Abstract
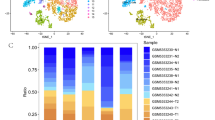
Although long noncoding RNAs (lncRNAs) have been implicated in the progression of prostate cancer (PCa), the functional roles of many of these molecules, especially in metastasis, remain poorly understood. To identify novel lncRNAs linked to PCa malignant phenotypes, we analyzed the differences in lncRNA expression between the highly metastatic PCa cell line (PC-3 M-1E8) and the poorly metastatic PCa cell line (PC-3 M-2B4) using transcriptome sequencing. This differential expression was confirmed by RT-qPCR, which showed that the lnc-ALX1-2 gene cluster (including lnc-ALX1-2:5, lnc-ALX1-2:7, lnc-ALX1-2:10) was significantly upregulated in PC-3 M-1E8 cells. Functional studies demonstrated that knockdown of the lnc-ALX1-2 gene cluster suppressed proliferation, migration, and invasion in PC-3 M-1E8 cells, with lnc-ALX1-2:10 showing the most prominent effect. Transcriptome analysis further revealed that lnc-ALX1-2:10 knockdown altered the expression of 194 genes related to both proliferation and migration. RT-qPCR and Western blot validated that lnc-ALX1-2:10 knockdown downregulated pro-tumorigenic factors (CCNE1, PDGFRA, ANGPT4) and EMT markers (N-cadherin, Snail, Vimentin), while upregulating ITGAL and E-cadherin. In vivo, lnc-ALX1-2:10 knockdown obviously decreased tumor volume but had no effect on mouse body weight, and molecular analysis of xenograft tumors confirmed consistent expression changes of key proteins as in vitro. Collectively, our findings identify lnc-ALX1-2:10 as a novel lncRNA that promotes aggressive phenotypes in PCa, highlighting its potential as a therapeutic target for metastatic disease.
Similar content being viewed by others
Data availability
The datasets generated by Forevergen Biosciences are available in NCBI with the ID: PRJNA753212(https://www.ncbi.nlm.nih.gov/bioproject/PRJNA753212).
References
Kratzer, T. B. et al. Prostate cancer statistics, 2025. CA. Cancer J. Clin. https://doi.org/10.3322/caac.70028 (2025).
Roobol, M. J., de Mansson, V. I. I., Godtman, M., Talala, R. A. & den Hond, K. M. European Study of Prostate Cancer Screening – 23-Year Follow-up. N. Engl. J. Med. 393 (17), 1669–1680 (2025).
Hoang, D. T., Iczkowski, K. A., Kilari, D., See, W. & Nevalainen, M. T. Androgen receptor-dependent and -independent mechanisms driving prostate cancer progression: Opportunities for therapeutic targeting from multiple angles. Oncotarget 8 (2), 3724–3745 (2017).
Shafi, A. A., Yen, A. E. & Weigel, N. L. Androgen receptors in hormone-dependent and castration-resistant prostate cancer. Pharmacol. Ther. 140(3), 223–38 (2013).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17(1), 47–62 (2016).
Ransohoff, J. D., Wei, Y. & Khavari, P. A. The functions and unique features of long intergenic non-coding RNA. Nat. Rev. Mol. Cell Biol. 19(3), 143–57 (2018).
Sanchez Calle, A., Kawamura, Y., Yamamoto, Y., Takeshita, F. & Ochiya, T. Emerging roles of long non-coding RNA in cancer. Cancer Sci. 109(7), 2093–100 (2018).
Tsagakis, I., Douka, K., Birds, I. & Aspden, J. L. Long non-coding RNAs in development and disease: Conservation to mechanisms. J. Pathol. 250(5), 480–495 (2020).
Yeh, C. F. et al. Expedition to the missing link: Long noncoding RNAs in cardiovascular diseases. J. Biomed. Sci. 27(1), 48. (2020).
Hua, J. T., Chen, S. & He, H. H. Landscape of noncoding RNA in prostate cancer. Trends Genet. 35(11), 840–51 (2019).
Wang, Z., Li, X., Liu, Z., Du, Y. & Wu, T. LncRNA HANR promotes the aerobic glycolysis in prostate cancer by stabilizing TPI1. Exp. Cell Res. 452(1), 114744 (2025).
Li, A. et al. The lncRNA MIR22HG suppresses prostate cancer cell proliferation, migration, and epithelial-mesenchymal transition via the miR-4428/PCDH9 axis. Transl. Cancer Res. 14(5), 3133–48 (2025).
Seitz, A. K. et al. Profiling of long non-coding RNAs identifies LINC00958 and LINC01296 as candidate oncogenes in bladder cancer. Sci. Rep. 7(1), 395. (2017).
