Abstract
Accurate early detection of clinically significant prostate cancer is crucial for improving patient outcomes. However, traditional diagnostic methods such as Digital Rectal Exam and Prostate-Specific Antigen (PSA) tests often lack the sensitivity and specificity needed for effective diagnosis. This study presents an AI-based approach for csPCa classification using MRI data, incorporating both the PI-CAI Challenge dataset and a newly compiled, diverse BIMCV Prostate dataset comprising over 9000 MRI sessions from 16 healthcare centers in the Valencian Region. The methodology includes a robust preprocessing pipeline, featuring prostate segmentation with a custom-trained nnUNet model, and utilizes a 3D variant of EfficientNet-B7. To ensure robustness, we employed a transfer learning strategy where five models pretrained on PI-CAI were fine-tuned on the BIMCV dataset and aggregated using a stacked meta-learner. This ensemble approach yielded a Receiver Operating Characteristic Area Under the Curve of 0.816 on the independent hold-out set, significantly outperforming a non-pretrained baseline (AUC 0.71). Furthermore, we demonstrated that synthesizing missing ADC maps using a mono-exponential model serves as an effective data augmentation strategy, preventing data loss without introducing domain shift. Interpretability techniques such as occlusion sensitivity and guided backpropagation were employed to provide insights into the model’s decision-making process, enhancing transparency. This research highlights the potential of AI-enhanced MRI techniques in advancing csPCa detection and diagnosis.
Similar content being viewed by others
Introduction
Prostate Cancer (PCa) represents a significant public health issue affecting men globally. In 2020, there were approximately 1.4 million new cases and around 375,000 deaths due to prostate cancer, underscoring its substantial impact on global health1. As the second most diagnosed cancer in men, PCa poses considerable economic and healthcare burdens. Within this context, identifying Clinically Significant Prostate Cancer (csPCa) is paramount. csPCa is characterized by a high likelihood of progression and symptomatic impact if left untreated, distinguishing it from lower-grade, indolent tumors2. Accurate classification is vital to balance the risks of overtreatment–which entails unnecessary side effects–against the dangers of undertreatment, where aggressive interventions such as surgery or radiation are delayed3,4.
Historically, detection has relied on Digital Rectal Exams (DRE) and Prostate-Specific Antigen (PSA) tests. While foundational, these methods suffer from limited specificity and sensitivity, often leading to unnecessary biopsies or missed diagnoses5. The advent of multi-parametric Magnetic Resonance Imaging (mpMRI) revolutionized diagnostics by combining anatomical and functional sequences to visualize tumor location, size, and aggressiveness6. Current guidelines recognize mpMRI’s utility not only in detection but also in staging and guiding biopsy protocols, helping to distinguish lesions requiring immediate intervention from those suitable for active surveillance7,8.
Despite these advancements, mpMRI-based diagnosis faces significant challenges regarding reproducibility and accuracy. The interpretation of MRI is inherently qualitative and prone to high inter-reader variability, influenced by radiologist experience, fatigue, or cognitive overload9,10. A critical bottleneck is the management of indeterminate findings, such as PI-RADS 3 lesions, where ambiguity complicates patient management strategies11. Furthermore, subtle or small significant tumors may still be missed due to the limitations of human visual perception or sampling errors during targeted biopsies.
The integration of Artificial Intelligence (AI) into medical imaging offers a robust solution to these limitations. Machine Learning (ML) and Deep Learning (DL) excel in analyzing complex volumetric data, identifying subtle patterns of malignancy that may elude human observers12,13. By providing objective, quantitative assessments, AI models can enhance the sensitivity and specificity of csPCa detection, standardize diagnostic workflows, and support clinical decision-making in cases of radiological uncertainty14,15.
In this work, we present a comprehensive approach to the classification of csPCa using advanced AI techniques applied to MRI data. We leverage two diverse datasets, including the newly collected BIMCV Prostate dataset, to develop and validate a deep learning model based on a 3D variant of EfficientNet-B7. Our methodology includes a robust preprocessing pipeline, incorporating prostate segmentation with a custom-trained nnUNet model, followed by extensive data augmentation and normalization processes. The model is initialized with weights pretrained on the Prostate Imaging: Cancer AI (PI-CAI) Challenge16 and fine-tuned on the BIMCV dataset. To ensure generalization, we implement an ensemble aggregation strategy using a stacked logistic-regression meta-learner. Additionally, we analyze the impact of synthesizing missing ADC maps and employ interpretability techniques such as occlusion sensitivity to provide transparent insights into the model’s decision-making process.
