Abstract
Modeling radiation-induced DNA damage is essential for understanding the relative biological effectiveness (RBE) of ionizing radiation. This study presents a simplified framework using the density-based spatial clustering of applications with noise (DBSCAN) algorithm and using simulated physical track structures to characterize DNA damage induced by 0.5–200 MeV protons. The model assumed that (i) energy deposition ≥ 17.5 eV induced DNA damage, (ii) at least two damage points within a distance ε formed a cluster, (iii) isolated damage points were treated as noises, and (iv) the cluster-to-noise ratio corresponded to the double strand break (DSB)-to-single strand break (SSB) yield ratio. From the clustering output, a new beam quality metric called Quality of Beam (QoB; clusters per particle per µm) and its normalized form (clusters per keV of deposited energy) were defined. For protons, normalized QoB exhibited a strong linear correlation with DSB yields, enabling direct estimation of DSB and SSB yields using a single conversion factor. Applying the same framework and model parameters to carbon ions revealed a similar linear relationship between normalized QoB and DSB yields up to LET values of 200 keV µm⁻¹, beyond which the overkill effect emerged. The normalized QoB qualitatively reproduced RBE–LET trends and offered a biologically meaningful alternative to conventional metrics such as LET. Compared to full water radiolysis modeling, the DBSCAN framework was computationally efficient and provided a robust method for assessing ion beam quality and predicting DNA damage yields.
Similar content being viewed by others
Introduction
Particle beam therapy using protons and carbon ions optimizes cancer treatment by utilizing the Bragg peak to target the tumor precisely while minimizing damage to surrounding healthy tissues. In clinical practice, the relative biological effectiveness (RBE) of protons, defined as the ratio of dose of a reference radiation to proton dose that produces the same biological effect, is commonly assumed to be 1.11, while the RBEs of carbon ions are based on biophysical models such as the local effect model (LEM)2 and the microdosimetric kinetic model (MKM)3. However, the variability in the RBE in proton therapy, especially at the distal region of the Bragg peak, has been observed in both cell experiments4 and clinical studies5, making it important to model the RBE more accurately, or using appropriate quantities for proton therapy optimization. In addition to physical dose (or constant RBE weighted dose) optimization, other beam qualities, such as, linear energy transfer (LET)6 and track ends7, have been introduced to reflect the biological effectiveness of protons. Also, in terms of RBE modeling, several approaches have been proposed, including phenomenological modeling8,9,10 and mechanistic modeling11. In particular, the latter approach aims to mimic the mechanisms of radiation-induced biological effects, starting from the spatial distribution of energy deposition at the atomic level to DNA damage induction and following biological processes. As such, modeling of radiation-induced DNA damage is usually the biological starting point for subsequent mechanistic modeling12.
Typically, DNA damage modeling involves Monte Calo track structure simulation to produce the spatial pattern of direct energy deposition and radiolytic species arising from all stages of water radiolysis (physical, physicochemical and chemical), commonly known as “radiation track structures”. Then, DNA damage is assumed based on the relative positions of direct energy deposition and free radicals to the DNA structure, as well as the threshold of energy deposition and the probability of free radicals to induce damage12,13,14. This approach, despite nearly realistically mimicking DNA damage induction by ionizing radiation, is computationally expensive as it requires a comprehensive simulation of water radiolysis, as well as the search through radiation track structures and DNA structures at the atomic level. The other approach of DNA damage modeling is to simulate only the physical stage of water radiolysis and use cluster analysis to group energy deposition points into clusters. By relating the geometrical size of the cluster to the DNA structure, and assuming energy deposition criteria for DNA damage induction, the yields of single strand breaks (SSBs) and double strand breaks (DSBs) can be predicted without the need to simulate all three stages of water radiolysis and the detailed structure of the DNA15,16,17,18,19.
In data mining, cluster analysis is a method of grouping data from a large dataset that have the same characteristics into the same group and describing how they are linked. Clustering algorithms are, for example, hierarchical clustering algorithm (connectivity-based clustering)20, k-means method or Lloyd’s algorithm (centroid-based clustering)21,22, and density-based clustering algorithms, including the Clustering Large Applications based on RANdomized Search (CLARANS) algorithm23 and the Density Based Spatial Clustering of Applications with Noise (DBSCAN) algorithm24,25. Although traditional clustering algorithms, including k-means and hierarchical methods, are suitable for datasets with well-defined, compact clusters, they may struggle with irregular spatial patterns of energy deposition produced by ionizing radiation, which often involves clusters of arbitrary shapes and sizes along with noises representing energy deposition points that do not lead to biologically significant damage. DBSCAN presents a powerful tool specifically designed to identify clusters of arbitrary shapes and sizes while effectively filtering out noises24, making it attractive for analyzing radiation track structures for DNA damage modeling.
In the past, Francis et al.15 introduced the application of DBSCAN for the calculation of SSB and DSB yields induced by 0.5–50 MeV protons. They assumed the probability of DNA damage induction as a linear function of energy deposited in the sensitive volume and introduced adjustable parameters: one representing the ratio of the sensitive volume to the total medium’s volume of 16% and the other being the maximum distance for cluster formation of 3.2 nm. Later, Matsuya et al.18,19 applied the clustering analysis for estimating SSB and DSB yields from x-ray, electron and proton beams, assuming a maximum distance for cluster formation of 3.4 nm and using adjustable parameters \(\:{k}_{\text{S}\text{S}\text{B}}\) and \(\:{k}_{\text{D}\text{S}\text{B}}\) for fitting the model to experimental SSB and DSB yields, respectively.
