Abstract
Physalis peruviana L. (Cape gooseberry) is susceptible to several pathogens, among which the sobemovirus physalis rugose mosaic virus (PhyRMV) is most prominent. This study investigated the anatomical, morphophysiological, and metabolic responses of P. peruviana to PhyRMV over the course of the infection cycle. P. peruviana plants were grown under greenhouse conditions (24 ± 2 ℃) and mechanically inoculated with buffer (mock) or PhyRMV inoculum. The local and systemic leaves were collected at 0, 3, 7, 14, 21, and 42 days after inoculation (DAI) and viral infection was confirmed by RT-qPCR. Morphological traits, including plant height, symptoms, and the Falker chlorophyll index, and histological changes were evaluated in PhyRMV-infected plants and compared to mock-inoculated controls. To characterize and compare their metabolic profile, the collected leaves were processed and subjected to gas chromatography-mass spectrometry (GC–MS). Yield was also assessed by quantifying the number and weight of fruits. PhyRMV could be detected in systemic leaves from 14 DAI onward, accompanied by an increase in viral load and the intensification of the symptoms, including mosaic, chlorosis, and leaf deformation from 21 DAI onward, which were associated with reduced plant height and chlorophyll contents. At 42 DAI, histological changes and increased starch accumulation were observed in the infected leaves, suggesting impaired photoassimilate transport. GC-MS revealed an accumulation of sucrose, pyruvate, and tricarboxylic acid (TCA) cycle metabolites, possibly due to the high energy demand associated with viral replication, indicating significant alterations in the functioning of the central metabolic pathways in systemic leaves in the late stage of infection at 42 DAI. Compounds such as glutamate, isoleucine, and malonate could potentially be involved in the activation of defense pathways, including the shikimic acid pathway. Metabolic networks demonstrated a loss of complexity in the infected plants, indicating a redirection of resources from primary metabolism to defense-associated mechanisms. Thus, fruit production was reduced by 31% in the infected plants, highlighting a negative impact of infection on the production potential. These results support management strategies based on the induction of plant defense mechanisms and reinforce the importance of the use of integrative approaches to understanding the physiological and biochemical impacts of viruses on crops.
Similar content being viewed by others
Introduction
Physalis peruviana L. (Cape gooseberry) is a fruit-bearing plant belonging to Solanaceae, native to the Andean region of South America whose cultivation has expanded in tropical and subtropical regions, including Brazil1. In the country, production is concentrated in the states of São Paulo, Santa Catarina, and Rio Grande do Sul, with increasing participation of small and medium growers2,3,4,5.
Despite its remarkable economic potential, the P. peruviana plant faces exacerbating phytosanitary problems, as it is susceptible to various pathogens, including viruses. Some of these viruses have already been identified in other commercial crops in Brazil; notably orthotospovirus tomato chlorotic spot virus (TCSV; Orthotospovirus tomatoflavi, family Tospoviridae)6 and groundnut ringspot virus (GRSV; Orthotospovirus arachianuli, family Tospoviridae)7 and, more recently, the Sobemovirus physalis rugose mosaic virus (PhyRMV; Sobemovirus PHYRMV, family Solemoviridae)2. A study has also identified mixed infections of two potyviruses (family Potyviridae), comprising potato virus Y (PVY; Potyvirus yituberosi) and pepper yellow mosaic virus (PepYMV; Potyvirus capsiflavi), and PhyRMV in Physalis8.
PhyRMV significantly damages the P. peruviana plant, causing significant reductions in plant height, leaf area, and fruit productivity9. Infected plants produce smaller fruits with reduced diameter and altered quality parameters, such as titratable acidity and Brix content, resulting in yield losses of up to up to 70%9,10. Although PhyRMV has been detected in seed samples, bioassays indicate that it is not seed-transmitted11. Nevertheless, the virus is easily transmitted mechanically, including contaminated pruning tools, leaf-to-leaf contact, and soil or soil extracts5. Though not yet reported, the presence of a biological vector cannot be ruled out.
Currently, there are no effective strategies for controlling PhyRMV infection, and management practices rely mainly on preventive measures, such as the use of virus-free plant material and sterile tools during cultivation. Therefore, exploring new management strategies is essential. Understanding the physiological, molecular, and metabolic responses of P. peruviana to PhyRMV infection, as well as the mechanisms underlying the virus–plant interaction, is crucial to provide insights that may support the development of more effective and sustainable control approaches. Plant defense mechanisms are complex and fundamental to their survival when under pathogen infection. When plants are infected, a series of signaling cascades is activated that interact with each other and initiate a metabolic program for the host defense12,13,14. These signals are coordinated with those from hormones and generate a defense response at the local and systemic levels. This enables the plant to defend itself against the pathogen at the site of infection initially and then systemically. A viral infection is considered successful when viruses produce virulence factors that are unaffected by the defense mechanisms of the plants15. Additionally, primary and secondary metabolites play essential roles in the defense process. They have specific functions that protect against biotic stress-causing factors and form an integrated and coordinated network that enables plants to respond appropriately to external threats16.
Primary metabolites play a fundamental role in plant survival as they are directly involved in vital processes such as growth, development, and reproduction17. These compounds include glucose, sucrose, starch, proteins, amino acids, lipids, and nucleic acids, which provide energy and structural support to the cell. In contrast, secondary metabolites, although not directly essential for survival, play a crucial role in the environmental adaptation of plants18. They are involved in growth, plant–plant communication, and, above all, defense against pathogens and herbivores19,20. These compounds included alkaloids, terpenoids, jasmonic acid and phenolic compounds (including flavonoids, salicylic acid, simple phenols, and tannins), the last being prominent due to its important role in the response of plants to pathogens21. The interaction between primary and secondary metabolites not only ensures plant development, but also their survival under different environmental conditions and when exposed to various pathogens.
Metabolomic approaches based on gas chromatography–mass spectrometry (GC–MS) enable comprehensive profiling of metabolites such as organic acids, sugars, amino acids, and phenolic compounds, which are closely associated with plant defense mechanisms. Changes in metabolic profiles following viral infection can negatively affect plant anatomy and morphophysiology. Therefore, this study aimed to evaluate the effects of PhyRMV infection on the anatomical, morphophysiological, and metabolic responses of P. peruviana. The results may contribute to the identification of metabolic biomarkers associated with viral infection and support biotechnological strategies, including genetic engineering, to enhance viral tolerance and promote more sustainable cultivation of this species.
Results
Symptoms and virus detection
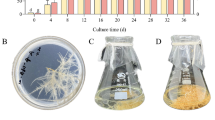
The severity of symptoms gradually increased in PhyRMV-inoculated P. peruviana plants throughout the course of infection cycle. At 42 days after inoculation (DAI), typical leaf symptoms such as mosaic, roughness, and deformation could be observed (Fig. 1B) in comparison to healthy (mock-inoculated) control plant (Fig. 1A). In addition, morphological changes were also detected. At 42 DAI, the infected plants showed more lateral branching than healthy ones. RT-qPCR identified the onset of infection by detecting an increase in PhyRMV copy number in local leaves 3 DAI compared to healthy plants and 0 DAI (Fig. 1C). A significant increase in viral copy number was also observed throughout the infection cycle up to 21 DAI (Fig. 1C). Infection of systemic leaves was observed from 14 DAI, with a gradual, marked increase up to 42 DAI (Fig. 1D).
