Abstract
Improvements in denitrifying bacteria since 1970 have optimized the activity of autotrophic denitrification and heterotrophic nitrification. Synchronous autotrophic biofilms, denitrification, and nitrification have enhanced nitrogen removal from industrial wastewater, but so far, this process relies on variables such as dissolved oxygen, pH, and carbon sources. The aerobic denitrification capability of the coking wastewater-derived Stutzerimonas stutzeri KA1 is quantified in the present investigation to enhance nitrogen removal potential under various environmental conditions. The strain was studied for its ability to isolate nitrate using sodium acetate as the carbon source, and the effectiveness of the microorganism was tested at different dissolved oxygen, pH, and C/N levels. Findings indicate that strain KA1 achieved nearly 100% nitrate elimination within 40 h at pH 6–10 and an optimal C/N ratio of 8, demonstrating resilience under both aerobic and anaerobic conditions. Strain KA1 showed robust denitrification across a wide range of dissolved oxygen levels (0-100%), including stable nitrate reduction under aerobic bulk conditions. Bioaugmentation experiments conducted in a Sequential Batch Reactor (SBR) confirmed that strain KA1 significantly enhanced nitrogen removal, particularly under saline wastewater conditions, exceeding the control systems. In conjunction with nitrogen elimination, the strain also demonstrates robust Chemical Oxygen Demand (COD) reduction, efficiently degrading organic pollutants in coking wastewater. Compared to other studies, Stutzerimonas stutzeri KA1 demonstrated higher denitrification efficiency and greater resistance to environmental stress, making it a scalable, cost-effective solution for denitrifying industrial wastewater. The results offer valuable insights into optimizing wastewater treatment parameters and can advance microbial biotechnology to foster sustainable environmental practices.
Similar content being viewed by others
Introduction
The 20th century was the century of capitalist entrenchment, with an unprecedented spurt in industrialization across the globe, and it was during this century that the world saw incredible improvements in humanity’s living standards over the last 120 years1, which was previously unimaginable. Nonetheless, this rapid industrialization has caused havoc to Earth’s ecosystem2. It was too late when mankind realized the harm it had done to the environment3. Although governments around the world are trying their best to mitigate environmental problems, the ongoing degradation process continues to worsen the situation4. Stakeholders take the problem of environmental degradation seriously in response to this crisis. All the adverse effects of industrialization are considered a formidable challenge that should be addressed immediately. Wastewater is a major industrial byproduct and a major source of pollution, contaminating freshwater with various pollutants5,6,7. Depending on the industry that emits the contaminants8, the consequences impact marine life and humans9. The most important solution to environmental pollution is industrial wastewater treatment. This will reduce pollution10 and protect marine ecosystems and human health11,12,13. In addition, it plays a crucial role in regulatory compliance14, saving valuable resources15, and sustainable industrial practice10. Treatment of wastewater is a pillar of responsible and ethical industrial activity16, which makes significant contributions to creating a cleaner, healthier, and sustainable future through human activity17.
Over the last 10 years, China has become the world leader in steel production and exports, increasing its crude steel production from 683 million metric tons in 2011 to an unbelievable 1.337 billion metric tons in 202118. This meteoric growth is attributed to the strength of local infrastructure-based consumption19 and competitive pricing in the global market18, supported by effective production practices20 and economies of scale21. The coke ovens play a central role in the main steel production process using the blast furnace method. These special ovens transform metallurgical coal into coke, a raw material for blast furnaces. Coke has two functions: it can serve as a reducing agent to obtain iron from iron ore22 and as a source of energy in the furnace stove23. Wastewater contaminated with pollutants is produced during the coking process. These are organic compounds that include polycyclic aromatic hydrocarbons (PAHs) and phenolic compounds24, nitrogen-containing organic compounds25, and heterocyclic compounds26.
The wastewater is also known to contain suspended solids, heavy metals, ammonia, thiocyanate, and cyanide, as determined in the research27,28. The effects of nitrogen-containing organic compounds29, inorganic nitrogen compounds like nitrate (NO3-) and nitrite (NO2-), as well as ammonia (NH3) and some of the heterocyclic compounds that are formed during the coking process are significantly harmful to human and marine organisms30,31. Nitrates are dangerous when present at high levels in drinking water, causing methemoglobinemia32, also known as Blue Baby Syndrome, in infants. Moreover, nitrates react with amines to form carcinogenic nitrosamines. (33, which are known to cause cancer. In the agricultural sector, when freshwater with high levels of nitrates and nitrites is used, nitrosamines accumulate in leafy greens and root vegetables, posing serious health risks to consumers34. Besides, dumping nitrogen-contaminated coking wastewater into freshwater bodies promotes the proliferation of harmful algal blooms35, affecting the aquatic food chain and endangering human and marine life. Ammonia, which is abundant in water, disrupts the salt balance of fish and invertebrates, affecting their growth and reproductive processes36. This disruption eventually threatens several delicate species in the marine ecosystems. Also, nitrogen-rich water bodies, particularly in rivers and coastal areas, are enriched, which causes eutrophication37. This uncontrolled increase in algae and aquatic plants, promoted by high nitrogen concentrations, leads to hypoxia38 when they die, eliminating dissolved oxygen and a major cause of death among marine organisms. Altogether, these toxic substances contribute to the environmental footprint, worsening the state of marine environments and increasing the impact on both human and marine environments39,40,41. Therefore, the necessity of treating coking wastewater for nitrogen-based compounds and other organic and inorganic pollutants cannot be overestimated, and heterotrophic denitrification is an effective and efficient option for this wastewater treatment42.
Denitrification is a central step in the nitrogen cycle that results in the release of gaseous end products of NO3−-N in soil and water. Nitrogen (N), is an oxidising agent, which is reduced in a sequence of four enzyme-mediated reactions involving nitrate reductase (NAR), nitrite reductase (NIR), nitric oxide reductase37, and nitrous oxide reductase (NOS)43,44. This enzymatic cascade ultimately leads to the formation of N2. The denitrification pathway can be succinctly represented as follows:
NO3 −→NO2 −→NO→N2O→N2.
Another significant milestone was recorded in 1972, when Arthrobacter sp. was isolated and shown to induce heterotrophic nitrification (HN) under natural conditions45. Later, in 1983, Thiosphaera pantotrova (now known as Paracoccus denitrificans), an expert heterotrophic nitrifier capable of performing autotrophic denitrification (AD), was discovered in the sludge of a wastewater treatment plant46. These discoveries prompted scientists to focus on HN and AD research, which led to a wide array of research. In 2003, HN perspective received another focus as researchers ventured into this field by growing mixed cultures of two or more strains, thus broadening the substrate and product range and increasing the technological uses of HN47. The HN view was further extended in 2005, as scientists theorize and actualize simultaneous nitrification and denitrification (SND) in the same reactor48. Later studies go further and develop into autotrophic denitrification biofilms (ADB), examining intricate interactions among variables, including carbon sources, temperature, pH, Dissolved Oxygen (DO), and other environmental factors, which significantly influence ADB denitrification activity. This development in the sphere has taken the practical implementation of ADB to a new level.