Based on the promising results of prior DBSCAN approaches, we developed a simple DBSCAN model with a newly proposed beam quality metric, the normalized Quality of Beam (QoB), defined as the number of clusters per unit deposited energy, to derive a single conversion factor for calculating both DSB and SSB yields. In this work, we present the details of the model and its application to therapeutic proton beams in the energy range of 0.5–200 MeV, comparing the results with experimental data and other simulation studies, and demonstrated the potential of normalized QoB to characterize proton beams through its correlation with DSB yields. In addition, we extended the same DBSCAN framework to carbon ions to assess its generalizability across ion species.
Methods
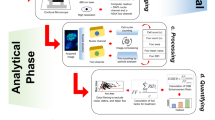
The flow diagram for this study is illustrated in Fig. 1 and detailed in the following sections.
The comprehensive workflow diagram of this work.
Geant4-DNA simulation
Monoenergetic proton and carbon ion track structures were simulated in liquid water using Geant4-DNA Monte Carlo track structure toolkit version 11.126, following the workflow illustrated in Fig. 1. The simulations were performed for proton energies ranging from 0.5 to 200 MeV (corresponding to linear energy transfer (LET) from 40.70 to 0.36 keV µm⁻¹, respectively), and carbon ion energies ranging from 2 to 100 MeV/n (corresponding to LET from 453.37 to 18.51 keV µm⁻¹, respectively). The LET larger than 10 keV µm⁻¹ was considered as high LET in this work.
Tables 1 and 2 summarize the simulation parameters for protons and carbon ions. The isotropic point sources were simulated with the starting point set at the center of a spherical water volume, defined as the working volume. The radius of the working volume was varied from 0.25 μm to 2.5 μm, and the number of simulated particle histories ranged from the order of 103 to 105, depending on the particle species and energy, to ensure sufficient statistics of energy deposition with a minimal change of particle energies inside the working volume. The Geant4-DNA physics constructor G4EmDNAPhysics_option4 was employed for physical interactions of protons and carbon ions with liquid water.
For each simulated particle history, the spatial coordinates and energy deposited at each interaction point in the physical stage of water radiolysis were recorded. This data was subsequently used for the clustering analysis explained in the next section. To validate the accuracy of the simulated track structures, LET and microdosimetric quantities27 of protons were calculated and compared with published data. Specifically, frequency-mean lineal energy \(\:\left( {{\bar{y}}}_{F} \right)\) and dose-mean lineal energy \(\:\left( {{\bar{y}}}_{D} \right)\) were calculated in cylindrical targets with the length and diameter of 2 nm, mimicking a short segment of the DNA28.
DBSCAN algorithm and DNA damage calculation
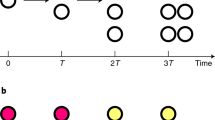
Illustration of the DBSCAN algorithm implemented in this work.
Figure 2 illustrates the DBSCAN algorithm implemented in this work. For each particle energy, all simulated particle tracks were examined once by the clustering analysis using the following three key parameters:
-
Energy deposition threshold (E thres), defined as the minimum energy required to induce a DNA strand break. The value of 17.5 eV was adopted from the publication of Nikjoo et al.29. Interaction points with energy deposition below Ethres were not considered in the clustering analysis.
-
Minimum Points (MinPts), defined as the minimum number of damage points to form a cluster within a given distance. MinPts of 2 was chosen based on the definition of a DSB as two closely spaced lesions.
-
Maximum distance for cluster formation (ε), defined as the maximum separation between two damage points to be considered a part of the same cluster. Isolated damage points were considered as noises.
The DBSCAN clustering analysis resulted in the number of clusters and noises. This information was used to calculate the following quantities:
-
Cluster-to-noise (C/N) ratio, interpreted in this work as the ratio of DSB to SSB yield (DSB/SSB ratio). This interpretation was supported by previous studies, demonstrating a correlation between clustered energy deposition and induction of complex DNA damage30,31. Specifically, closely spaced energy deposition points within a DNA molecule increased the probability of two or more strand breaks occurring in close proximity, leading to DSB formation. Conversely, isolated energy depositions were more likely to induce single, isolated strand breaks, resulting in SSBs.
-
Quality of Beam (QoB), defined as the average number of clusters formed per unit path length per particle. This metric provided a measure of the clustering potential of the beam and had the unit of particle− 1 μm− 1.
-
Normalized QoB, calculated as the QoB divided by LET, thus normalizing the clustering density to energy deposited per unit path length. The normalized QoB had the unit of clusters keV− 1, enabling the comparison of beam qualities over different particle energies.
For each particle species and energy, we treated every simulated particle history as an independent replicate. Uncertainties were calculated to reflect the stochastic variance from the finite number of particle histories rather than systematic effects from physics models or DBSCAN parameterization. For LET, we computed the track-averaged value per history and summarized results with the sample mean. The 95% confidence interval (CI) of the LET was obtained based on the mean and its standard error using the normal approximation. This approach was considered appropriate due to the large number of histories involved. For QoB, we calculated the cluster yield per unit path length for each history and reported the mean with 95% CIs in the same manner as for LET. For normalized QoB, we formed the ratio per history (QoB divided by that history’s LET) to avoid bias from dividing two separate means. Then, these ratios were averaged over particle histories, and the 95% CI was derived from the mean and standard error of the per-history ratio. The C/N ratio was computed globally for each beam energy from the total number of clusters divided by the total number of noises, aggregated over all histories. Since the C/N ratio was a single global ratio rather than a per-history statistic, the statistical error and CI were not calculated.
