Abstract
Understanding how large carnivores partition dietary resources is essential for assessing intra-guild competition and informing conservation strategies. In this study, we used DNA metabarcoding of scats to quantify and compare the diets of sympatric African lions (Panthera leo) and spotted hyenas (Crocuta crocuta) across wet and dry seasons in the Greater Etosha Landscape of Namibia. Across 98 scat samples (lion = 69; spotted hyena = 29), we identified 19 vertebrate prey species. Overall, large ungulates dominated both carnivores’ diets. For lions, the most frequent prey items included gemsbok (Oryx gazella), common eland (Taurotragus oryx), plains zebra (Equus quagga burchelli), and blue wildebeest (Connochaetus taurinus). For spotted hyenas, the most frequent prey items were plains zebra, gemsbok, springbok (Antidorcas marsupialis), and black rhinoceros (Diceros bicornis bicornis). Dietary niche breadth was not significantly different between species, though lions exhibited the broadest across both seasons, while diet composition was similar between species and seasons. However, the smaller sample size for spotted hyenas may limit full characterization of their diet and influence measures of overlap and niche breadth. These results suggest a moderate diet overlap and limited resource partitioning both within and among these large carnivore species across seasons, likely facilitated by opportunistic scavenging and kleptoparasitism. Both species exhibited broader dietary niche breadths during the wet season, likely reflecting increased prey availability and dispersion. Ongoing monitoring of carnivore diets using molecular tools, which provides a more accurate and comprehensive identification of diet items than manual sorting, will be essential for detecting changes in resource use and interspecific interactions in response to shifting environmental conditions and anthropogenic pressures.
Similar content being viewed by others
Introduction
Large carnivores play critical roles in maintaining ecosystem health and diversity1. Yet, despite multifaceted conservation initiatives, large carnivore populations are declining globally due to anthropogenic pressures2,3,4 including human-mediated conflicts due to their large home range requirements and the expanding human footprint5. As population footprint expands, large carnivores are becoming increasingly restricted to protected areas6, which can lead to higher levels of interspecific interactions and intra-guild competition. Thus, monitoring large carnivore populations, especially where intra-guild competition is likely to be high, is imperative for conservation efforts7,8.
In regions with well-preserved predator guilds, large carnivore species often occupy overlapping geographic ranges and compete for prey resources. Interspecific competition can drive community structure and function, and affect the distribution, demographic patterns, and behavioral strategies of co-occurring species9,10,11. Because large carnivore populations are primarily shaped by the abundance and availability of their main food items12,13, prey represents a critical resource that coexisting carnivores can partition to reduce competition12,14. Intra-guild competition intensity can vary spatio-temporally due to changes in prey availability, which can fluctuate due to a multitude of biotic and abiotic factors such as seasonality, droughts, climate change, or disease outbreaks15,16,17. While numerous studies have investigated carnivore competition and resource partitioning, data is lacking in certain areas of sub-Saharan Africa, where largely intact carnivore guilds exist. Understanding how sympatric carnivores partition dietary resources in these systems is thus essential to understand carnivore ecology18 and inform conservation practices through vegetation and prey species management.
Some of the most diverse and multifaceted terrestrial assemblages of large carnivores occur in sub-Saharan Africa, with several species of large carnivores occurring sympatrically including African lions (Panthera leo) (hereafter lions), spotted hyenas (Crocuta Crocuta), cheetahs (Acinonyx jubatus), leopards (Panthera pardus), and brown hyenas (Parahyaena brunnea). Lions and spotted hyenas are two of the most numerous large carnivores in this region where they overlap extensively in distribution19 and diel activity20,21,22. To facilitate coexistence, lions and spotted hyenas have evolved disparate behavioral, social, and foraging strategies23,24. Lions are ambush predators, who hide under vegetation and pursue medium-large ungulate prey23,25, yet al.so hunt in open areas26 and scavenge opportunistically27,28. Conversely, spotted hyenas are cursorial predators, typically chasing prey and using endurance for hunting success29,30 but also readily scavenge carcasses when available28,31. Additionally, spotted hyenas do successfully hunt calves or juvenile large ungulate species29,32,33.
Lions and spotted hyenas engage in kleptoparasitism; spotted hyenas lose more of their kills to lions than lions lose to spotted hyenas23, though this kind of intra-guild competition is highly variable based on pride and clan size and structure34. With overlapping diel activity patterns, kleptoparasitism, and opportunistic scavenging, dietary overlap is expected to be high between lions and spotted hyenas with substantial variation across their shared geographic range. Previous studies report broad dietary niches for spotted hyenas35, with similar36 or narrower12 niche breadths compared to lions. However, spotted hyenas may reduce their niche overlap by preying on different prey size via different sex or age classes23. Additionally, top prey items vary across studies for both species (e.g., Hayward36,, and the role of seasonality in shaping dietary niche breadth and overlap remains poorly understood35.
Few studies have employed high-throughput sequencing methods like DNA metabarcoding to address these questions. DNA metabarcoding has been shown to have higher sensitivity and greater taxonomic resolution relative to conventional dietary studies that rely on morphological identification of undigested remains in scats37,38). Morphological identification can introduce additional biases into dietary results through misidentification of closely related prey items, lack of detection for prey items with highly digestible tissues, and underestimating prey items (Monterosso et al., 2019). The conflicting findings in previous dietary studies, along with the limited application of molecular techniques, underscore the importance of assessing carnivore diet partitioning, especially between lions and spotted hyenas, to quantify dietary overlap in a rich carnivore community. Moreover, while little is known about the effects of seasonality on dietary niche overlap among large carnivores, understanding these dynamics is critical for advancing our knowledge of intra-guild resource partitioning and coexistence in shared landscapes.
Our goal in this study was to use DNA metabarcoding to quantify dietary patterns of sympatric lions and spotted hyenas in a dryland ecosystem in southern Africa. Specifically, our objectives were to (1) quantify the vertebrate diet composition of sympatric lions and spotted hyenas; (2) quantify seasonal variation in diet composition within and between each species; (3) assess dietary niche breadth for each species across seasons; and (4) evaluate interspecific dietary overlap to determine if there is evidence of niche partitioning and competition. We also formulated a series of predictions. First, we predicted that both species would have higher diversity in their diet during the wet season due to increased availability of young herbivores during the birthing season for many ungulate species, and animals dispersing across the landscape and away from predictable waterhole locations, forcing carnivores to be less selective for prey species. Second, based on scavenging activity and previous studies3635;, we predicted that spotted hyenas would have higher dietary niche breadths than lions during both seasons. Lastly, we predicted that both species would have high dietary overlap during both seasons because of kleptoparasitism and opportunistic scavenging.