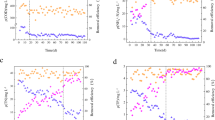
Several studies have described the removal of nitrogen from municipal and industrial wastewater via heterotrophic denitrification, providing useful information. Li and Lu49, one of the earlier researchers, utilized Nanofiltration concentrate in a sequencing batch reactor to treat coking wastewater, involving a biological denitrification process, with subsequent removal efficiencies of 47.6% for Chemical Oxygen Demand (COD), 61.1% Total Nitrogen (TN), and 94.6% nitrate. The pivotal microorganisms driving denitrification belonged to the genera Thauera, Hyphomicrobium, Methyloversatilis, Hydrogenophaga, Ignavibacterium, Rubrivivax, and Parvibaculum.Notably, the denitrifying genes narG, nirS, nirK, and nosZ were investigated, with nirS emerging as the most efficient gene, quantified at 104–105 copies/ng in DNA, underscoring its importance in denitrification. In a parallel, Li, et al50. elucidated the prowess of sequencing batch reactors in eliminating nitrate from reverse osmosis concentrate originating from coking wastewater with the following remarkable nitrate removal efficiencies: 79.5% for Chemical Oxygen Demand (COD), 90.5% for Total Nitrogen (TN), and a staggering 93.1% for nitrate. The authors investigated the microbial community using advanced molecular methods and identified several key genera, including Hyphomicrobium, Thauera, Methyloversatilis, and Rhodobacter. At the same time, Zheng, et al51. studied the emissions of NO2 and NO, and used a biological aerated filter with high NO2 and NO elimination, achieving average removal efficiencies of 92.8% and 73.3% of NH3-N and COD, respectively. In the meantime, Yuan, et al52. investigated the application of Moving Bed Biofilm Reactors (MBBRs) in denitrification. They found that a range of carriers, such as polyethylene, polypropylene, polyurethane foam, and haydite, could be used under the following conditions: pH of 7.2 to 8.0, temperature ranged between 24 and 26 o C, a hydraulic retention time (HRT) of 12 h, and a final external methanol dosage of 25.5 mg L− 1. Polyethylene was found to be the most effective denitrifying carrier in MBBRs, with impressive removals of 44.9 ± 19.1% for Total Nitrogen (TN) and 83.4 ± 13.0% for nitrate. These academic efforts highlight the processes and strategies for nitrogen removal from wastewater with different types and chemical characteristics, providing significant insights into the symbiotic interactions between microorganisms and novel reactor configurations to achieve optimal denitrification results.
The purpose of the paper is to explain a niche aspect of wastewater treatment, which is the coking wastewater treatment and introduce more effective ways of its treatment. Even though the taxonomic classification of Stutzerimonas stutzeri has been reinstated within the Pseudomonas stutzeri complex, which is commonly reported in denitrification experiments, its strain-specific aerobic performance in denitrifying high-strength coking wastewater has not been described before. In particular, strain KA1 demonstrates exceptional nitrate removal efficiency, tolerance to a wide range of dissolved oxygen levels, and stable performance under saline conditions, distinguishing it from previously reported Pseudomonas stutzeri strains.
The gap this study addresses is that the performance of denitrification under varying dissolved oxygen conditions in the treatment of nitrogen-rich coking wastewater has not been adequately described at the strain level. Whereas heterotrophic and aerobic denitrification has been extensively studied in municipal and industrial wastewater systems, little has been done in the strain-specific nitrate removal behavior of Stutzerimonas stutzeri in high-strength coking wastewater matrices. Secondly, the isolation and characterization of Stutzerimonas stutzeri strain KA1 and its nitrate removal activity under the investigation of various carbon sources, including methanol, sodium citrate, glucose, sodium acetate, ethanol and sodium succinate, with the discovery of Sodium acetate as the most effective carbon source, which reached a 100% NO3 -N removal rate in 40 h. Thirdly, the effects of the carbon-to-nitrogen (C/N) ratio and dissolved oxygen are determined to be optimized during denitrification. Fourthly, the paper has also determined the optimal pH range for nitrogen removal by Stutzerimonas stutzeri, demonstrating that this bacterium is robust across a wide pH range. The results provide experimentally supported information on the efficiency of nitrogen removal and operational stability in the coking wastewater setting, which may have practical implications for bioaugmentation-based wastewater treatment without exaggerating the novelty at the species level.
Materials and methods
Origin of bacterial strain and preparation of culture media
The water sample was collected from the anoxic zone of Dongfang Zhengda coking wastewater and used to isolate aerobic denitrifying bacteria. A denitrification medium (DM) was prepared using distilled water with the following composition per liter: sodium acetate (anhydrous) 1.71 g, KNO3 0.6 g, K2HPO4 1.6 g, MgSO4•7H2O 0.10 g, CaCl2 0.02 g, FeSO4•7H2O 0.005 g, and a trace-element solution 0.10 ml. The trace-element solution contained (per liter): MgSO4•H2O 0.0344 g, H3BO30.05 g, ZnCl2 0.07 g, Na2MoO4•2H2O0.0726 g, CaCl2•2H2O 0.02 g, NiCl2•6H2O 0.024 g, CoCl2•6H2O 0.08 g, FeSO4•7H2O 1.00 g. For plate cultivation, agar (20 g L⁻¹) and bromothymol blue solution (1 mL of 1%) were added to the DM. All media were adjusted to an initial pH of 7.0–7.5.0.5 before sterilization. Sterilization was performed at 121 °C for approximately 90 min to prevent contamination and ensure consistent experimental conditions.
Screening and isolation of strain KA1
A meticulous experimental procedure introduced 10 ml of coking wastewater into a conical flask, which was then precisely blended with 90 ml of sterilized Denitrification Medium (DM). This inoculated medium was enriched and cultivated at 30 degrees Celsius with agitation at 120 rpm. After that, the culture medium was refrigerated at 4 °C for preservation. The bacterial solution was then inoculated at 30 °C and 120 rpm until diverse colonies were visible. Subsequently, a cell suspension of 10 ml was obtained from the enriched medium and acclimated in another 90 ml of sterile DM under the same temperature and agitation conditions. This acclimation process has three stages to cultivate the prevailing strain, specifically for nitrate nitrogen degradation. In addition, a careful dilution series was initiated: 1 ml of the domesticated bacteria suspension was added to 9 ml of sterile purified water, resulting in a concentration gradient of 10 − 1. Then 1 ml of the cell dilution obtained was again added to 9 ml of sterilized purified water to obtain successive gradients of 10 − 2, 10 − 5, 10 − 6, 10 − 7, 10 − 8, 10 − 9, etc. The diluted cell suspension was immediately evenly dispersed on agar plates, and the cultures were incubated in a constant-temperature biochemical incubator (30 C). Individual colonies with different morphologies were picked based on their visible appearance. A rigorous separation and purification of these colonies was then performed by streaking them onto a solid medium plate. This carefully taken purification process was carried out three times to ensure that only pure, uncontaminated colonies were obtained, thereby assuring the scientific integrity of the experiment.
Bioaugmentation for treating coking water in SBR by aerobic denitrification bacteria
The experiments were performed using activated sludge from the Beijing Xiaojiahe Sewage Treatment Plant, with an initial concentration of about 15,700 mg/L. Three identical sequential batch reactors were operated: SBR1 (control), SBR2, and SBR3 (bioaugmented reactors). Each reactor had a working volume of 300 mL, which consisted of 150 mL of activated sludge and 150 mL of synthetic coking wastewater.
