Abstract
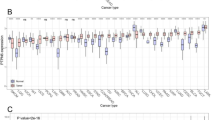
The role of the prostaglandin F2 receptor negative regulator (PTGFRN) in tumor biology remains incompletely understood. This study aimed to perform a comprehensive pan-cancer analysis to elucidate the functions of PTGFRN in tumor progression and its potential immunomodulatory effects. Utilizing data from the cancer genome atlas (TCGA) and the genotype-tissue expression (GTEx) project, we analyzed PTGFRN expression profiles, genetic alterations (including mutations, copy number variations, and DNA methylation), and its prognostic significance across multiple cancer types. Pathway enrichment analysis was conducted using the R package “clusterProfiler.” The correlation between PTGFRN expression and immune cell infiltration levels within the tumor microenvironment was assessed via the TIMER2 database. PTGFRN was significantly overexpressed in a wide range of cancers, and its elevated expression was consistently associated with poorer patient prognosis. Furthermore, pan-cancer analysis revealed that PTGFRN expression is linked to an immunosuppressive tumor microenvironment, showing a positive correlation with immunosuppressive cells such as cancer-associated fibroblasts and a negative correlation with anti-tumor effector cells like CD8⁺ T cells. Functional validation in lung adenocarcinoma (LUAD) cells confirmed that PTGFRN acts as an oncogene, enhancing proliferative, migratory, and invasive capabilities. Our findings establish PTGFRN as a potential prognostic biomarker across cancer types. Its overexpression is indicative of an immunosuppressive tumor microenvironment, positioning PTGFRN as a promising therapeutic target for cancer immunotherapy.
Similar content being viewed by others
Data availability
The datasets analyzed during the current study are available in the TIMER2.0(http://timer.cistrome.org/),UCSC(https://xenabrowser.net/), UALCAN(https://ualcan.path.uab.edu/analysis-prot.html), GEPIA2(http://gepia2.cancer-pku.cn/analysis), cBioPortal(https://www.cbioportal.org/), CPTAC database (https://proteomics.cancer.gov/programs/cptac), CCLE(https://sites.broadinstitute.org/ccle), TISIDB(cis.hku.hk/TISIDB/) StarBase(https://rnasysu.com/encori/), and GSCALite(https://guolab.wchscu.cn/GSCA/#/).
References
Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 12 (1), 31–46 (2022).
Mala, U., Baral, T. K. & Somasundaram, K. Integrative analysis of cell adhesion molecules in glioblastoma identified prostaglandin F2 receptor inhibitor (PTGFRN) as an essential gene. BMC Cancer. 22 (1), 642 (2022).
Ding, Y. et al. EWI2 and its relatives in Tetraspanin-enriched membrane domains regulate malignancy. Oncogene 42 (12), 861–868 (2023).
Marquez, J. et al. Effect of PTFGRN Expression on the Proteomic Profile of A431 Cells and Determination of the PTGFRN Interactome. ACS Omega. 9 (12), 14381–14387 (2024).
Aguila, B. et al. The Ig superfamily protein PTGFRN coordinates survival signaling in glioblastoma multiforme. Cancer Lett. 462, 33–42 (2019).
Li, T. et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 48 (W1), W509–W514 (2020).
Charoentong, P. et al. Pan-cancer Immunogenomic Analyses Reveal Genotype-Immunophenotype Relationships and Predictors of Response to Checkpoint Blockade. Cell. Rep. 18 (1), 248–262 (2017).
Ru, B. et al. TISIDB: an integrated repository portal for tumor-immune system interactions. Bioinformatics 35 (20), 4200–4202 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47 (W1), W556–W560 (2019).
Chandrashekar, D. S. et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia 25, 18–27 (2022).
Suhas, V., Vasaikar, P., Straub, J., Wang, B. & Zhang LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res. 46 (Issue D1), D956–D963 (2018).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42 (Database issue), D92–D97 (2014).
Liu, C. J. et al. GSCA: an integrated platform for gene set cancer analysis at genomic, pharmacogenomic and immunogenomic levels. Brief. Bioinform. 24 (1), bbac558 (2023).
Chen, H. W. et al. Prostaglandin F2 receptor inhibitor overexpression predicts advanced who grades and adverse prognosis in human glioma tissue. Chin. J. Physiol. 65 (2), 93–102 (2022).
