Abstract
Reliable links between genes and diseases are central to biomedical research; however, many computational methods overlook the semantic and hierarchical layers of ontologies, missing indirect relationships and producing shallow association scores. We propose an ontology-driven framework for gene–disease association mining that integrates hierarchical knowledge from the Gene Ontology and Disease Ontology. Our text-mining pipeline processes PubMed text by cleaning, annotating, and extracting sentence-level co-occurrences of biomarker-related terms. We evaluated and compared well-known association rule mining algorithms, namely Apriori, FP-Growth, and Eclat, and applied a tie-aware rank-based transformation to correct for non-normal distributions of association scores. The resulting Athar Semantic Enriched Association (ASEA) score combines entity-specific associations with Hierarchical Ontology Associations, with an enhanced Apriori variant showing superior performance in capturing direct and indirect associations. Benchmarking against the Comparative Toxicogenomics Database, ASEA detected 17 high-grade associations (30.4% more than Apriori and Eclat, 88.9% more than FP-Growth). In total, ASEA produced 185 associations, compared with 217 for Apriori, 166 for Eclat, and 71 for FP-Growth. Among these, 21 belong to high-confidence databases (Case 1), 28 are supported by substantial literature, but not yet high-confidence (Case 2), 39 have low/intermediate database support with no strong literature (Case 3), and 22 are purely speculative (Case 4), including 12 particularly novel associations absent from the curated resources. Overall, this framework provides a transparent and extensible pipeline for biomedical knowledge discovery, combining statistical co-occurrence with ontology-driven enrichment to retrieve established knowledge and generate reliable predictions for precision medicine and hypothesis-generation.
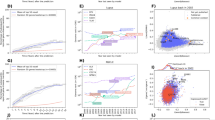
Similar content being viewed by others
Data availability
The dataset and codes of the proposed model are publicly available at https://github.com/atharnaqash/assocation-miner.
References
Jensen, L. J., Saric, J. & Bork, P. Literature mining for the biologist. Nat. Rev. Genet. 7(2), 119–129 (2006).
Tam, V. et al. Benefits and limitations of genome-wide association studies. Nat. Rev. Genet. 20(8), 467–484 (2019).
Zhou, Y., Yang, Q., Zhao, C., Li, Z. & Wang, Z. Deep learning for bioinformatics: From raw data to predictive models. Bioinformatics 34(5), 837–844 (2018).
Huang, Q. et al. Machine learning in biomedical informatics: A survey. Biomed. Res. Int. 2018, 1–15 (2018).
Yang, Q. et al. Integrating multi-source data for enhanced gene-disease association mining. BMC Genomics 19, 562 (2018).
Zhu, Y., Song, M., Chen, C., Liu, D. & Zhao, H. Advances in biomedical literature mining for disease gene discovery. Brief Bioinform. 22, bbaa057 (2020).
Campos, D. P., Oliveira, A. & De Maio, N. Efficient data mining techniques in biomedical literature. BioData Min. 12, 1–15 (2019).
Wei, C.-H., Allot, A., Leaman, R. & Lu, Z. PubTator Central: Automated concept annotation for biomedical full text articles. Nucleic Acids Res. 47(W1), W587–W593. https://doi.org/10.1093/nar/gkz389 (2019).
Tan, P.-N., Kumar, V. & Srivastava, J. Selecting the right objective measure for association analysis. In Proceedings of the 8th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 32–41. ACM (2002). https://doi.org/10.1145/775047.775053.
Church, K. W. & Hanks, P. Word association norms, mutual information, and lexicography. Comput. Linguist. 16(1), 22–29 (1990).
Agrawal, R., Imieliński, T. & Swami, A. Mining association rules between sets of items in large databases. In Proceedings of the ACM SIGMOD International Conference on Management of Data, 207–216 (1993).
Zhou, Y., Wang, X. & Zhang, L. Application of Apriori algorithm in medical data mining. Front. Public Health. 10, 912273. https://doi.org/10.3389/fpubh.2022.912273 (2022).
Han, J., Pei, J. & Yin, Y. Mining frequent patterns without candidate generation. In ACM Sigmod Record, 1–12 (2000).