The SBRs operated on a 12-hour cycle, consisting of an 8-hour aerobic reaction stage, a 3-hour settling period, and a 1-hour supernatant replacement phase. Strain KA1 was first cultured in denitrification medium at 30 °C and 120 rpm for 24 h. Cells were then collected by centrifugation at 10,000 rpm for 5 min. The bacterial suspension was added to SBR2 and SBR3 at volumes of 7.5 mL and 15.0 mL, respectively, while SBR1 was maintained without bacterial addition. Reactor operation continued until nitrate concentrations in the effluent stabilized, indicating successful bioaugmentation.
Assessment of COD Reduction
The chemical oxygen demand is high in industrial, marine, and coking wastewater. The steps for the COD experiment are as follows: The researcher prepares the enrichment medium by mixing KNO3 0.6 g/L, sodium acetate 1.71 g/L, MgSO4·7H2O 0.1 g/L, K2HPO4 1.6 g/L, CaCl2 0.002 g/L, FeSO4 · 7H2O 0.005 g/L, and a trace element solution 0.1 mL, with a pH of 7.8–8.0.8.0. Sterilize 100 mL of the enrichment medium in an anaerobic bottle for 2 h. at 121 °C. After sterilization, cool the culture for 20 min at the sterilization table. After cooling, 10 mL of stored bacteria (kept at 4 °C) was added to the medium. After adding, put the anaerobic bottle in the shaker at 30 °C for 24–48 h. After 2 days, take 4 bottles and put 100 mL of coking wastewater in each bottle (2 bottles contain only 100 mL of coking wastewater, and the other 2 contain 100 mL of coking wastewater and 0.0134 g Sodium Acetate). Put these four bottles in the sterilization pot. After sterilization and cooling, a 0-hour sample was collected into centrifuge sampling tubes. Then, add 10 mL of bacteria to each bottle grown in the shaker; put the leftover bacteria in the refrigerator and store them at 4 °C. Take 2.5 m of water sample into the digestive tube, add 1.5 mL of digestive fluid and 3.5 mL of catalytic fluid. Tighten the digestive tubes with caps, shake evenly, and put the digestive tubes in the heating Digester at 150 °C for 2 h. Cool the digestive tubes at room temperature. Take five 50 mL conical flasks and add the digestive tube mixture to each. Add 500 µl of the indicator. The color is titrated with ammonium ferrous sulfate (titration solution) and ranges from green to blue to grey. I recorded the milliliters of Ammonium ferrous Sulfate used in the titration. (A) Use UV water for calibration and record the millesimal number (B) (blank sample) for ammonium ferrous sulfate solution calibration. Ferrous ammonium Sulfate must be calibrated before use. Add another 5 mL of UV water, then 5 mL of concentrated sulfuric acid. Add 3 mL concentrated sulfuric acid and K2Cr2O7 solution (0.2452 g → 100 mL), 5 mL. Add 500 µl indicator titration record millimeter. The researcher collects samples every 12 h for 3 days and measures COD using the method described above.
Chemical oxygen demand (COD) was determined using the dichromate digestion method followed by titration with ammonium ferrous sulfate (FAS). The FAS solution was standardized prior to analysis, and COD values were determined as the difference in titrant volume consumed by the blank and the wastewater sample after digestion. All COD measurements were carried out using the same procedure to ensure consistency across experiments.
Identification and characterization of the strain
The strain’s genetic material was extracted using a DNA extraction kit (MPbio, America) according to the manufacturer’s instructions. The 16 S ribosomal DNA (rDNA) gene was then amplified using the polymerase chain reaction (PCR) methodology, with universal primers specific to 16 S rDNA (27 F: 5’-AGAGTTTGATCMTGGCTCAG, 1492R: 5’-TACGGYTACCTTGTTACGACTT-3’) as previously described53. The PCR experiment involved pre-denaturation at 94 °C for 3 min, denaturation at 94 °C for 30 s, annealing at 54 °C for 30 s, and an extension step at 72 °C for 1 min and 30 s, followed by a final extension step at 72 °C for 10 min. Meiji Biomedical Technology Co., Ltd, then sequenced the amplified PCR products. The sequencing data were obtained and analyzed using the BLAST tool (https://www.ncbi.nlm.nih.gov/BLAST/Blast.cgi), which enabled in-depth analysis of the genetic sequence and allowed accurate strain determination. The 16 S rRNA gene sequence of strain KA1 was amplified and used for taxonomic identification by comparing it with reference sequences in the NCBI database using the BLAST algorithm. Moreover, the sequences were subjected to complete Blast analysis in the National Center for Biotechnology Information (NCBI) database. Phylogenetic analysis was conducted using the neighbor-joining method based on aligned 16 S rRNA gene sequences in MEGA version 11, with bootstrap analysis performed using 1,000 replicates to evaluate branch robustness.
Analysis of performance of denitrification strain
The aerobic denitrification capacity of the isolated strain was measured by a shaking flask experiment. The strain (3 ml) was inoculated into 30mL denitrification medium and cultured at 30 °C and 120 rpm. Meanwhile, samples were collected every few hours and centrifuged at 12,000 RPM for 1 min to obtain the supernatant for detecting the removal of nitrogen-containing substances. Single-factor experiments (carbon sources, DO, salinity, pH, and C/N) were conducted to investigate the effects of various culture conditions on the strain’s denitrification capability. In all single-factor experiments, the strain stored in the 4 °C refrigerator needed to be activated in advance. The specific steps were as follows: removed 3 ml of the required bacterial solution from the refrigerator and placed it into the sterilized 30 ml DM, then incubated in the shaker until the bacterial solution was cloudy.
This concoction was placed in a shaker at 120 rpm and incubated until the bacterial solution reached a cloudy consistency, indicating it was ready for experimentation. Six widely used carbon substrates were systematically examined to identify optimal carbon sources for nitrogenous wastewater treatment, including methanol, sodium citrate, glucose, sodium acetate, ethanol, and sodium succinate. These compounds were used singly as the sole carbon sources to assess their impact on the strain’s denitrification potential. The experiments were initiated with an initial Carbon-to-Nitrogen (C/N) ratio of 5, and the nitrate-nitrogen concentration was maintained at 100 mg/L. The effect of DO on nitrogen removal was assessed by incrementing oxygen to 0%, 10%, 20%, 30%, 50%, and 100%. At the same time, to achieve a more appropriate pH and C/N range, the nitrogen removal rates at pH 6, 7, 7.5, 8, 9, and 10, and C/N 2, 4, 5, 6, 8, and 10 were investigated.
In all single-factor experiments, activated bacterial cultures were inoculated into sterile denitrification medium and incubated at 30 °C with shaking at 120 rpm. Each experimental condition was performed in duplicate, and measurements were taken from two parallel samples. Final results are presented as average values obtained from the replicate experiments.
Data analysis
The collected data were thoroughly analyzed in Microsoft Excel 2010, and graphical representations were generated using Origin 8.5, ensuring an extensive and inclusive evaluation of the experimental outcomes. The consequences stemming from these experiments are logically accessible as the mean and standard deviation (SD) of the mean, providing a robust statistical basis for the conclusion.