The contributions of this work are threefold: the creation and utilization of the multi-center BIMCV Prostate dataset, the development of a highly accurate AI model for csPCa classification using a transfer learning and ensemble aggregation strategy, and the implementation of interpretability techniques to enhance clinical trust. The rest of the article is structured as follows: The Results section presents model performance and interpretability analysis. The Discussion contextualizes the findings within the broader field, and the Methods section details the datasets, preprocessing, and training procedures.
Results
Model pretraining with PI-CAI
The PI-CAI pretraining produced a useful initialization but exhibited substantial fold-wise variability. Figure 1 shows the cross-fold ROC and precision–recall curves for the PI-CAI 5-fold models. The mean ROC-AUC across PI-CAI folds was 0.62 (SD 0.12) and the mean PR-AUC was 0.40 (SD 0.17). Individual fold ROC-AUCs ranged from 0.47 to 0.77, and precision–recall AUCs ranged from 0.16 to 0.59, demonstrating instability in some folds (Fold 4 accuracy on PI-CAI was 36.1%). The per-fold classification metrics are summarized in Table 1.
Performance of the 5 PI-CAI pretraining folds.
Model training and aggregation on BIMCV
Given the fold-wise variability observed during PI-CAI pretraining, PI-CAI models were retained strictly as initializations. Each of the five PI-CAI folds was fine-tuned independently on the BIMCV training data. To ensure robust inference and mitigate dependence on any single initialization, we aggregated the five fine-tuned models on the BIMCV hold-out set using two validation-calibrated schemes: (i) a weighted logit ensemble, where weights were assigned proportional to the BIMCV validation ROC-AUC of each fold (Eq. 2 and Eq. 3); and (ii) a stacked logistic-regression meta-learner trained on the per-model logit margins (Eq. 4). Crucially, all aggregation parameters and decision thresholds were fitted exclusively on the BIMCV validation split.
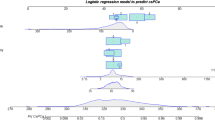
Aggregation strategies substantially improved discrimination performance compared to the single-model baseline. On the independent BIMCV hold-out set, the stacked logistic-regression meta-learner achieved the highest discrimination with an ROC-AUC of 0.816 (95% CI 0.774–0.858) and a PR-AUC of 0.806. The weighted logit ensemble yielded an ROC-AUC of 0.802 (95% CI 0.758–0.846) and a PR-AUC of 0.758. Both aggregation methods outperformed the non-pretrained single-model baseline, which attained an ROC-AUC of 0.711 (95% CI 0.659–0.762). DeLong tests confirmed the high statistical significance of these improvements, yielding \(p < 0.0001\) for both the stacked model (\(p = 9.10 \times 10^{-8}\)) and the ensemble (\(p = 2.19 \times 10^{-5}\)) relative to the baseline.
The ROC and Precision-Recall curves illustrating these performance gains are presented in Fig. 2a
Performance comparison on the BIMCV hold-out set: Stacked logistic-regression meta-learner, Weighted logit ensemble, and Non-pretrained baseline.
Table 2 details the class-wise classification metrics at the default decision threshold. Notably, the weighted logit ensemble proved particularly effective for screening purposes, achieving the highest sensitivity (Recall) for clinically significant prostate cancer (csPCa) at 0.83, combined with an overall accuracy of 0.75. The stacked meta-learner provided a robust alternative with the highest AUC and balanced metrics.
Impact of synthesized ADC maps
In the BIMCV dataset, ADC maps were missing for a subset of examinations and were retrospectively synthesized from DWI using the mono-exponential model (Eq. 1). To quantify whether the use of synthesized ADC introduced a distributional shift or bias, we compared two five-fold cross-validation training regimes: (i) a DER model trained on the full dataset (including both native and synthesized ADC maps), and (ii) a NODER model trained strictly on the subset of sessions with native ADC maps. Both models utilized T2w, DWI, and ADC as inputs and were evaluated on the subset of cases with native ADC acquisitions.