Symptom assessment and quantification of the PhyRMV load. (A) Health Physalis peruviana plants (mock-inoculation) 42 days after inoculation (DAI). (B) P. peruviana plants 42 DAI with PhyRMV; symptoms of blistering, leaf deformation, and mosaic. (C) The smaller graph represents the number of viral copies detected in the local leaves of the healthy (− PhyRMV) and infected (+ PhyRMV) plants at 00, 03, 07, and 14 DAI; the larger graph presents the number of viral copies detected in the local leaves of healthy (− PhyRMV) and infected (+ PhyRMV) plants over 00, 03, 07, 14, and 21 DAI. (D) The smaller graph represents the number of viral copies detected in the systemic leaves of healthy (− PhyRMV) and infected (+ PhyRMV) plants at 00, 03, 07, 14, and 21 DAI; the larger graph presents the number of viral copies detected in the local leaves of healthy (− PhyRMV) and infected (+ PhyRMV) plants at 00, 03, 07, 14, 21, and 42 DAI. *Significant differences from the controls at P ≤ 0.05 assessed by the t-test (n = 3 biological replicates).
Anatomical characterization of the leaves of P. peruviana plants infected by PhyRMV
In general, toluidine blue-stained sections of local leaves (mock-inoculated) and systemic leaves (young healthy) of control plants showed a epidermal cell layer on their adaxial and abaxial surfaces, respectively (Fig. 2). In addition, the leaves bore tector trichomes (Fig. 2D); while stomata predominated the abaxial surface, they were only sporadic on the adaxial surface (Fig. 2A). The mesophyll consisted of a single layer of palisade parenchyma cells (elongated and juxtaposed) and 3–4 layers of lacunar parenchymatous cells with obvious intercellular spaces (Fig. 2A–D). Collateral vascular bundles were mainly observed in healthy leaves (Fig. 2A–D).
Cross sections of the local and systemic leaves of healthy (− PhyRMV–mock-inoculation) and infected (+ PhyRMV) plants stained with toluidine blue and Lugol’s iodine at 42 days after inoculation. (A, B) The blade of a healthy local leaf stained with toluidine blue. (C) The blade of a healthy local leaf pictured with Lugol’s iodine, (D, E) The blade of a healthy systemic leaf stained with toluidine blue, (F) The blade of a healthy systemic leaf stained with Lugol’s iodine, (G, H) The blade of a local leaf from a PhyRMV-infected plant stained with toluidine blue, (I) The blade of a local leaf from an infected plant stained with Lugol’s iodine, (J, K) The blade of a systemic leaf from a PhyRMV-infected plant stained with toluidine blue, (L) The blade of a systemic leaf from an infected plant was stained with Lugol’s iodine.
In the inoculated local leaves, the epidermal cells reduced in size and collapsed (Fig. 2G), while the chlorenchyma cells (palisade and lacuna cells) collapsed with enlarged intercellular spaces and tissue disorganization (Fig. 2G,H) and increased size of the nuclei (Fig. 2H). In addition, structural alterations were also observed in the vascular bundle cells in leaves locally-inoculated with PhyRMV (Fig. 2G,H), which could affect the photoassimilate flux. Changes could be observed in the mesophyll cell structure of infected systemic leaves only to a lesser extent (Fig. 2J,K). In this case, the chlorophyll parenchyma cells, particularly the palisade parenchyma, demonstrated hyperplasia (Fig. 2J,K). Compared with those from healthy plants, the cross sections of infected leaves, stained with Lugol’s reagent, demonstrated an increase in the number of starch granules (Fig. 2I,L).
Effects of PhyRMV infection on the productivity of plants
The negative impacts of PhyRMV infection manifested as a significant reduction in plant height after 21 and 42 DAI (Fig. 3A). One characteristic symptom of PhyRMV-infected leaves was a mosaic in which areas with a normal green color alternated with chlorotic ones. To assess the decline in chlorophyll levels in these tissues, the total chlorophyll indices of the healthy and PhyRMV-infected plants at 21 and 42 DAI were compared. The results revealed that the virus adversely impacted the total chlorophyll index (Fig. 3B). PhyRMV-infected plants showed delayed flowering and bud opening compared to the healthy plants (at 42 DAI, data not shown). A reduction in the number of fruits was visually evident in the infected plants; a remarkable 31% at 100 DAI, indicating potential crop damage (Fig. 3C,D).
(A) Average height of the healthy (-PhyRMV–mock-inoculation) and infected (+ PhyRMV) plants at 0, 21, and 42 days after inoculation (DAI). (B) The total Falker Chlorophyll Index of the local and systemic leaves of healthy (− PhyRMV–mock-inoculation) and infected (+ PhyRMV) plants at 21 DAI. (C) Number of fruits from virus-free (− PhyRMV) and infected (+ PhyRMV) plants. (D) The morphology and appearance of the fruits collected from the healthy (− PhyRMV) and infected (+ PhyRMV) plants at 100 DAI. *Significant differences from controls at P ≤ 0.05 determined by employing the t-test (n = 3 biological replicates).
Primary leaf metabolism in the PhyRMV-infected plants over the course of infection cycle
Considering the effects of PhyRMV infection in P. peruviana, as demonstrated in the previous sections, and as a strategy to further understand how PhyRMV influences host physiological and biochemical processes, we analyzed the metabolic profiles of local and systemic leaves from plants inoculated with buffer or PhyRMV at 3, 7, 14, 21, and 42 DAI. Based on KEEG (Kyoto Encyclopedia of Genes and Genomes) ID, retention time, and mass fragmentation pattern, 49 differentially expressed metabolites (DEMs) were identified, which included amino acids, organic acids, carbohydrates, and others (Table S1). PhyRMV infection induced significant changes (p < 0.05) in the levels of several metabolites. In the local and systemic leaves, 8 and 11 metabolites demonstrated temporal variations, respectively (Table S1A). However, no significant changes were detected in the mock- vs. PhyRMV-infected local leaves (Table S1B). In total, 15 DEMs (p < 0.05) were identified in the local and systemic leaves of infected and non-infected plants: five amino acids, three organic acids, four citric acid cycle intermediates, three sugars, and their derivatives (Table S1). Most of these metabolites did not show any significant differences (ns) between the local and systemic leaves. However, the levels of certain compounds, such as citric acid, fructose, glutarate, glutaric acid, glycine, isocitrate, malate, putrescine, salicylic acid, shikimate, and sorbitol, were altered remarkably (p < 0.05), indicating a temporal modulation of the metabolic response (Table S1).
A multivariate principal component analysis (PCA) of the normalized GC/MS data was performed to assess the metabolic variations between the local and systemic leaves of plants inoculated with PhyRMV at different time points. The PCA diagrams showed that the PCA scores in the local leaves of the healthy (mock) and PhyRMV-infected plants overlapped (up to 21 DAI) without forming fully distinct clusters (Fig. 4). The samples were divided into two principal component groups, PC1 and PC2/PC4, at all times (Fig. 4). Although the treatment values overlapped, PC1 explained the largest proportion of total data variation at all time points analyzed: 3 (47.9%), 7 (54.1%), 14 (49.4%), and 21 (52.4%) DAI. PC4 explained the maximal contributions at 3 DAI (10.4%); PC2 at 7 (22.3%), 14 (33.5%), and 21 (18.5%) DAI (Fig. 4).
Principal component analysis (PCA) of raw GC–MS data was performed for metabolite profiling. Raw metabolite levels were normalized to ribitol internal standard and sample fresh weight. PCA was conducted separately for local leaves (upper panel) and systemic leaves (lower panel) collected from healthy (red) and PhyRMV-infected (green) plants at 3, 7, 14, 21, and 42 days after inoculation (DAI). The percentages of variation explained by PC1 and PC2 are indicated on the corresponding axes. Analyses were carried out using the MetaboAnalyst platform (n = 3 biological replicates).
