Abstract
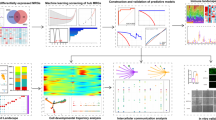
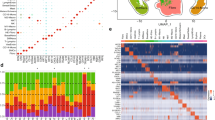
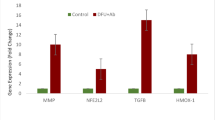
Diabetic foot ulcer is a severe complication of diabetes, characterized by impaired wound healing and immune dysregulation. Although Astragalus membranaceus (AM) has been widely used and reported to exert beneficial effects in diabetic complications, its underlying mechanisms of action in DFU remain incompletely understood. This study integrated single-cell RNA sequencing (scRNA-seq) data from a public bioinformatic cohort (GSE245703; 4 non-diabetic foot ulcer (NFU) and 5 diabetic foot ulcer (DFU) samples) with network pharmacology to explore potential molecular mechanisms by which AM may be involved in DFU pathology. scRNA-seq analysis identified ten major cell types within the DFU microenvironment and revealed significant macrophage heterogeneity. Network pharmacology identified 14 active compounds in AM and their predicted targets, some of which overlapped with macrophage-associated differentially expressed genes. Molecular docking suggested strong binding affinities between selected AM compounds and macrophage-associated hub genes. qPCR validation in a clinical cohort (6 NFU and 9 DFU patients) confirmed differential expression of several candidate hub genes overlapping with predicted AM targets. Collectively, these results provide a single-cell–resolved, systems-level framework that links AM components to macrophage-associated molecular processes in DFU, offering a hypothesis-generating basis for future functional and translational studies.
Similar content being viewed by others
Data availability
The scRNA-seq data used in this study were obtained from the Gene Expression Omnibus database under accession number GSE245703 and are available at the following URL: https://identifiers.org/geo: GSE245703.
References
Yang, L., Rong, G. C. & Wu, Q. N. Diabetic foot ulcer: Challenges and future. World J. Diabetes. 13, 1014 (2022).
Aljohary, H., Murad, M. A., Alfkey, R. & Elgohary, S. Stepping up to the Challenge: Confronting the Global Burden of Diabetic Foot Disease. in Diabetic Foot - Advanced Methods of Management (Raghav, A., ed.) (IntechOpen, 2025). https://doi.org/10.5772/intechopen.1012471
Artasensi, A., Pedretti, A., Vistoli, G. & Fumagalli, L. Type 2 diabetes mellitus: A review of multi-target drugs. Molecules 25, 1987 (2020).
Armstrong, D. G. et al. Five year mortality and direct costs of care for people with diabetic foot complications are comparable to cancer. J. Foot Ankle Res. 13, 16 (2020).
Wang, F. et al. Recent advances in the adjunctive management of diabetic foot ulcer: Focus on noninvasive technologies. Med. Res. Rev. 44, 1501–1544 (2024).
Zamanifard, M. et al. Healing of diabetic foot ulcer with topical and oral administrations of herbal products: A systematic review and meta-analysis of randomized controlled trials. Int. Wound J. 21, e14760 (2024).
Dawi, J. et al. Diabetic foot ulcers: Pathophysiology, immune dysregulation, and emerging therapeutic strategies. Biomedicines 13, 1076 (2025).
Gan, J. et al. Immune microenvironment and fibroblast subpopulation in diabetic wound healing. Iscience 28, 113440 (2025).
Aitcheson, S. M., Frentiu, F. D., Hurn, S. E. & Edwards, K. Murray, R. Z. Skin wound healing: Normal macrophage function and macrophage dysfunction in diabetic wounds. Molecules 26, 4917 (2021).
Song, J., Wu, Y., Chen, Y., Sun, X. & Zhang, Z. Epigenetic regulatory mechanism of macrophage polarization in diabetic wound healing. Mol. Med. Rep. 31, 1–20 (2025).
Nazari, M., Taremi, S., Elahi, R., Mostanadi, P. & Esmeilzadeh, A. Therapeutic properties of M2 macrophages in chronic wounds: an innovative area of biomaterial-assisted M2 macrophage targeted therapy. Stem Cell. Rev. Rep. 21, 390–422 (2025).
Chen, L. et al. Single-cell RNA sequencing in the context of neuropathic pain: Progress, challenges, and prospects. Translational Research: J. Lab. Clin. Med. 251, 96–103 (2023).
Xiong, B. B. et al. Macrophage polarization in disease therapy: Insights from astragaloside IV and cycloastragenol. Front Pharmacol. 16, (2025).