Methods
Study area
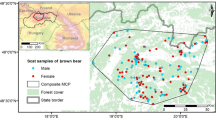
This study was conducted in the Greater Etosha Landscape (GEL) encompassing Etosha National Park (“Etosha”), the Ongava Game Reserve (“Ongava”), and King Nehale communal conservancy (“King Nehale”) in north-central Namibia (Fig. 1)39. Etosha is a 22,900 km2 fenced protected area, although the fence is permeable to wildlife40,41. Ongava is a fenced 300 km2 private game reserve on the southern border of Etosha. King Nehale is a 508 km2 communal conservancy located in the Oshikoto region, bordering the northeast corner of Etosha. One main source of income for people living in the conservancy is keeping livestock, including cattle, goats, and sheep42. The GEL is located within the semi-arid weather zone with an average annual rainfall of 380 mm43,44). Despite fencing around Etosha, the prey and predator guild composition in all these areas is very similar and together they form part of the same larger scale ecosystem, including movement of carnivores between Ongava and Etosha, and more so between King Nehale and Etosha40. The main vegetation types in Etosha are grasslands, steppe, shrubland, Mopane (Colophospermum mopane) tree savanna, and mixed tree savanna45. The landcover on Ongava is classified as Karstveld, with Mopane shrubland and woodland being the dominant vegetation type (~ 70%) and other areas with savannah-like vegetation (~ 30%46,. During the wet season (November-April), rainfall provides additional temporal water sources, aside from the permanent artificial water sources across the landscape47,48.
Location of collected fecal samples from African lions (Panthera leo, red dots) and spotted hyenas (Crocuta Crocuta, black dots) in the Greater Etosha Landscape, Namibia. Collection occurred between October 2021 and March 2024.
Both Etosha and Ongava are tourist destinations for wildlife-based viewing and support a broad diversity of carnivore and ungulate species [e.g., black-faced impala (Aepyceros melampus petersi), springbok (Antidorcas marsupialis), blue wildebeest (Connochaetus taurinus), plains zebra (Equus quagga burchelli), gemsbok (Oryx gazella)], including megaherbivores [e.g, African elephant (Loxodonta africana), southern Angolan giraffe (Giraffa giraffa angolensis), southern black rhinoceros (Diceros bicornis bicornis)]. However, prey species’ composition and abundance differ between Ongava and Etosha, and across the different regions of Etosha (west, central, and east)49,50. Across both sites, potential prey species richness for mammals >5 kg is approximately 21 wild species and three domestic livestock species. In 2008, the estimated spotted hyena population in Etosha was 339 \(\pm\) 17651 and in 2022, the estimated lion population in Etosha was 350 (MEFT, 2022). Data are unavailable for population sizes in Ongava.
Sample collection
We collected 164 dried or fresh fecal samples [spotted hyena (n = 33) and lion (n = 132)] from October 2021 through March 2024 around Etosha, Ongava, and King Nehale. When divided by location, we collected 74 samples in Etosha [spotted hyena (n = 28; dry season = 20, wet season = 8) and lion (n = 46; dry season = 40, wet season = 6)], three samples in King Nehale [spotted hyena (n = 2 in the wet season) and lion (n = 1 in the dry season)], and 88 samples in Ongava [spotted hyena (n = 3 in the dry season) and lion (n = 85; dry season = 77, wet season = 8)]. Most samples (n = 140) were collected opportunistically from the ground while driving along roads, around waterholes, and near carnivore kill sites within our study area (approved by Namibian Ministry of Environment, Forestry, and Tourism under permit #AN202101004). For these samples, whole fecal samples were collected into a paper bag and dried naturally before being subsampled for DNA extraction, which included both the external surface and internal matrix to capture both consumer and prey DNA. Additionally, 24 samples [lions (n = 19), spotted hyenas (n = 5)] were collected during carnivore immobilizations as part of ongoing research (Appendix S1; Text S1). These samples were manually retrieved from the immobilized animal, placed into an open plastic vial and stored in a paper bag for natural drying. Due to the small amount collected directly from immobilized animals, these samples were not subsampled. All immobilizations were performed by veterinarians registered with the Namibian Veterinary Council and the Ministry of Environment, Forestry, and Tourism and were approved by the Namibian Ministry of Environment, Forestry, and Tourism (permit #AN202101004) as well as the University of Georgia Institutional Animal Care And Use Committee under protocols A2024 05–009-06 and A2021 04–013-Y3-A11.
Laboratory analyses and data curation
All samples were shipped from Namibia to the United States in July 2024 and DNA was extracted from feces using DNeasy PowerSoil Pro DNA Kits (QIAGEN) in August 2025. Jonah Ventures, LLC conducted DNA amplification, sequencing, and bioinformatics for DNA metabarcoding (Appendix 1; Text S2). This technique analyzes DNA within scats using high-throughput sequencing using small, highly variable universal primers to identify prey species and confirm predator species52,53). To characterize prey DNA from carnivore fecal samples, we amplified a portion of the 12S rRNA gene using the primer pair 12SVertF (forward 5′- ACTGGGATTAGATACCCYACTATG − 3′) and 12SVertR (reverse 5′- GAGRRTGACGGGCGGTDT − 3′)54. Both forward and reverse primers contained a 5’ adaptor sequence to allow for subsequent indexing and Illumina sequencing. Exact sequence variants (ESVs) were identified using the UNOISE3 algorithm55 implemented in VSEARCH (Rogens et al., 2016), retaining only those with ≥ 4 read counts and ≤ 1 expected error. We assigned taxa of ESVs using a custom reference database consisting of publicly available sequences on NCBI GenBank56 and Jonah Ventures voucher sequences records, accepting the reference taxonomy for any taxonomic level with > 90% agreement within 1% of the top hits. We discarded any reads of taxa not occurring in the study region, and to limit the effects of contamination, only prey species that consisted of more than 1% of total prey sequences were retained57. Because host DNA typically dominates the sequencing dataset58, we were able to confirm predator species for each sample.
Data analyses
We conducted all analyses and produced figures using R version 4.1.1 (R Core Team59,. We summarized the data with both frequency of occurrence (FOO) and relative read abundance (RRA) at the species level, and calculated prey species richness for each carnivore species by season (wet and dry). To calculate FOO, we divided the number of occurrences of a prey species by the total number of carnivore scat samples and expressed FOO as a percentage. We calculated RRA for each prey species as the read count (the number of DNA sequence reads) for that the given prey species divided by the total number of reads in the carnivore scat sample, also expressed as a percentage.