For DNA damage modeling, we used the DSB/SSB ratios and the absolute DSB yields reported by Kundrát et al.13 to fine tune our model. First, we optimized the maximum distance for cluster formation (ε) so that the calculated C/N ratios reproduced the reported DSB/SSB ratios. We used the optimized ε value to calculate the normalized QoB and found that the normalized QoB exhibited a strong linear correlation with the published DSB yields. Thus, the slope of the linear fit served as a single conversion factor (\(\:\text{C}\text{F}\)) for determination of the DSB yield, and by dividing the DSB yield by the C/N ratio, the SSB yield could be calculated, as follows
All DBSCAN calculations were performed using the scikit-learn machine learning library32 in Python 3.10.
After completing DNA damage modeling for protons, we applied the same DBSCAN framework to carbon ions to evaluate its generalizability. To maintain methodological consistency, we fixed the clustering parameters to those validated for protons (Ethres = 17.5 eV, MinPts = 2, and ε = 2.1 nm).
Code availability
The custom codes for Geant4-DNA particle track simulation and the Python-based DBSCAN analysis are available via GitHub at https://github.com/radsimbio/DBSCAN. To ensure stability, the specific version used in this study is permanently archived on Zenodo at https://doi.org/10.5281/zenodo.18364916.
Results
Validation of simulated proton track structures
The accuracy of the simulated proton track structures was assessed by comparing the calculated LET and microdosimetric quantities with published data. Figure 3 shows the comparison of simulated LET, frequency-mean lineal energy \({{\bar{y}}}_{F}\), and dose-mean lineal energy \({{\bar{y}}}_{D}\) with those obtained from the PSTAR database for LET33 and the simulations of Liamsuwan et al. for microdosimetric quantities34. Our results agreed generally well with the published data, indicating accurate generation of proton energy deposition patterns. The largest discrepancy observed was 23% for relatively low LET protons.
DNA damage yields
Figure 4a compares the best fit of the calculated C/N ratios with the DSB/SSB ratios reported by Kundrát et al.13. To determine the optimal maximum distance for cluster formation (ε), we systematically varied ε from 1.0 nm to 10.0 nm. At smaller ε values (e.g., 1.0 nm), the calculated C/N ratios showed pronounced discrepancies compared with the reference DSB/SSB ratios, while larger ε values (approaching 10.0 nm) tended to overestimate clustering. The best agreement was obtained at ε = 2.1 nm. This value was subsequently used for calculating the normalized QoB.
Figure 4b shows the linear relationship between the normalized QoB (calculated with ε= 2.1 nm) and the DSB yields reported by Kundrát et al.13, with a high correlation coefficient (R2 = 0.997). The slope of the linear fit was (1.1498 ± 0.019)×10− 10 keV Gy− 1 Da− 1, which was subsequently used as a conversion factor to calculate absolute DSB and SSB yields, as described by Eqs. (1)-(2).
When compared with other simulations15,19,35,36, a consistent linear correlation was observed between normalized QoB and DSB yields (R² > 0.930). However, variation in the absolute DSB yield values reported by those studies led to differences in the slope of the linear fits, ranging from (0.7678 ± 0.123)×10⁻¹⁰ keV Gy− 1 Da− 1for Mokari et al.36 to (2.8012 ± 0.220)×10⁻¹⁰ keV Gy− 1 Da− 1for Matsuya et al.19.
Comparison of the calculated C/N ratios with the DSB/SSB ratios obtained from the detailed DNA damage simulation of Kundrát et al.13 (a) and the relationship between normalized Quality of Beam (QoB) (using 2.1 nm as the optimal value of the maximum cluster radius, ε) and simulated DSB yields reported by different investigators15,19,35,36 (b).
The resulting SSB and DSB yields are shown in Fig. 5, compared with those obtained from the experimental data and other simulations (both detailed simulations including all stages of water radiolysis and clustering analyses)19,35,37,38,39,40,41,42,43. In this figure, most of the experimental data used different types of cell lines39,40,41,42,43. Only Fulford et al.38 and Leloup et al.37 used plasmid DNA as the targets. The detailed simulations shown for comparison were also based on different cell lines or targets, i.e. the simulations of Friedland et al.35 were for human fibroblasts, Kundrát et al.13 for human lymphocytes, and Mokari et al.36 for 216 bp B-DNA. The simulations of Francis et al.15 and Matsuya et al.19 were based on clustering analyses mentioned earlier in the Introduction. As expected, both our calculated SSB and DSB yields agreed well with the simulation of Kundrát et al.13, which were used to fine tune the model. A notable exception was seen at 200 MeV, where our result showed an increase in the SSB and DSB yields, similar to the experimental results of Leloup et al.37. The discrepancies observed between the different simulations and experimental data were likely due to the different cell lines or targets used for the simulations and measurements.