Jiang, X., Zhang, J. & Huang, Y. Tetraspanins in cell migration. Cell. Adh Migr. 9 (5), 406–415 (2015).
Colin, S. et al. A truncated form of CD9-partner 1 (CD9P-1), GS-168AT2, potently inhibits in vivo tumour-induced angiogenesis and tumour growth. Br. J. Cancer. 105 (7), 1002–1011 (2011).
Guilmain, W. et al. CD9P-1 expression correlates with the metastatic status of lung cancer, and a truncated form of CD9P-1, GS-168AT2, inhibits in vivo tumour growth. Br. J. Cancer. 104 (3), 496–504 (2011).
Aguila, B. et al. The Ig superfamily protein PTGFRN coordinates survival signaling in glioblastoma multiforme. Cancer Lett. 462, 33–42 (2019).
Marquez, J. et al. Effect of PTFGRN Expression on the Proteomic Profile of A431 Cells and Determination of the PTGFRN Interactome. ACS Omega. 9 (12), 14381–14387 (2024).
Aliazis, K. et al. The tumor microenvironment’s role in the response to immune checkpoint blockade. Nat. Cancer. 6 (6), 924–937 (2025).
Huang, G. et al. Tumor suppressor miR-33b-5p regulates cellular function and acts a prognostic biomarker in RCC. Am. J. Transl Res. 12 (7), 3346–3360 (2020).
Rius, F. E. et al. Genome-wide promoter methylation profiling in a cellular model of melanoma progression reveals markers of malignancy and metastasis that predict melanoma survival. Clin. Epigenetics. 14 (1), 68 (2022).
Zhang, S. et al. Truncated PD1 Engineered Gas-Producing Extracellular Vesicles for Ultrasound Imaging and Subsequent Degradation of PDL1 in Tumor Cells. Adv. Sci. (Weinh). 11 (12), e2305891 (2024).
Janes, P. W., Vail, M. E., Ernst, M. & Scott, A. M. Eph Receptors in the Immunosuppressive Tumor Microenvironment. Cancer Res. 81 (4), 801–805 (2021).
Zhang, P. et al. Integrating machine learning and single-cell analysis to uncover lung adenocarcinoma progression and prognostic biomarkers. J. Cell. Mol. Med. 28 (13), e18516 (2024).
Gong, Z. et al. Machine learning identifies TIME subtypes linking EGFR mutations and immune states in lung adenocarcinoma. NPJ Digit. Med. 8 (1), 796 (2025).
El-Gamal, D. et al. PKC-β as a therapeutic target in CLL: PKC inhibitor AEB071 demonstrates preclinical activity in CLL. Blood 124 (9), 1481–1491 (2014).
Yakkala, P. A., Penumallu, N. R., Shafi, S. & Kamal, A. Prospects of Topoisomerase Inhibitors as Promising Anti-Cancer Agents. Pharmaceuticals (Basel). 16 (10), 1456 (2023).
Marquardt, V., Theruvath, J. & Pauck, D. etal. Tacedinaline (CI-994), a class I HDAC inhibitor, targets intrinsic tumor growth and leptomeningeal dissemination in MYC-driven medulloblastoma while making them susceptible to anti-CD47-induced macrophage phagocytosis via NF-kB-TGM2 driven tumor inflammation. J. Immunother Cancer. 11 (1), e005871 (2023).
Thomas, A. & Pommier, Y. Targeting Topoisomerase I in the Era of Precision Medicine. Clin. Cancer Res. 25 (22), 6581–6589 (2019).
Acknowledgements
The datasets for this study can be found in the public databases TCGA, GTEx, TIMER 2.0, GDSA, and cBioportal.
Funding
The present study was supported by the Hebei Medical Science Research Project (No. 20210980).
Author information
Authors and Affiliations
Contributions
Zhao Qi and Dong tianjian performed bioinformatics analysis, and Zhao Qi, Sheng qingyu, Duan zhuoning, Wang xiangming and Dong tinajian co-authored and completed the manuscript. All authors gave final approval to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mu, N., Dong, T., Sheng, Q. et al. Pan-cancer analysis reveals the oncogenic and immunomodulatory roles of PTGFRN across human cancers. Sci Rep (2026). https://doi.org/10.1038/s41598-026-41027-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-41027-y