Zaki, M. J., Hsiao, C.-T., et al. Eclat: A new algorithm for fast discovery of association rules. In Proceedings of the eighth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 326–331 (2001).
Lee, J. et al. BioBERT: A pre-trained biomedical language representation model for biomedical text mining. IEEE Access 8, 67834–67842 (2020).
Zhang, Y. et al. Attention mechanisms in BioBERT for gene-disease association extraction. J. Mach. Learn. Med. 8(1), 23–35 (2021).
Gene Ontology Consortium. Gene Ontology (2025).
D. Ontology, Disease Ontology. http://purl.obolibrary.org/obo/d.owl
G. O. Consortium. The Gene Ontology resource: Enriching a GOld mine. Nucleic Acids Res. 49(D1), D325–D334 (2021).
Wang, X., Zhang, M., Yu, G., Li, W. & Li, Y. Ontology-guided clustering for gene-disease relationship identification. J. Biomed. Semantics 12(1), 14–23 (2021).
Shapiro, S. S. & Wilk, M. B. An analysis of variance test for normality (complete samples). Biometrika 52(3–4), 591–611. https://doi.org/10.1093/biomet/52.3-4.591 (1965).
Anderson, T. W. & Darling, D. A. A test of goodness of fit. J. Am. Stat. Assoc. 49(268), 765–769. https://doi.org/10.1080/01621459.1954.10501232 (1954).
Lehmann, E. L. Nonparametrics: Statistical Methods Based on Ranks (Springer, 1998).
Groza, T. et al. Ontology-based annotation and integration of rare disease data for precision medicine. NPJ Genom. Med. 1(1), 1–7 (2015).
Li, P., Zhou, X., Wang, C. & Wang, J. Dynamic ontologies for real-time gene-disease prediction. J. Comput. Biol. 29(4), 315–327 (2022).
Kim, Y., Cho, H. & Lee, D. Enhancing Gene Ontology for precise gene-disease association mining. Nat. Commun. 10(1), 2534 (2019).
Disgenet, Ed. DisgeNET Organization. http://www.disgenet.org/web/DisGeNET/menu
Davis, A. P. et al. Comparative Toxicogenomics Database’s 20th Anniversary: Update 2025. Nucleic Acids Res. 53(D1), D1328–D1334. https://doi.org/10.1093/nar/gkae883 (2025).
Wahidi, N. & Ismailova, R. Association rule mining algorithm implementation for e-commerce in the retail sector. J. Appl. Res. Technol. Eng. 5(2), 63–68. https://doi.org/10.4995/jarte.2024.20753 (2024).
Kallay, P. & Mihoc, T. D. Comparative analysis of frequent pattern mining algorithms. Acta Univ. Sapientiae Inform. https://doi.org/10.1007/s44427-025-00008-1 (2025).
Li, T., Liu, F., Chen, X. & Ma, C. Web log mining techniques to optimize Apriori association rule algorithm in sports data information management. Sci. Rep. 14(1), 24099. https://doi.org/10.1038/s41598-024-74427-z (2024).
Diaz-Garcia, J. A., Ruiz, M. D. & Martin-Bautista, M. J. A survey on the use of association rules mining techniques in textual social media. Artif. Intell. Rev. 56(2), 1175–1200. https://doi.org/10.1007/s10462-022-10196-3 (2023).
Shawkat, M., Badawi, M., El-ghamrawy, S., Arnous, R. & El-desoky, A. An optimized FP-growth algorithm for discovery of association rules. J. Supercomput. 78(4), 5479–5506. https://doi.org/10.1007/s11227-021-04066-y (2022).
Spasic, I., He, Q., Wang, H. & De Meo, P. Text mining and ontologies in biomedicine. Brief. Bioinform. 6(3), 246–256 (2005).
Hanisch, D., Fundel, K., Mevissen, H.-T., Zimmer, R. & Fluck, J. Prominer: Rule-based protein and gene entity recognition. BMC Bioinform. 6, 1–13 (2005).