The PCA diagrams of systemic leaves revealed that the PCA scores of healthy (mock) and PhyRMV-infected plants differed at day 42 DAI (Fig. 4). The samples collected at all time points were divided into two main components, PC1 and PC2 (Fig. 4). At all points, PC1 explained the largest proportion of total data variation, at 3 (58.8%), 7 (58.5%), 14 (56.3%), 21 (49.4%), and 42 (51.3%) DAI. It was followed by PC2, which contributed at 3 (16.3%), 7 (20.5%), 14 (16.3%), 21 (20%), and 42 (27.3%) DAI (Fig. 4). The primary metabolites responsible for PCA cleavage in the systemic leaves at 42 DAI were isocitrate, malate, and glutamate in the infected plants, in contrast to ribose, xylose, fructose, glycolate, galactonic acid, and maltose in the healthy ones (Figure S1).
PhyRMV infection triggered conspicuous changes in the primary metabolism, affecting several metabolic pathways crucial for growth and stress responses. Among these, the most significantly affected were glycolysis, the citric acid cycle, and those involved in the biosynthesis of amino acids such as alanine, valine, serine, and glycine (Fig. 5). The temporal redistribution of metabolites suggested that the plants adapted their metabolism as the infection progressed, either to limit damage or in response to the exploitation of energy resources (Fig. 5). At 42 DAI, sucrose, pyruvate, shikimate, citrate, malate, fumarate, isocitrate, and glutaric acid were significantly upregulated in contrast to ornithine, asparagine, and galactonic acid in the systemic leaves, indicating a restructuring of the energy levels-associated metabolism (Fig. 5).
Changes in the levels of the primary metabolites induced in Physalis peruviana by PhyRMV infection. The local and systemic leaves were subjected to mock- or PhyRMV-inoculation treatments and harvested at 03, 07, 14, 21, and 42 days after inoculation (DAI). The TCA cycle was constructed using the GC–MS data, post-normalization through a volcano plot analysis conducted utilizing the MetaboAnalyst software. The content of each metabolite is represented by the color scale of the legend. Red and blue squares indicate metabolites with increased or decreased levels in local (upper squares) and systemic (lower squares) leaves from PhyRMV-infected plants, compared to their respective mock controls. The dashed lines indicate non-sequential metabolic reactions (n = 3 biological replicates).
As the changes in primary metabolite levels of the systemic leaves were statistically significant, especially at 42 DAI (Fig. 5), the metabolic networks were analyzed based on their correlations (Fig. 6). Such analysis in systemic leaves revealed remarkable changes in metabolite composition and associations in response to PhyRMV infection (Fig. 6A). In systemic leaves, the metabolic networks demonstrated lower density and heterogeneity at 42 DAI in infected plants compared to the healthy ones (Fig. 6B). The decline in density reflects a decrease in the number and complexity of inter-metabolite connections, indicating a simplification of the network (Fig. 6A).
Correlation-based metabolic networks in systemic leaves from plants subjected to mock-inoculation (blue nodes) or PhyRMV infection (green nodes) at 42 days after inoculation (DAI). Networks were constructed from GC–MS-derived metabolite profiling data. Nodes represent individual metabolites, and edges indicate significant correlations between metabolites. Node size and edge thickness correspond to the number of connections and the magnitude of the debiased sparse partial correlation (DSPC) coefficient (r), respectively. Red edges indicate positive correlations, and blue edges indicate negative correlations. Networks were generated using the MetScape plugin in Cytoscape, applying correlation thresholds of r > 0.9 or r < − 0.9. Network density and heterogeneity were quantified using NetworkAnalyzer in Cytoscape.
However, a global reconfiguration of metabolism was observed, possibly reflecting the activation of pathways related to the stress response and antiviral defense in later stages of infection (Fig. 6A). Despite an apparent structural abridgment of the network, the quantity and strength of positive and negative correlations between metabolites elevated markedly. Such a change suggests that, though the network became less complex, the remaining pathways gained tighter correlations, reflecting a possibly highly coordinated and less flexible stress response focused on plant–pathogen interactions and mounting a defense response. However, the uninfected plants showed a greater network density and heterogeneity at 42 DAI (Fig. 6B,C). Additionally, robust connections were observed between the citric acid cycle metabolites, such as sucrose, glucose phosphate, and citric acid, which are vital for energy metabolism (Fig. 6A).
Discussion
Higher viral load and symptom severity were positively correlated
Stress response involves the integration of molecular, physiological, biochemical, and metabolic reactions; however, they vary temporally22. The results presented show that with the progression of PhyRMV infection, the viral load increased, directly causing cell, anatomical, morphological, and metabolic changes, and hence reducing productivity in P. peruviana . The infection progressed rapidly, with the virus replication detected in the local leaves as early as 3 DAI. The confirmation of systemic infection after 14 DAI, with a significant increase in viral load observed up to 42 DAI, directly correlated with the intensification of symptoms such as mosaic, chlorosis, and leaf deformation, a phenomenon frequently observed in plant–virus interactions23. In addition to foliar symptoms, the architecture of the infected plants changed remarkably, characterized by an increase in lateral branching and a reduction in height (Figs. 1B and 3A).
These effects suggest that PhyRMV could interferes with growth-related hormonal pathways such as auxin and cytokinin signaling, a phenomenon also described in other pathosystems involving viruses24. The reduction in chlorophyll content emphasizes the negative impact of the virus on photosynthesis rate while confirming the presence of chlorotic leaf spots, which ultimately delayed flowering and reduced the production potential, as evidenced by a 31% decreased number of fruits at 100 DAI. These data suggest the potential of PhyRMV to cause high damage and losses in the P. peruviana crop or in other Solanaceae members that may also be infected in the field. These results align with the data presented in other sobemovirus, such as rice yellow mottle virus (Sobemovirus RYMV; family Solemoviridae)25, southern bean mosaic virus (Sobemovirus SBMV; family Solemoviridae)26, tomato spotted wilt virus (TSWV, Orthotospovirus tomatomaculae, family Tospoviridae)27, and with PhyRMV2,9, supporting the notion that infection intensity directly influences plant physiology and production performance.
Leaves of PhyRMV-infected plants showed an altered histology and increased starch accumulation
Viruses are biotrophic pathogens that depend on the host machinery to complete their life cycle. The increase in virus accumulation associated with systemic spread can lead to direct cell-level damage, such as changes in organelle and whole-cell morphology, affecting energy production and photoassimilate distribution13. Therefore, for further investigation, the anatomies of systemic and local leaves of healthy (mock-inoculated) and PhyRMV-infected plants at 42 DAI were analyzed using an optical microscope.
Virus infection led to severe changes in local and systemic leaves. In the inoculated leaves, a collapse of the epidermal and mesophyll cells, tissue disorganization, and nuclear enlargement were observed, indicating intense metabolic activity most probably related to viral replication. Changes in vascular bundles presumably affect photoassimilate transport and starch accumulation within the infected leaves, as confirmed by Lugol’s staining. Such a pattern is a characteristic of biotic stresses where sucrose transport to the storage organs is blocked, leading to localized carbohydrate accumulation28,29. These results suggest that PhyRMV directly interferes with the host physiology and diverts resources toward viral replication.