Xu, X. et al. The long noncoding RNA LINC02820 promotes tumor growth and metastasis through regulating MYH9 expression in esophageal squamous cell carcinoma. MedComm (2020) 6(6), e70218 (2025).
Wang, J. et al. Novel long noncoding RNA LINC02820 augments TNF signaling pathway to remodel cytoskeleton and potentiate metastasis in esophageal squamous cell carcinoma. Cancer Gene Ther. 30 (2), 375–387 (2023).
Yao, W. et al. ALX1 promotes migration and invasion of lung cancer cells through increasing snail expression. Int. J. Clin. Exp. Pathol. 8 (10), 12129–12139 (2015).
Yuan, H. et al. ALX1 induces snail expression to promote epithelial-to-mesenchymal transition and invasion of ovarian cancer cells. Cancer Res. 73(5), 1581–90 (2013).
Wang, H. et al. Akt mediates metastasis-associated gene 1 (MTA1) regulating the expression of E-cadherin and promoting the invasiveness of prostate cancer cells. PLoS One 7(12), e46888 (2012).
Yang, W. et al. TMTP1, a novel tumor-homing peptide specifically targeting metastasis. Clin. Cancer Res. 14(17), 5494–502 (2008).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D7 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–51 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30 (2000).
Teo, M. Y., Rathkopf, D. E. & Kantoff, P. Treatment of advanced prostate cancer. Annu. Rev. Med. 70, 479–99 (2019).
Wu, J. et al. Long noncoding RNA LINC01296 is associated with poor prognosis in prostate cancer and promotes cancer-cell proliferation and metastasis. OncoTargets Ther. 10, 1843–52 (2017).
Hejazi, M. et al. LINC00162 silencing enhances sorafenib sensitivity and inhibits thyroid cancer cells progression through modulation of MAPK signaling and apoptosis. Sci. Rep. 15(1), 29726 (2025).
Alobid, S. Targeting the PI3K/AKT/mTOR signaling pathway in prostate cancer: Molecular dysregulation, therapeutic advances, and future directions. Saudi Pharm. journal: SPJ : official publication Saudi Pharm. Soc. 34 (1), 2 (2026).
Jin, C. et al. NUF2 activated by YY1 promotes prostate cancer malignancy via p38/MAPK signaling axis and serves as a therapeutic target. Biochem. Pharmacol. 237, 116919 (2025).
Ustach, C. V. et al. A novel signaling axis of matriptase/PDGF-D/ss-PDGFR in human prostate cancer. Cancer Res. 70 (23), 9631–9640 (2010).
Wang, X. et al. Dysregulation of long non-coding RNA SNHG12 alters the viability, apoptosis, and autophagy of prostate cancer cells by regulating miR-195/CCNE1 axis. Int. J. Clin. Exp. Pathol. 12(4), 1272–83 (2019).
Pencik, J. et al. STAT3/LKB1 controls metastatic prostate cancer by regulating mTORC1/CREB pathway. Mol. Cancer. 22 (1), 133 (2023).
Lin, T. H. et al. Apelin facilitates integrin alphavbeta3 production and enhances metastasis in prostate cancer by activating STAT3 and inhibiting miR-8070. Int. J. Biol. Sci. 21 (9), 4117–4128 (2025).
Sternberg, C. et al. Cell-autonomous IL6ST activation suppresses prostate cancer development via STAT3/ARF/p53-driven senescence and confers an immune-active tumor microenvironment. Mol. Cancer. 23 (1), 245 (2024).
Funding
This research was supported by Natural Science Foundation of Xiamen, China (Project No. 3502Z202373098, Xinjun Wang), Natural Science Foundation of Fujian Province (Grant Nos. 2023J011592, Xinjun Wang and 2024J011333 Guangcheng Luo), and Major Project (2024GZL-ZD03, Guangcheng Luo) and and Youth Project (2024GZL-QN103, Zhangqun Li) under the Xiamen Health High-Quality Development Science and Technology Program.
Author information
Authors and Affiliations
Contributions
Conceptualization, Funding acquisition: LGC, WXJ and LZQ; Data curation, Formal analysis, Validation: ZQ and LZQ; Methodology: ZB; Investigation: WXQ; Project administration, Writing - original draft: WXJ; Resources, Software, Visualization: BY; Supervision, Writing - review & editing: LGC.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
Animal experiment was approved by the committee of Guangzhou Forevergen Biosciences (Guangzhou, China, approval number: IACUC-G16057). We confirm that all methods were performed in accordance with the relevant guidelines and regulations. The study is reported in accordance with ARRIVE guidelines.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, X., Zong, Q., Bu, Y. et al. lnc-ALX1-2:10 is a novel regulator that enhances proliferation, migration and invasion in prostate cancer cells. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42299-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42299-0