Liu, B., Zhang, S., Tang, L. & Guo, J. Dictionary-based entity recognition in text mining. J. Biomed. Inform. 61, 108–118 (2016).
Smith, B., Williams, J. & Schulze-Kremer, S. Gene Ontology and the meaning of ‘function’. Bioinformatics 23(11), 1–6 (2007).
Noy, N. F. & McGuinness, D. L. Ontology development for the Semantic Web. Commun. ACM 45(2), 5–26 (2001).
Kumar, A., Smith, B., Borgelt, C., Ester, M. & Feldman, R. Text mining and ontologies for identifying associations. Brief. Bioinform. 6(3), 256–278 (2005).
Chen, J., Zhang, S., Huang, X., Huang, T. & Cai, Y.-D. Hybrid CNN-RNN model for gene-disease association mining. J. Biomed. Inform. 107, 103467 (2020).
Sharma, R., Kumar, P. & Gupta, R. Graph neural networks for gene-disease link prediction. Bioinformatics 38(3), 662–670 (2022).
Ali, A., Mohan, J., Nadaf, T., Ravishankar, H. & R, D. K. Bioinformatics-driven discovery of signaling pathways and genes influencing cervical cancer. SN Comput. Sci. https://doi.org/10.1007/s42979-024-03347-6 (2024).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28(11), 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53(D1), D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Ramachandra, H. V., Ali, A., Ambili, P. S., Thota, S. & Asha, P. N. An optimization on bicluster algorithm for gene expression data. In 2023 4th IEEE Global Conference for Advancement in Technology (GCAT), 1–6 (2023). https://doi.org/10.1109/GCAT59970.2023.10353373.
Xue, J., Wang, B., Ji, H. & Li, W. H. RT-Transformer: Retention time prediction for metabolite annotation to assist in metabolite identification. Bioinformatics https://doi.org/10.1093/bioinformatics/btae084 (2024).
Wang, Y. et al. Integrative graph-based framework for predicting circRNA drug resistance using disease contextualization and deep learning. IEEE J. Biomed. Health Inform. 29(11), 7932–7944. https://doi.org/10.1109/JBHI.2024.3457271 (2025).
Shi, W., Zhang, Y., Sun, Y. & Lin, Z. Function-genes and disease-genes prediction based on network embedding and one-class classification. Interdiscip. Sci. 16(4), 781–801. https://doi.org/10.1007/s12539-024-00638-7 (2024).
Xu, L. et al. Fine-tuning BERT for gene-disease association extraction using domain-specific ontologies. Artif. Intell. Med. 113, 102007 (2022).
Ha, J. DeepWalk-based graph embeddings for miRNA–disease association prediction using deep neural network. Biomedicines https://doi.org/10.3390/biomedicines13030536 (2025).
Ha, J. Graph convolutional network with neural collaborative filtering for predicting miRNA-disease association. Biomedicines https://doi.org/10.3390/biomedicines13010136 (2025).
Ha, J. SVDTI: Stacked variational autoencoder with SMILES-based drug representations for identifying drug-target interaction. Neurocomputing 661, 131837. https://doi.org/10.1016/j.neucom.2025.131837 (2026).
Ha, J. LncRNA expression profile-based matrix factorization for predicting lncRNA- disease association. IEEE Access 12, 70297–70304. https://doi.org/10.1109/ACCESS.2024.3401005 (2024).
Kim, K. & Ha, J. GMFLDA: improved prediction of lncRNA-disease association via graph convolutional network. IEEE Access 13, 85330–85341. https://doi.org/10.1109/ACCESS.2025.3568461 (2025).
Ha, J. Transfer learning with BioBERT embeddings for lncRNA–disease association prediction. IEEE. Trans. Comput. Biol. Bioinform. 22(6), 3463–3475. https://doi.org/10.1109/TCBBIO.2025.3628675 (2025).
Lin, C. H. et al. A disease-specific language representation model for cerebrovascular disease research. Comput. Methods Programs Biomed. https://doi.org/10.1016/j.cmpb.2021.106446 (2021).