Contrary to the concern that synthesized data might introduce noise, the model trained with the inclusion of synthesized ADCs (DER) consistently outperformed the model trained on native ADCs alone (NODER). Pooled across all native validation cases (\(n=1,050\)), the DER model achieved an ROC–AUC of 0.779 (95% CI 0.750–0.807) compared to 0.717 (95% CI 0.686–0.749) for the NODER model. This represents a significant performance gain of \(\Delta\)AUC \(= 0.062\) (paired DeLong test \(p < 0.001\)).
We further stratified the performance of the DER model by the origin of the ADC map (native vs. synthesized) to detect potential distributional shifts. The DER model achieved an ROC–AUC of 0.779 on cases with native ADC and 0.795 (95% CI 0.763–0.827) on cases with synthesized ADC. The difference in performance between native and synthesized subsets was not statistically significant (paired t-test across folds, \(p=0.25\)). These results indicate that synthesized ADC maps serve as an effective data augmentation strategy, improving generalization on real clinical acquisitions without introducing a measurable domain shift; the full subgroup analysis is summarized in Table 3.
Model interpretability
Interpretability techniques such as occlusion sensitivity and guided backpropagation were employed to gain insights into the deep learning model’s decision-making process. These techniques help to understand which parts of the input images are most influential in the model’s predictions, thereby providing transparency and trust in the model’s functionality.
Occlusion sensitivity
Occlusion Sensitivity involves systematically occluding parts of the input image and observing the changes in the model’s prediction confidence. By sliding a small occluding patch across different regions of the image, we can generate a sensitivity map that highlights the areas most critical for the model’s decision17. In the context of prostate cancer detection, this technique is particularly effective for zonal localization. As the occlusion patch is larger than typical textural features, this method indicates the general anatomical region or slice stack that drives the classification, rather than delineating pixel-perfect boundaries. Figure 3 illustrates this zonal focus, where the model’s confidence drops significantly when the prostate region is occluded.
Occlusion Sensitivity maps for the BIMCV dataset. The heatmaps demonstrate the model’s zonal focus on the prostate gland. The red/blue overlays correspond to regions where occlusion causes the most significant drop in prediction confidence, aligning with the anatomical location of the prostate.
Guided backpropagation
Guided Backpropagation visualizes the gradient of the output with respect to the input image, modifying the backward pass to allow only positive gradients to flow through ReLU activations. This results in sharper saliency maps that highlight specific pixel-level features contributing to the prediction18. Unlike Occlusion Sensitivity, which highlights broad zones, Guided Backpropagation is sensitive to fine-grained textural patterns. As shown in Figure 4, this technique highlights specific signal intensities within the lesion area across T2-weighted, ADC, and DWI sequences, confirming the model relies on relevant tumor features rather than background artifacts.
Guided Backpropagation results for a PI-RADS 3 patient. (a) T2w, (b) ADC, (c) DWI, and (d) Saliency maps. The dashed red lines indicate the expert-annotated lesion. The saliency maps (d) show high activation within the lesion boundaries, demonstrating the model’s ability to identify relevant textural features associated with malignancy.
Lesion localization analysis
To validate the model’s ability to correctly identify lesion locations without explicit segmentation supervision, we performed a post-hoc localization study on a subset of 125 MRI sessions from 48 patients. For these cases, bounding box ground truth (GT) segmentations of csPCa lesions were manually annotated by expert urologists under radiological guidance. We generated both Occlusion Sensitivity and Guided Backpropagation maps for each session and qualitatively assessed their spatial concordance with the GT masks.
We observed a strong spatial concordance in 110 of the 125 sessions (88%). The analysis revealed the complementary nature of the two methods: Occlusion Sensitivity consistently identified the anatomical zone and slice stack of the lesion (zonal localization), while Guided Backpropagation successfully highlighted the specific textural morphology of the tumor (feature localization). This confirms that, despite being trained on patient-level labels (“weak supervision”), the model effectively learns to spatially prioritize clinically significant lesions rather than relying on background artifacts.
