNet pre-training, and two-step transfer learning. We test two-step transfer learning on Q-band classification (BioImLab dataset19) and G-band classification (CIR dataset10), where each dataset serves as intermediate domain for the other. We evaluate 11 architecture families spanning traditional CNNs (ResNet-34/50/101, ResNeXt-50/101, VGG-16/19) and modern architectures (ConvNeXt-Tiny, Swin-T, ViT-S/16, MobileNetV3-L) across two classification tasks: Q-band classification (BioImLab dataset, relatively challenging) and G-band classification (CIR dataset, higher quality). Our findings show that two-step transfer learning provides substantial benefits for modern architectures on challenging datasets, while all methods approach saturation on high-quality datasets. To enable reproducibility, we employ two open datasets and make our source code and scripts available, allowing other researchers to validate and potentially build upon our findings using larger datasets.
Results
We report findings from 66 classifiers (11 architectures spanning classical CNNs to modern attention-based models \(\times\) 3 training approaches \(\times\) 2 classification tasks) evaluated via 5-fold cross-validation. For each classifier configuration, we train five independent models (one per fold) and report performance as the mean and standard deviation across the five held-out validation folds. Macro-F1 score, which incorporates per-class precision and recall (see “Materials and methods”, Eqs. 1–4), serves as the primary metric for comparing training approaches. Results are organized into four subsections: domain similarity analysis, quantitative performance comparison, architecture-dependent patterns, and qualitative case studies.
Domain similarity motivates intermediate domain selection
To assess whether Q-band and G-band datasets are suitable intermediate domains for each other, we computed Maximum Mean Discrepancy (MMD) distances between feature embeddings extracted from ImageNet-pretrained ConvNeXt-Tiny architectures. MMD is a kernel-based metric suitable for comparing high-dimensional feature distributions20,21, requiring no density estimation or distributional assumptions (see “Materials and methods”). Table 1 presents the pairwise MMD\(^{2}\) values with 95% confidence intervals.
The results show that chromosome images from different staining techniques (G-band and Q-band) share greater similarity with each other (MMD\(^{2}\) = 0.148) than either has with natural images from ImageNet (MMD\(^{2}\) = 0.310 and 0.264, respectively). Despite different staining techniques, the CIR and BioImLab datasets are substantially closer to each other than either is to ImageNet, supporting their use as intermediate domains for each other in two-step transfer learning.
Two-step transfer learning improves performance for modern architectures on challenging datasets
To evaluate the utility of intermediate domains, we trained 66 classifiers (11 architectures \(\times\) 3 training approaches \(\times\) 2 classification tasks) using 5-fold cross-validation. Table 2 presents the comprehensive performance comparison across Macro-F1, Accuracy, and AUC-OvR metrics.
Training from scratch reveals architecture-dependent failure modes. Modern architectures (ConvNeXt-Tiny, Swin-T, ViT-S/16) completely fail to converge on these small-scale datasets, with Macro-F1 scores particularly low on G-band classification (31–76%). In contrast, traditional CNNs (ResNet/ResNeXt series) still achieve 90–95% performance, while VGG architectures reach 80–89% when trained from scratch. Pre-training, whether using ImageNet pre-training or two-step transfer learning, is essential for modern architectures and elevates their performance to 93–98%.
Comparing pre-training approaches. On G-band classification (higher-quality dataset), two-step transfer learning achieves comparable or slightly better performance than ImageNet pre-training across ResNet families and modern architectures (+0.1 to +0.7 percentage points in Macro-F1), while showing degradation on ResNeXt-50 and VGG-16/19. Notably, several modern architectures (ConvNeXt-Tiny, Swin-T, MobileNetV3-L) achieve perfectly consistent performance across all five validation folds (zero standard deviation) with Macro-F1 reaching 97–98%, suggesting that these models have reached a performance ceiling on this dataset, leaving minimal room for further improvement from alternative pre-training strategies. On Q-band classification (relatively lower-quality dataset), traditional architectures show no gain or slight degradation with two-step transfer learning compared to ImageNet pre-training. However, modern architectures consistently outperform ImageNet pre-training when using two-step transfer learning, with gains of +0.8 to +3.3 percentage points in Macro-F1.