To evaluate differences in vertebrate prey composition (beta diversity) between carnivore species and seasons, we performed non-metric multidimensional scaling (NMDS). Analyses were conducted separately for wet and dry seasons using Bray–Curtis dissimilarities derived from binary presence–absence matrices, where each sample with identified prey was represented by a vector of prey species detections. NMDS ordinations were computed in three dimensions (k = 3) with 1000 permutations (trymax = 1000) to improve solution stability, and we used stress values assess fit. We generated plots where arrows represent prey species most associated with carnivore-specific patterns in prey use and samples cluster apart in ordination space if carnivore diets are compositionally distinct.
We used two complementary approaches to evaluate dietary similarity between and within carnivore species. First, we applied a non-parametric Analysis of Similarities (ANOSIM) with binary Jaccard dissimilarities to test whether prey composition within carnivore scats differed between lions and hyenas within each season. Because ANOSIM evaluates overall compositional differences but does not quantify the degree of overlap, we also calculated Pianka’s index of niche overlap60 using FOO, with values ranging from 0 (no overlap) to 1 (complete overlap) based on the proportional use of each prey species. To account for unequal sample sizes, we generated 95% bootstrap confidence intervals by stratified resampling of carnivore scats for both species (2,000 iterations). Together, these analyses allowed us to assess both whether diets differed significantly between carnivore species and the extent of dietary niche overlap, providing a more complete picture of dietary similarity across seasons. We used the vegan package in R61 to conduct the NMDS and ANOSIM and spaa package in R62 to determine Pianka’s index.
We calculated dietary niche breadth (Ba) for each carnivore species in both seasons by using Hurlbert’s standardized version of Levin’s measure63,64 for FOO proportions of prey. To account for differences in sample size across carnivore species and seasons, we applied a bootstrap approach with 2,000 iterations. For each iteration, all samples were resampled with replacement within each carnivore species–season group, and standardized niche breadth was recalculated. Broader dietary niche breadth is indicated by a higher Ba while a narrow dietary niche breadth is indicated by a low Ba (ranging from 0 to 1). We did this overall with all data pooled, and separately for each study site [Etosha (including King Nehale) versus Ongava].
To evaluate completeness of our sampling efforts and prey richness, we generated sample-based rarefaction and extrapolation curves for each carnivore species and season using the specaccum function in the vegan package (Oksanen et al., 2022). Scat samples were converted to presence–absence matrices of prey taxa, and 1,000 random permutations were used to estimate mean prey richness and associated 95% confidence intervals. We further estimated asymptotic prey species richness with the Chao1 estimator65; specpool function), a non-parametric approach that accounts for incomplete sampling by inferring the total number of species based on the frequency of rare species observed only once or twice. We then plotted these estimates (± standard error) alongside rarefaction curves to assess whether observed prey detections approached estimated richness. For all analyses we pooled data from all study sites to generate broad inferences about carnivore diet, and we also visually examined diet composition between Etosha and King Nehale versus Ongava.
Results
Diet overview
From the 164 samples that were sequenced, 98 had detectable vertebrate prey items and were included in analysis [spotted hyena (n = 29; dry season = 17, wet season = 12) and lion (n = 69; dry season = 54, wet season = 15)]. Overall, we detected 9 families and 17 genera encompassing 19 vertebrate prey species (Table 1). For spotted hyenas, the total read count was 127,640 with a mean read count of 3868 per sample (± 4461 SD). For lions, the total read count was 107,965 with a mean read count of 1367 per sample (± 2733 SD).
Spotted hyena diet
From 29 sequenced spotted hyena samples, plains zebra was the most frequently consumed prey species in the dry season (FOO = 29.4%) followed by gemsbok (Oryx gazella) and springbok (FOO = 17.7% each). In the wet season, spotted hyenas equally consumed plains zebras and gemsbok (FOO = 18.8% each, Fig. 2). Overall, prey species richness was 10 and included two domestic livestock species, cow (Bos taurus; n = 2) and goat (Capra hircus; n = 2). Both domestic prey species were found in samples near or outside of northeast Etosha (Fig. 1). ANOSIM revealed no seasonal differences in diet composition for spotted hyenas (Table 2).
Seasonal composition of vertebrate prey species detected in the diets of African lions (Panthera leo) and spotted hyenas (Crocuta crocuta) in the Greater Etosha Landscape based on DNA metabarcoding. Bars represent the proportion of fecal samples in which each prey species was detected, calculated as the number of unique samples per species and season divided by the total number of samples for that carnivore and season. Results are shown separately for the dry season (green) and wet season (grey), and prey species are ordered by overall frequency of occurrence from left to right. Only prey species with ≥ 1% relative read abundance were included.
When we separated out by study site [Etosha and King Nehale (n = 30; dry season = 20, wet season = 10) versus Ongava (n = 3 in the dry season)], during the dry season, plains zebra (FOO = 33%) and springbok (FOO = 20%) were the most consumed prey items in Etosha (Fig. S1) while gemsbok and kudu (Tragelaphus strepsiceros) were the only consumed prey items in Ongava (FOO = 50% each) (Fig. S2). During the wet season, spotted hyenas primarily consumed plains zebra and gemsbok in Etosha (FOO = 25% each) (Fig. S1).
Lion diet
From 69 sequenced samples, lions consumed a wide variety of species but primarily large ungulates such as gemsbok (FOO = 25.4%), followed by blue wildebeest and eland (Taurotragus oryx) (FOO = 13.6% each), then plains zebra (FOO = 10.2%) during the dry season. In the wet season lions consumed eland, gemsbok, and plains zebra equally (FOO = 15% each), followed by black-faced impala, blue wildebeest, giraffe, and hare species (FOO = 10% each, Fig. 2). Overall, prey richness was 16 and ANOSIM revealed no significant seasonal differences in diet composition for lions (Table 2).
Separated out by study site [Etosha and King Nehale (n = 47; dry season = 41, wet season = 6) versus Ongava (n = 85; dry season = 77, wet season = 8)], during the dry season, lions consumed springbok (FOO = 21%) most frequently in Etosha, followed by giraffe, plains zebra, eland, red hartebeest, and black rhinoceros equally (FOO = 12%) (Fig. S1). During the dry season on Ongava, lions consumed gemsbok most frequently (FOO = 35%), followed by blue wildebeest (FOO = 20%), eland (FOO = 15%), and plains zebra (FOO = 10%) (Fig. S2). During the wet season in Etosha, lions primarily consumed giraffe and plains zebra equally (FOO = 28%) followed by eland, red hartebeest, blue wildebeest, kudu, and python species equally (FOO = 14%) (Fig. S1). During the wet season on Ongava, lions consumed gemsbok, eland, black-faced impala, and hare spp. equally (FOO = 25%), followed by blue wildebeest and plains zebra (FOO = 12.5%) (Fig. S2).