Discussion
PI-CAI pretraining produced a useful initialization but displayed substantial fold-wise instability. Cross-fold PI-CAI performance had mean ROC-AUC 0.62 (SD 0.12) and mean PR-AUC 0.40 (SD 0.17), with individual fold ROC-AUCs from 0.47 to 0.77 and a low-performing fold (Fold 4 accuracy 36.1%). This variability motivated a transfer strategy that retained all five PI-CAI initializations and deferred aggregation and calibration to the BIMCV stage. The PI-CAI results highlight that pretraining alone can produce uneven per-fold behaviour and should not be interpreted as a final inference strategy without downstream validation and aggregation.
Fine-tuning on BIMCV and performing BIMCV-level cross-fold aggregation substantially improved and stabilised performance. Using the BIMCV validation split to fit and select aggregation schemes we obtained on the BIMCV hold-out: stacked logistic-regression meta-learner ROC-AUC 0.816 (PR-AUC 0.806, accuracy reported in Table 2), weighted logit ensemble ROC-AUC 0.802 (PR-AUC 0.758), and a non-pretrained single-model baseline ROC-AUC 0.711 (PR-AUC 0.707). Aggregation removed dependence on any single PI-CAI fold and yielded consistent gains in discrimination and recall (see Fig. 2a, band Table 2). These results justify using validation-set calibrated ensembling when pretraining exhibits high fold variability.
However, the importance of model interpretability cannot be overstated. Techniques such as Guided Backpropagation not only enhance the transparency of the model’s decision-making process but also provide valuable visual assistance to medical professionals (Figure 4). By highlighting specific image regions that drive predictions, these techniques help radiologists understand the model’s rationale, potentially identifying regions of interest (ROIs) that require further investigation. This capability is particularly valuable in high-volume clinical settings, where radiologist fatigue and heavy workloads can lead to cognitive overload and diagnostic errors. In this context, the proposed tool functions not as a replacement for human expertise, but as a vigilant assistant or “second reader,” drawing attention to suspicious areas that might otherwise be overlooked. Furthermore, unlike fully supervised segmentation models that require labor-intensive, pixel-level manual annotations for training, our approach delivers effective lesion localization using only patient-level labels. This additional layer of insight is crucial for building trust in AI systems, ensuring they can be effectively integrated into clinical workflows as robust decision-support tools.
We explicitly addressed the concern that synthesizing missing ADC maps might introduce a distributional shift or bias detrimental to model performance. Our results strongly refute this hypothesis. On the strict subset of subjects with native ADC acquisitions, the model trained with the inclusion of synthesized maps (DER) significantly outperformed the model restricted to native data (NODER), yielding a performance gain of \(\Delta\)AUC \(= 0.062\) (\(p < 0.001\)). This finding indicates that excluding sessions with missing modalities is counterproductive; instead, physically motivated synthesis (via the mono-exponential model) acts as an effective data augmentation strategy, enhancing the model’s generalization capabilities on real, native acquisitions. Furthermore, within the DER model, we observed no statistically significant difference in performance between native and synthesized validation subsets (\(p=0.25\)), confirming that the synthesized maps preserve the semantic features necessary for csPCa detection without introducing a measurable domain gap.
Several limitations remain. First, this work evaluates classification at the session/patient level; lesion-level sensitivity and specificity require radiologist-verified lesion maps and lesion-wise labels. Future work should integrate radiologist-annotated lesion masks and report lesion-wise detection metrics (sensitivity at fixed false-positive rates, lesion-precision, lesion-F1) to quantify clinical utility more precisely. Second, while aggregation improves hold-out metrics, external validation on independent cohorts from other regions is required to establish generalisability. Third, the present study focuses on classification performance; prospective integration into biopsy planning or radiologist decision support remains to be demonstrated. We recommend a reader study and a closed-loop workflow pilot to evaluate the model’s impact on biopsy allocation, radiologist workflow, and clinical decision making.
In summary, retaining multiple PI-CAI initializations and performing BIMCV-level aggregation stabilises inference and improves discrimination for csPCa classification. Interpretability and subgroup analyses increase transparency. The next steps are lesion-level benchmarking, prospective reader studies and external cohort validation to move toward clinical translation.