Architecture-dependent benefit patterns reflect task difficulty and architectural design
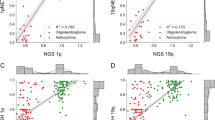
To understand the factors underlying the observed performance differences, we examined the relationship between model complexity and transfer learning gains. Table 3 lists the parameter counts and computational costs (GFLOPs) for all 11 architectures evaluated in this study. Figure 2 shows the change in Macro-F1 scores (two-step transfer learning compared to ImageNet pre-training) plotted against model parameters. On G-band classification, most architectures show minimal changes (within ± 1 percentage point), with some traditional architectures showing slight degradation. On Q-band classification, modern architectures show larger positive gains (+0.8 to +3.3 percentage points), while traditional architectures show minimal or negative changes.
Change in Macro-F1 score (percentage points) from two-step transfer learning compared to ImageNet pre-training, plotted against model complexity (log\(_{10}\)(#parameters)) for (a) G-band and (b) Q-band classification tasks. Error bars indicate bootstrap 95% confidence intervals (see “Materials and methods”, Statistical analysis).
The results reveal no simple linear relationship between model capacity and the benefit from two-step transfer learning. MobileNetV3-L, which has the fewest parameters among modern architectures (4.18M), achieves the largest gain on Q-band classification (+3.3 percentage points), whereas parameter-heavy models such as VGG-16 (134.37M parameters) show degradation on both tasks (-2.5 and -0.5 percentage points on G-band and Q-band, respectively).
Qualitative case studies reveal refined attention patterns
To provide qualitative evidence for the quantitative findings, we examined attention patterns using Gradient-weighted Class Activation Mapping (Grad-CAM)22 across the three training approaches. Figure 3 presents three representative cases: MobileNetV3-L on Q-band classification (exhibiting the largest performance gain from two-step transfer learning), ViT-S/16 on G-band classification (where two-step transfer learning recovers meaningful feature localization despite from-scratch training failing to converge), and VGG-16 on G-band classification (where two-step transfer learning shows degradation).
Grad-CAM visualizations for three representative architectures under the three training approaches (from-scratch, ImageNet pre-training, and two-step transfer learning). For each sample, the leftmost image shows the original chromosome, followed by Grad-CAMs corresponding to the three training approaches.
For MobileNetV3-L on Q-band classification, the from-scratch classifier focuses primarily on chromosome edges, capturing only shape information. ImageNet pre-training shifts attention toward banding regions on the long arm but remains insensitive to the short arm. Two-step transfer learning achieves comprehensive coverage of both long and short arms, with attention systematically distributed along chromosome structures. For ViT-S/16 on G-band classification, the from-scratch classifier fails to converge, producing uninformative attention maps despite high activation intensity. ImageNet pre-training introduces severe artifacts with unstable attention patterns. Two-step transfer learning concentrates attention on both arms while reducing spurious activations. For VGG-16 on G-band classification, ImageNet pre-training performs best, with attention covering both long and short arms. However, both two-step transfer learning and from-scratch classifiers fail to attend to the short arm.
To understand the relationship between attention distribution and misclassification, we examined three failure cases on G-band classification (Fig. 4). Case (a) shows a ConvNeXt-Tiny misclassification on an image with visible cropping artifacts, likely resulting from incomplete extraction of overlapping chromosomes during preprocessing. The Grad-CAM for the predicted label concentrates on edges or erroneous regions, while attention under the true label remains diffuse. Cases (b) and (c) present VGG-16 failures, where Grad-CAMs for predicted labels show unfocused attention spreading across chromosomes, and even true labels fail to elicit attention toward discriminative banding regions.