Zhen, Z. et al. Astragalus polysaccharide improves diabetic ulcers by promoting M2-polarization of macrophages to reduce excessive inflammation via the β-catenin/ NF-κB axis at the late phase of wound-healing. Heliyon 10, e24644 (2024).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2019).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Nogales, C. et al. Network pharmacology: curing causal mechanisms instead of treating symptoms. Trends Pharmacol. Sci. 43, 136–150 (2022).
Li, T., Wei, D., Wang, J. & Gao, L. Revealing the Multi-Target Mechanisms of Fespixon Cream in Diabetic Foot Ulcer Healing: Integrated Network Pharmacology, Molecular Docking, and Clinical RT-qPCR Validation. Curr. Issues Mol. Biol. 47, 485 (2025).
Li, J. et al. Exploring the high-quality ingredients and mechanisms of da chuanxiong formula in the treatment of neuropathic pain based on network pharmacology, molecular docking, and molecular dynamics simulation. Biomed. Pharmacother. 178, 117195 (2024).
Jin, Q. et al. A network pharmacology to explore the mechanism of astragalus membranaceus in the treatment of diabetic retinopathy. Evidence-Based Complementary Alternative Med. 8878569 (2020).
Daina, A., Michielin, O. & Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 7, 42717 (2017).
Kanehisa, M., Sato, Y., Furumichi, M., Morishima, K. & Tanabe, M. New approach for understanding genome variations in KEGG. Nucleic Acids Res. 47, D590–D595 (2019).
Li, C. et al. A Systems pharmacology approach for identifying the multiple mechanisms of action for the Rougui-Fuzi herb pair in the treatment of cardiocerebral vascular diseases. Evid Based Complement Alternat Med 5196302 (2020).
Zhao, S. et al. Single-cell RNA sequencing reveals a new mechanism of endothelial cell heterogeneity and healing in diabetic foot ulcers. Biol. Direct. 20, 34 (2025).
Chen, R. & Zou, L. Combined analysis of single-cell sequencing and bulk transcriptome sequencing reveals new mechanisms for non-healing diabetic foot ulcers. PLOS One. 19, e0306248 (2024).
Liu, Z. et al. Spatiotemporal single-cell roadmap of human skin wound healing. Cell. Stem Cell. 32, 479–498e8 (2025).
Asunama, A. S. R., Nambi, N., Radhakrishnan, L., Prasad, M. K. & Ramkumar, K. M. Neutrophil migration is a crucial factor in wound healing and the pathogenesis of diabetic foot ulcers: Insights into pharmacological interventions. Curr. Vasc. Pharmacol. 23, 98–112 (2025).
Armstrong, D. G., Tan, T. W., Boulton, A. J. M. & Bus, S. A. Diabetic foot ulcers: A review. JAMA 330, 62–75 (2023).
McDermott, K., Fang, M., Boulton, A. J. M., Selvin, E. & Hicks, C. W. Etiology, epidemiology, and disparities in the burden of diabetic foot ulcers. Diabetes Care. 46, 209–221 (2023).
Dong, Y. et al. Single-cell RNA-seq in diabetic foot ulcer wound healing. Wound Repair. Regeneration. 32, 880–889 (2024).
Ma, J. et al. Single-cell RNA-seq analysis of diabetic wound macrophages in STZ-induced mice. J. Cell. Communication Signal. 17, 103–120 (2023).
Lin, C. W. et al. New horizons of macrophage immunomodulation in the healing of diabetic foot ulcers. Pharmaceutics 14, 2065 (2022).
Liu, Y. et al. Single-cell RNA sequencing reveals the impaired epidermal differentiation and pathological microenvironment in diabetic foot ulcer. Burns Trauma. 13, tkae065 (2025).
Mir, R. A. et al. Molecular Pathways and Immunometabolic Processes. in Macrophages - Molecular Pathways and Immunometabolic Processe IntechOpen, (2024). https://doi.org/10.5772/intechopen.1007012
Pandey, S., Anshu, T., Maharana, K. C. & Sinha, S. Molecular insights into diabetic wound healing: focus on wnt/β-catenin and MAPK/ERK signaling pathways. Cytokine 191, 156957 (2025).
Wu, X. et al. Macrophage polarization in diabetic wound healing. Burns Trauma. 10, tkac051 (2022).
Tratnig-Frankl, M. et al. Hepatocyte growth factor modulates corneal endothelial wound healing in vitro. Int. J. Mol. Sci. 25, 9382 (2024).
