Abstract
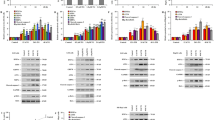
Chronic intermittent hypoxia (CIH), a hallmark feature of obstructive sleep apnea (OSA), is strongly implicated in the development of hepatic injury. However, the cellular mechanisms underlying CIH-induced liver dysfunction remain poorly understood. This study aimed to systematically characterize hepatic cellular responses to CIH at single-nucleus resolution. A rodent model of CIH was established and subjected to histological and transcriptomic analyses. Single-nucleus RNA sequencing (snRNA-seq) was performed to delineate transcriptional alterations across liver cell populations. Differentially expressed genes (DEGs) and pathway enrichment analyses were conducted to identify key biological processes and signaling networks affected by CIH exposure. CIH exposure resulted in marked hepatic injury characterized by spotty necrosis and prominent infiltration of inflammatory cells. SnRNA-seq identified ten major liver cell types with stable composition but revealed extensive transcriptional reprogramming across multiple hepatic subpopulations. CIH suppressed PPAR signaling and fatty acid metabolism in hepatocytes and hepatic stellate cells and activated AMPK and PI3K–Akt pathways related to stress response and fibrogenic processes. Mononuclear phagocytes showed upregulation of NF-κB signaling and complement/coagulation cascades. Endothelial cells exhibited changes in genes associated with cytoskeletal organization and tight junctions. T cell subpopulations displayed altered expression of genes involved in metabolic regulation and endoplasmic reticulum stress. This study provides the first single-nucleus transcriptomic atlas of the liver under CIH, revealing cell type–specific molecular remodeling across hepatocytes, stromal, and immune cells. These findings elucidate the complex cellular interplay associated with CIH-induced hepatic injury and offer novel insights into potential therapeutic targets for OSA-related liver dysfunction.
Similar content being viewed by others
Data availability
The single-nucleus RNA sequencing datasets generated and analyzed in this study have been deposited in the NCBI Sequence Read Archive (SRA) under accession number PRJNA1345096 and are publicly available at: [https://www.ncbi.nlm.nih.gov/sra/PRJNA1345096](https:/www.ncbi.nlm.nih.gov/sra/PRJNA1345096) .
References
Benjafield, A. V. et al. Estimation of the global prevalence and burden of obstructive sleep apnoea: a literature-based analysis. Lancet Respir Med. 7, 687–698. https://doi.org/10.1016/S2213-2600(19)30198-5 (2019).
Mesarwi, O. A., Loomba, R. & Malhotra, A. Obstructive Sleep Apnea, Hypoxia, and Nonalcoholic Fatty Liver Disease. Am. J. Respir Crit. Care Med. 199, 830–841. https://doi.org/10.1164/rccm.201806-1109TR (2019).
Wang, L. et al. Association of Obstructive Sleep Apnea with Nonalcoholic Fatty Liver Disease: Evidence, Mechanism, and Treatment. Nat. Sci. Sleep. 16, 917–933. https://doi.org/10.2147/NSS.S468420 (2024).
Savransky, V. et al. Chronic intermittent hypoxia causes hepatitis in a mouse model of diet-induced fatty liver. Am. J. Physiol. Gastrointest. Liver Physiol. 293, G871–877. https://doi.org/10.1152/ajpgi.00145.2007 (2007).
Savransky, V. et al. Chronic intermittent hypoxia predisposes to liver injury. Hepatology 45, 1007–1013. https://doi.org/10.1002/hep.21593 (2007).
Chen, L. D. et al. Effect of chronic intermittent hypoxia on gene expression profiles of rat liver: a better understanding of OSA-related liver disease. Sleep. Breath. 24, 761–770. https://doi.org/10.1007/s11325-019-01987-0 (2020).
Kim, N., Kang, H., Jo, A., Yoo, S. A. & Lee, H. O. Perspectives on single-nucleus RNA sequencing in different cell types and tissues. J. Pathol. Transl Med. 57, 52–59. https://doi.org/10.4132/jptm.2022.12.19 (2023).
Hicks, S. C., Townes, F. W., Teng, M. & Irizarry, R. A. Missing data and technical variability in single-cell RNA-sequencing experiments. Biostatistics 19, 562–578. https://doi.org/10.1093/biostatistics/kxx053 (2018).