Grad-CAM visualizations for three misclassified chromosome samples. Each row shows one misclassification case: the left column is the input image, the middle column shows Grad-CAM for the predicted (incorrect) label, and the right column for the true label.
Discussion
The use of intermediate domains in transfer learning for medical imaging has been underexplored despite the availability of related datasets from diverse imaging modalities and staining techniques. This work shows that two-step transfer learning can improve chromosome image classification by leveraging intermediate domains from related staining techniques. We demonstrate that classifiers achieve performance gains when intermediate domain similarity is high and target data quality is limited. Our experimental design across Q-band (BioImLab) and G-band (CIR) datasets reveals that modern architectures benefit most from two-step transfer learning on the more challenging Q-band dataset, while all architectures approach saturation on the higher-quality G-band dataset. The first objective of this work was to evaluate whether intermediate domains from related staining techniques provide performance gains over direct ImageNet transfer. The smaller MMD distance between Q-band and G-band datasets compared to either dataset’s distance from ImageNet supports the premise that these datasets are suitable candidates for two-step transfer learning. The second objective focused on identifying architecture-dependent responses to intermediate domain knowledge. Our findings reveal that the benefit depends on task difficulty and architectural design: modern architectures with attention mechanisms exploit intermediate domain knowledge more effectively than traditional CNNs, particularly when target data is challenging. The lack of correlation between model parameters and performance gain is exemplified by lightweight MobileNetV3-L achieving the largest benefit, whereas parameter-heavy VGG-16 shows degradation. This contrast confirms that architectural design determines the effectiveness of intermediate domain transfer.
Grad-CAM visualizations provide evidence for these architecture-dependent patterns. In cases where two-step transfer learning provides performance gains (e.g., MobileNetV3-L on Q-band), the attention becomes more concentrated on chromosome-specific banding patterns compared to ImageNet pre-training. Conversely, in cases where two-step transfer learning provides no benefit or degradation (e.g., VGG-16 on G-band), attention patterns show minimal differences across training approaches. In misclassification cases, visual defects such as cropping artifacts disrupt attention patterns regardless of training approach.
The moderately sized datasets reflect realistic constraints in medical imaging, where acquiring labeled chromosome images is labor-intensive and requires expert annotation. Data quality issues observed in misclassification cases, such as cropping artifacts from incomplete extraction of overlapping chromosomes, highlight inherent challenges in real-world chromosome image preprocessing.
These findings advance our understanding of transfer learning in medical imaging and reveal a general principle applicable across scientific domains: when target data is scarce or challenging, related but distinct information sources can refine decision boundaries and improve generalization. This principle extends to any domain where labeled data is costly but related data sources are available. The success of intermediate domain learning suggests that medical imaging researchers can leverage related datasets to complement ImageNet pre-training. The inclusion of intermediate domains refines regions where target data is limited, providing a more comprehensive understanding of domain-specific features. For automated karyotyping system deployment, our results demonstrate that deep learning classifiers can achieve 93-98% Macro-F1 using moderately sized datasets (2,986-5,474 images) with appropriate transfer learning strategies, while misclassification analysis reveals that preprocessing quality (cropping artifacts from overlapping chromosomes) represents a fundamental constraint that cannot be overcome by training strategies alone, establishing realistic performance expectations and quality requirements for clinical applications.
Conclusion
This study demonstrates that two-step transfer learning using intermediate domains from related staining techniques can improve chromosome image classification, with benefits depending on architecture choice and target data characteristics. Modern architectures achieve +0.8 to +3.3 percentage point gains in Macro-F1 on the challenging Q-band dataset, while traditional CNNs and high-quality datasets show minimal improvement. These findings suggest that practitioners should consider intermediate domain transfer when working with limited or challenging medical imaging data, particularly when using modern architectures. Future work may explore optimal intermediate domain selection criteria and extend this approach to other biomedical imaging modalities.