The nitrate removal efficiency was calculated using the following equation:
Nitrate removal efficiency (%) = [(C₀ − Cₜ)/C₀] × 100.
where C₀ is the initial concentration of nitrate-nitrogen (mg L⁻¹) in the medium, and Cₜ represents the nitrate–nitrogen concentration (mg L⁻¹) measured at time t during the experiment, this was the same equation used to assess nitrate removal performance under various experimental conditions.
Results and discussion
Isolation and identification of aerobic denitrifiers
After coking, wastewater was added to a denitrifying medium for enrichment. Pure bacterial cultures were isolated by gradient dilution and plate streaking. A BLAST comparison found that the strain showed high homology with a collection of bacteria of the genus Stutzerimonas, and the strain was named Stutzerimonas Stutzeri. In particular, the strain was named Stutzerimonas stutzeri, a species that is known to have denitrifying properties. To support these results, 16 S ribosomal DNA (rDNA) sequences from the isolated strain and from evolutionary and functionally related denitrification strains were submitted for analysis. The ensuing phylogenetic tree construction, as illustrated in Fig. 1, definitively established the strain’s classification as Stutzerimonas stutzeri and strengthened its denitrifying nature as a microbial taxon. Based on the 16 S rRNA gene sequence, strain KA1 showed the greatest similarity to representatives of the Stutzerimonas stutzeri lineage, confirming its species-level classification.
Neighbor-joining phylogenetic tree based on partial 16 S rRNA gene sequences showing the taxonomic position of strain KA1 relative to closely related reference strains. Bootstrap values (1,000 replicates) greater than 50% are shown at branch nodes. The scale bar indicates nucleotide substitutions per site.
A controlled experiment was conducted to investigate the greater nitrogen removal capacity of the isolated strain in coking wastewater. The strain was cultured in a liquid medium containing potassium nitrate as the nitrogen source and sodium acetate as the carbon source. The cultivation was conducted under constant conditions of 30 degrees Celsius and 120 revolutions per minute. Over a specified period, nitrate and nitrite levels in the medium were monitored at specified intervals. The results showed distinct patterns across strains. Strain 1 exhibits a slow growth rate and achieves a low rate of NO3– N removal of merely 51.28% after 30 h, as shown in Fig. 2. On the other hand, strain 2 shows promising progress in NO3–N removal efficiency, rising from 29.18% to an astonishing 92.45% over 18–32 h. Remarkably, in strain 3, the concentration of NO3–N in the aerobic denitrification medium quickly decreases from 128.73 mg/l to 63.17 mg/l between the 18th and 32nd hours. In strain 4, the NO3–N removal rates were about 45% and 51% after 30 h, respectively. Strain 5, on the other hand, showed outstanding performance, achieving a total NO3–N removal rate within 20 h. On the contrary, a tremendous progress was achieved by strain 2, as the NO3–N removal efficiency increased from 29.18% to an impressive 92.45% during the 18–32 h. It is interesting to note here that in strain 3, the concentration of NO3–N in the aerobic denitrification medium drops rapidly from 128.73 mg/l to 63.17 mg/l during 18th to 32nd hours. In strain 4, the NO3–N removal rates were about 45% and 51% after 30 h, respectively. Strain 5, on the other hand, showed exceptional performance, achieving a total NO3-N removal rate of over 20 h. Contrastingly, strain 6 had a slow rate of NO3–N removal (54% at 30 h of cultivation). Scrupulous data analysis revealed that strains 2 and 5 were the most capable of utilizing carbon sources, thereby demonstrating the highest efficiencies in NO3- -N removal. These results underscore the importance of strain selection and highlight the critical role of specific microbial strains in optimizing denitrification processes in complex coking wastewater scenarios.
Aerobic denitrification performance of strain KA1.
Effects of different factors on aerobic denitrification by the strain
Carbon sources
In heterotrophic denitrifying bacteria, carbon is the main source of cell growth and metabolism, and it plays a key role in nitrogen elimination54. To attain this goal, heterotrophic denitrifiers use NO3 -N and NO2 -N as electron acceptors. In contrast, organic carbon sources serve as electron donors, facilitating the reduction of nitrogen compounds during electron transfer. Microbial growth and enzyme activity differ with different carbon sources because of their distinctive structures, and therefore, nitrogen removal efficiency is not uniform across all carbon sources. Figure 3 explains the effect of different carbon sources on the treatment of the nitrogen-bearing wastewater using strain KA1. Observations revealed that when methanol served as the sole carbon source, the strain exhibited sluggish growth, resulting in a NO3–-N removal rate of only 70% after 60 h (Fig. 3a and b). Contrastingly, the medium’s NO3–-N concentration decreased from 122.5 mg/l to 34.43 mg/l between 26 and 60 h when glucose was used as the sole carbon source. Consequently, the NO3–-N removal efficiency surged from 13% to 75% during the same period. In media supplemented with ethanol, the NO3–-N removal rates were approximately 75% and 90% after 26 h, respectively. They were strikingly using sodium acetate as the sole carbon source, resulting in a remarkable 100% NO3–-N removal rate within 40 h. However, with sodium citrate and sodium succinate as the sole carbon sources, the NO3–-N removal rate ranged from 95% to 100%, but did not reach complete denitrification.
Figure 3 illustrates that the strain exhibited nitrite accumulation traits when methanol, sodium citrate, glucose, sodium acetate, ethanol, and sodium succinate were used as sole carbon sources. However, nitrite concentration varied depending on the carbon source. Use of methanol and glucose as the sole carbon sources resulted in low nitrite production, keeping nitrite levels below 1 mg/L during denitrification. During denitrification, bacterial growth was greatest at 4 h (21.19 mg/l) and 26 h (81.82 mg/l) in media containing sodium acetate and sodium citrate, respectively. On the same note, ethanol- and sodium succinate-based denitrification media demonstrated peak NO2 -N at 40 h (60.79 mg/l) and 60 h (80.79 mg/l), respectively. When critically analyzed, it was evident that Sodium Acetate was the most effective carbon source, with the highest NO3 -N removal efficiency, followed by glucose, sodium citrate, and finally sodium succinate. Compared with other solvents, Methanol exhibits the lowest NO3- -N removal rate. This conclusion emphasized that carbon source structure and molecular composition can significantly affect nitrogen removal efficiency55. Overall, the use of carbon sources that are more readily digestible and contain small molecules facilitates efficient nitrogen removal56. In addition, the inequalities in the metabolic routes, which were correlated to certain carbon sources, explained the difference in the nitrite accretion. For example, sodium acetate was directly converted to acetyl-CoA by denitrifying bacteria, bypassing the tricarboxylic acid (TCA) cycle and avoiding conversion to NADH2, an essential source of microbial energy. The resulting energy deficit during the NO3 -N reduction to NO2 -N leads to nitrite accumulation57. Instead, certain carbon sources, such as methanol, were challenging because of the lag period in denitrification and failure to enter the TCA cycle, which interferes with efficient reduction of nitrogen compounds58. These results clarify the complexity of the interactions among the choice of carbon source, microorganism metabolism, and the capacity to denitrify materials, which is highly informative for enhancing denitrification methods in multifaceted wastewater treatment systems.