Haque, A., Engel, J., Teichmann, S. A. & Lonnberg, T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 9, 75. https://doi.org/10.1186/s13073-017-0467-4 (2017).
Minati, M. A., Fages, A., Dauguet, N., Zhu, J. & Jacquemin, P. Optimized nucleus isolation protocol from frozen mouse tissues for single nucleus RNA sequencing application. Front. Cell. Dev. Biol. 11, 1243863. https://doi.org/10.3389/fcell.2023.1243863 (2023).
Gao, R. et al. Nanogrid single-nucleus RNA sequencing reveals phenotypic diversity in breast cancer. Nat. Commun. 8, 228. https://doi.org/10.1038/s41467-017-00244-w (2017).
Li, X. Y. et al. Single-nucleus RNA sequencing uncovers metabolic dysregulation in the prefrontal cortex of major depressive disorder patients. Sci. Rep. 15, 7418. https://doi.org/10.1038/s41598-025-92030-8 (2025).
Lau, S. F., Cao, H., Fu, A. K. Y. & Ip, N. Y. Single-nucleus transcriptome analysis reveals dysregulation of angiogenic endothelial cells and neuroprotective glia in Alzheimer’s disease. Proc. Natl. Acad. Sci. U S A. 117, 25800–25809. https://doi.org/10.1073/pnas.2008762117 (2020).
Lee, S. H. T. et al. Single nucleus RNA-sequencing integrated into risk variant colocalization discovers 17 cell-type-specific abdominal obesity genes for metabolic dysfunction-associated steatotic liver disease. EBioMedicine 106, 105232. https://doi.org/10.1016/j.ebiom.2024.105232 (2024).
Chen, J. et al. 2-Methoxyestradiol attenuates lung injury induced by chronic intermittent hypoxia via inhibiting the HIF1-alpha/SLC7A11 pathway. Sci. Rep. 15, 17601. https://doi.org/10.1038/s41598-025-02675-8 (2025).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods. 16, 1289–1296. https://doi.org/10.1038/s41592-019-0619-0 (2019).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Krolow, G. K., Garcia, E., Schoor, F., Araujo, F. B. S. & Coral, G. P. Obstructive sleep apnea and severity of nonalcoholic fatty liver disease. Eur. J. Gastroenterol. Hepatol. 33, 1104–1109. https://doi.org/10.1097/MEG.0000000000001920 (2021).
Lian, N. et al. Risk Factors of Nonalcoholic Fatty Liver Disease and Liver Fibrosis in Non-Obese Patients with Obstructive Sleep Apnea. Nat. Sci. Sleep. 14, 2143–2149. https://doi.org/10.2147/NSS.S388203 (2022).
Jiang, Y. B., Huang, Z. W., Lin, X. J., Luo, J. M. & Chen, L. D. Association between self-reported sleep apnea and biomarkers of liver injury: Evidence from National Health and Nutrition Examination Survey. Med. (Baltim). 103, e39393. https://doi.org/10.1097/MD.0000000000039393 (2024).
Feng, S. Z. et al. An experimental research on chronic intermittent hypoxia leading to liver injury. Sleep. Breath. 15, 493–502. https://doi.org/10.1007/s11325-010-0370-3 (2011).
Song, K., Zhang, Y., Ga, Q., Bai, Z. & Ge, R. L. High-altitude chronic hypoxia ameliorates obesity-induced non-alcoholic fatty liver disease in mice by regulating mitochondrial and AMPK signaling. Life Sci. 252, 117633. https://doi.org/10.1016/j.lfs.2020.117633 (2020).
Liang, H. & Song, K. Comprehensive metabolomics and transcriptomics analysis reveals protein and amino acid metabolic characteristics in liver tissue under chronic hypoxia. PLoS One. 18, e0291798. https://doi.org/10.1371/journal.pone.0291798 (2023).
Mooli, R. G. R. et al. Hypoxia via ERK Signaling Inhibits Hepatic PPARalpha to Promote Fatty Liver. Cell. Mol. Gastroenterol. Hepatol. 12, 585–597. https://doi.org/10.1016/j.jcmgh.2021.03.011 (2021).