Primary metabolism in the leaves of PhyRMV-infected plants is altered throughout the infection cycle
A metabolite profiling approach was utilized to characterize the patterns of metabolic changes within the Physalis peruviana–PhyRMV pathosystem. GC-MS revealed significant alterations in primary metabolism throughout the infection cycle. Statistical analyses identified 15 DEMs (p < 0.05; Table S1). Among these, compounds related to glycolysis, the citric acid cycle, and amino acid biosynthesis were predominant (Fig. 6). Although there were no statistically significant differences in the occurrence of primary metabolites between the local and systemic leaves at the onset of PhyRMV infection, such changes were observed at 42 DAI, suggesting a dynamic and programmed alteration of host metabolism in response to the late stage of viral infection (Fig. 6). Changes in plant primary metabolites have been observed for other plant viral pathosystems in a host-specific manner. Plants infected with the cucurbit chlorotic yellows virus (CCYV; Crinivirus, family Closteroviridae) demonstrated changes in the levels of metabolites related to the flavonoid, lipid, and amino acid biosynthesis pathways at 7 and 14 DAI30. However, the most pronounced metabolic changes were observed at the late stage of infection (42 DAI), likely reflecting cumulative virus replication, sustained activation of antiviral defence pathways, and progressive metabolic reprogramming associated with symptom development and virus-induced senescence.
The PCA results clearly indicated the temporal dynamics during the plant metabolic responses to PhyRMV infection (Fig. 4). Up to 21 DAI, the responses between the local and systemic leaves of infected and healthy plants overlapped, indicating the localized and subtle nature of the changes observed (Fig. 4). This result suggests that plants can maintain metabolic homeostasis post-inoculation, even with a high viral load at 21 DAI. However, as the infection progressed, a clear separation could be observed in the systemic leaves between treatments at 42 DAI, as indicated by PC1 (Fig. 4). These results demonstrate that metabolic reprogramming was triggered by viral load accumulation (Fig. 1D). The robust contribution of glutamate, isoleucine, and malonate (Figure S1) potentially involved in redirection of the metabolism toward plant defense-associated pathways. The amino acid glutamate links carbon and nitrogen metabolism in plants31. It is converted into α-ketoglutarate, a component of the citric acid cycle, and provides energy for maintaining cell processes under stress. Additionally, it is essential for the synthesis of γ-aminobutyric acid (GABA), a precursor for the biosynthesis of polyamines such as putrescine, which plays a critical role in stress responses32. In this study, putrescine concentrations differed markedly throughout the infection process, both in the local and systemic leaves (Table S1). Tobacco plants (cv. “Samsun NN”) infected with the tobacco mosaic virus (TMV; Tobamovirus tabaci, family Virgaviridae) showed an enhancement in the contents of polyamines, including putrescin, during a hypersensitive response (HR)33. This compound was present in the living cells surrounding necrotic lesions, suggesting its importance in limiting the movement of viral particles.
At 42 DAI, sucrose levels increased significantly in the systemic leaves of PhyRMV-infected plants (Fig. 5), related to starch accumulation within the leaf tissue, as indicated by anatomical analyses (Fig. 2). In the infected leaves, the mesophyll cells were disorganized, and the vascular cells (xylem and phloem) were impaired, directly affecting the remobilization of photoassimilates to the sink organs. In contrast, a significant elevation in cytosolic sucrose levels, especially at 42 DAI is associated with an increase in the availability of a carbon source for energy metabolism. The increased sucrose content suggests an enhancement in the glycolytic pathway, thereby supporting viral replication13, as a high viral load was observed at 42 DAI (Fig. 1). A few glycolytic pathway-associated enzymes are translocated to specific sites within the cell that are associated with viral propagation, increasing the ATP pool34,35.
Furthermore, an increase in the intermediary compounds of carbohydrate metabolism, in addition to supplying carbon sources for cells36, also indicates a modulation of the plant defense response-associated metabolic pathways12,37. The cytosolic accumulation of sucrose correlated with an increase in pyruvate levels, which was confirmed by our study (Fig. 5). The pyruvate then enters through the mitochondria and acts as a source of TCA metabolites. In our study, the TCA metabolites, including citrate, isocitrate, fumarate, and malate, increased significantly in the systemic leaves at 42 DAI (Fig. 5). This phenomenon could be related to the high-energy-consuming activities required during the viral infection of plants. It also suggests a possible metabolic reprogramming, in later stages of infection, to provide precursors for the biosynthesis of other biomolecules, such as amino acids, lipids, nucleotides, and secondary metabolites in response to biotic stress13. In addition, Jiang et al. (2023) found that the occurrence of mosaic symptoms in maize leaves infected with sugarcane mosaic virus (SCMV; Potyvirus sacchari, family Potyviridae) is related to the mitochondrial accumulation of malate, mediated by high pyruvate orthophosphate dikinase (PPDK) activity, which in turn causes the accumulation of reactive oxygen species within the mitochondria (mROS)12.
Plants infected by viruses reprogram their metabolism to limit the spread of the pathogen by producing secondary metabolites with antiviral and antioxidant properties13. For instance, the leaves of cucumber plants infected with CCYV exhibited spatiotemporal changes in secondary metabolite accumulation at 7 and 15 DAI30. In this study, the shikimate levels in the systemic leaves of plants infected with PhyRMV increased remarkably by 42 DAI. This observation suggested an activation of shikimic acid metabolism, correlated with biosynthesis of aromatic amino acids such as phenylalanine, tyrosine, and tryptophan is an essential component of the role of secondary metabolism in plant defense. This pathway functions in the chloroplast and links carbohydrate metabolism with the biosynthesis of aromatic compounds via phosphoenolpyruvate38.
Metabolite network analysis revealed a substantial reorganization of correlations among the metabolites (Fig. 6). At 42 DAI, the healthy plants exhibited complex integrated metabolic networks with greater density, heterogeneity, and inter-metabolite correlations, which helped maintain plant homeostasis (Fig. 6). These networks have several central hubs, including carbohydrates (sucrose, glucose, raffinose, mannitol, and sorbitol), organic acids (fumarate, pyruvate, and citrate), and essential amino acids (alanine and β-alanine). These metabolite groups are associated with specific functions during energy production and growth, indicating that healthy plants allocate energetic resources to related metabolic pathways. In contrast, plants infected at 42 DAI showed simplified metabolic networks with lower densities and heterogeneity (Fig. 6). This finding indicates a reduction in the complexity of the inter-metabolite correlations, as shown by the positive (red lines) and negative (blue lines), which exhibited an antagonistic relationship compared with healthy plants, as indicated by an increase in negative correlations. Evidently, at 42 DAI, the systemic leaves of infected plants reprogram metabolic pathways to activate the defense responses. Metabolic hubs such as isoleucine, glutamate, GABA, threonine, valine, and putrescine were the most prominent (Fig. 6A), suggesting a strong involvement of amino acid metabolism in the infection response. The metabolic hub Glutamate and its downstream metabolite GABA indicate metabolic and signaling reprogramming associated with stress-induced Ca2⁺ signaling, carbon–nitrogen balance, and systemic defense responses32. Together, these metabolomic changes indicate that late-stage infection is characterized by a coordinated adjustment of amino acid and polyamine metabolism that contributes to defense signaling, redox homeostasis, and metabolic resilience during virus–plant interactions.
The data from this study showed that infection disrupts the cooperation between primary metabolic processes in plants, which impairs the coordination among physiological responses as the resources are diverted toward defense mechanisms in response to the virus. Stress tolerance in plants involves maintaining a balance between the contents of the primary and secondary metabolites18,39,40. In addition, studies also highlight the critical role of primary metabolism in the acclimatization of plants to biotic stresses 41,42,43. Our results showed that up to 21 DAI, the infected plant attempts to adapt its metabolism and maintain metabolic homeostasis. However, after 42 DAI, an evident metabolic response occurs, in which the plant tries to activate defense-related pathways, which are, however, ineffective, and the physiological status declines, as indicated by the proposed model (Fig. 7). The results presented, therefore, provide insights into the mechanisms of plant–virus interactions in this pathosystem and their impacts on the metabolism and physiology of the host plant. These results support the development of future management strategies, such as the use of plant protection products. Furthermore, these results emphasize the importance of studies that integrate physiological, anatomical, and metabolic approaches to fully understand the actual impact of virus infections in crop species.