Dietary niche breadth
Overall, during the wet and dry seasons, lions exhibited broader dietary niche breadth [Ba: wet = 0.45 (0.27–0.54); dry = 0.4 (0.32–0.56)] than spotted hyenas [Ba: wet = 0.38 (0.21–0.48)]; dry = 0.28 (0.15–0.39)] though there were no significant differences between species. Both species had broader dietary niche breadths during the wet season compared to the dry season (Fig. 3), with no significant differences between seasons for either species. In Etosha, lions exhibited broader dietary niche breadth than spotted hyenas in the dry season (Ba: lions = 0.5, hyenas = 0.29), but spotted hyenas had slightly broader dietary niche breadths in the wet season (Ba: lions = 0.44, hyenas = 0.47). Lions had broader dietary niche breadths in the dry season, while spotted hyenas were broader in the wet season (Fig. S3). In Ongava, lions had similar dietary niche breadths across seasons (Ba: dry = 0.417, wet = 0.414), and broader dietary niche breadth compared to spotted hyenas in the dry season (Ba = 0.09) (Fig. S4).
Seasonal variation in standardized niche breadth (Hurlbert’s B′) for African lions (Panthera leo) and spotted hyenas (Crocuta crocuta) in the Greater Etosha Landscape, in both the dry (May-October) and wet (November-April) seasons. Bars represent mean dietary breadth based on frequency of occurrence (FOO) of prey species in fecal samples, with error bars showing 95% confidence intervals from 2,000 bootstrap iterations.
Interspecific dietary variation
We found no differences in dietary composition for either carnivore species overall (all data pooled) or across seasons. The NMDS ordination for dietary overlap (Fig. 4) had a stress value of 0 in the wet season and < 0.001 in the dry season. While a stress value of or near 0 is uncommon, it can occur in small or low-dimensional datasets but is under the threshold value (stress < 0.2) as suggested by Clarke66 for an interpretable ordination. Pairwise ANOSIM comparisons revealed no significant differences (p > 0.05) in diet composition between lions and spotted hyenas within either season or overall. R statistics were close to zero or negative, indicating strong dietary overlap among these carnivores across seasons and overall (Table 2). Pianka’s index revealed moderate dietary overlap between lions and spotted hyenas overall (O = 0.75). Seasonal analyses showed that overlap between species was similar between the dry season (O = 0.68) and the wet season (O = 0.66). Within species, lions exhibited greater dietary similarity across wet and dry seasons (O = 0.84) compared to spotted hyenas (O = 0.77) (Table 3).
Non-metric multidimensional scaling plot showing beta diversity in vertebrate prey species (12 S rRNA) detected in the diet of African lions (Panthera leo), and spotted hyenas (Crocuta crocuta) in the Greater Etosha Landscape, by season (dry = May-October and wet = November-April). Points represent individual predator scat samples. The position of each point in ordination space reflects the multivariate composition of prey in that sample, with closer points indicating more similar prey profiles. Black arrows represent the direction and strength of correlation between specific prey species and the NMDS axes. Species labels at the arrow tips indicate which prey taxa most strongly influence variation in carnivore diet composition along NMDS axes.
Rarefaction curves
Rarefaction curves revealed that prey species richness in lion scats approached asymptotic levels in the dry season, where observed richness (15 taxa) closely matched Chao1 estimates (17.0 ± 2.6). In the wet season, lion richness was lower (10 taxa, 11.0 ± 1.7), suggesting some rare prey were likely not captured. Spotted hyena curves indicated lower observed richness overall (dry = 8, wet = 9 taxa), with estimates suggesting additional undetected prey species, particularly in the dry season (Chao1 = 15.5 ± 11.0), though uncertainty was high due to small sample size (Fig. 5).
Sample-based rarefaction and extrapolation curves of prey species richness detected in lion (Panthera leo) and spotted hyena (Crocuta crocuta) scat samples during dry (green) and wet (black) seasons. Solid lines represent mean richness estimates with shaded areas indicating 95% confidence intervals based on 1,000 random permutations. Triangles show observed richness, and vertical error bars represent Chao1 asymptotic richness estimates (± SE).
Discussion
Determining how large carnivores partition dietary resources is critical for understanding intra-guild interactions and coexistence. Using DNA metabarcoding, we characterized the diets of lions and spotted hyenas in the GEL and found substantial dietary similarity between both species, regardless of the season. Despite differences in prey species richness and niche breadth between carnivore diets, there were no significant seasonal or interspecific differences in prey species composition. These findings suggest that in this landscape, diet composition alone may not strongly differentiate resource use between lions and spotted hyenas, and that any niche partitioning may occur through prey age or sex selection, spatial segregation, or other behavioral mechanisms not captured by prey identity alone. Collectively, these data contribute to our overall understanding of dietary patterns of sympatric lions and spotted hyenas in a semi-arid ecosystem.
Species with broad dietary niche breadths are typically considered dietary generalists while those with narrow dietary niche breadths are considered specialists67. In our study, lions had a wider dietary niche breadth during both seasons compared to spotted hyenas; however, our findings demonstrate that lions and spotted hyenas exhibit broadly similar diets with consistently high overlap, suggesting that these two apex predators rely heavily on a shared prey base. When comparing dietary niche breadth between species, lions and spotted hyenas are reported to have similar dietary niche breadths, which aligns with our results12,36; reviewed in23. The comparison between seasons supported our prediction that both lions and spotted hyenas had broader dietary niche breadths in the wet season compared to the dry. This difference in seasonality has been previously documented in African mammal diets35 perhaps because some prey species are more abundant and more easily accessible68,69 during the birthing season or because rainfall leads to prey dispersing across the landscape. As a result, predators can no longer rely on high prey densities near predictable waterhole locations and may be less selective, consuming whatever prey they encounter. However, because spotted hyena sample sizes were lower than those for lions, estimates of hyena dietary niche breadth should be interpreted with caution.