The effect of carbon source on NO3−-N concentration (a), NO3−-N remove efficiency (b) NO₂⁻-N concentration (c) Nitrate Removal Efficiency.
Effect of C/N
Carbon is pivotal in cell growth and nitrate reduction processes, with aerobic denitrification intricately linked to heterotrophic denitrification. In this experiment, Sodium succinate was used as the carbon source, participating actively in the tricarboxylic acid cycle. The concentration of NO3−-N was pre-established, while the C/N ratio was manipulated by adjusting the carbon source concentration, resulting in C/N mass ratios of 2, 4, 5, 6, 8, and 10, respectively. It is a 30-hour Experiment, and we can take nitrate and nitrite samples every 2 h.
According to the diagram, the denitrification rate at a C/N ratio of 8 is the fastest. After 12 h of culture, the nitrate concentration in the medium had dropped to 0 mg/L. However, when the C/N ratio was 2, the nitrate concentration in the medium decreased from the initial 137.35 mg/L to 90.28 mg/L after 30 h of shaker culture. This is because if the carbon source is insufficient, there will not be sufficient electron flow to support bacterial growth, and the corresponding denitrification efficiency will also be low. Moreover, we can see that the nitrogen removal rate increased rapidly with increasing C/N ratio. This is mainly because the more abundant the initial carbon source, the faster bacteria grow. However, at a C/N ratio of 10, the denitrification rate decreased, but the change was not noticeable.
This is because when the carbon source provided is far higher than the bacteria’s demand, the carbon source is no longer a limiting factor, the bacteria’s growth and metabolic activity are stable, and the denitrification and nitrogen removal efficiency do not change much. In addition, it can be seen from Fig. 4 (a) and (b) that under different C/N ratios, the nitrite concentration was deficient, indicating that the activity of nitrite reducates of the strain was very high. The nitrite produced by bacterial nitrate reductase was also rapidly reduced under the very high activity of nitrite reductase, as shown in Fig. 4 (a) and (b).
(a) C/N ratios on NO3−-N concentration and NO3−-N removal efficiency, (b)and NO₂⁻-N concentration and NO₂⁻-N removal efficiency in the aerobic denitrifier, Stutzerimonas stutzeri KA1.
Effect of DO
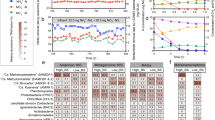
Dissolved oxygen is a complex parameter that significantly influences the growth patterns of denitrifying strains. The different strains exhibit distinct growth and denitrification capabilities due to variable dissolved oxygen concentrations59. The concentration of dissolved oxygen was controlled by bubbling air into the serum flask and then varying the amount of oxygen. The experiments were conducted with sodium acetate as the sole carbon source. The medium was inoculated with 100 mg/L of nitrate and a C/N ratio of 8, incubated at 120 RPM and 30 C/O for 32 h (Fig. 5). Nitrate removal under anaerobic conditions (0% oxygen volume ratio) was rapid. At 36 h, the NO3–N level dropped to 1.12 mg/L, and the resulting percentage removal of nitrate was impressive at 99.12%. Nevertheless, the accretion of nitrite was insignificant, and it was − 0.322 mg/L after 32 h. Negative values of NO2 -N in the plots indicate concentrations below the level of detection and are not directly interpreted as negative concentrations, but rather as complete or near complete removal of nitrite. Denitrification performance was consistent across different dissolved oxygen concentrations. The percent removal of NO3, when the ratio of oxygen volume was 10% and 20%, was 98.34% and 99%, respectively. At the same time, in these circumstances, the concentration of NO2 -N was − 0.322 mg/L and 0.316 mg/L after 32 h. The 50% oxygen volume ratio achieved 91% nitrate removal after 16 h and 98% after 24 h, whereas the 30% oxygen volume ratio achieved 91% after 16 h and 98% after 24 h. Interestingly, additional increases in dissolved oxygen content failed to significantly affect the rate of nitrate removal, which remained constant at 100%. Despite the strain KA1 showing a good ability to remove nitrate even in the presence of 100% dissolved oxygen, genetic screening of nitrate reductase genes (e.g., napA or narG) or enzyme activity tests were not performed in the study. The resultant reduction in nitrate under fully aerobic bulk environments can therefore result from aerobic denitrification facilitated by periplasmic nitrate reductase or from a decrease in nitrate under anoxic microsites within the inner regions of bacterial aggregates/flocs. Previous studies have demonstrated that high dissolved oxygen at the bulk scale does not necessarily preclude localized oxygen-limited microenvironments that support denitrification activity, and similar mechanistic ambiguity has been widely acknowledged in aerobic denitrification research60,61,62,63. Typically, denitrification is believed to occur strictly under anaerobic conditions because oxygen inhibits electron transfer to NO₂⁻-N and NO3−-N. However, the data presented in Figs. 5, 6 indicate that increased dissolved oxygen concentrations did not inhibit denitrification. On the contrary, they seemed to promote it. This observation is consistent with previously reported aerobic denitrification–like behavior; however, the underlying enzymatic mechanisms were not directly examined in this study64,65,66,67.
Reduction of nitrate (a) and nitrite (b) Stutzerimonas Stutzeri with different dissolved oxygen levels(c) Nitrate Removal Efficiency.
Effect of pH
The pH factor plays a pivotal role in the denitrification process of aerobic denitrifying bacteria. The pH of the environment greatly influences the absorption of nutrients by microorganisms, altering the charge of the plasma membrane and, in turn, affecting their life activities. PH also regulates the activity of enzymes involved in microbial metabolism; excessively low or high pH can pose a danger to microbial growth68. Under typical conditions, both acidic (pH < 5) and alkaline (pH > 10) environments inhibit the growth of aerobic denitrifying bacteria69. Maoxia Chen isolated HN-02, identified as Aeromonas sp., from activated sludge in a lab-scale CASS reactor. This strain exhibited remarkable acid and alkali resistance, demonstrating robust denitrification performance even at pH values of 2.3 and 1170. In liquid culture medium, nitrate and nitrite concentrations were measured at various time points under different pH conditions. Strain KA1 exhibits extraordinary aerobic denitrification capabilities within a pH range of 6 to 10. The NO3–-N removal rate reaches nearly 100% within this pH range, with denitrification rates remaining consistent across pH 6–9. Exclusively, at pH 6, the NO3-N concentration decreased from 115.35 mg/L to 1.5 mg/L by 42 h. Likewise, at pH 7, the nitrate deletion rate was 98% after 42 h of culture. Furthermore, denitrification rates accelerated with increasing initial pH of the medium, indicating the bacteria’s ability to thrive in alkaline environments and to play a denitrification role; they exhibit facultative basophilic characteristics. However, at pH 10, the denitrification rate slowed slightly. Conspicuously, the NO3–– N concentration decreased from 98.6 mg/L to 89.731 mg/L at 18 h. However, after 50 h, the nitrate removal rate increased to 97%. In summary, the pH range of 6–10 was advantageous to the efficient removal of NO3−-N by strain KA-1. Considering the strain’s growth patterns, the optimal pH range for nitrogen removal was determined to be 6–10 for strain KA1.