Ha, J. & Park, S. NCMD: Node2vec-based neural collaborative filtering for predicting MiRNA-disease association. IEEE/ACM Trans. Comput. Biol. Bioinform. 20(2), 1257–1268. https://doi.org/10.1109/TCBB.2022.3191972 (2023).
Wang, C., Li, Y. & Chen, J. Text mining and knowledge graph construction from geoscience literature legacy: A review. Geosci. Front. 13(5), 101211. https://doi.org/10.1016/j.gsf.2022.101211 (2022).
Ahmed, K., Wang, E., Van den Broeck, G. & Chang, K.-W. Leveraging Unlabeled data for entity-relation extraction through probabilistic constraint satisfaction. In Proceedings of the 2021 Conference on Empirical Methods in Natural Language Processing (EMNLP 2021), 1–15 (2021). https://arxiv.org/abs/2103.11062
Chen, M., Tian, Y., Chang, K.-W., Skiena, S. & Zaniolo, C. Co-training embeddings of knowledge graphs and entity descriptions for cross-lingual entity alignment. arXiv preprint arXiv:1806.06478 (2018)
Zhang, Y. et al. KenDTI: An ensemble model for predicting drug-target interaction by integrating multiple data sources. IEEE Access 9, 100953–100963. https://doi.org/10.1109/ACCESS.2021.3092654 (2021).
Dhade, P. & Shirke, P. Federated learning for healthcare: A comprehensive review. MDPI 59(1), 230. https://doi.org/10.3390/2673-4591/59/1/230 (2024).
Rebholz-Schuhmann, D., Kirsch, H. & Couto, F. M. Text-mining solutions for biomedical knowledge discovery. Brief. Bioinform. 8(5), 358–370 (2007).
Kim, S., Lee, J. & Kang, J. Attention-based models for gene-disease prediction from unstructured biomedical text. IEEE Access 9, 12345–12356 (2021).
Hristovski, D., Peterlin, B., Mitchell, J. A. & Humphrey, S. M. Using literature-based discovery to identify disease candidate genes. Int. J. Med. Inform. 79(8), 522–529. https://doi.org/10.1016/j.ijmedinf.2010.05.002 (2010).
Wei, C.-H., Kao, H.-Y. & Lu, Z. PubTator: A web-based text mining tool for assisting biocuration. Nucleic Acids Res. 41(W1), W518–W522. https://doi.org/10.1093/nar/gkt441 (2013).
Boudellioua, I. et al. Semantic prioritization of novel causative genomic variants. PLoS Comput. Biol. 13(4), e1005500. https://doi.org/10.1371/journal.pcbi.1005500 (2017).
U. S. N. L. of M. for Biotechnology Information, Ed., NCBI Pubmed Database. https://www.ncbi.nlm.nih.gov/pubmed/
Bravo, Á., Piñero, J., Queralt-Rosinach, N., Rautschka, M. & Furlong, L. I. A knowledge-driven approach to extract disease-related biomarkers. Biomed Res. Int. 2014, 253128 (2014).
Ashburner, M. et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Hahsler, M., Gruen, B. & Hornik, K. Introduction to arules—A computational environment for mining association rules and frequent item sets. J. Stat. Softw. 14(15), 1–27 (2007).
Han, J., Kamber, M. & Pei, J. Data Mining: Concepts and Techniques 3rd edn. (Morgan Kaufmann, 2012).
Tan, P.-N., Steinbach, M., Karpatne, A. & Kumar, V. Introduction to Data Mining (Pearson, 2018).
Han, J., Kamber, M. & Pei, J. Data Mining: Concepts and Techniques (Morgan Kaufmann, 2011).
Alao, D., et al. Using association rules for ontology enrichment. In Proceedings of the 1st International Workshop on Knowledge Discovery and Knowledge Graphs (KDKG 2021), in CEUR Workshop Proceedings, vol. 2904, pp. 229–239 (2021). https://ceur-ws.org/Vol-2904/29.pdf
Razali, N. M. et al. Power comparisons of Shapiro-Wilk, Kolmogorov-Smirnov, Lilliefors, and Anderson-Darling tests. J. Stat. Model. Anal. 2(1), 21–33 (2011).