Proton beam qualities
Table 1 summarizes the proton beam characteristics obtained from the Monte Carlo track structure simulation and the DBSCAN model, including the LET, the C/N ratios, the Quality of Beam (QoB), and the normalized QoB. All these quantities generally decreased with the increasing proton energy, reflecting the decreasing density of energy deposition points. The exception was seen for the normalized QoB at 100–200 MeV, where the normalized QoB increased as the proton energy increased, similar to the increase of the SSB and DSB yields shown in Fig. 5.
The relationship between the newly proposed beam quality, the normalized QoB, with the commonly used beam quality, the LET, is shown in Fig. 6. From this plot, we found a non-linear relationship between both beam qualities. Again, the increase in the normalized QoB at the lowest LET proton (200 MeV protons) was observed in accordance with the increase in the SSB and DSB yields for protons of this energy.
The relationship between the LET and the normalized QoB for protons. The inset shows the zoomed-in view for the LET values not exceeding 5 keV µm⁻¹.
Carbon ions beam qualities
Table 2 summarizes the characteristics of carbon ions obtained from the Monte Carlo track structure simulation and the same DBSCAN framework used for protons. As the carbon ion energy decreased, the LET increased markedly from 18.51 to 453.37 keV µm⁻¹, and the QoB rose monotonically with the LET from 3.17 to 154.21 particle⁻¹ µm⁻¹. In contrast, the normalized QoB exhibited a non-monotonic relationship with the LET, namely, it increased with the LET, reached a maximum near 200 keV µm⁻¹, and declined at higher LET values (> 200 keV µm⁻¹), as shown in Fig. 8. The C/N ratio also increased with the LET, reflecting enhanced clustering at lower energies. Collectively, these findings indicated that while the QoB (absolute cluster yield per unit track length) scaled positively with the LET, the normalized QoB (cluster yield per unit deposited energy) peaked at intermediate LET values, indicating an optimal energy deposition range for cluster formation.
Figure 7a compares the calculated C/N ratios (calculated with ε= 2.1 nm) for carbon ions with the simulated results of Kundrát et al.13. The calculated C/N ratios systematically exceeded the reference DSB/SSB ratios across all simulated carbon ion energies, indicating an overestimation of clustering with the current parameterization optimized for protons. Figure 7b plots the normalized QoB (calculated with ε= 2.1 nm) against the simulated DSB yield reported by Kundrát et al.13. A strong linear association was observed for LET values from 18.5 to 160 keV µm⁻¹ (R² = 0.999), above which the relationship deviated from linearity with the graph turning point at ~ 200 keV µm⁻¹. In the linear regime (up to 160 keV µm⁻¹), the fitting function was y = 6.816 × 10− 12 + (4.357 ± 0.025)×10− 11 x, with the slope shallower than that for protons of (1.1498 ± 0.019)×10− 10 keV Gy− 1 Da− 1, as shown in Fig. 4b.
Comparison of the calculated C/N ratios (ε = 2.1 nm) (symbols) with the DSB/SSB ratios obtained from the simulation of Kundrát et al.13 (the dashed line) for carbon ions (a), and the relationship between the normalized Quality of Beam (QoB) and the DSB yields reported by Kundrát et al.13 for carbon ions (b). The solid red line in the panel (b) is the trend line obtained from linear fit.
Figure 8 shows the normalized QoB–LET relationship for both protons and carbon ions. As shown previously in Fig. 6, protons exhibited a non-linear but monotonic rise in normalized QoB with LET, including a slight increase at the lowest-LET condition (200 MeV protons), consistent with the accompanying increase in SSB and DSB yields. In contrast, carbon ions displayed a steeper, non-monotonic trend with LET. For carbon ions, normalized QoB increased rapidly with LET, reached a maximum near ~ 200 keV µm⁻¹, and declined at higher LET. At the same LET values above approximately 10 keV µm⁻¹, carbon ions yielded a higher normalized QoB.
RBE against LET was also plotted in Fig. 8. The RBE values were obtained from the experimental data of V79 cells at the 10% survival of Sørensen et al.44. The ordinate values for RBE were an order of magnitude higher than that of normalized QoB. Interestingly, the two datasets shared a similar shape: an initial rise, a broad maximum around 100–300 keV µm⁻¹, and a decline at higher LET. This qualitative agreement suggested that the normalized QoB captured track structure features that covaried with the RBE.
The relationship of the normalized Quality of Beam (QoB) for protons and carbon ions (left ordinate) and the experimental relative biological effectiveness (RBE) for V79 cells at the 10% survival44 (right ordinate) against the linear energy transfer (LET).
Discussion
The simulated proton track structures in this study were validated by comparing the calculated LET and microdosimetric quantities with the published data (Fig. 3), supporting the accuracy of the Geant4-DNA toolkit for simulating proton energy deposition patterns in liquid water. However, a notable discrepancy of up to 23% was observed in the LET values for low LET protons, particularly in the energy range of 50 to 200 MeV. This discrepancy could be attributed, at least in part, to the choice of the water sphere diameter used in our simulation. The water sphere radius (0.25–2.5 μm) employed as the scoring volume in this study were chosen to potentially include all secondary electrons generated by the primary particles. For high energy (low LET) protons, energetic secondary electrons could be produced, which could travel further beyond the water sphere volume. Although a larger water sphere would have improved the LET accuracy for relatively low LET protons, the computational time would have been largely increased for the clustering process. For example, increasing the radius of the working volume to 25 μm resulted in a 9.5% increase in LET for 200 MeV protons, while the computational time rose by an order of magnitude (data not shown). To balance LET accuracy with computational feasibility, we limited the radius of the water sphere to 2.5 μm for proton energies above 20 MeV.