The results are consistent with moderate dietary overlap between both carnivore species, as indicated by Pianka’s index (O = 0.75). This pattern was generally similar across seasons, with only minor fluctuations between wet and dry periods. Apart from domestic livestock species, all prey species detected in the spotted hyena diet were also found in the lion diet. Extensive dietary overlap may indicate occurrences of kleptoparasitism by lions and spotted hyenas, which previously have been reported34. Additionally, lions and spotted hyenas are documented competitors that consume similar prey items12,36 and exhibit comparable temporal activity patterns20,21,22. Lions and spotted hyenas have been documented partitioning spatially70,71, though some studies have found a high degree of spatial overlap23,24. Further, hunting styles can differ between the two predators; lions conceal behind vegetation to pursue prey23, yet al.so hunt in open areas26, while spotted hyenas typically chase prey in more open areas29,30 but also readily scavenge carcasses when available28,31. Thus, our findings indicate that lions and spotted hyenas in Etosha may partition their prey resources by age or sex class, or spatially to co-exist and we recommend further studies to quantify prey partitioning across demographic classes and spatial scales to better understand mechanisms of niche differentiation and coexistence among large carnivores.
In a review of spotted hyena diet across several countries, Hayward36 determined that gemsbok was the most frequently consumed prey species where it occurred, and plains zebra and springbok were typically consumed infrequently. Conversely, other studies50,72 found plains zebra and springbok were the main prey species for spotted hyenas in some areas of Etosha, while Berry73determined wildebeest and plains zebra were the main prey species in Etosha. Fester et al.74, found gemsbok and springbok were the main prey items in the Namib desert of southwest Namibia. In our study, spotted hyenas most frequently consumed plains zebras, gemsbok, and springbok, respectively. As such, spotted hyenas may target gemsbok but also consume plains zebras and springbok in the GEL due to their high abundance and availability compared to other ungulates. For example, in 2015, the Etosha density of gemsbok was estimated at 0.22 individuals/km2 while plains zebra was 0.77 individuals/km2 and springbok were 0.56 individuals/km249. As plains zebra and springbok are grazers75, and gemsbok are mixed feeders that primarily graze76,77, these results also align with Patterson et al.71, who found that spotted hyenas in Etosha select for areas of high grass cover during both seasons. Collectively, these studies indicate that in the GEL, spotted hyenas are likely hunting in open, grassy areas and primarily consuming grazing prey species.
The range of prey items for lions showed no strong selection for either browsers or grazers, as both (browsers: giraffe; mixed: eland, gemsbok; grazers: plains zebra, blue wildebeest75, were found as lion top prey species. Our results align with findings from Hayward and Kerley25 who found lions’ primary prey species were gemsbok, blue wildebeest, giraffe, eland, and zebra across several countries and Berry and73 who found lions’ primary prey items in Etosha were gemsbok, wildebeest, and plains zebra. Consistent with other findings, our study shows that lions likely consume small prey opportunistically25,78,79. This supports the optimal foraging theory, which suggests that a predator can distinguish among prey of differing profitability and typically select the most profitable types80. Large-bodied grazers were consumed more during the dry season in our study, which may indicate that lions were opting for higher caloric returns from large prey species to sustain themselves during higher periods of stress25. Additionally, in the dry season herbivores congregate at fixed waterhole locations in the GEL, increasing predator access to a higher density of potential prey. Conversely, in the wet season lions appear to consume fewer species, with several at higher frequencies relative to the dry season (e.g., hare spp., black-faced impala, giraffe), potentially reflecting opportunistic foraging or scavenging due to greater prey dispersion.
We found three spotted hyena samples with domestic livestock species (cow and goat). All samples were collected near or outside of Etosha in the northeast area (King Nehale communal conservancy) where livestock are abundant and spotted hyenas often leave the park and enter onto communal land40,41,71, sometimes resulting in retaliatory killings of carnivores after livestock depredation events81,82,83. Unfortunately, there is little information on the extent and demography of spotted hyenas killed in anthropogenic areas surrounding Etosha, but local farmers report frequent sightings of spotted hyenas on their farms84. No lion samples contained livestock species; while livestock depredation by lions has been reported around Etosha81, they are less likely to leave the park and cross the fence relative to spotted hyenas40,41 into King Nehale. Instead, the southern periphery of Etosha is a more common area for human-lion conflicts and livestock depredation events by lions81,85 where no samples were collected for this study. These findings contribute to our overall understanding of livestock depredation in a high human-carnivore conflict area.
The rarefaction analyses highlight important differences in sampling completeness between lions and spotted hyenas, which help contextualize niche overlap between lions and spotted hyenas, as well as niche breadth estimates. We could not detect prey items in 66 samples due to predator DNA swamping, which occurs when host DNA is present in much higher concentrations than prey DNA86. Host DNA swamping was especially prevalent in lion samples. While degradation of scat samples before collection can increase host DNA dominance87,88, both lion and spotted hyena samples were primarily collected opportunistically, and exact scat age was unknown. Thus, differences in digestive physiology and feeding behavior may influence host DNA amplification bias and contribute to interspecific variation in prey detection rates89. Further research is needed to assess how such physiological and behavioral traits affect host DNA swamping and prey DNA degradation in dietary metabarcoding studies, particularly among apex carnivores like lions and spotted hyenas. Additionally, future metabarcoding studies may improve prey detection by explicitly optimizing sequencing depth, particularly for rare or highly degraded prey DNA. For lions, dry-season rarefaction curves approached asymptotic levels, and Chao1 estimates suggested that only a small number of prey species were likely undetected, indicating that our dietary characterization for lions is robust and provides a reliable picture of dry season prey use. In contrast, the rarefaction curve for spotted hyenas showed lower observed richness and substantially higher Chao1 estimates, particularly in the dry season where uncertainty in prey species detection was large, reflecting potential under-sampling. This limitation suggests that while lions and spotted hyenas appear to have high dietary overlap, the true extent of spotted hyena prey use may be broader than captured in this study. Of the 24 potential mammalian prey species in the system >5 kg, lions consumed 13 and hyenas consumed 10, indicating that both predators utilized only a subset of the available prey community. Consequently, the high dietary overlap we observed may partially reflect more complete sampling of lion diets compared to spotted hyenas. Nonetheless, the combination of dietary niche overlap and breadth results indicates that both species rely heavily on shared prey resources, with lions exhibiting more stable and well-characterized prey use across seasons. Future work that includes expanded spotted hyena sampling could help clarify whether spotted hyena dietary breadth converges more strongly with or diverges from that of lions under different seasonal conditions.
We found notable differences in frequently consumed prey species and dietary niche breadth for both lions and spotted hyenas between study sites, likely reflecting variation in prey abundance and community composition within the sampled areas. Although both Etosha and Ongava are protected areas, they are managed by different entities and are different sizes, which may influence wildlife distributions and prey availability. However, the small number of spotted hyena samples from Ongava (n = 2) limited our ability to accurately characterize their dietary niche breadth at that site. Additionally, because sampling did not encompass the entire extent of Etosha, we were unable to provide a comprehensive synthesis of lion and spotted hyena diets across the entire GEL. Despite these limitations, this study provides valuable insights into the diets of lions and spotted hyenas where dietary data, particularly derived from high-resolution DNA metabarcoding, are lacking.