Son, G., Hines, I. N., Lindquist, J., Schrum, L. W. & Rippe, R. A. Inhibition of phosphatidylinositol 3-kinase signaling in hepatic stellate cells blocks the progression of hepatic fibrosis. Hepatology 50, 1512–1523. https://doi.org/10.1002/hep.23186 (2009).
Zhao, X. K. et al. Focal Adhesion Kinase Regulates Hepatic Stellate Cell Activation and Liver Fibrosis. Sci. Rep. 7, 4032. https://doi.org/10.1038/s41598-017-04317-0 (2017).
Hijazi, N., Shi, Z. & Rockey, D. C. Characterization of focal adhesion proteins in rodent hepatic stellate cells. Histochem. Cell. Biol. 158, 325–334. https://doi.org/10.1007/s00418-022-02123-y (2022).
Wang, Y., Nakajima, T., Gonzalez, F. J. & Tanaka, N. PPARs as Metabolic Regulators in the Liver: Lessons from Liver-Specific PPAR-Null Mice. Int. J. Mol. Sci. 21 https://doi.org/10.3390/ijms21062061 (2020).
Sunami, Y. et al. Hepatic activation of IKK/NFkappaB signaling induces liver fibrosis via macrophage-mediated chronic inflammation. Hepatology 56, 1117–1128. https://doi.org/10.1002/hep.25711 (2012).
El Kasmi, K. C. et al. Macrophage-derived IL-1beta/NF-kappaB signaling mediates parenteral nutrition-associated cholestasis. Nat. Commun. 9, 1393. https://doi.org/10.1038/s41467-018-03764-1 (2018).
Cui, Y. et al. The role of AMPK in macrophage metabolism, function and polarisation. J. Transl Med. 21, 892. https://doi.org/10.1186/s12967-023-04772-6 (2023).
Kaczara, P. et al. Liver sinusoidal endothelial cells rely on oxidative phosphorylation but avoid processing long-chain fatty acids in their mitochondria. Cell. Mol. Biol. Lett. 29, 67. https://doi.org/10.1186/s11658-024-00584-8 (2024).
Patten, D. A. et al. Human liver sinusoidal endothelial cells promote intracellular crawling of lymphocytes during recruitment: A new step in migration. Hepatology 65, 294–309. https://doi.org/10.1002/hep.28879 (2017).
Zhou, Y. et al. Angiotensin II depends on hippo/YAP signaling to reprogram angiogenesis and promote liver fibrosis. Cell. Signal. 123, 111355. https://doi.org/10.1016/j.cellsig.2024.111355 (2024).
Dunkelberger, J. R. & Song, W. C. Complement and its role in innate and adaptive immune responses. Cell. Res. 20, 34–50. https://doi.org/10.1038/cr.2009.139 (2010).
Zhang, W. & Cao, X. Unfolded protein responses in T cell immunity. Front. Immunol. 15, 1515715. https://doi.org/10.3389/fimmu.2024.1515715 (2024).
Acknowledgements
The authors thank the Singleron Biotechnologies for their help with the single-cell RNA sequencing analysis.
Funding
This work was supported by Natural Science Foundation of Fujian Province (2022J011474), Young people training project from Fujian Province Health Bureau (2022GGA053), and Startup Fund for Scientific Research of Fujian Medical University (2024QH1687), and grant PDA202202 for scientific research of Zhangzhou Affiliated Hospital of Fujian Medical University.
Author information
Authors and Affiliations
Contributions
LDC supervised and designed this project. WSH, CQW, and JFH performed the animal experiment. JML and CDY prepared samples for snRNA-seq. WSH, LDC and LL analyzed the snRNA-seq data. YZH performed pathology analysis. WSH and CQW wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
This study was approved by the approved by the Animal Ethics Committee of Fujian Medical University (IACUC FJMU 2022-0031). All methods were carried out in accordance with relevant guidelines and regulations. All methods are also reported in accordance with ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, WS., Wang, CQ., Huang, YZ. et al. Single-nucleus RNA sequencing uncovers cell type-specific alterations in OSA-related liver injury. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42236-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42236-1