As expected, the proton-induced DSB and SSB yields calculated in this study (Fig. 5) were generally consistent with the detailed simulations by Kundrát et al.13, particularly at proton energies below 200 MeV. The overall trends in DNA damage yields also qualitatively aligned with other detailed simulation studies, suggesting that our simplified approach effectively captured the key features of proton-induced DNA damage.
The observed increase in SSB and DSB yields at 200 MeV may reflect statistical fluctuations in the track density and cluster formation at high energies, where energy deposition events were more sparsely distributed. To assess statistical convergence, we increased the number of proton histories by ca. fivefold for 1 and 200 MeV protons. The resulting LET, QoB, and normalized QoB values remained within the 95% confidence intervals of the original simulations, as shown in Table 1. Furthermore, the Monte track structure simulations included over 10 million interaction points for each proton energy and more than 50 million for each carbon ion energy, providing a statistically robust foundation for the DBSCAN analysis.
It is to note that the agreement between simulations and experimental data depended also on cell lines or targets of interest. As shown in Fig. 5, even using the same suite of simulation codes, i.e. PARTRAC, the results of Friedland et al.35. for human fibroblasts and Kundrát et al.13 for human lymphocytes differed significantly from each other. While our DSB yields were generally lower than those observed in plasmid DNA experiments and slightly higher than those in cell line experiments, these differences could be mitigated by adjusting the empirically derived conversion factor. A slightly higher conversion factor would bring our results to closer agreement to the plasmid DNA data, while a slightly lower factor would improve the agreement with other cell line experiments. Our model may not give the ultimate solution for DNA damage modeling as the model parameters need to be adjusted to suit cell lines or targets of interest. Nevertheless, the proposed framework of how to calculate SSB and DSB yields based on direct energy deposition of radiation beams and the linear relationship between the normalized QoB and DSB yields found in this work provided a simple method to calculate radiation-induced SSB and DSB yields.
It is important to note that our clustering approach differed from previous studies using DBSCAN to analyze DNA damage. Francis et al.15 and Bernal et al.16 used the maximum distance for cluster formation (ε) of 3.2 nm and 3.3 nm, respectively, combined with a fixed probability for interaction points to fall within a sensitive region (0.16 in Francis et al.15 and 0.2 in Bernal et al.16) and a variable probability for energy deposition to induce damage. Matsuya et al.18,19 used 3.4 nm as the value of ε and did not assign energy deposition threshold for damage induction, but used individual conversion factors, \(\:{k}_{\text{S}\text{S}\text{B}}\) and \(\:{k}_{\text{D}\text{S}\text{B}}\), to convert the number of clusters or noises per energy deposited to the DSB and SSB yields, respectively. From these publications, the value of ε in the range of 3.2–3.4 nm was assumed to follow the commonly accepted definition of a DSB to be two breaks on the opposite strands with the distance not exceeding 10 base pairs (equivalent to 3.4 nm). However, since each DBSCAN model did not consider damages induced by free radicals (indirect effect) explicitly, each model had to use other model parameters to fine tune the model to experimental data, i.e. the probability for interaction points to fall within a sensitive region as used by Francis et al.15 and Bernal et al.16, and the factors \(\:{k}_{\text{S}\text{S}\text{B}}\) and \(\:{k}_{\text{D}\text{S}\text{B}}\) as used by Matsuya et al.18,19.
In our modeling approach, we obtained an optimal value of ε to be 2.1 nm, comparable to the DNA diameter, and assumed that every interaction point with energy deposition of at least 17.5 eV caused DNA damage. The parameter ε should be interpreted as an effective clustering radius that accounted for both direct and indirect effects within this simplified framework, rather than representing the physical migration distance of reactive species. This interpretation was supported by the fact that only direct energy deposition during the physical stage of water radiolysis was considered, without explicit modeling of chemical reactions or diffusion processes. The proximity of ε to the DNA diameter suggested that clustering at this scale was sufficient to reproduce observed DSB/SSB ratios, while implicitly incorporating the contribution of indirect effects into a unified parameter. Using this ε value, the normalized QoB consistently demonstrated a strong linear correlation with DSB yields reported in multiple simulation datasets (Fig. 4b), supporting its reliability for modeling DNA damage clustering induced by protons.
Another key consideration in DNA damage modeling was the energy deposition threshold required to induce damage. The fixed threshold of 17.5 eV used in this study may differ from models that employed energy-dependent probabilities for damage induction 14,35. Nevertheless, this value has been validated against experimental data 45,46, and, as demonstrated in this work, provided a robust basis for evaluating DNA damage and beam quality within the DBSCAN clustering framework.
Also, the assumption that the cluster-to-noise (C/N) ratio directly corresponded to the DSB/SSB ratio, despite being relatively simplified, provided a valuable means of connecting the spatial clustering of energy deposition to the type of DNA damage. Not only that, the linear relationship between normalized QoB and DSB yields enabled us to calculate SSB and DSB yields using a single conversion factor. This simplification allowed for a more streamlined calculation while still capturing the essential relationship between energy deposition clustering and DNA damage induction.