Methods
This study employs a comprehensive methodology to develop and evaluate AI models for classifying csPCa. Leveraging two primary datasets–the PI-CAI Challenge Dataset and the BIMCV Prostate Dataset–the approach includes extensive data preprocessing, model training, and validation techniques. The preprocessing pipeline involves prostate segmentation using a custom-trained nnUNet model, followed by normalization and data augmentation to ensure model robustness. A 3D variant of EfficientNet-B7 was used for classification, optimized with Cross-Entropy Loss and the Adadelta optimizer. The model was pretrained on the PI-CAI dataset and then fine-tuned on the BIMCV dataset, using a 5-fold cross-validation strategy to enhance generalization. All code is available at BIMCV-CSUSP/BIMCV-Prostate-Classification.
Datasets
The first dataset used in this work is the Public Training and Development Dataset from the Prostate Imaging: Cancer AI (PI-CAI) Challenge16. This specific subset comprises 1,500 fully anonymized prostate MRI cases, which were made available for research purposes under a non-commercial license. The dataset includes exams from men suspected of having csPCa, typically due to elevated PSA levels or abnormal digital rectal exam (DRE) findings. It contains 425 csPCa-positive cases, recorded by a team of trained investigators and expert radiologists. Each exam in this dataset includes essential clinical variables such as patient age, prostate volume, PSA level, and PSA density, as well as acquisition details like scanner manufacturer and diffusion b-value. The MRI scans themselves consist of axial, sagittal, and coronal T2W, high b-value DWI, and ADC maps. AI-derived lesion annotations and whole-gland segmentations are available for all 1,500 cases, generated using a semi-supervised learning strategy. This dataset serves as the foundation for developing and evaluating the AI models aimed at improving prostate cancer detection and diagnosis in this work.
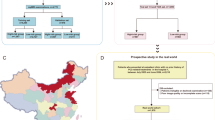
The second dataset used in this work, which represents a significant contribution, is the anonymized BIMCV Prostate Dataset19. This dataset includes 8,441 subjects and 9,341 MRI sessions, collected from 16 different health centers, thereby ensuring a diverse and representative dataset. Such diversity enhances the generalizability of the AI models developed, as they incorporate a wide range of imaging practices and patient demographics. The dataset predominantly consists of T2-weighted (T2W) images (62.97%), followed by Diffusion-Weighted Imaging (DWI) images (15.49%), and Apparent Diffusion Coefficient (ADC) maps (21.53%). A critical selection criterion for this study was to focus on sessions that included all three image modalities (T2W, DWI, and ADC), resulting in 4,700 sessions. These were further divided into training and validation sets based on the health centers, with images from three different centers set aside for validation, ensuring independent training and validation sets. This division yielded 537 images for validation and 4,163 images for training.
To organize and manage the imaging data, the BIMCV Prostate Dataset was structured using the Medical Imaging Data Structure (MIDS) framework20, which categorizes data into subject-level, session-level, and modality-specific folders, accompanied by metadata files detailing imaging protocols, patient demographics, and clinical annotations. The dataset includes contributions from various MRI machines, predominantly GE (66.7%), followed by Philips (25.1%) and Siemens (8.13%). To address the issue of incomplete ADC maps in some sessions, the ADC values were calculated from the available DWI images using the standard formula (see Equation 1)21.
where \(S_{b0}\) and \(S_{b1}\) are the signal intensities of the DWI images acquired with b-values \(b0\) and \(b1\), respectively. These generated ADC maps were saved in the derivatives folder of the MIDS structure, ensuring the dataset remained comprehensive and useful for developing AI models.
Preprocessing
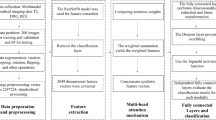
Prostate segmentation precedes all downstream steps. We trained nnUNet on a combined corpus (Prostate15822, PI-CAI, plus 164 manually segmented T2w BIMCV cases) using 5-fold cross-validation and ensembled fold outputs to obtain robust gland masks. A bounding box around the prostate was expanded by 20 pixels in all directions to include local context. Cropped volumes were linearly registered to the T2w modality and resampled to a fixed input grid of \(128\times 128\times 32\). Intensity scaling uses min–max normalization to [0, 1] for T2w and DWI. ADC channels are clipped at the 0.5 and 99.5 foreground percentiles then standardized by channel-wise z-score to preserve physical units. Augmentations applied during training included random rotations (\(\pm 10^{\circ }\)), flips, isotropic zooms (0.9–1.1), additive Gaussian noise and contrast jitter. The full preprocessing pipeline is summarized in Figure 5.