(a) Reduction of nitrate (b)Nitrate Removal Efficiency (c) Reduction of nitrite by Stutzerimonas Stutzeri with different pH levels.
Bio augmentation
Bioaugmentation technology is increasingly widely used for the treatment of nitrogen-containing wastewater. In the current experiments, pre-screened aerobic denitrifiers at different concentrations and initial active sludge were inoculated in an SBR for the treatment of Coking wastewater. The nitrogen-degradation performance of a bacterial bioaugmentation system with different concentrations and a common microbial system in treating high-salt wastewater was compared.
Figure 7 shows the variation trends of nitrate and nitrite in SBR1, SBR2, and SBR3 over 10 operating cycles. At the initial stage of the reaction, NO3–N decreased in both the original system and the two enhanced systems. After the first cycle, the NO3−-N removal rates of the three reaction systems were 78%, 75% and 72%, respectively. At this time, the NO3−-N concentration of SBR1 was higher than that of SBR2 and SBR3. Similarly, after the second cycle, the NO3−-N removal rate in SBR1 reached 80%, which was higher than those in the other two systems (77% and 79%). This indicated that significant denitrification occurred in all three systems. At the initial stage of the reaction, nitrate reduction in the original system was as evident as in the two enhanced systems, indicating that the original system contained a certain amount of denitrifying bacteria. When there was a lot of readily degradable organic matter in the three systems, the existing denitrifying bacteria could contribute to NO3−-N removal.
During continuous reactor operation, the NO3–N removal rate in SBR1 continues to decline. The nitrate removal rate decreased from 86% in the fourth cycle to 72% in the tenth cycle. This may be due to effects on denitrifying bacteria in the original system. However, due to the addition of aerobic denitrification bacteria in the two enhanced systems, the NO3−-N removal rates in the SBR2 and SBR3 systems were gradually higher than those in the SBR1 system from the third cycle. Supplementary aerobic denitrifying bacteria could survive and reproduce more quickly under harsh conditions in response to environmental changes. Moreover, the added bacteria were halophilic, which have strong tolerance to salinity. Therefore, the NO3−-N removal rates of SBR2 and SBR3were always higher than those of SBR1 under seawater salinity impact.
In the tenth cycle, the nitrate removal rates for SBR1, SBR2, and SBR3 were 72%, 87%, and 92%, respectively. Besides, because the supplemental concentration of aerobic denitrification strain was slightly higher in SBR3 system than that of SBR2 at the beginning, theNO3−N removal rate of SBR3 was faster thanSBR2. At the same time, the nitrite accumulation in the enhanced system was lower than that in the original system, suggesting a new application for the strain in nitrogen removal wastewater.
(a) Reduction of nitrate (b)Nitrate Removal Efficiency (c) Reduction of nitrite by Stutzerimonas Stutzeri with different pH levels.
COD experiment by Stutzerimonas stutzeri
High COD levels indicate a high concentration of oxidizable organic matter, which can deplete oxygen in the water and harm aquatic life. Figure 8 shows the COD concentration in coking wastewater over time, including controls (C1 and C2) and treatment samples (S1 and S2). COD measures the amount of organic pollutants in water, and a decrease in COD indicates their removal71. In the initial phase, from 0 to 20 h, C1 exhibits a COD concentration that starts around 400 mg/L and decreases to about 250 mg/L over the first 20 h. There is a moderate decrease in this control, likely due to natural degradation. Just as C1, C2 begins at approximately 400 mg/L but fakes a greater reduction to approximately 200 mg/L in 20 h. This implies that C2 can be a conditionally slightly different control or an intrinsically varied control, raising the question of its potential. S1 and S2 samples begin at approximately 400 mg/L and decline more slowly than the controls, reaching approximately 300 mg/L. This slower reduction in COD suggests that the treatment process is just beginning to take effect. In the Mediate Phase, from 20 to 60 h, C1 and C2, both controls, show a steady decline in COD concentration, reaching around 100 mg/L and 80 mg/L, respectively, by 60 h. The consistent decrease in COD in the controls indicates that natural or baseline processes reduce the organic load72.
The samples S1 and S2 show a more pronounced reduction in COD during this phase, reaching around 150 mg/L and 140 mg/L, respectively. Compared to the controls, the sharp decline in COD suggests that the treatments applied to S1 and S2 effectively degrade the organic pollutants, leaving the audience impressed by the results. Subsequently, in the last Phase, from 60 to 110 h. The controls C1 and C2 continue to reduce COD but at a dilatory rate, with concentrations nearing zero by 110 h. This shows that most of the organic pollutants in the controls have been degraded over time. The treatment samples S1 and S2 show a faster decrease in COD, with both approaching zero COD by 100 h. The more rapid reductions in S1 and S2 compared to the controls highlight the treatments’ effectiveness in accelerating the breakdown of organic matter, convincing the audience of their efficacy. Compared with the controls, C2 shows slightly better performance in reducing COD, possibly due to different conditions or natural variations. Overall, the Figure shows that the treatments applied to samples S1 and S2 significantly enhance COD reduction in coking wastewater compared to natural degradation in the controls C1 and C2.
Reduction of COD in coking wastewater by Stutzerimonas stutzeri.
Conclusion
This paper has isolated and identified the Stutzerimonas stutzeri strain KA1 from coking wastewater and assessed its outstanding aerobic denitrification performance under different environmental conditions. The strain achieved almost 100% nitrate removal efficiency within 40 h when sodium acetate was used as the carbon source, especially at a C/N ratio of 8 and a pH of 6–10. The strain exhibited a high nitrate removal rate across a broad range of dissolved oxygen conditions, suggesting that its denitrification activity remains stable under aerobic bulk conditions. The study also examined the potential of bioaugmentation with strain KA1 for treating coking wastewater in Sequential Batch Reactors (SBRs), in addition to denitrification. It was found that the bioaugmentation process significantly enhanced nitrate removal efficiency, and that bioaugmented systems performed better than control systems. The improved bacteria proved resistant to environmental and salinity conditions, confirming bioaugmentation as a viable method for enhancing wastewater treatment. The decrease in COD was also a considerable result. Strain KA1 effectively reduced COD levels in coking wastewater and accelerated the decomposition of organic pollutants. The bioaugmented systems demonstrated higher COD reduction rates than natural degradation processes, indicating that the strain Stutzerimonas stutzeri KA1 can treat industrial wastewater through a cost-effective, efficient, and scalable approach to nitrogen removal and COD reduction. The research provides useful insights into maximizing key parameters, such as carbon sources, pH, dissolved oxygen, and bioaugmentation applications, to advance the field of microbial biotechnology and environmental sustainability.
Data availability
All data generated or produced have been given in the manuscript.
References
Mackey, J. & Sisodia, R. Conscious capitalism, with a new preface by the authors: Liberating the heroic spirit of business (Harvard Business Review, 2014).
Moore, J. W. Sugar and the expansion of the early modern world-economy: Commodity frontiers, ecological transformation, and industrialization. Review (Fernand Braudel Center), 409–433 (2000).
Romm, J. J. Climate change: What everyone needs to know (Oxford University Press, 2022).