A plant–virus interaction model was constructed using the physalis rugose mosaic virus (PhyRMV) and Physalis peruviana pathosystem. (1) P. peruviana plants were mechanically inoculated with PhyRMV 45 days after transplanting (DAT). (2) The viral load increased in the infected plants at 3 days after inoculation (DAI), and it increased further until 21 DAI. (3) The virus spread throughout the plant; in the case of PhyRMV, it is not yet known whether it moves through the xylem or the phloem or both. (4) An increase in viral load was observed in the systemic outlets at 14 DAI, with an exponential enhancement up to 42 DAI. (5) Blistering, leaf deformation, and mosaic symptoms, as well as a reduction in plant size and total chlorophyll contents, were observed at ~ 21 DAI. (6) Anatomical changes were observed in the epidermal cells of the localized leaves at 42 DAI, with reduced size and collapse of chlorophyll parenchyma cells and structural changes in vascular tissue cells, impairing the photoassimilate flux due to elevated starch accumulation. In systemic leaves from plants infected with the virus, anatomical changes such as hyperplasia of the chlorophyll parenchyma cells, especially the palisade parenchyma cells, and increased starch accumulation were also observed. In addition, the glycolytic metabolic pathway was significantly upregulated at 42 DAI, which was reflected in a marked elevation in the levels of the metabolites, sucrose and pyruvate. Additionally, the contents of shikimate, an important precursor for the synthesis of secondary metabolites which form a part of the plant defense response, also increased. Analysis of metabolic networks in the systemic leaves at 42 DAI revealed remarkable changes in metabolite composition and correlations in response to PhyRMV infection. Overall, a global reconfiguration of primary metabolism was observed, possibly reflecting the activation of metabolic pathways related to stress responses and antiviral defense mechanisms. (7) The metabolic alterations observed at 42 DAI suggest a trade-off between plant growth and biotic stress defense, which in turn reduced the fruit set by 31% at 100 DAI.
Materials and methods
Plant material and growth conditions
Physalis peruviana fruits were collected from a commercial production field in Lages, Santa Catarina, Brazil (27° 48′ 57″ S, 50° 19′ 34″ W). The seeds were extracted and stored at the Plant Virology Laboratory, Santa Catarina State University (UDESC), Lages, Brazil. Seeds from healthy P. peruviana plants were germinated; at 30 days after germination, the seedlings were transplanted into 5 L pots containing a 3:1 soil–Maxfertil substrate mixture. The Ernani® liquid fertilizer (6% N, 7% P, and 8% K2O) was applied every 15 days at 10 mL/10 L of water per the manufacturer’s recommendations. The plants were infected with PhyRMV, isolated from Lages, SC, using a PhyRMV:BR:SC:01:2 (GenBank: MN782300). They were grown in a greenhouse at Santa Catarina State University, Brazil (latitude: − 27.792484; longitude: − 50.301851) at a controlled temperature of 24 ± 2 ℃ in 25 L pots with a 3:1 soil–Maxfertil® commercial substrate mixture serving as the inoculum source. The cultivation conditions were maintained during the entire development phase of the plants; all experiments were carried out under the same conditions.
Inoculation of Physalis leaves with PhyRMV, experimental design and sample collection
The virus inoculum is maintained in Physalis peruviana plants under protected conditions in a greenhouse, using aphid-proof cages. The initial inoculum was screened by high-throughput sequencing (HTS) and by PCR or RT-PCR using universal primers targeting the genera Begomovirus, Orthotospovirus, Crinivirus, and Tobamovirus; only the sobemovirus PhyRMV was detected.
Infected leaves were used to prepare the plant extract. Immediately after collection, 1 g of leaves was macerated in 5 mL of 0.02 M sodium phosphate buffer (pH 7.0) supplemented with 0.02 M sodium sulfite, with the addition of carborundum to promote microlesions on the leaf surface. The extract was then filtered through gauze and immediately used for mechanical inoculation of healthy plants. The virus was inoculated 45 days after transplanting (DAT), when the P. peruviana plants bore three true leaves.
A total of 36 Physalis peruviana plants were inoculated and divided into two equal groups (mock-inoculated and PhyRMV-infected, n = 18 per group). PhyRMV-infected plants were maintained at a distance of approximately 4 m from non-infected plants, with no direct contact between treatments. At each sampling time point (3, 7, 14, 21, and 42 days after inoculation, DAI), local (inoculated) and systemic (young) leaves were collected from a separate group of plants for each time point. Collected tissue was immediately frozen in liquid nitrogen to preserve metabolites and prevent degradation. Biological replicates were defined as independent individual plants, with three biological replicates per treatment per time point used for each analysis. The same plants were not monitored throughout the experiment, as repeated leaf collection would have caused tissue damage and compromised subsequent analyses.
Extraction of genetic material (RNA)
To detect the presence of the virus, ~ 50 mg of P. peruviana leaves with or without PhyRMV symptoms were homogenized separately in liquid nitrogen, and the total RNA was extracted using TRI reagent (Sigma Aldrich), according to the manufacturer’s instructions. The RNA integrity and quantity were checked using the 1% agarose gel electrophoresis and a NanoDrop 2000 spectrophotometer (Thermo Fisher). These analyses were performed at the Plant Virology Laboratory of the Center for Agricultural and Veterinary Sciences of the State University of Santa Catarina (UDESC).
cDNA synthesis
For cDNA synthesis, 3 µg of the total RNA, 0.1 µM of the reverse primer SOBDR2 (Table S2), and sufficient RNase-free water to obtain a final volume of 15 µL were initially added to the microtube. The tube was incubated at 70 ℃ for 5 min and at 2 ℃ for 10 min. Then, 3 µL of RNAse-free water, 5 µL of 5 × MMLV buffer, 1 µL of dNTPs (10 µM), and 1 µL of 200u/µL RT-MMLV enzyme (Promega) were added to the tube, which was incubated at 42 ℃ for 1 h.
Polymerase chain reaction (PCR)
At all sampling times, virus detection in plants infected with PhyRMV was evaluated by PCR (Figure S2) using primers specific to the genus Sobemovirus (Table S2). PCR was performed utilizing the GoTaq® Flexi DNA Polymerase (Promega) following the manufacturer’s instructions. The generic primers for Sobemoviruses, SOBDR2 and SOBDF22, were used (Table S2). The thermocycler (Applied Biosystems) was programmed as follows: preheating at 94 ℃ for 5 min; 40 cycles of 30 s at 94 ℃ (denaturation), 1 min at 58 ℃ (annealing), and 1 min at 72 ℃ (extension); and final extension for 7 min at 72 ℃. The PCR product was subjected to 1.0% agarose gel electrophoresis, stained with GelRed (Biotium Inc., CA, USA), visualized under UV light, and photographed.
Designing of qPCR primers and cloning of the viral coat protein-encoding gene
The coat protein gene (CP) sequence (MN782300) was downloaded from the GenBank database and used as a template to design the primers with the Primer3 Plus program (https://www.primer3plus.com/index.html). The specificity of the primers was validated in silico using the Nucleotide BLAST tool (https://blast.ncbi.nlm.nih.gov/Blast.cgi). The sequences of the primers targeting the partial CP coding region are listed in Table S2. For cloning, ~ 50 mg of the tissue from an infected plant was sampled and used for RNA extraction, followed by cDNA synthesis as described previously. The primers PhRMV_CP(F) and PhRMV_CP(R) were used (Table S2). The following program was employed: preheating at 95 ℃ for 2 min; 35 cycles of 1 min at 95 ℃ (denaturation), 30 s at 55 ℃ (annealing), and 30 s at 72 ℃ (extension); and a final extension for 7 min at 72 ℃. The PCR products were then separated on a 2% agarose gel electrophoresis, stained with GelRed (Biotium), and visualized under UV light. The bands of the expected size were excised and purified using the Wizard® SV Gel and PCR Clean-Up System Kit (Promega) per the manufacturer’s instructions.