Our study contributes to the area of emerging research using DNA metabarcoding analysis to compare diets of large, sympatric carnivores. The results indicate considerable dietary similarity between lions and spotted hyenas across both wet and dry seasons, though limitations in sample size, particularly for spotted hyenas, preclude strong conclusions about the extent of dietary overlap or niche partitioning. Our findings suggest that prey availability and opportunistic feeding behavior may be the primary drivers of diet composition in this well-preserved carnivore guild. Despite differences in dietary niche breadth, both species showed high overlap in key prey taxa, perhaps due to opportunistic scavenging and kleptoparasitism. These findings emphasize the importance of maintaining abundant ungulate populations to support diverse predator communities. Importantly, the detection of domestic livestock in spotted hyena diets highlights the potential for human–wildlife conflict, emphasizing the need for continued management interventions around Etosha. Further diet monitoring is essential for detecting shifts in predator–prey dynamics in response to environmental change, prey population fluctuations, or increased anthropogenic pressures. Future research should also explore the spatial and temporal aspects of scavenging and interspecific interactions to better inform adaptive management and conservation strategies in protected areas and surrounding multi-use landscapes.
Data availability
The datasets generated and analysed during the current study are available in the NCBI Sequence Read Archive repository at [https://www.ncbi.nlm.nih.gov/sra/PRJNA1357869](https:/www.ncbi.nlm.nih.gov/sra/PRJNA1357869) .
References
del Rio, C. M. et al. The importance of large carnivores to healthy ecosystems. Endanger. Species Update. 18 (5), 202–210 (2001).
Di Marco, M. et al. A retrospective evaluation of the global decline of carnivores and ungulates. Conserv. Biol. 28, 1109–1118 (2014).
Parmesan, C. Ecological and evolutionary responses to recent climate change. Annu. Rev. Ecol. Evol. Syst. 37, 637–669 (2006).
Ripple, W. J. et al. Status and ecological effects of the world’s largest carnivores. Science 343, 1241484 (2014).
Carbone, C. & Gittleman, J. L. A common rule for the scaling of carnivore density. Science 295, 2273–2276 (2002).
Mills, M. G. & Harvey, M. African Predators (Struik, 2001).
Burgar, J. M., Burton, A. C. & Fisher, J. T. The importance of considering multiple interacting species for conservation of species at risk. Conserv. Biol. 33 (3), 709–715 (2019).
Linnell, J. D. & Strand, O. Interference interactions, co-existence and conservation of mammalian carnivores. Divers. Distrib. 6 (4), 169–176 (2000).
Chase, J. M. et al. The interaction between predation and competition: a review and synthesis. Ecol. Lett. 5 (2), 302–315 (2002).
Chesson, P. & Kuang, J. J. The interaction between predation and competition. Nature 4567219, 235–238 (2008).
Vanak, A. T. et al. Moving to stay in place: behavioral mechanisms for coexistence of African large carnivores. Ecology 94 (11), 2619–2631 (2013).
Hayward, M. W. & Kerley, G. I. Prey preferences and dietary overlap amongst Africa’s large predators. South. Afr. J. Wildl. Res. 38 (2), 93–108 (2008).
Karanth, K. U., Nichols, J. D., Kumar, N. S., Link, W. A. & Hines, J. E. Tigers and their prey: predicting carnivore densities from prey abundance. Proceedings of the National Academy of Sciences, 101(14): 4854–4858. (2004).
Schoener, T. W. Field experiments on interspecific competition. Am. Nat. 122 (2), 240–285 (1983).
Duncan, C., Chauvenet, A. L., McRae, L. M. & Pettorelli, N. Predicting the future impact of droughts on ungulate populations in arid and semi-arid environments. PloS one. 7 (12), e51490 (2012).
Walther, G-R. et al. Ecological responses to recent climate change. Nature 416, 389–395 (2002).
Zidon, R., Garti, S., Getz, W. M. & Saltz, D. Zebra migration strategies and anthrax in Etosha National Park. Namibia Ecosphere. 8 (8), e01925 (2017).
Radloff, F. G. T. & du Toit, J. T. Large predators and their prey in a southern African savanna: a predator’s size determines its prey range size. J. Anim. Ecol. 73, 410–423 (2004).
Hatton, I. A. et al. The predator-prey power law: Biomass scaling across terrestrial and aquatic biomes. Science 3496252, aac6284 (2015).
Hayward, M. W. & Hayward, G. J. Activity patterns of reintroduced lion Panthera leo and spotted hyaena Crocuta crocuta in the Addo Elephant National Park, South Africa. Afr. J. Ecol. 45 (2), 135–141 (2007).
Mills, M. G. & Biggs, H. C. Prey apportionment and related ecological relationships between large carnivores in Kruger National Park. In: Dunstone, N. & Gorman, M.L. Zoological Society of London Symposia 65: 253–268 (1993).
Patterson, J. R., Ndlovu, N., Beasley, J. C. & Périquet, S. Effects of human presence on African mammal waterhole attendance and temporal activity patterns. J. Zool. 325 (3), 210–223 (2024).
Périquet, S., Fritz, H. & Revilla, E. The Lion King and the Hyaena Queen: large carnivore interactions and coexistence. Biol. Rev. 90 (4), 1197–1214 (2015).
Swanson, A., Arnold, T., Kosmala, M., Forester, J. & Packer, C. In the absence of a landscape of fear: How lions, hyenas, and cheetahs coexist. Ecol. Evol. 6 (23), 8534–8545 (2016).
Hayward, M. W. & Kerley, G. I. Prey preferences of the lion (Panthera leo). J. Zool. 267 (3), 309–322 (2005).
Stander, P. E. Cooperative hunting in lions: the role of the individual. Behav. Ecol. Sociobiol. 29, 445–454 (1992).
Amorós, M., Gil-Sánchez, J. M., López‐Pastor, B. D. L. N. & Moleón, M. Hyaenas and lions: how the largest African carnivores interact at carcasses. Oikos 129 (12), 1820–1832 (2020).
Moleón, M., Sánchez-Zapata, J. A., Sebastián‐González, E. & Owen‐Smith, N. Carcass size shapes the structure and functioning of an African scavenging assemblage. Oikos 124 (10), 1391–1403 (2015).
Kruuk, H. The Spotted Hyena: A Study of Predation and Social Behavior (University of Chicago Press, 1972).
Mills, M. G. Kalahari Hyaenas: the Comparative Behavioural Ecology of Two Species (Allen & Unwin, 1990).