Seddon, N. et al. Understanding the value and limits of nature-based solutions to climate change and other global challenges. Philos. Trans. R. Soc. B Biol. Sci. 375, 20190120 (2020).
Mushtaq, N., Singh, D. V., Bhat, R. A., Dervash, M. A. & Hameed, O. Freshwater contamination: sources and hazards to aquatic biota. In Fresh water pollution dynamics and remediation 27–50 (2020).
Saleem, F., Saleem, M. A., Khalil, U. & Maqsood, M. S. Plastic waste: Environmental impact, innovative solutions, and pathways to sustainable management. J. Life Soc. Sci. 2025, 36. https://doi.org/10.64013/bbasrjlifess.v2025i1.36 (2025).
Zahid, A. et al. Chlorophyll degradation under smog exposure: unveiling the molecular and ecological consequences. Bulletin of Biological and Allied Sciences Research 96–96 (2025). (2025).
Qadri, R. & Faiq, M. A. Freshwater pollution: effects on aquatic life and human health. In Fresh water pollution dynamics and remediation 15–26 (2020).
Sonone, S. S., Jadhav, S., Sankhla, M. S. & Kumar, R. Water contamination by heavy metals and their toxic effect on aquaculture and human health through food chain. Lett. Appl. NanoBioSci. 10, 2148–2166 (2020).
Saravanan, A. et al. Effective water/wastewater treatment methodologies for toxic pollutants removal: Processes and applications towards sustainable development. Chemosphere 280, 130595 (2021).
Ashar, A., Afshan, N., Aslam, T., Noor, S. & Bhutta, Z. A. Treatment of Wastewater and Its Reuse. In Advanced Oxidation Processes for Wastewater Treatment 213–222 (CRC Press, 2022).
Ali, J. Water scarcity in Pakistan: Reasons, impact and curative measures. Journal of Physical, Biomedical and Biological Sciences 2025, 42. https://doi.org/10.64013/jpbab.v2025i1.42 (2025).
Hammad, M., Shafiq, M., Batool, A. & Sherazi, S. Mechanisms of action and signaling pathways involved in abiotic stress elicitation. Bull. Biol. Allied Sci. Res. 116. https://doi.org/10.64013/bbasr.v2026i1.116 (2026).
Riffat, R. & Husnain, T. Fundamentals of wastewater treatment and engineering (Crc, 2022).
Singh, R. L., Singh, R. P., Gupta, R. & Singh, R. Advances in biological treatment of industrial waste water and their recycling for a sustainable future (Springer, 2019).
Aljerf, L. Data of thematic analysis of farmer׳ s use behavior of recycled industrial wastewater. Data brief. 21, 240–250 (2018).
Shashi, Centobelli, P., Cerchione, R. & Mittal, A. Managing sustainability in luxury industry to pursue circular economy strategies. Bus. Strategy Environ. 30, 432–462 (2021).
Song, L. China: Steel Industry. In Encyclopedia of Mineral and Energy Policy 142–152 (Springer, 2023).
Ji, J., Zou, Z. & Tian, Y. Energy and economic impacts of China’s 2016 economic investment plan for transport infrastructure construction: An input-output path analysis. J. Clean. Prod. 238, 117761 (2019).
Zhu, X., Li, H., Chen, J. & Jiang, F. Pollution control efficiency of China’s iron and steel industry: Evidence from different manufacturing processes. J. Clean. Prod. 240, 118184 (2019).
Gielen, D., Saygin, D., Taibi, E. & Birat, J. P. Renewables-based decarbonization and relocation of iron and steel making: A case study. J. Ind. Ecol. 24, 1113–1125 (2020).
Safarian, S. To what extent could biochar replace coal and coke in steel industries?. Fuel https://doi.org/10.1016/j.fuel.2023.127401 (2023).
Rosado, D. J. M. et al. Energetic analysis of reheating furnaces in the combustion of coke oven gas, Linz-Donawitz gas and blast furnace gas in the steel industry.. Appl. Therm. Eng. 169, 114905 (2020).
Mosca, S., Guerriero, E., Torelli, G. N. & Tramontana, G. & Rotatori, M.
Biswas, J.
Chen, L., Xu, Y. & Sun, Y. Combination of coagulation and ozone catalytic oxidation for pretreating coking wastewater. Int. J. Environ. Res. Public Health https://doi.org/10.3390/ijerph16101705 (2019).
Wen, G. et al. Aerobic denitrification performance of strain Acinetobacter johnsonii WGX-9 using different natural organic matter as carbon source: Effect of molecular weight.. Water Res. 164, 114956 (2019).
Zheng, M. et al. N2O and NO emission from a biological aerated filter treating coking wastewater: Main source and microbial community. J. Clean. Prod. https://doi.org/10.1016/j.jclepro.2018.12.182 (2019).
Ditto, J. C., Machesky, J. & Gentner, D. R. Analysis of reduced and oxidized nitrogen-containing organic compounds at a coastal site in summer and winter. Atmospheric Chem. Physics (2022).
Fouad, F. A., Youssef, D. G., Shahat, F. M. & Abd El-Ghany, M. N. Role of Microorganisms in Biodegradation of Pollutants. In Handbook of Biodegradable Materials 221–260 (Springer, 2023).
Abada, A. et al. Aerobic bacteria produce nitric oxide via denitrification and promote algal population collapse.. ISME J. 17, 1167–1183 (2023).
Patel, N. et al. Nitrate contamination in water resources, human health risks and its remediation through adsorption: A focused review.. Environ. Sci. Pollut. Res. Int. 29, 69137–69152 (2022).
Zahra, N., Saeed, M. K. & Raza, M. H. Nitrosamines: Incredibly unsafe contaminants in different food commodities.. Chem. Int. 9, 27–36 (2023).
Kiani, A. et al. Accumulation and human health risk assessment of nitrate in vegetables irrigated with different irrigation water sources- transfer evaluation of nitrate from soil to vegetables. Environmental research, 112527 (2021).
McKindles, K. M., Frenken, T., McKay, R. M. L. & Bullerjahn, G. S. Binational Efforts Addressing Cyanobacterial Harmful Algal Blooms in the Great Lakes. In The Handbook of Environmental Chemistry (2020).
Liao, W. et al. Toxicity mechanisms and bioavailability of copper to fish based on an adverse outcome pathway analysis. J. Environ. Sci. 127, 495–507 (2022).
Kelly, N. E., Guijarro-Sabaniel, J. & Zimmerman, R. Anthropogenic nitrogen loading and risk of eutrophication in the coastal zone of Atlantic Canada. Estuar. Coast. Shelf Sci. https://doi.org/10.1016/j.ecss.2021.107630 (2021).
Paerl, H. W., Otten, T. G. & Kudela, R. M. Mitigating the expansion of harmful algal blooms across the freshwater-to-marine continuum.. Environ. Sci. Technol. 52(10), 5519–5529 (2018).
Aa, I., Op, A., Ujj, I. & Mt, B. A critical review of oil spills in the Niger Delta aquatic environment: Causes, impacts, and bioremediation assessment. Environ. Monit. Assess. https://doi.org/10.1007/s10661-022-10424-x (2022).
Heil, C. & Muni-Morgan, A. in Frontiers in Ecology and Evolution.