The purified amplicons were cloned into pGEM-T Easy Vector (Promega) using T4 DNA Ligase (Promega), following the manufacturer’s instructions. The reaction was prepared on ice and incubated in a thermocycler at 4 ℃ for 16 h. The vector was then used to transform the Escherichia coli DH5α competent cells via electroporation using an Eppendorf Evaporator device, the factory pre-setting P1 (10 ng cells at 1 ℃). The mixture was added with 8 mL of Luria-Bertani broth (LB) medium, and the bacteria were incubated with shaking at 225 rpm for 5 h at 37 ℃.
Then, the bacteria-containing medium was centrifuged at 12,000g for 15 s to pellet the cells. They were then resuspended and plated on LB agar containing ampicillin (100 µg mL−1), IPTG (10 µL of a 100 mM solution), and X-GAL (16 µL of a 20 mg mL−1 solution). The plates were incubated at 37 ℃ for 16 h. The white colored colonies (positive clones) were selected for PCR-based confirmation of transformation. The confirmed colonies were selected and grown for 14 h at 37 ℃ with shaking at 200 rpm in LB broth. The plasmid DNA was extracted using the Wizard® Genomic DNA Purification Kit (Promega, USA), following the protocol provided. The product (plasmid containing the cloned CP fragment) was used to generate a standard amplification curve for PhyRMV:CP, which was then employed for the absolute quantification of the virus. For viral load analysis, standard curves were prepared using serial dilutions of plasmid DNA containing the complete CP coding region of PhyRMV (101–107 copies of the virus per reaction). The standard curves were generated by regression analysis of cycle threshold (Ct) values for each replicate of a given dilution against the logarithm of the amount of DNA in each dilution. To determine the absolute quantification of the number of viral copies in the different cultivars, 200 ng of cDNA, prepared as described previously, was used in reactions containing sets of virus-specific primers.
Quantification of the viral load by RT-qPCR
The RT-qPCR assay was performed in 96-well MicroAmp™ Optical reaction plates (Applied Biosystems, Warrington, UK). The reaction mix contained: 5 µL of 2X SYBR® Green PCR Master Mix (Applied Biosystems), 2 µL of 0.4 µM of each primer: PhRMV_CP(F) and PhRMV_CP(R), and 1 µL of 100 ng/µL cDNA, totaling 10µL. Reactions were performed in triplicate using a 7500 Real-Time PCR System (Applied Biosystems™) thermocycler. The program was as follows: initial denaturation at 95 ℃ for 10 min; 35 cycles of 95 ℃ for 15 s and 60 ℃ for 15 s. The plate consisted of a negative control (ultrapure water + RT-qPCR mix), the test mixes, and 11 mixes each corresponding to a specific concentration used for plotting the standard curve. The data were analyzed using the 7500 v2.0.6 software (Applied Biosystems) by determining the Ct value. The average of three technical triplicates for each reaction was calculated. The standard curve was generated by plotting the Ct values obtained against the log value of the RNA copy number (Ct vs. the log of the standard sample amount). Regression analyses were applied to the standard curves using Excel software (Microsoft, WA, USA). The virus copy number was calculated based on the mean Ct value and the linear regression equation from the standard curve.
Analysis of the total chlorophyll content (Falker Chlorophyll Index [ICF]), plant height, and number of fruits
The total chlorophyll content was analyzed using clorofiLOG (Falker, Porto Alegre, Brazil) at 21 DAI. Plant height (cm) was measured with a tape measure at 0, 21, and 42 DAI. The number of fruits was determined by collecting the fruits at 150 DAI.
Morphological and anatomical analyses
Leaves were sampled at 42 DAI44. After 24 h of collection, the samples were prewashed with phosphate buffer, dehydrated in an ethanol series, and stored in 70% ethanol. They were then dehydrated in an ethanol series and embedded in hydroxyethyl methacrylate resin (Historesin; Leica, Heidelberg, Germany). Cross sections, 5 µm thick, were prepared with a 1508 R rotary microtome (Ciencor Scientific, São Paulo, Brazil), stained with 0.05% toluidine blue in 0.1 M phosphate buffer (pH 6.8)45 and mounted in Acrilex® colorless stained glass enamel46. Observations and photographic documentation were performed employing a BX61 motorized microscope (Olympus, Tokyo, Japan) attached with a DP-72 CCD digital camera (Olympus) at the Plant Anatomy Laboratory of the Instituto Federal Goiano, Rio Verde Campus, Goiás, Brazil. Lugol’s iodide was added to detect the starch in the samples47.
GC–MS-based metabolic profiling
A previously described protocol was adapted to obtain the metabolite profile48. Local and systemic leaves of plants inoculated with buffer or PhyRMV were collected at 0, 3, 7, 14, 21, and 42 DAI, flash-frozen in liquid nitrogen, and then stored in an ultrafreezer at − 80 ℃. The samples were lyophilized at –50 ℃ and macerated in liquid nitrogen until a fine powder was obtained. The material was then weighed, 50 mg of it was then added to 700 μL of ice-cold methanol and 30 μL of 0.2 mg mL ribitol solution, used as an internal standard in the analysis for relative quantification, followed by vortex homogenization. Subsequently, the samples were stirred at 70 °C for 15 min at 350 RPM, and centrifuged at 12,000g and 4 ℃ for 10 min. Next, 375 μL of ice-cold chloroform and 750 μL of ultrapure MilliQ water were added, and vortex mixed. Immediately, the samples were centrifuged at 4000 RPM for 15 min at 4 ℃, and aliquots of the upper phase formed after the process, which corresponded to the polar phase, were collected. Of each, 150 μL was aliquoted for derivatization and analyzed using GC–MS. Of each sample, 5 μL was aliquoted into a single microtube to prepare a control, from which 150 μL was aliquoted. Finally, all samples were speed vacuum dried for 16 h.
For derivatization, preceding injection into the GC, 20 μL of methoxyamine hydrochloride dissolved in pyridine (20 mg mL−1) was added to the samples. The microtubes were then vortexed at 37 ℃ for 2 h at 550 RPM, followed by spin-drying. Next, 35 μL of N-methyl-N-trimethylsilyl-trifluoroacetamide (MSTFA) was added, and shaken at 37 ℃ for 30 min at 550 RPM. Immediately, 55 μL of the sample was transferred to the vials and injected. The metabolites were analyzed with a QP2010 GC–MS (Shimadzu, Tokyo, Japan) by injecting 1 μL of derivatized sample composed of a RTX-5MS capillary column; 30 m × 0.25 mm × 0.25 μm (Restek, PA, USA), with helium gas at a flow rate of 1.2 mL min−1 in split 5 mode (sample–gas dilution of 1:5). The injection and ion source temperatures were 230 ℃ and 250 ℃, respectively. The run parameters were set to 80 ℃ for 4 min, followed by a gradual heating at 10 ℃ min−1 until a maximum of 315 ℃ was reached and maintained for 8 min. The metabolites were ionized and fragmented by electron impact at 70 eV and subsequently analyzed in a quadrupole, generating spectra between 40 and 700 m/z in full scan mode. The chromatograms obtained were examined, and each metabolite was identified using the Xcalibur™ 2.2 software (Thermo Fisher Scientific, MA, USA) with quality control as a parameter. For this process, a library of previously detected metabolites and the GOLM molecule database (http://gmd.mpimp-golm.mpg.de/; Max Planck Institute of Molecular Plant Physiology, Potsdam, Germany) were used for comparing the chromatography retention times and the fragmentation patterns available in them as a reference with those obtained via MS. Relative quantification was based on the ratio between the areas of the detected peak and the internal standard peak (Ribitol), the result of which was divided by the sample mass. The metabolites were then identified and classified using the Kyoto Gene and Genome Encyclopedia (KEGG) database. These analyses were performed at the Plant Metabolism Laboratory of the Federal University of Ceará (UFC), Ceará, Brazil.