Another interesting point of the proposed modeling approach was that it introduced the Quality of Beam (QoB), defined as the average number of clusters per proton per µm, which exhibited a dependence on proton energy, and thus LET (Table 1). Furthermore, the normalized QoB emerged as a promising tool for proton beam quality assessment as it was linearly correlated with DSB yields (Fig. 4b), in contrast to the non-linear relationship between the LET and DSB yields implied by Fig. 6.
To assess the generalizability of the DBSCAN framework, we applied the same clustering parameters optimized for protons (Ethres = 17.5 eV, MinPts = 2, and ε = 2.1 nm) to carbon ions. For carbon ions, the QoB increased monotonically with LET, while the normalized QoB peaked around 200 keV µm⁻¹ and declined at higher LET values (Table 2 and Fig. 8). Although a strong linear relationship between normalized QoB and DSB yields was still observed in the LET range of 18.5–160 keV µm⁻¹ (Fig. 7b), the slope of this relationship was ca. 2.5 times lower than that observed for protons. Additionally, the calculated C/N ratios for carbon ions consistently exceeded the reported DSB/SSB ratios (Fig. 7a), indicating that the proton-based parameterization overestimated clustering for carbon ions and limited the accuracy of SSB yield predictions. These findings indicated that DBSCAN parameters optimized for protons could not be directly applied to carbon ions. Instead, ion-specific model parameters, such as, ε, Ethres, MinPts, and other tuning parameters, would be required to accurately model DNA damage induced by heavy ions. Nevertheless, the strong linear correlation between normalized QoB and DSB yields observed for carbon ions with LET below 200 keV µm⁻¹ (Fig. 7b) suggested that within this LET range, the current DBSCAN framework may still provide reasonable estimates of DNA damage clustering.
A notable feature observed in the carbon ion data was the distinct peak in normalized QoB near 200 keV µm⁻¹, reflecting the overkill effect, an aspect not captured by LET alone. At very high LET, excess energy was deposited in regions already containing damage, causing new ionizations to merge with existing clusters rather than forming new ones. Consequently, the number of clusters per unit deposited energy (normalized QoB) decreased with increasing LET. The shape of the normalized QoB-LET curve (Fig. 8) effectively captured this overkill behavior, indicating that normalized QoB could reveal meaningful physical and biological characteristics of ion tracks beyond what LET could convey.
Furthermore, the normalized QoB-LET curve for both protons and carbon ions exhibited a pattern similar to published RBE–LET data for V79 cells at the 10% survival (Fig. 8). For a given LET, normalized QoB values were lower than RBE values by approximately a factor of 10. This consistent offset suggested that normalized QoB may serve as a more biologically relevant indicator of radiation effectiveness than LET alone as it captured key features of DNA damage clustering that correlated with biological outcomes.
In terms of computational efficiency, the DBSCAN-based DNA damage modeling relied solely on the physical stage of water radiolysis, resulting in simulation times approximately 2–10 times shorter than those required for full three-stage radiolysis modeling, depending on the particle pathlength. In addition to speed, normalized QoB offered advantages over conventional metrics such as LET, as it captured the spatial clustering of energy deposition events at the nanometer scale rather than compressing track structure into a single average value. This spatial sensitivity explained the stronger and more consistent correlation between normalized QoB and DSB yields, and its ability to reproduce the overkill effect observed at high LET, while LET alone cannot adequately represent.
Several limitations of this study should be acknowledged. First, the analysis was restricted to protons and carbon ions. Within the current DBSCAN framework and parameter settings, normalized QoB showed a linear correlation with DSB yields in the absence of the overkill effect, and qualitatively matched RBE trends over the full LET range for both ion types. However, accurate prediction of absolute SSB and DSB yields required ion-specific DBSCAN parameters. Further validation with additional particle species will be necessary to strengthen the generalizability of this approach.
Second, each DBSCAN-identified cluster was assumed to represent a single DSB, regardless of its complexity. While this may be a reasonable approximation for protons, complex lesions are more frequent at high LET and are associated with slower repair kinetics, potentially contribute to increased RBE47. A modified DBSCAN model that incorporates damage complexity should be explored for integration with subsequent repair kinetic modeling.
Finally, although the current DBSCAN framework may lose accuracy in predicting absolute DNA damage yield at very high LET (Fig. 7a), and comparisons with retrospective datasets should be interpreted with caution (Fig. 5), normalized QoB derived solely from physical track structures demonstrated biological relevance (Fig. 8). Future studies should investigate its relationship with RBE for various biological endpoints, cell lines, and ion species to further establish its utility.
Conclusions
This study presents a simplified framework for characterizing early DNA damage induced by 0.5–200 MeV protons using Monte Carlo simulated physical track structures and a DBSCAN clustering model. The generalizability of the framework was evaluated using carbon ions, representing high LET radiation. The model employed a fixed energy deposition threshold of 17.5 eV for damage induction and equated the cluster-to-noise ratio for protons to the DSB/SSB ratio. The minimum clustering radius (ε) was optimized based on proton data, and a new beam quality metric, the normalized Quality of Beam (QoB), was introduced. Normalized QoB showed a strong linear correlation with DSB yields for protons and for carbon ions with LET below 200 keV µm− 1. This newly introduced beam quality metric successfully reproduced the trend of RBE across the full LET range, including the overkill effect. For protons, the DBSCAN framework enabled direct estimation of DSB and SSB yields using normalized QoB and a single conversion factor. For heavier ions, ion-specific parameters are required to ensure accurate predictions of absolute DNA damage yields. Nonetheless, normalized QoB offered a more biologically meaningful measure of radiation quality than conventional metrics such as LET.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author upon reasonable request.