Prostate segmentation
The nnUNet segmentation stage used a 3D U-Net backbone with automatic configuration per nnUNet defaults and was trained with 5-fold cross-validation. Training data comprised 119 Prostate158 images22, 1,500 PI-CAI images, and 164 manual BIMCV T2w segmentations. Final segmentations are ensemble averages of the five folds, followed by post-processing to generate the prostate bounding box used for cropping.
Preprocessing Pipeline applied to prostate MRI images for csPCa classification. Linear affine registration to align all images to the T2-weighted (T2w) modality, min-max normalization applied to T2w and DWI images to scale intensity values between 0 and 255, 0.5-99.5 percentile normalization applied to ADC maps to clip extreme values and standardize the intensity range, and Z-score normalization to standardize the intensity values using the mean and standard deviation.
Deep learning model
We adapt EfficientNet-B7 to 3D by converting 2D convolutions, pooling and MBConv blocks into their 3D counterparts while preserving the compound scaling prescription23. Inputs are three-channel volumes (T2w, DWI, ADC) concatenated along the channel axis. Training minimizes categorical cross-entropy and uses the Adadelta optimizer24 with initial learning rate 1.0, decay \(\rho =0.95\) and \(\epsilon =1\times 10^{-7}\). Training used mixed precision where available and batch sizes selected to fit a 32GB GPU. PI-CAI pretraining ran as 5-fold cross-validation; all five fold models were kept as independent initializations and then fine-tuned separately on BIMCV training data. No PI-CAI-level ensembling or calibration was performed.
Transfer learning and ensemble aggregation
To mitigate the fold-wise variability observed during PI-CAI pretraining and improve generalization on the BIMCV dataset, we implemented a transfer learning strategy that retains the diversity of the pretrained models. Instead of selecting a single “best” fold, we used all five PI-CAI-initialized models as a committee. Each model i (where \(i \in \{1,\dots ,5\}\)) was fine-tuned independently on the BIMCV training set.
At inference time, we aggregated the predictions from these five models using two distinct strategies to produce a single, robust prediction for each sample x. Let \(\textbf{z}_i(x)=(z_{i,0}(x),z_{i,1}(x))^\top\) denote the raw output logits from model i for the negative and positive classes, respectively.
Strategy 1: weighted logit ensemble
This approach computes a weighted average of the logits from all five models. The contribution of each model is weighted proportional to its performance on the BIMCV validation split. Specifically, if \(a_i\) is the ROC-AUC achieved by model i on the validation set, we compute its normalized weight \(w_i\) as:
The ensemble logit vector \(\textbf{s}(x)\) is then calculated as the linear combination of individual logits:
Final probability scores are obtained by applying the softmax function to \(\textbf{s}(x)\).
Strategy 2: stacked generalization (Meta-learner)
To further refine the decision boundary, we trained a secondary “meta-learner” using Logistic Regression. The input features for this meta-learner are the logit margins from the five fine-tuned models. The logit margin \(m_i(x) = z_{i,1}(x) - z_{i,0}(x)\) represents the raw confidence score of model i for the positive class.
We construct a feature vector \(\textbf{m}(x)=[m_1(x),\dots ,m_M(x)]^\top\) containing the margins from all folds. The meta-learner then predicts the probability of clinically significant prostate cancer, \(p_{\textrm{stack}}(x)\), as follows:
The parameters \(\beta _0\) and \(\varvec{\beta }\) were fitted on the BIMCV validation split using \(\ell _2\) regularization and balanced class weights to handle potential class imbalance. Importantly, all aggregation parameters (weights \(w_i\) and regression coefficients \(\varvec{\beta }\)) and decision thresholds were determined exclusively on the BIMCV validation set to prevent data leakage from the test set.