Watts, H. E. & Holekamp, K. E. Ecological determinants of survival and reproduction in the spotted hyena. J. Mammal. 90, 461–471 (2009).
Holekamp, K. E., Smale, L., Berg, R. & Cooper, S. M. Hunting rates and hunting success in the spotted hyena (Crocuta crocuta). J. Zool. 242 (1), 1–15 (1997).
Salnicki, J., Teichmann, M., Wilson, V. & Murindagomo, F. Spotted hyaenas, Crocuta crocuta, prey on new-born elephant calves in Hwange National Park. Zimbabwe Koedoe. 44 (2), 79–83 (2001).
Trinkel, M. & Kastberger, G. Competitive interactions between spotted hyenas and lions in the Etosha National Park, Namibia. Afr. J. Ecol. 43 (3), 220–224 (2005).
Vissia, S., Virtuoso, F. A., Bouman, A. & van Langevelde, F. Seasonal variation in prey preference, diet partitioning and niche breadth in a rich large carnivore guild. Afr. J. Ecol. 61 (1), 141–152 (2023).
Hayward, M. W. Prey preferences of spotted hyena (Crocuta crocuta) and degree of dietary overlap with the lion (Panthera leo). J. Zool. 270, 666–614 (2006).
Stein, E. D., Martinez, M. C., Stiles, S., Miller, P. E. & Zakharov, E. V. Is DNA barcoding actually cheaper and faster than traditional morphological methods: Results from a survey of freshwater bioassessment efforts in the United States? PLoS One, 9(4), e95525 (2014).
Galan, M. et al. Metabarcoding for the parallel identification of several hundred predators and their prey: Application to bat species diet analysis. Mol. Ecol. Resour. 18 (3), 474–489 (2018).
Turner, W. C. et al. Africa’s drylands in a changing world: Challenges for wildlife conservation under climate and land-use changes in the Greater Etosha Landscape. Global Ecol. Conserv. 38, e02221 (2022).
Melton, M. H. Drivers of movement and resource selection of wildlife living in a protected area in sub-Saharan Africa. Master’s thesis. Athens, GA: University of Georgia. (2024).
Naha, D. et al. Fencing affects movement patterns of two large carnivores in Southern Africa. Front. Ecol. Evol. 11, 1031321 (2023).
Nakanyala, J., Heita, J., Lewis, E., Siyambango, N. & Lendelvo, S. Local farmers’ perceptions of human-wildlife conflicts in the King Nehale Conservancy, Namibia. J. Stud. Humanit. Social Sci. 11 (1&2), 99–116 (2022).
De Beer, Y., Kilian, W., Versfeld, W. & Van Aarde, R. J. Elephants and low rainfall alter woody vegetation in Etosha National Park, Namibia. J. Arid Environ. 64 (3), 412–421 (2006).
Atlas of Namibia Team. Atlas of Namibia: its land, water and life (Namibia Nature Foundation, 2022).
Le Roux, C. J. G., Grunow, J. O., Bredenkamp, G. J., Morris, J. W. & Scheepers, J. C. A classification of the vegetation of the Etosha National Park. South. Afr. J. Bot. 54 (1), 1–10 (1988).
Berry, C. & Loutit, B. Trees and Shrubs of the Etosha National Park 3rd edn (Namibia Scientific Society, 2002).
Berezin, J. L., Odom, A. J., Hayssen, V. & O’Connell-Rodwell, C. E. A snapshot into the lives of elephants: camera traps and conservation in Etosha National Park. Namibia Divers. 15 (11), 1146 (2023).
Engert, S. Spatial variability and temporal periodicity of rainfall in the Etosha National Park and surrounding areas in northern Namibia. Madoqua 1997(1), 115–120 (1997).
Kilian, W. & Kolberg, H. Aerial survey of Etosha National Park (Internal Report of Ministry of Environment and Tourism, 2015).
Trinkel, M. Prey selection and prey preferences of spotted hyenas Crocuta crocuta in the Etosha National Park, Namibia. Ecol. Res. 25 (2), 413–417 (2010).
Trinkel, M. A keystone predator at risk? Density and distribution of the spotted hyena (Crocuta crocuta) in the Etosha National Park, Namibia. Can. J. Zool. 87 (10), 941–947 (2009).
Pompanon, F. et al. Who is eating what: Diet assessment using next generation sequencing. Mol. Ecol. 21 (8), 1931–1950 (2012).
Xiong, M. et al. 2107. Molecular dietary analysis of two sympatric felids in the Mountains of Southwest China biodiversity hotspot and conservation implications. Sci. Rep., 7: 41909 (2017).
Evans, N. T. et al. Quantification of mesocosm fish and amphibian species diversity via environmental DNA metabarcoding. Mol. Ecol. Resour. 16 (1), 29–41 (2016).
Edgar, R. C. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing 081257 (BioRxiv, 2016a).
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Wheeler, D. L. GenBank Nucleic Acids Res., 1(33): 34–38. (2005).
Shively, K. A. et al. Metabarcoding reveals striking dietary variation in a reintroduced mesocarnivore (Journal of Mammalogy, 2025).
Forin-Wiart, M. A. et al. Evaluating metabarcoding to analyse diet composition of species foraging in anthropogenic landscapes using Ion Torrent and Illumina sequencing. Sci. Rep. 8 (1), 17091 (2018).
R Core Team. R: A language and environment for statistical computing (Version 4.4.0) [Computer software]. R Foundation for Statistical Computing. (2024). https://www.R-project.org/
Pianka, E. R. The structure of lizard communities. Annu. Rev. Ecol. Syst. 4, 53–74 (1973).
Oksanen, J. et al. (2025). _vegan: Community Ecology Package_. R package version 2.6–10.
Zhang, J. spaa: Species Association Analysis. R package version 0.2.2. (2016). https://CRAN.R-project.org/package=spaa
Hurlbert, S. H. The measurement of niche overlap and somerelatives. Ecology 59 (1), 67–77 (1978).
Smith, E. P. Niche breadth, resource availability, and inference. Ecology 63 (6), 1675–1681 (1982).
Chao, A. Nonparametric estimation of the number of classes in a population. Scandinavian J. Stat., 265–270 (1984).
Clarke, K. R. Non-parametric multivariate analyses of changes in community structure. Aust. J. Ecol. 18 (1), 117–143 (1993).
Carvalho, J. C. & Cardoso, P. Decomposing the causes for niche differentiation between species using hypervolumes. Front. Ecol. Evol. 8, 243 (2020).