Network analysis
Metabolites correlation (r) was calculated using the Lasso Penalty method, assisted by the Correlation Calculator software49. Metabolic networks are constructed utilizing the MetScape plugin in CytoScape software (https://apps.cytoscape.org/apps/metscape). The Debiased Sparse Partial Correlation (DSPC) between two nodes was represented by correlation coefficients in the range − 0.9 > r > 0.9. Only strongs correlations were considered in the network. Metabolic network nodes were considered hubs if they were above the average connectivity of the network, indicating greater influence within the network. Network parameters such as network heterogeneity and network density were obtained using the NetworkAnalyzer on Cytoscape®, as described earlier50.
Experimental design and multivariate statistical analyses
The experiments were conducted employing a completely randomized design. ANOVA with an F-test was performed using SAS® OnDemand for Academics (https://www.sas.com/en_in/software/on-demand-for-academics.html) to analyze the chlorophyll total content at 21 days, as well as the number of fruits and plant height at 42 days.
For metabolomics, the files with the compounds of interest were processed employing the MetaboAnalyst 6.0 web interface (http://www.metaboanalyst.ca). The data were normalized by cube root transformation and automatic scaling (centered mean divided by the standard deviation of each variable), resulting in a statistically more appropriate normal distribution. The principal component analysis (PCA) was performed to identify the differences between the two treatments at each time point separately for the local and systemic leaves. The results were subjected to ANOVA to obtain their statistical significance, followed by Tukey’s test (P < 0.05) to identify the statistical significance of the variations between the mock and PhyRMV groups of local and systemic leaves, regardless of time. The fold change threshold of 2.0 generated by the Volcano plot was used to construct the citric acid pathway.
Data availability
All data generated or analyzed during this study are included in this published article. The raw datasets are available from the corresponding author on reasonable request.
References
Fischer, G., Almanza-Merchán, P. J. & Miranda, D. Importancia y cultivo de la uchuva (Physalis peruviana L.). Rev. Bras. Frutic. 36, 01–15. https://doi.org/10.1590/0100-2945-441/13 (2014).
Fariña, A. E. et al. Molecular and biological characterization of a putative new sobemovirus infecting Physalis peruviana. Arch Virol. 164, 2805–2810. https://doi.org/10.1007/s00705-019-04374-y (2019).
Muniz, J. et al. General aspects of physalis cultivation. Cienc. Rural 44, 964–970. https://doi.org/10.1590/S0103-84782014005000006 (2014).
Rufato, L. et al. Aspectos técnicos da cultura da physalis. Lages: CAV/UDESC; Pelotas: UFPel, 101p, (2008).
Kraide, H. D. et al. Transmission of Physalis rugose mosaic virus via pruning, leaves contact and soil. Trop. Plant Pathol. 50, 21. https://doi.org/10.1007/s40858-025-00691-7 (2025).
Eiras, M. et al. First report of a tospovirus in a commercial crop of Cape gooseberry in Brazil. New Disease Rep. 25, 25. https://doi.org/10.5197/j.2044-0588.2012.025.025 (2012).
Esquivel, A. F. et al. First report of Groundnut ring spot virus on Physalis peruviana in Brazil. Plant Dis. 102, 1468. https://doi.org/10.1094/PDIS-10-17-1684-PDN (2018).
Esquivel-Fariña, A. et al. Occurrence of natural infection of Physalis peruviana with potato virus Y and pepper yellow mosaic virus in Brazil. J. Plant Pathol. 104, 1315–1318. https://doi.org/10.1007/s42161-021-00999-8 (2022).
Gorayeb, E. S. et al. Damage quantification in Physalis peruviana L. infected by the new putative sobemovirus physalis rugose mosaic virus. Trop. Plant Pathol. 45, 476–483. https://doi.org/10.1007/s40858-020-00354-9 (2020).
Kraide, H. D. et al. Effects of physalis rugose mosaic and groundnut ringspot viruses, in single and double infections, on the development, yield and postharvest fruit characteristics of Physalis peruviana plants. Plant. Pathol. 72, 1104–1110. https://doi.org/10.1111/ppa.13733 (2023).
Savi, A. et al. Near-complete genome sequence and seed transmission evaluation of Physalis rugose mosaic virus from southern Brazil. Cienc. Rural 51, e20200702. https://doi.org/10.1590/01038478cr20200702 (2021).
Jiang, T. & Zhou, T. Unraveling the mechanisms of virus-induced symptom development in plants. Plants 12, 2830. https://doi.org/10.3390/plants12152830 (2023).
Jiang, T. et al. Reprogrammed plant metabolism during viral infections: Mechanisms pathways and implications. Mol. Plant Pathol. 26, e70066. https://doi.org/10.1111/mpp.70066 (2025).
Singh, A. et al. Influence of phosphorus on the performance of cowpea (Vigna unguiculata (L) Walp.) varieties in the Sudan savanna of Nigeria. Agric. Sci. 2, 313–317. https://doi.org/10.4236/as.2011.23042 (2011).
Brosset, A. & Blande, J. D. Volatile-mediated plant–plant interactions: Volatile organic compounds as modulators of receiver plant defence, growth, and reproduction. J. Exp. Bot. 73, 511–528. https://doi.org/10.1093/jxb/erab487 (2022).
De Souza, L. P., Alseekh, S., Brotman, Y. & Fernie, A. R. Network-based strategies in metabolomics data analysis and interpretation: From molecular networking to biological interpretation. Expert Rev. Proteom. 17(4), 243–255. https://doi.org/10.1080/14789450.2020.1766975 (2020).
Sulpice, R. et al. Starch as a major integrator in the regulation of plant growth. Proc. Natl. Acad. Sci. USA 106(25), 10348–10353. https://doi.org/10.1073/pnas.0903478106 (2009).
Cardoso, L. L., Freire, F. B. S. & Daloso, D. M. Plant metabolic networks under stress: A multi-species/stress condition meta-analysis. J. Soil Sci. Plant Nutr. 23, 4–21. https://doi.org/10.1007/s42729-022-01032-2 (2023).
Erb, M. & Kliebenstein, D. J. Plant secondary metabolites as defenses, regulators, and primary metabolites: The blurred functional trichotomy. Plant Physiol. 184(1), 39–52. https://doi.org/10.1104/pp.20.00433 (2020).
Piasecka, A., Jedrzejczak-Rey, N. & Bednarek, P. Secondary metabolites in plant innate immunity: Conserved function of divergent chemicals. New Phytol. 206, 948–964. https://doi.org/10.1111/nph.13325 (2015).
Vicente, M. R. & Plasencia, J. Salicylic acid beyond defence: Its role in plant growth and development. J. Exp. Bot. 62(10), 3321–3338. https://doi.org/10.1093/jxb/err031 (2011).
Galviz, Y., Souza, G. M. & Lüttge, U. The biological concept of stress revisited: Relations of stress and memory of plants as a matter of space–time. Theor. Exp. Plant Physiol. 34, 239–264. https://doi.org/10.1007/s40626-022-00245-1 (2022).
HULL, R. Plant Virology. 5. ed. Norwich: Minihane, C., 2013.