Materials and methods
Dataset
The BioImLab dataset19 includes 5,474 Q-band chromosome images from 119 normal human karyotypes, captured under fluorescence microscopy at the University of Padova, Italy. The CIR dataset10 contains 2,986 G-band chromosome images from 65 normal human karyotypes, captured under optical microscopy at Guangdong Women and Children Hospital, China. Both datasets contain 24 chromosome classes (autosomes 1-22 and sex chromosomes X and Y), with naturally balanced class distributions reflecting typical human karyotypes (approximately two instances per class per metaphase spread, except sex chromosomes which appear once per cell).
Due to differences in dye affinities and staining characteristics between Q-banding and G-banding techniques, the two datasets exhibit complementary banding patterns23. Specifically, AT-rich regions appear as bright fluorescent bands in Q-banding but as dark bands in G-banding, while GC-rich regions show the opposite pattern. BioImLab images exhibit lower resolution and less pronounced banding patterns compared to CIR images. This complementary relationship makes the two datasets biologically related yet visually distinct, providing a suitable test case for evaluating intermediate domain transfer learning.
We use stratified 5-fold cross-validation on both datasets. Since each metaphase spread contains chromosomes from a complete cell, the class distribution is naturally balanced across metaphase spreads. Stratified splitting ensures that this biological balance is preserved across training and validation folds, reflecting the realistic class proportions that would be encountered in clinical karyotyping. All images are resized to 256 \(\times\) 256 pixels by first padding the shorter side with black pixels (value 0) to achieve square dimensions, then applying bilinear interpolation. Data augmentation (random horizontal/vertical flips and Gaussian blurring) is applied only to training sets, while validation sets remain unaugmented.
Domain similarity analysis
We selected Maximum Mean Discrepancy (MMD)20 for domain similarity analysis because it offers several advantages for comparing high-dimensional feature distributions: (1) it operates directly on sample embeddings without requiring density estimation, unlike Kullback-Leibler divergence which can be unreliable in high-dimensional spaces; (2) as a kernel-based metric, it captures complex distributional differences beyond first-order statistics; and (3) it has well-established theoretical properties with known convergence rates.
We used an unbiased MMD estimator with a Gaussian RBF kernel, where the bandwidth parameter was selected via the median heuristic. Feature embeddings were extracted from the final pooling layer (before the classification head) of an ImageNet-pretrained ConvNeXt-Tiny architecture. For each dataset pair (ImageNette2-16024, BioImLab, CIR), MMD distances were computed using all available samples, with 95% confidence intervals estimated via bootstrap resampling (1,000 iterations). Domain similarity rankings were consistent when using CORAL (CORrelation ALignment) distance25 as an alternative metric (see Supplementary Information). Complete mathematical formulations are provided in Supplementary Information.
Classification task
Our classification task involves two distinct tasks: Q-band classification and G-band classification. Each task is a 24-class classification problem, and cross-entropy is employed as the loss function. This study uses the BioImLab dataset for Q-band classification and the CIR dataset for G-band classification. Classifiers are trained and validated for each classification task under a unified 5-fold cross-validation setting.
Training approaches and architectures
Three training approaches are evaluated for both Q-band and G-band classification tasks: training from scratch, ImageNet pre-training, and two-step transfer learning.
-
Training from scratch: Classifiers are initialized with random weights and trained directly on the target dataset (BioImLab for Q-band or CIR for G-band).
-
ImageNet pre-training: Classifiers are initialized with ImageNet pre-trained weights and fine-tuned on the target dataset.
-
Two-step transfer learning: Classifiers are first fine-tuned on the intermediate domain dataset starting from ImageNet weights, then fine-tuned on the target domain dataset.
Table 4 summarizes the training approaches for both classification tasks.