Shi, Z., Yao, C., Shui, Y., Li, S. & Yan, H. Research progress on the mechanism of angiogenesis in wound repair and regeneration. Front. Physiol. 14, 1284981 (2023).
Okonkwo, U. A. & DiPietro, L. A. Diabetes and wound angiogenesis. Int. J. Mol. Sci. 18, 1419 (2017).
Deepika & Maurya, P. K. Health benefits of quercetin in age-related diseases. Molecules 27, 2498 (2022).
Nazarnezhad, S., Nasiri, S. N., Gorgani, S. & Aghajanloo, B. & Seyedasli, N. Quercetin-loaded biocomposites for wound healing applications: from therapeutic efficacy to delivery platforms. Phytochem Rev. (2025).
Abid, H. M. U. et al. Wound-healing and antibacterial activity of the quercetin–4-formyl phenyl boronic acid complex against bacterial pathogens of diabetic foot ulcer. ACS Omega. 7, 24415–24422 (2022).
Zhang, Z., Wang, L., Li, X., Miao, Y. & Li, D. Integrating network pharmacology, molecular docking and experimental validation to explore the pharmacological mechanisms of quercetin against diabetic wound. Int. J. Med. Sci. 21, 2837–2850 (2024).
Zhang, L. J. et al. New isoflavonoid glycosides and related constituents from astragali radix (astragalus membranaceus) and their inhibitory activity on nitric oxide production. J. Agric. Food Chem. 59, 1131–1137 (2011).
Luo, W. et al. Bulk and single-cell transcriptome analyses unravel gene signatures of mitochondria-associated programmed cell death in diabetic foot ulcer. J. Cell. Mol. Med. 28, e70319 (2024).
Wei, H. et al. Structures of p53/BCL-2 complex suggest a mechanism for p53 to antagonize BCL-2 activity. Nat. Commun. 14, 4300 (2023).
Tang, Y. B. et al. Non-coding RNAs: Role in diabetic foot and wound healing. World J. Diabetes. 13, 1001–1013 (2022).
Hu, J. et al. Long noncoding RNA GAS5 regulates macrophage polarization and diabetic wound healing. J. Invest. Dermatol.. 140, 1629–1638 (2020).
Yan, C. et al. Emerging roles of long non-coding RNAs in diabetic foot ulcers. Dabetes, Metabolic Syndrome and Obesity: Targets Ther. 14, 2549–2560 (2021).
He, W. et al. The role of p53 in regulating chronic inflammation and PANoptosis in diabetic wounds. Aging Disease. 16, 373–393 (2024).
Wang, X. et al. Identification and verification of ferroptosis-related genes in diabetic foot using bioinformatics analysis. Int. Wound J. 20, 3191–3203 (2023).
Rai, V. Transcriptomics revealed differentially expressed transcription factors and MicroRNAs in human diabetic foot ulcers. Proteomes 12, 32 (2024).
Chen, J. et al. Targeting matrix metalloproteases in diabetic wound healing. Front. Immunol. 14, (2023).
Chang, M. & Nguyen, T. T. Strategy for treatment of infected diabetic foot ulcers. Acc. Chem. Res. 54, 1080–1093 (2021).
Zhang, W. Q. et al. Effect of matrix metalloproteinases on the healing of diabetic foot ulcer: A systematic review. J. Tissue Viability. 32, 51–58 (2023).
Fu, K. et al. Role of matrix metalloproteinases in diabetic foot ulcers: Potential therapeutic targets. Front. Pharmacol. 13, 1050630 (2022).
Funding
This study was supported by the Yunnan Provincial Science and Technology Department (202301BE070001-039), and the Project for Establishing a Flagship Department of Integrated Chinese and Western Medicine.
Author information
Authors and Affiliations
Contributions
X.L. conceived and designed the study, performed data analysis, and drafted the manuscript. C.H. and Y.D. contributed to methodology and validation. G.Z. and N.C. supervised the study and revised the manuscript. Y.N., Y.L. and R.Z. collected samples. N.C. and Y.Y. ainistered the project and provided resources. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Informed consent
Informed consent was obtained from all subjects involved in the study.
Institutional Review Board Statement
The study was conducted in accordance with the Declaration of Helsinki and approved by the Ethics Committee of Anning First People’s Hospital (Approval No: Lun Shen 2025-026 (Other)-01).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, X., Dong, Y., Huang, C. et al. Mechanism of action of Astragalus membranaceus for treating diabetic foot ulcers based on single-cell RNA sequencing data and network pharmacology. Sci Rep (2026). https://doi.org/10.1038/s41598-026-41921-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-41921-5