Yin, D., et al., Can large language models reliably extract human disease genes from full-text scientific literature? (2025). https://doi.org/10.1101/2025.07.27.667022.
Yang, H. et al. EnrichDO: A global weighted model for Disease Ontology enrichment analysis. Gigascience 14, 1021. https://doi.org/10.1093/gigascience/giaf021 (2025).
Jiang, T. et al. GENEasso: A curated resource of credible disease–gene associations across complex diseases from GWAS summary statistics. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaf1097 (2025).
Cheung, W.A., Ouellette, B.F., & Wasserman, W. W. Compensating for literature annotation bias when predicting novel drug-disease relationships through Medical Subject Heading Over-representation Profile (MeSHOP) similarity (2012). http://www.biomedcentral.com/1755-8794/6/S2/S3
Raber, J. et al. CD4+ T cells support hippocampal neurogenesis. Nat. Commun. 5 (2014).
Ohguro, N. et al. Erythropoietin and neovascular glaucoma. Invest. Ophthalmol. Vis. Sci. 53(8), 5278–5285. https://doi.org/10.1167/iovs.12-9794 (2012).
Oliveira, A. M. et al. USP6 gene rearrangement not in chondroblastoma. Am. J. Pathol. 179(5), 1777–1783 (2011).
Gao, M. et al. Identifying genetic signatures associated with oncogene-induced replication stress in osteosarcoma and screening for potential targeted drugs. Biochemical Genetics 62, 1690-1715 (2024).
Zhao, Y. et al. NOS2 expression and prognosis in chondrosarcoma. Clin. Cancer Res. 16(15), 3877–3885 (2010).
Coutinho, L. L. et al. NOS2 and COX-2 Co-expression promotes cancer progression: a potential target for developing agents to prevent or treat highly aggressive breast cancer. Int. J. Mol. Sci. 25, 6103 (2024).
Yang, I. V. & Schwartz, D. A. Epigenetics of idiopathic pulmonary fibrosis. Translational Research 165, 48-60 (2015).
Pandita, V. et al. Salivary mucin 4 levels in subjects with oral potentially malignant disorders and oral squamous cell carcinoma. Gulhane Medical Journal (2024).
Senevirathna, K. et al. Diagnostic potential of salivary IL-1β, IL-8, SAT, S100P, and OAZ1 in oral squamous cell carcinoma, oral submucous fibrosis, and oral lichen planus based on findings from a Sri Lankan cohort. Scientific Reports 14, 27226 (2024).
Khor, G. H. et al. DNA methylation profiling revealed promoter hypermethylation-induced silencing of p16, DDAH2 and DUSP1 in primary oral squamous cell carcinoma. International journal of medical sciences 10, 1727 (2013).
Schoenmakers, E. F. P. M. et al. Fusion of AHRR-NCOA2 in soft tissue tumors: Molecular and clinicopathologic analysis. Am. J. Surg. Pathol. 36(2), 182–190. https://doi.org/10.1097/PAS.0b013e31823c39a2 (2012).
Oliveira, A. M. et al. Gene fusion causes USP6 overexpression and fibroblast proliferation in fibromas. Mod. Pathol. 34(7), 1277–1286. https://doi.org/10.1038/s41379-021-00810-7 (2021).
de Jorge, E. et al. Role of CFHR1 in lymphoma treatment response. Blood 119(26), 6348–6357. https://doi.org/10.1182/blood-2012-02-413559 (2012).
Zhang, X. et al. GLT8D1 amplifies tumor aggressiveness in mucosal melanoma. Oncotarget 10(40), 4000–4014. https://doi.org/10.18632/oncotarget.27060 (2019).
Qiu, Y. et al. FOXK2 as an oncogenic driver in endometrial carcinoma. Gynecol. Oncol. 158(1), 206–214. https://doi.org/10.1016/j.ygyno.2020.05.023 (2020).
Sato, N. et al. FBXO32 silencing promotes tumor aggressiveness in endometrial carcinoma. Int. J. Cancer 134(2), 335–344. https://doi.org/10.1002/ijc.28349 (2014).