Pereira, L. M., Owen-Smith, N. & Moleón, M. Facultative predation and scavenging by mammalian carnivores: seasonal, regional and intra-guild comparisons. Mammal Rev. 44 (1), 44–55 (2014).
Trinkel, M. Climate variability, human wildlife conflict and population dynamics of lions Panthera leo. Naturwissenschaften 100 (4), 345–353 (2013).
Evers, E. E. et al. Varying degrees of spatio-temporal partitioning among large carnivores in a fenced reserve, South Africa. Wildl. Res. 49 (5), 477–490 (2022).
Patterson, J. R. et al. Effects of tourism on seasonal movements and fine-scale habitat selection of African lions and spotted hyenas in Etosha National Park, Namibia (Global Ecology and Conservation, 2025). e03681.
Gasaway, W. C., Mossestad, K. T. & Standers, P. E. Food acquisition by spotted hyaenas in Etosha National Park, Namibia: predation versus scavenging. Afr. J. Ecol. 29 (1), 64–75 (1991).
Berry, H. H. Abnormal levels of disease and predation as limiting factors for wildebeest in the Etosha National Park. Madoqua, 1981(4): 241–253. (1981).
Fester, K. S. M., Hockings, G., van Vuuren, R. J. & van Vuuren, M. Spotted hyaena Crocuta crocuta feeding ecology and selectivity of large herbivorous prey in the Namib desert. Ecol. Evol. 11 (9), 3672–3678 (2021).
Kingdon, J. The Kingdon field guide to African mammals (Bloomsbury Publishing, 2015).
Lehmann, D. et al. Spatiotemporal responses of a desert dwelling ungulate to increasing aridity in North-eastern Namibia. J. Arid Environ. 179, 104193 (2020).
Sponheimer, M. et al. Diets of southern African Bovidae: stable isotope evidence. J. Mammal. 84 (2), 471–479 (2003).
Barnardo, T. et al. Opportunistic feeding by lions: non-preferred prey comprise an important part of lion diets in a habitat where preferred prey are abundant. Mammal Res. 65 (2), 235–243 (2020).
Davidson, Z. et al. Seasonal diet and prey preference of the African lion in a waterhole-driven semi-arid savanna. PloS One. 8 (2), e55182 (2013).
Krebs, J. R. Optimal foraging: decision rules for predators. In Behavioural ecology: an evolutionary approach: 23–63 (eds Krebs, J. R. & Davies, N. B.) (Blackwell Scientific, 1978).
Goelst, C. E. Assessing patterns of spatial occurrence and human-carnivore conflict for African lions (Panthera leo) in and around Etosha National Park, Namibia. Master’s thesis. New York City, NY: Columbia University. (2018).
Naha, D., Kilian, J. W., Goelst, C., Cloete, C. & Beasley, J. C. Anthropogenic mortality risk of lions depends on a combination of environmental, climatic, and cultural factors within a Sub-Saharan African ecosystem. Global Ecol. Conserv. 59, e03514 (2025).
Stander, P. E. An analysis of the spatial, temporal, and demographic patterns of lions destroyed along the borders of Etosha National Park over a 20-year period. In Unpublished Internal Report (Ministry of Environment and Tourism, 2004).
Lendelvo, S. M., Angula, M. N. & Mfune, J. K. E. Indigenous knowledge used in the management of human–wildlife conflict along the borders of the Etosha National Park. Indigenous Knowl. Namibia. 16, 219–240 (2019).
Trinkel, M., Fleischmann, P. H. & Slotow, R. Electrifying the fence or living with consequences? Problem animal control threatens the long-term viability of a free-ranging lion population. J. Zool. 301, 41–50 (2016).
Shi, Y., Hoareau, Y., Reese, E. M. & Wasser, S. K. Prey partitioning between sympatric wild carnivores revealed by DNA metabarcoding: a case study on wolf (Canis lupus) and coyote (Canis latrans) in northeastern Washington. Conserv. Genet. 22 (2), 293–305 (2021).
Oehm, J., Juen, A., Nagiller, K., Neuhauser, S. & Traugott, M. Molecular scatology: how to improve prey DNA detection success in avian faeces? Mol. Ecol. Resour. 11, 620–628 (2011).
Thuo, D. et al. Food from faeces: Evaluating the efficacy of scat DNA metabarcoding in dietary analyses. PLoS One. 14 (12), e0225805 (2019).
Cuff, J. P. et al. The predator problem and PCR primers in molecular dietary analysis: swamped or silenced; depth or breadth? Mol. Ecol. Resour. 23 (1), 41–51 (2023).
Edgar, R. C. UCHIME2: improved chimera prediction for amplicon sequencing. BioRxiv, 12, 074252. (2016b).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet 17 (1), 10–12 (2011).
Oxford Nanopore Technologies. Dorado basecalling software (Version 0.7.1). GitHub. (2024). https://github.com/nanoporetech/dorado
Smuts, G. L., Whyte, I. J. & Dearlove, T. W. A mass capture technique for lions. Afr. J. Ecol. 15 (1), 81–87 (1977).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Acknowledgements
We thank the Ongava Game Reserve for providing some equipment and staff assistance. We thank the field technicians for their work in helping to collect and process scat/fecal samples. Thank you to the University of Georgia and the US Department of Energy Office of Environmental Management for funding assistance.
Funding
This work was supported by University of Georgia and the US Department of Energy Office of Environmental Management Award Number DE-EM0005228 to the University of Georgia Research Foundation. Disclaimer: This manuscript was prepared as an account of work sponsored by an agency of the United States Government. Neither the United States Government nor any agency thereof, nor any of their employees, makes any warranty, express or implied, or assumes any legal liability or responsibility for the accuracy, completeness, or usefulness of any information disclosed, or represents that its use not infringe privately owned rights. Reference herein to any specific commercial product, process, or service by trade name, trademark, manufacturer, or otherwise does not constitute or imply its endorsement, recommendation, or favoring by the United States Government or any agency thereof. The views and opinions of the authors expressed herein do not necessarily state or reflect those of the United States Government or any agency thereof.
Author information
Authors and Affiliations
Contributions
Jessica Patterson, James Beasley, and Stephanie Periquet-Pearce contributed to the study conception and design. Material preparation and data collection were performed by all authors. Date analyses, the first draft, and revisions of the manuscript were completed by Jessica Patterson. All authors commented on previous versions of the manuscript and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Patterson, J.R., Périquet-Pearce, S., Melton, M.H. et al. Revealing seasonal dietary niche overlap among sympatric large carnivores using DNA metabarcoding. Sci Rep 16, 13590 (2026). https://doi.org/10.1038/s41598-026-43423-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-026-43423-w