Herath, S. S. & Satoh, S.
Liu, Y., Ma, S., Yang, Y. & Lv, Y. Characteristics of a Heterotrophic Nitrifier Consortium and Its Application in Coking Wastewater. Water Air & Soil. Pollution 234 (2023).
Lloyd, D. Aerobic denitrification in soils and sediments: From fallacies to factx.. Trends Ecol. Evol. 8(10), 352–356 (1993).
Robertson, L. A. & Kuenen, J. G. Aerobic denitrification: A controversy revived. Arch. Microbiol. 139, 351–354 (2004).
Verstraete, W. & Alexander, M. Heterotrophic nitrification by Arthrobacter sp. J. Bacteriol. 110, 955–961 (1972).
Kuenen, J. G., Robertson, L. A. & v. Gemerden, H. Microbial interactions among aerobic and anaerobic sulfur-oxidizing bacteria. Adv. Microb. Ecol. 8, 1–59 (1985).
Marazioti, C., Kornaros, M. E. & Lyberatos, G. Kinetic modeling of a mixed culture of Pseudomonas denitrificans and Bacillus subtilis under aerobic and anoxic operating conditions. Water Res. 37(6), 1239–1251 (2003).
Samuelsson, P.
Li, E. & Lu, S. Denitrification processes and microbial communities in a sequencing batch reactor treating nanofiltration (NF) concentrate from coking wastewater. Water Sci. Technol. 76, 3289–3298 (2017).
Li, E. et al. Investigation into the nitrate removal efficiency and microbial communities in a sequencing batch reactor treating reverse osmosis concentrate produced by a coking wastewater treatment plant. Environ. Technol. 39, 2203–2214 (2018).
Zheng, M. et al. N2O and NO emission from a biological aerated filter treating coking wastewater: Main source and microbial community. J. Clean. Prod. 213, 365–374 (2019).
Yuan, Q. et al. Comparison of the MBBR denitrification carriers for advanced nitrogen removal of wastewater treatment plant effluent. Environ. Sci. Pollut. Res. 22, 13970–13979 (2015).
Duan, J., Fang, H., Su, B., Chen, J. & Lin, J. Characterization of a halophilic heterotrophic nitrification–aerobic denitrification bacterium and its application on treatment of saline wastewater. Bioresour. Technol. 179, 421–428. https://doi.org/10.1016/j.biortech.2014.12.057 (2015).
Wang, H., Chen, N., Feng, C. & Deng, Y. Insights into heterotrophic denitrification diversity in wastewater treatment systems: Progress and future prospects based on different carbon sources. Sci. Total Environ. 780, 146521 (2021).
Her, J.-J. & Huang, J.-S. Influences of carbon source and C/N ratio on nitrate/nitrite denitrification and carbon breakthrough. Bioresour. Technol. 54, 45–51 (1995).
Wu, X., Yu, Z., Yuan, S., Tawfik, A. & Meng, F. An ecological explanation for carbon source-associated denitrification performance in wastewater treatment plants. Water Res. 247, 120762 (2023).
Elefsiniotis, P. & Li, D. The effect of temperature and carbon source on denitrification using volatile fatty acids. Biochem. Eng. J. 28, 148–155 (2006).
Louzeiro, N. R., Mavinic, D. S., Oldham, W. K., Meisen, A. & Gardner, I. S. Methanol-induced biological nutrient removal kinetics in a full-scale sequencing batch reactor. Water Res. 36, 2721–2732 (2002).
Zheng, H.-Y. et al. Characterization of a marine origin aerobic nitrifying–denitrifying bacterium. J. Biosci. Bioeng. 114, 33–37 (2012).
Lloyd, D. Aerobic denitrification in soils and sediments: from fallacies to factx. Trends Ecol. Evol. 8, 352–356 (1993).
Robertson, L. & Kuenen, J. Aerobic denitrification—Old wine in new bottles?. Antonie Van Leeuwenhoek 50, 525–544 (1984).
Ali, J. et al. Comparative nutritional and minerals composition of azadirachta indica, eucalyptus globules and melia azedarach. Journal of Life and Social Sciences 39 (2024). (2025).
Afridi, R. et al. Gray water to green gold: Characterization and potential of polyhydroxyalkanoate-producing microbes from industrial effluents. Bulletin of Biological and Allied Sciences Research 105. https://doi.org/10.64013/bbasr.v2025i1.105 (2025).
Patureau, D., Bernet, N., Delgenès, J. P. & Moletta, R. Effect of dissolved oxygen and carbon–nitrogen loads on denitrification by an aerobic consortium. Appl. Microbiol. Biotechnol. 54, 535–542. https://doi.org/10.1007/s002530000386 (2000).
Hammad, M., Talib, U., Sher, A., Shafiq, M. & Sherazi, S. Climate-resilient horticulture through genomic tools: A decade of genome-wide association studies (GWAS) applications amidst a changing climate. Bulletin of Biological and Allied Sciences Research 115. https://doi.org/10.64013/bbasr.v2026i1.115 (2026).
Kayani, R. et al. Chlorophytum comosum-mediated iron nanoparticles: an eco-friendly approach for antimicrobial and dye degradation applications. Bulletin of Biological and Allied Sciences Research 94–94 (2025). (2025).
Safdar, R., Imran, M., Mushtaq, E. & Nasir, A. From toxicology to technology: Human health risks of mycotoxins in the food chain and current approaches to their detection and control. Bulletin of Biological and Allied Sciences Research 114. https://doi.org/10.64013/bbasr.v2026i1.114 (2026).
Zumft, W. G. Cell biology and molecular basis of denitrification. Microbiol. Mol. Biol. Rev. 61, 533–616 (1997).
Hao, Z.-L., Ali, A., Ren, Y., Su, J.-F. & Wang, Z. A mechanistic review on aerobic denitrification for nitrogen removal in water treatment. Sci. Total Environ. 847, 157452 (2022).
Chen, M. et al. Impact resistance of different factors on ammonia removal by heterotrophic nitrification–aerobic denitrification bacterium Aeromonas sp. HN-02. Bioresour. Technol. 167, 456–461 (2014).
Modi, S. et al. Recent and emerging trends in remediation of Methylene blue dye from wastewater by using zinc oxide nanoparticles. Water 14, 1749 (2022).
Musa, M. A., Idrus, S., Hasfalina, C. M. & Daud, N. N. N. Effect of organic loading rate on anaerobic digestion performance of mesophilic (UASB) reactor using cattle slaughterhouse wastewater as substrate. Int. J. Environ. Res. Public Health 15, 2220 (2018).
Author information
Authors and Affiliations
Contributions
The authors would like to extend their sincere appreciation to Ongoing Research Funding Program (ORF-2026–165), King Saud University, Riyadh, Saudi Arabia. KN and KA carried out the study, and AS and QA helped collect the data. AS and QA helped set the article sequence. KA, DA, MS, UT, AB, and QA carried out the final revisions to the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Naseer, K., Ashfaq, K., Shamim, A. et al. Isolation of aerobic denitrifying bacteria Stutzerimonas stutzeri and its application in coking wastewater treatment. Sci Rep 16, 8717 (2026). https://doi.org/10.1038/s41598-026-43338-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-43338-6