Amary, M. F. et al. HEY1–NCOA2 fusion as a hallmark for osteoblastoma. Nat. Commun. 9(1), 1–10. https://doi.org/10.1038/s41467-018-03833-5 (2018).
Landa, J. et al. ACVR2A mutations in bone tumors. J. Bone Oncol. 8, 28–33. https://doi.org/10.1016/j.jbo.2017.07.002 (2017).
Amary, M. F. et al. FOS is the most commonly altered gene in classic osteoblastoma, driving proliferation. Nat. Commun. 11, 1187. https://doi.org/10.1038/s41467-020-14945-4 (2020).
Kaur, R. et al. Role of CXCL10 in mastoiditis and related conditions. J. Infect. Dis. 196(11), 1626–1633. https://doi.org/10.1086/523110 (2007).
Szabo, G. et al. Key player in inflammatory response in mastoiditis: CXCL8/IL-8. Cytokine 72(2), 150–156. https://doi.org/10.1016/j.cyto.2015.02.003 (2015).
Flesher, D. L. et al. GTF2B and lupus nephritis: Gene transcription effects. Arthritis Rheumatol. 64(11), 3802–3810. https://doi.org/10.1002/art.34679 (2012).
Makishima, H. et al. CBL mutation leads to uncontrolled growth in chronic myelomonocytic leukemia. Blood 137(8), 1097–1108. https://doi.org/10.1182/blood.2020008069 (2021).
Naureckiene, S. et al. NPC2 mutations and Niemann-Pick disease type C2. Mol. Genet. Metab. 71(1–2), 65–74. https://doi.org/10.1006/mgme.2000.3076 (2000).
Smith, L. B. et al. ZMYND15 mutations linked to azoospermia and macrozoospermia. Hum. Genet. 143(5), 793–803. https://doi.org/10.1007/s00439-024-02564-8 (2024).
Dalbeth, N. et al. Minor role of AP1B1 in inflammatory response in gout. Rheumatol. Int. 25(3), 207–212 (2005).
Vasilevsky, N. A. et al. Mondo: integrating disease terminology across communities. Genetics https://doi.org/10.1093/genetics/iyaf215 (2025).
Bodenreider, O. The Unified Medical Language System (UMLS): Integrating biomedical terminology. Nucleic Acids Res. https://doi.org/10.1093/nar/gkh061 (2004).
Hamosh, A., Scott, A. F., Amberger, J. S., Bocchini, C. A. & McKusick, V. A. OMIM: Online mendelian inheritance in man. Nucleic Acids Res. 33(suppl_1), D514–D517 (2005).
Hewett, M. et al. PharmGKB: The Pharmacogenetics Knowledge Base (2002). http://www.nigms.nih.gov/
Milacic, M. et al. The reactome pathway knowledgebase 2024. Nucleic Acids Res. https://doi.org/10.1093/nar/gkad1025 (2024).
Oughtred, R. et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. https://doi.org/10.1002/pro.3978 (2021).
Szklarczyk, D. et al. The STRING database in 2021: Customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 49(D1), D605–D612. https://doi.org/10.1093/nar/gkaa1074 (2021).
Acknowledgements
Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R384), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through the Large Group Project under Grant Number (RGP.2/702/46).
Funding
Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2026R384), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia. The authors extend their appreciation to the Deanship of Research and Graduate Studies at King Khalid University for funding this work through the Large Group Project under Grant Number (RGP.2/702/46).
Author information
Authors and Affiliations
Contributions
MA.Q., Data Creation, Implementation, methodology, and Writing. M.A., Supervision, writing, and validation. J. U proofreading, writing, and Supervision. HS.H writing, visualization. A. R., interpretation, Writing, and Visualization. W. A., Writing, Interpretation, and Implementation HK. A., Supervision, funding, and Proof-Reading. HA. M., formal analysis, writing, and resources. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Naqash, M.A., Amin, M., Uddin, J. et al. Ontology-driven association rule mining for biomedical entity relationships: integrating hierarchical knowledge to improve gene-disease discovery. Sci Rep (2026). https://doi.org/10.1038/s41598-026-42584-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-026-42584-y