Evaluation and implementation details
Model performance was evaluated on the independent BIMCV hold-out set using Area Under the Receiver Operating Characteristic curve (ROC-AUC) and Area Under the Precision-Recall curve (PR-AUC). Additionally, we report class-wise Precision, Recall, and F1-score at the optimal decision threshold determined during validation. Confidence intervals (95% CI) for AUC values were estimated using DeLong’s method, and statistical comparisons between models were conducted using paired DeLong tests; p-values \(<0.05\) were considered statistically significant.
To ensure the robustness of the model against data imputation strategies, we conducted specific subgroup analyses comparing performance on sessions with native ADC acquisitions versus those with synthesized ADC maps (as detailed in the Results section).
The deep learning models were implemented using PyTorch and the MONAI 1.2.0 framework. The logistic regression meta-learner was implemented using scikit-learn with parameters ‘class_weight=’balanced’‘, ‘penalty=’l2’‘, and ‘C=1.0‘. All experiments were conducted on NVIDIA GPUs using mixed-precision training. Random seeds, dataset splits, and preprocessing parameters were fixed to ensure reproducibility. The complete workflows for PI-CAI pretraining and BIMCV fine-tuning are illustrated in Figures 6 and 7, respectively.
PICAI Pretraining Workflow. The preprocessing and prostate zone segmentation steps are applied, followed by concatenation of the image modalities (T2w, DWI, ADC) and training using 5-fold cross validation. All five fold models were retained for subsequent fine-tuning on the BIMCV Prostate dataset..
BIMCV Prostate Dataset Training Workflow. The pretrained models from PI-CAI are fine-tuned on the BIMCV dataset. The dataset is divided into training and validation sets by center. The preprocessing, prostate zone segmentation, and model training steps are illustrated. At inference, outputs from the five BIMCV models are aggregated on the hold-out set..
Data availability
The code used in this study is publicly available on GitHub at BIMCV-CSUSP/BIMCV-Prostate-Classification. The datasets generated and analyzed during the current study are available in the Zenodo repository under the accession number 1325431819. A detailed description of the dataset and its collection process can be found on the BIMCV website at https://bimcv.cipf.es/bimcv-projects/prostate/. Please note that the data is under restricted access, and access must be requested through the Zenodo platform.
References
Ferlay, J. et al. Global cancer observatory: Cancer today (2020).
Matoso, A. & Epstein, J. I. Defining clinically significant prostate cancer on the basis of pathological findings. Histopathology 74, 135–145. https://doi.org/10.1111/his.13712 (2019).
Dall’Era, M. A. et al. Active surveillance for prostate cancer: A systematic review of the literature. Eur. Urol. 62, 976–983 (2012).
Tosoian, J. J., Carter, H. B., Lepor, A. & Loeb, S. Active surveillance for prostate cancer: Current evidence and contemporary state of practice. Nat. Rev. Urol. 13, 205–215 (2016).
Radtke, J. P., Teber, D., Hohenfellner, M. & Hadaschik, B. A. The current and future role of magnetic resonance imaging in prostate cancer detection and management. PubMed 4, 326–41. https://doi.org/10.3978/j.issn.2223-4683.2015.06.05 (2015).
Lomas, D. J. & Ahmed, H. U. All change in the prostate cancer diagnostic pathway. Nat. Rev. Clin. Oncol. 17, 372–381. https://doi.org/10.1038/s41571-020-0332-z (2020).
Boesen, L. Multiparametric mri in detection and staging of prostate cancer. Danish Medical Bulletin (Online) 64 (2017).
Thompson, J. et al. The diagnostic performance of multiparametric magnetic resonance imaging to detect significant prostate cancer. J. Urol. 195, 1428–1435. https://doi.org/10.1016/j.juro.2015.10.140 (2016).
Stec, N., Arje, D., Moody, A. R., Krupinski, E. A. & Tyrrell, P. N. A systematic review of fatigue in radiology: Is it a problem?. Am. J. Roentgenol. 210, 799–806. https://doi.org/10.2214/ajr.17.18613 (2018).
Fütterer, J. J. et al. Can clinically significant prostate cancer be detected with multiparametric magnetic resonance imaging? A systematic review of the literature. Eur. Urol. 68, 1045–1053. https://doi.org/10.1016/j.eururo.2015.01.013 (2015).