We evaluate 11 architectures spanning traditional CNNs and modern architectures. Traditional CNNs include VGG-16 and VGG-1926, ResNet-34, ResNet-50, and ResNet-10127, and ResNeXt-50 and ResNeXt-10128. Modern architectures include ConvNeXt-Tiny29, Swin Transformer (Swin-T)30, Vision Transformer (ViT-S/16)31, and MobileNetV3-Large32. This selection spans three dimensions: architectural era (2015–2022), design paradigm (convolutional, attention-based, and efficient architectures), and computational budget (4M to 192M parameters; Table 3), ensuring generalizability across architectures commonly used in medical image analysis. In total, we evaluate 66 classifier configurations (11 architectures \(\times\) 3 training approaches \(\times\) 2 classification tasks).
Training procedure
Classifier training and inference are conducted on a computer equipped with an Intel Xeon W-2245 CPU (16x 4.7GHz) and an NVIDIA RTX A6000 GPU (48GB VRAM), running Ubuntu 22.04 OS with 64GB RAM. Our implementation primarily employs the PyTorch framework for both training and evaluation. For each classifier, we fine-tune all parameters on the training set. The ImageNet pre-training weights are sourced from public checkpoints implemented in PyTorch. For the training from scratch approach, we randomly initialize the parameters of each neural network layer.
All classifiers are trained using cross-entropy loss with architecture-specific hyperparameters optimized for each model family. Training generally employs a two-stage optimization strategy combining Adam and SGD optimizers, with total training duration of 80 epochs. Complete details on batch sizes, learning rates, weight decay, and optimization schedules for each architecture are provided in Supplementary Information.
For each architecture-training approach combination, we train five independent classifiers corresponding to the five cross-validation folds. During training, we monitor accuracy at the end of each epoch and save the checkpoint with the highest validation accuracy for each fold. Each trained classifier is then evaluated on its corresponding held-out validation fold. Reported performance metrics represent the mean and standard deviation across the five validation fold results, where the standard deviation reflects fold-to-fold variability in the dataset rather than classifier instability.
Performance metrics
We evaluate classification performance using three metrics: Macro-F1 score, accuracy, and AUC-OvR (Area Under the Curve for One-vs-Rest). For each architecture-training approach combination, these metrics are computed on each of the five held-out validation folds, and we report the mean and standard deviation across the five folds.
Macro-F1 score. For each class j, precision measures the proportion of correct predictions among all samples predicted as class j:
where \(TP_j\) is the number of true positives for class j, and \(FP_j\) is the number of false positives (samples incorrectly predicted as class j). Recall measures the proportion of class j samples that are correctly identified:
where \(FN_j\) is the number of false negatives (class j samples incorrectly predicted as other classes).
The F1 score for each class j is the harmonic mean of its precision and recall, balancing both metrics:
The Macro-F1 score then averages these per-class F1 scores across all 24 chromosome classes:
where \(N=24\).
Accuracy measures the overall proportion of correctly classified samples:
where POP is the total number of samples in the validation set.
AUC-OvR (Area Under the ROC Curve for One-vs-Rest). For each class j, we compute the Area Under the Receiver Operating Characteristic Curve (AUC) by treating it as the positive class and all remaining 23 classes as the negative class:
where TPR (True Positive Rate) and FPR (False Positive Rate) are computed across all classification thresholds. The AUC-OvR is then averaged across all classes:
Statistical analysis
To compare the effectiveness of two-step transfer learning against ImageNet pre-training, we compute paired differences in Macro-F1 scores across the five cross-validation folds for each architecture-task combination. For each pair of classifiers trained with the two approaches on the same fold, we calculate the difference: \(\text {Macro-F1}_{\text {two-step}} - \text {Macro-F1}_{\text {ImageNet}}\).
For each architecture-task combination with n paired observations (one per fold), we estimate the mean difference and 95% confidence intervals using bootstrap resampling with 10,000 iterations. The bootstrap approach draws n samples with replacement from the observed differences, computes the mean for each resample, and derives confidence intervals from the 2.5th and 97.5th percentiles of the bootstrap distribution. We report architecture-task combinations with five paired folds to ensure statistical reliability.