References
Paganetti, H. et al. Report of the AAPM TG-256 on the relative biological effectiveness of proton beams in radiation therapy. Med. Phys. 46, e53–e78 (2019).
Scholz, M., Kellerer, A. M., Kraft-Weyrather, W. & Kraft, G. Computation of cell survival in heavy ion beams for therapy. Radiat. Environ. Biophys. 36, 59–66 (1997).
Hawkins, R. B. A microdosimetric-kinetic model of cell death from exposure to ionizing radiation of any LET, with experimental and clinical applications. Int. J. Radiat. Biol. 69, 739–755 (1996).
Paganetti, H. Relative biological effectiveness (RBE) values for proton beam therapy. Variations as a function of biological endpoint, dose, and linear energy transfer. Phys. Med. Biol. 59, R419–R472 (2014).
Underwood, T. S. A. et al. A systematic review of clinical studies on variable proton Relative Biological Effectiveness (RBE). Radiother. Oncol. 175, 79–92 (2022).
Giantsoudi, D. et al. Linear Energy Transfer-Guided Optimization in Intensity Modulated Proton Therapy: Feasibility Study and Clinical Potential. Int. J. Radiat. Oncol. 87, 216–222 (2013).
McIntyre, M., Wilson, P., Gorayski, P. & Bezak, E. A systematic review of LET-guided treatment plan optimisation in proton therapy: Identifying the current state and future needs. Cancers 15, 4268 (2023).
Kalholm, F., Grzanka, L., Toma-Dasu, I. & Bassler, N. Modeling RBE with other quantities than LET significantly improves prediction of in vitro cell survival for proton therapy. Med. Phys. 50, 651–659 (2023).
Wedenberg, M., Lind, B. K. & Hårdemark, B. A model for the relative biological effectiveness of protons: The tissue specific parameter α/β of photons is a predictor for the sensitivity to LET changes.. Acta Oncol. 52, 580–588 (2013).
McNamara, A. L., Schuemann, J. & Paganetti, H. A phenomenological relative biological effectiveness (RBE) model for proton therapy based on all published in vitro cell survival data.. Phys. Med. Biol. 60, 8399–8416 (2015).
McMahon, S. J. & Prise, K. M. Mechanistic modelling of radiation responses. Cancers 11, 205 (2019).
Nikjoo, H. et al. Radiation track, DNA damage and response - A review.. Rep. Prog. Phys. 79, 116601 (2016).
Kundrát, P. et al. Analytical formulas representing track-structure simulations on DNA damage induced by protons and light ions at radiotherapy-relevant energies.. Sci. Rep. 10, 15775 (2020).
Sakata, D. et al. Fully integrated Monte Carlo simulation for evaluating radiation induced DNA damage and subsequent repair using Geant4-DNA.. Sci. Rep. 10, 20788 (2020).
Francis, Z., Villagrasa, C. & Clairand, I. Simulation of DNA damage clustering after proton irradiation using an adapted DBSCAN algorithm. Comput. Methods Programs Biomed. 101, 265–270 (2011).
Bernal, M. A. et al. Track structure modeling in liquid water: A review of the Geant4-DNA very low energy extension of the Geant4 Monte Carlo simulation toolkit.. Physica Medica 31, 861–874 (2015).
Yoshii, Y., Sasaki, K., Matsuya, Y. & Date, H. Cluster analysis for the probability of DSB site induced by electron tracks. Nucl. Instrum. Methods Phys. Res. B Beam Interact. Mater. Atoms 350, 55–59 (2015).
Matsuya, Y. et al. Modeling of yield estimation for DNA strand breaks based on Monte Carlo simulations of electron track structure in liquid water. J. Appl. Phys. 126, 124701 (2019).
Matsuya, Y., Kai, T., Parisi, A., Yoshii, Y. & Sato, T. Application of a simple DNA damage model developed for electrons to proton irradiation. Phys. Med. Biol. 67, 215017 (2022).
Zepeda-Mendoza, M. L. & Resendis-Antonio, O. Hierarchical Agglomerative Clustering. In Encyclopedia of Systems Biology 886–887 (Springer New York, 2013).
Lloyd, S. Least squares quantization in PCM. IEEE Trans. Inf. Theory. 28, 129–137 (1982).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: A K-means clustering algorithm. Applied Statistics 28, 100–108 (1979).
Ng, R. T. & Han, J. Efficient and Effective Clustering Methods for Spatial Data Mining. In Proceedings of the 20th International Conference on Very Large Data Bases, 144–155 (Morgan Kaufmann Publishers Inc., 1994).
Ester, M., Kriegel, H. P., Sander, J., Xu, X. A. & Density-Based Algorithm for Discovering Clusters in Large Spatial Databases with Noise. In Proceedings of the Second International Conference on Knowledge Discovery and Data Mining, 226–231 (AAAI Press, 1996).