Schoots, I. G. Mri in early prostate cancer detection: how to manage indeterminate or equivocal pi-rads 3 lesions?. Transl. Androl. Urol. 7, 70–82. https://doi.org/10.21037/tau.2017.12.31 (2018).
Belue, M. J. & Turkbey, B. Tasks for artificial intelligence in prostate MRI. Eur. Radiol. Exp. https://doi.org/10.1186/s41747-022-00287-9 (2022).
Luo, R., Zeng, Q. & Chen, H. Artificial intelligence algorithm-based MRI for differentiation diagnosis of prostate cancer. Comput. Math. Methods Med. 1–10, 2022. https://doi.org/10.1155/2022/8123643 (2022).
Hötker, A. M., Mutten, R. D., Tiessen, A., Konukoglu, E. & Donati, O. F. Improving workflow in prostate MRI: Ai-based decision-making on biparametric or multiparametric MRI. Insights Imaging https://doi.org/10.1186/s13244-021-01058-7 (2021).
Sushentsev, N. et al. Comparative performance of fully-automated and semi-automated artificial intelligence methods for the detection of clinically significant prostate cancer on MRI: A systematic review. Insights Imaging https://doi.org/10.1186/s13244-022-01199-3 (2022).
Saha, A. et al. Artificial intelligence and radiologists at prostate cancer detection in MRI — the PI-CAI challenge. In Medical Imaging with Deep Learning, short paper track (2023).
Zeiler, M. D. & Fergus, R. Visualizing and understanding convolutional networks. In Fleet, D., Pajdla, T., Schiele, B. & Tuytelaars, T. (eds.) Computer Vision - ECCV 2014, 818–833 (Springer International Publishing, Cham, 2014).
Springenberg, J. T., Dosovitskiy, A., Brox, T. & Riedmiller, M. A. Striving for simplicity: The all convolutional net. CoRR abs/1412.6806 (2014).
Alzate-Grisales, J. A. & de la Iglesia Vaya, M. Bimcv-prostate-dataset v1. Zenodo. https://doi.org/10.5281/zenodo.13254318 (2024).
Saborit-Torres, J. M. et al. Beyond the brain: Mids extends bids to multiple modalities and anatomical regions. Stud. Health Technol. Inf. https://doi.org/10.3233/shti220488 (2022).
Koh, D.-M. & Collins, D. J. Diffusion-weighted MRI in the body: Applications and challenges in oncology. Am. J. Roentgenol. 188, 1622–1635. https://doi.org/10.2214/AJR.06.1403 (2007) (PMID: 17515386).
Adams, L. C. et al. Prostate158 - an expert-annotated 3t MRI dataset and algorithm for prostate cancer detection. Comput. Biol. Med. 148, 105817. https://doi.org/10.1016/J.COMPBIOMED.2022.105817 (2022).
Tan, M. & Le, Q. EfficientNet: Rethinking model scaling for convolutional neural networks. In Chaudhuri, K. & Salakhutdinov, R. (eds.) Proceedings of the 36th International Conference on Machine Learning, vol. 97 of Proceedings of Machine Learning Research, 6105–6114 (PMLR, 2019).
Zeiler, M. D. Adadelta: an adaptive learning rate method. arXiv preprint arXiv:1212.5701 (2012).
Acknowledgements
This work is funded by the Spanish Ministry of Economic Affairs and Digital Transformation (Project MIA.2021.M02.0005 TARTAGLIA, from the Recovery, Resilience, and Transformation Plan financed by the European Union through Next Generation EU funds). TARTAGLIA takes place under the R&D Missions in Artificial Intelligence program, which is part of the Spain Digital 2025 Agenda and the Spanish National Artificial Intelligence Strategy.
Author information
Authors and Affiliations
Contributions
J.A.A.G. conceived the experiments, J.A.A.G. and A.M.R. conducted the experiments, C.R.T., A.N.B., F.G.G., and M.P.T. analyzed the results. M.I.V. supervised the project. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The author(s) declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Alzate-Grisales, J.A., Mora-Rubio, A., Peán-Teruel, M. et al. Clinically significant prostate cancer detection with deep learning in a multi-center magnetic resonance imaging study. Sci Rep 16, 10976 (2026). https://doi.org/10.1038/s41598-026-42214-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-42214-7