Class activation map visualization
We generate class activation maps (CAMs) using the Gradient-weighted Class Activation Mapping (Grad-CAM) method22. Grad-CAM computes importance weights for each feature map in a target convolutional layer based on the gradient of the class score with respect to the feature maps. These weights are used to create a weighted combination of the feature maps, followed by ReLU activation to retain only positive contributions. The resulting activation map is normalized to the range [0, 1] and upsampled to the input image resolution using bilinear interpolation. We overlay this map onto the original chromosome image using a heatmap colormap (jet).
For correctly classified samples, we generate CAMs using the ground-truth class. For misclassified samples, we generate two CAMs: one using the predicted (incorrect) label and another using the true label.
Data availability
Datasets analyzed in this study are publicly available: the BioImlab dataset is accessible at https://www.kaggle.com/datasets/arifmpthesis/bioimlab-chromosome-data-set-for-classification, and the CIR dataset can be found at https://github.com/CloudDataLab/CIR-Net.
Code availability
The code and experimental results supporting this study are publicly available at https://github.com/MuscleOne/chromosome_TL. The repository includes training scripts, evaluation code, and model configurations for all classifier configurations evaluated in this work. There are no restrictions on code access or reuse.
References
Bickmore, W. A. Karyotype analysis and chromosome banding. Encycl. Life Sci. (2001).
Roizen, N. J. & Patterson, D. Down’s syndrome. Lancet 361, 1281–1289 (2003).
Arber, D. A. et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood J. Am. Soc. Hematol. 127, 2391–2405 (2016).
Munot, M. V. Development of computerized systems for automated chromosome analysis: Current status and future prospects. Int. J. Adv. Res. Comput. Sci. 9, 782–791 (2018).
Rahman, T. et al. Transfer learning with deep convolutional neural network (CNN) for pneumonia detection using chest X-ray. Appl. Sci. 10, 3233 (2020).
Alzubaidi, L. et al. Novel transfer learning approach for medical imaging with limited labeled data. Cancers 13, 1590 (2021).
Ray, I., Raipuria, G. & Singhal, N. Rethinking imageNet pre-training for computational histopathology. In 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC) 3059–3062 (IEEE, 2022).
Qin, Y. et al. Varifocal-net: A chromosome classification approach using deep convolutional networks. IEEE Trans. Med. Imaging 38, 2569–2581 (2019).
Xiao, L. et al. DeepACEv2: Automated chromosome enumeration in metaphase cell images using deep convolutional neural networks. IEEE Trans. Med. Imaging 39, 3920–3932 (2020).
Lin, C. et al. Cir-net: Automatic classification of human chromosome based on inception-resnet architecture. IEEE ACM Trans. Comput. Biol. Bioinform. 19, 1285–1293 (2020).
Lin, C. et al. ChromosomeNet: A massive dataset enabling benchmarking and building basedlines of clinical chromosome classification. Comput. Biol. Chem. 100, 107731 (2022).
Lin, C. et al. Mixnet: A better promising approach for chromosome classification based on aggregated residual architecture. In 2020 International Conference on Computer Vision, Image and Deep Learning (CVIDL) 313–318 (IEEE, 2020).
Kornblith, S., Shlens, J. & Le, Q. V. Do better imagenet models transfer better? In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition 2661–2671 (2019).
Raghu, M., Zhang, C., Kleinberg, J. & Bengio, S. Transfusion: Understanding transfer learning for medical imaging. Adv. Neural Inf. Process. Syst. 32 (2019).
Ke, A., Ellsworth, W., Banerjee, O., Ng, A. Y. & Rajpurkar, P. CheXtransfer: performance and parameter efficiency of ImageNet models for chest X-Ray interpretation. In Proceedings of the Conference on Health, Inference, and Learning 116–124 (2021).