Islam, W. et al. Plant defense against virus diseases; growth hormones in highlights. Plant Signal. Behav. https://doi.org/10.1080/15592324.2019.1596719 (2019).
Kouassi, A. Y., Djedatin, G., Zangue, R. & Almouni, A. Rice Yellow Mottle Virus (RYMV): A review. Viruses 16(11), 1707. https://doi.org/10.3390/v16111707 (2024).
Kareem, K. & Taiwo, M. Interactions of viruses in Cowpea: Effects on growth and yield parameters. Virol. J. 4, 15. https://doi.org/10.1186/1743-422X-4-15 (2007).
Tiberini, A., Cillo, F., Gentili, A. & Bertin, S. Tomato spotted wilt virus (Orthotospovirus tomatomaculae), a cyclically occurring threat to crop production worldwide. Ann. Appl. Biol. 186(2), 93–114. https://doi.org/10.1111/aab.12977 (2025).
Schoelz, J. E., Harries, P. A. & Nelson, R. S. Intracellular transport of plant viruses: Finding the door out of the cell. Mol. Plant. 4(5), 813–831. https://doi.org/10.1093/mp/ssr070 (2011).
Cornara, L., Mandrone, M. & Smeriglio, A. Biotic and abiotic stressors in plant metabolism. Int. J. Mol. Sci. 25(1), 121. https://doi.org/10.3390/ijms25010121 (2023).
Zhang, Z. et al. Widely targeted analysis of metabolomic changes of Cucumis sativus induced by cucurbit chlorotic yellows virus. BMC Plant Biol. 22, 158. https://doi.org/10.1186/s12870-022-03555-3 (2022).
Seifi, H. S., Van Bockhaven, J., Angenon, G. & Höfte, M. Glutamate metabolism in plant disease and defense: Friend or foe?. Mol Plant Microbe Interact 26(5), 475–485. https://doi.org/10.1094/MPMI-07-12-0176-CR (2013).
Caspi, Y. et al. Why did glutamate, GABA, and melatonin become intercellular signalling molecules in plants?. Elife 12, e83361. https://doi.org/10.7554/eLife.83361 (2023).
Torrigiani, P. et al. Polyamine synthesis and accumulation in the hypersensitive response to TMV in Nicotiana tabacum. New Phytol. 135, 467–473. https://doi.org/10.1046/j.1469-8137.1997.00669.x (1997).
Nagy, P. D. & Lin, W. Taking over cellular energy-metabolism for TBSV replication: The high ATP requirement of an RNA virus within the viral replication organelle. Viruses 12, 56. https://doi.org/10.3390/v12010056 (2020).
Liu, Y., Lin, W. & Nagy, P. D. Proviral and antiviral roles of phosphofructokinase family of glycolytic enzymes in TBSV replication. Virology 599, 110190. https://doi.org/10.1016/j.virol.2024.110190 (2024).
Fernie, A. R., Carrari, F. & Sweetlove, L. J. Respiratory metabolism: Glycolysis, the TCA cycle and mitochondrial electron transport. Curr. Opin. Plant Biol. 7(3), 254–261. https://doi.org/10.1016/j.pbi.2004.03.007 (2004).
Kogovšek, P. et al. Primary metabolism, phenylpropanoids and antioxidant pathways are regulated in potato as a response to potato virus Y infection. PLoS ONE 11, e0146135. https://doi.org/10.1371/journal.pone.0146135 (2016).
Lu, X. et al. The shikimate pathway regulates programmed cell death. J. Genet. Genom. 49(10), 943–951. https://doi.org/10.1016/j.jgg.2022.02.001 (2022).
Bechtold, U. & Field, B. Molecular mechanisms controlling plant growth during abiotic stress. J. Exp. Bot. 11, 2753–2758. https://doi.org/10.1093/jxb/ery157 (2018).
Cai, J. & Aharoni, A. Amino acids and their derivatives mediating defense priming and growth tradeoff. Curr. Opin. Plant Biol. https://doi.org/10.1016/j.pbi.2022.102288 (2022).
Liu, M. et al. Growth–defense trade-off regulated by hormones in grass plants growing under different grazing intensities. Physiol. Plant. 166(2), 553–569. https://doi.org/10.1111/ppl.12802 (2019).
Costa, I. R. S. et al. Metabolomic exhibits different profiles and potential biomarkers of Vitis spp. co-cultivated with Fusarium oxysporum for short, medium, and long times. Physiol. Plant. 175(3), e13918. https://doi.org/10.1111/ppl.13918 (2023).
Feng, S. et al. The glutamate receptor plays a role in defense against Botrytis cinerea through electrical signaling in tomato. Appl. Sci. 23, 11217. https://doi.org/10.3390/APP112311217 (2021).
Karnovisk, M. J. A. A formaldehydeglutaraldehyde fixative of high osmolarity for use in electron microscopy. J. Cell Biol 27, 137–138 (1965).
O’Brien, T. P., Feder, N. & McCully, M. E. Polychromatic staining of plant cell walls by toluidine blue. Protoplasma 59, 368–373 (1964).
de Paiva, J. G. A., Fank-de-Carvalho, S. M., Magalhães, M. P. & Graciano-Ribeiro, D. Verniz vitral incolor 500®: Uma alternativa de meio de montagem economicamente viável. Acta Bot. Bras. 20, 257–264. https://doi.org/10.1590/S0102-33062006000200002 (2006).
JENSEN, W. A. Botanical Histochemistry: Principles and Practice. W.H. Feeman & Co., San Francisco (1962).
Lisec, J. et al. Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat. Protocols 1, 387–396 (2006).
Basu, S. et al. Sparse network modeling and metscape-based visualization methods for the analysis of large-scale metabolomics data. Bioinformatics 33, 1545–1553. https://doi.org/10.1093/bioinformatics/btx012 (2017).
Assenov, Y., Ramírez, F., Schelhorn, S.-E., Lengauer, T. & Albrecht, M. Computing topological parameters of biological networks. Bioinformatics 24, 282–284. https://doi.org/10.1093/bioinformatics/btm554 (2008).
Acknowledgements
This work was financially supported by the Fundação de Amparo à Pesquisa e Inovação do Estado de Santa Catarina (FAPESC—Grant no. 2024TR001588 to G.C.M), the Santa Catarina Federal Institute of Education, Science and Technology (IFSC/PROPPI Grants no. 02/2024). M.H.S are grateful to Santa Catarina State University (PROMOP), J.R.B and N.B.F to FAPESC, M.E.S.S to IFSC, and H.H.C. and F.N.S (304883/2025-5) to CNPq, for the scholarships. We are grateful to the Plant Metabolism Laboratory (LabPlant) at the Federal University of Ceará for their partnership.
Funding
This work was financially supported by the Fundação de Amparo à Pesquisa e Inovação do Estado de Santa Catarina (FAPESC—Grant no. 2024TR001588 to G.C.M).
Author information
Authors and Affiliations
Contributions
GCM and FNS conceived and designed experiments. MHS, JP, JRB, NBF and MESS performed the greenhouse experiments; MHS performed RNA extractions, cDNA synthesis, greenhouse experiments and data analysis; MHS and JRB performed cloning; MHS, JP and NBF performed fluorescence analysis; MHS, IMCP and HHC performed metabolic analysis; VCD performed anatomical analysis; GCM and FNS supervised, coordinated all the experiments and prepared the draft of the manuscript. A.B and E.E.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Studnicka, M.H., Bianchini, J.R., Felisberto, N.B. et al. Morphophysiological disorders and metabolic reprogramming in Physalis peruviana infected with the physalis rugose mosaic virus. Sci Rep 16, 9015 (2026). https://doi.org/10.1038/s41598-026-39107-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-39107-0