Ester, M., Kriegel, H. P., Sander, J. & Xu, X. Density-Connected Sets and their Application for Trend Detection in Spatial Databases. In Proceedings of the Third International Conference on Knowledge Discovery and Data Mining, 10–15 (AAAI Press, 1997).
Incerti, S. et al. Geant4-DNA example applications for track structure simulations in liquid water: A report from the Geant4-DNA project. Med. Phys. 45, e722–e739 (2018).
ICRU Report 36, Microdosimetry. (International Commission on Radiation Units and Measurements, 1983).
Chaibura, S., Liamsuwan, T. & Autsavapromporn, N. Simple radiation-induced DNA damage modeling approach for proton therapy. AIP Conf. Proc. 3210, 050002 (2024).
Nikjoo, H., O’Neill, P., Terrissol, M. & Goodhead, D. T. Quantitative modelling of DNA damage using Monte Carlo track structure method. Radiat. Environ. Biophys. 38, 31–38 (1999).
Mladenova, V., Mladenov, E., Stuschke, M. & Iliakis, G. DNA damage clustering after ionizing radiation and consequences in the processing of chromatin breaks. Molecules 27, 1540 (2022).
Rezaee, M. & Adhikary, A. The effects of particle LET and fluence on the complexity and frequency of clustered DNA damage. DNA 4, 34–51 (2024).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
ICRU Report 49, Stopping Power and Ranges for Protons and Alpha Particles. (International Commission on Radiation Units and Measurements, 1984).
Liamsuwan, T., Uehara, S., Emfietzoglou, D. & Nikjoo, H. Physical and biophysical properties of proton tracks of energies 1 kev to 300 MeV in water. Int. J. Radiat. Biol. 87, 141–160 (2011).
Friedland, W., Jacob, P., Bernhardt, P., Paretzke, H. G. & Dingfelder, M. Simulation of DNA damage after proton irradiation. Radiat. Res. 159, 401–410 (2003).
Mokari, M., Alamatsaz, M. H., Moeini, H. & Taleei, R. A simulation approach for determining the spectrum of DNA damage induced by protons. Phys. Med. Biol. https://doi.org/10.1088/1361-6560/aad7ee (2018).
Leloup, C. et al. Evaluation of lesion clustering in irradiated plasmid DNA. Int. J. Radiat. Biol. 81, 41–54 (2005).
Fulford, J., Nikjoo, H., Goodhead, D. T. & O’Neill, P. Yields of SSB and DSB induced in DNA by Al K ultrasoft X-rays and α-particles: Comparison of experimental and simulated yields. Int. J. Radiat. Biol. 77, 1053–1066 (2001).
Frankenberg, D. et al. Induction of DNA double-strand breaks by 1 H and 4 He ions in primary human skin fibroblasts in the LET range of 8 to 124 keV/mm. Radiat. Res. 151540. (1999).
Belli, M., Cherubini, R. & Dalla Vecch, M. DNA DSB induction and rejoining in V79 cells irradiated with light ions: A constant field gel electrophoresis study. Int. J. Radiat. Biol. 76, 1095–1104 (2000).
Belli, M. et al. DNA fragmentation in mammalian cells exposed to various light ions. Adv. Space Res. 27, 393–399 (2001).
Campa, A. et al. DNA DSB induced in human cells by charged particles and gamma rays: Experimental results and theoretical approaches. Int. J. Radiat. Biol. 81, 841–854 (2005).
Botchway, S. W., Stevens, D. L., Hill, M. A., Jenner, T. J. & O’Neill, P. Induction and rejoining of DNA double-strand breaks in Chinese hamster V79-4 cells irradiated with characteristic aluminum K and copper L ultrasoft X rays. Radiat. Res. 148317. (1997).
Sørensen, B. S., Overgaard, J. & Bassler, N. In vitro RBE-LET dependence for multiple particle types. Acta Oncol. 50, 757–762 (2011).
Charlton, D. E. & Humm, J. L. A method of calculating initial DNA strand breakage following the decay of incorporated 125 I. Int. J. Radiat. Biol. 53, 353–365 (1988).
Martin, R. F. & Haseltine, W. A. Range of radio chemical damage to DNA with decay of iodine-125. Science 213, 896–898 (1981).
Nikitaki, Z. et al. Key biological mechanisms involved in high-LET radiation therapies with a focus on DNA damage and repair. Expert Rev. Mol. Med. 24, e15 (2022).
Acknowledgements
SC acknowledges the scholarship of the Development and Promotion of Science and Technology Talent Project (DPST).
Funding
This work was supported by the Thematic Funding of Princess Srisavangavadhana College of Medicine, Chulabhorn Royal Academy of the Fiscal Year 2022 and Chulabhorn Royal Academy (Fundamental Fund by National Science Research and Innovation Fund (NSRF): fiscal year 2025) (FRB680064/0240 Project code 206665).
Author information
Authors and Affiliations
Contributions
Both authors contributed to the study conception and design. The study supervision was done by TL. Simulation and model development were done by SC. Both authors contributed to analysis and interpretation of the data, as well as drafting, critically reviewing and approving the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chaibura, S., Liamsuwan, T. Predicting DNA damage yields and assessing beam quality for protons and carbon ions using a DBSCAN algorithm. Sci Rep 16, 10327 (2026). https://doi.org/10.1038/s41598-026-40571-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-40571-x