Lopez-Tiro, F. et al. Boosting kidney stone identification in endoscopic images using two-step transfer learning. In Mexican International Conference on Artificial Intelligence 131–141 (Springer, 2023).
Lopez-Tiro, F. et al. Improving automatic endoscopic stone recognition using a multi-view fusion approach enhanced with two-step transfer learning. In Proceedings of the IEEE/CVF International Conference on Computer Vision 4165–4172 (2023).
Alzubaidi, L. et al. Towards a better understanding of transfer learning for medical imaging: A case study. Appl. Sci. 10, 4523 (2020).
Poletti, E., Grisan, E. & Ruggeri, A. Automatic classification of chromosomes in q-band images. In 2008 30th Annual International Conference of the IEEE Engineering in Medicine and Biology Society 1911–1914 (IEEE, 2008).
Gretton, A., Borgwardt, K. M., Rasch, M. J., Schölkopf, B. & Smola, A. A kernel two-sample test. J. Mach. Learn. Res. 13, 723–773 (2012).
Long, M., Cao, Y., Wang, J. & Jordan, M. Learning transferable features with deep adaptation networks. In Proceedings of the 32nd International Conference on Machine Learning, Vol. 37 of Proceedings of Machine Learning Research (eds. Bach, F. & Blei, D.) 97–105 (PMLR, 2015).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Int. J. Comput. Vis. 128, 336–359. https://doi.org/10.1007/s11263-019-01228-7 (2020).
Schreck, R. R. & Distèche, C. M. Chromosome banding techniques. Curr. Protoc. Hum. Genet. (1994).
Howard, J. Imagenette: A smaller subset of 10 easily classified classes from imagenet (2019).
Sun, B. & Saenko, K. Deep coral: Correlation alignment for deep domain adaptation. In Computer Vision – ECCV 2016 Workshops 443–450 (Springer International Publishing, 2016). https://doi.org/10.1007/978-3-319-49409-8_35.
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 25 (2012).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 770–778 (2016).
Xie, S., Girshick, R., Dollár, P., Tu, Z. & He, K. Aggregated residual transformations for deep neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition 1492–1500 (2017).
Liu, Z. et al. A convnet for the 2020s. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 11966–11976. https://doi.org/10.1109/CVPR52688.2022.01167 (2022).
Liu, Z. et al. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV) 10012–10022. https://doi.org/10.1109/ICCV48922.2021.00986 (2021).
Dosovitskiy, A. et al. An image is worth 16 \(\times\) 16 words: Transformers for image recognition at scale. In International Conference on Learning Representations. https://doi.org/10.48550/arXiv.2010.11929 (2021).
Howard, A. et al. Searching for MobileNetV3. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV) 1314–1324. https://doi.org/10.1109/ICCV.2019.00140 (2019).
Acknowledgements
The authors appreciate Mr. Xiaolu Yan at China Agricultural University, Beijing, for his critical revision of this study.
Funding
This work is funded by Macao Polytechnic University (Grant No. RP/FCA-14/2023), with the permission number s/c fca.e0fc.544d.1 from Macao Polytechnic University.
Author information
Authors and Affiliations
Contributions
T.C., C.X., W.Z., and T.L., X.H. contributed to the conception. T.C., W.K., T.L., and K.L. designed the study and the structure of the manuscript. T.C., C.X., and Y.L. implemented the experiments. T.C. analysed the results and drafted the manuscript. All authors contributed to the critical revision of the manuscript and approved the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study protocol was reviewed and approved by the Institutional Review Board (IRB) (Approval ID: MPU-FCA-202312041677), and the informed written consent from the participants was waived due to the use of deidentified data from publicly available databases.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, T., Xie, C., Zhang, W. et al. ImageNet pre-training and two-step transfer learning in chromosome image classification. Sci Rep 16, 7572 (2026). https://doi.org/10.1038/s41598-026-38662-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-38662-w